BIOL214 Exam 2 - Heritable Variation (ch 5+6)
1/101
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
102 Terms
Important Molecules for Evolution
3 kinds of molecules are important for evolution
proteins
DNA
RNA
Proteins
chains of amino acids
can act as enzymes, structural support, regulate passage of substances across cell membrane, immune function, and coordinate signaling pathways
Organelles Involved in Protein Manufacture
ribosomes
golgi apparatus
endoplasmic reticulum
Protein Production
transcription: in the nucleus results in DNA → RNA
translation: at the ribosomes results in RNA → proteins
After Transcription…
RNA travels from nucleus → ribosome
proteins are made in ribosomes (some bound to rough ER, some float in the cytoplasm)
Endoplasmic Reticulum
filled with membranes
smooth ER: contains enzymes that produce lipids
rough ER: contains ribosomes that produce many types of proteins
Golgi Apparatus
proteins are finalized and packaged in the golgi apparatus
proteins are finished in vesicles (small bubbles of membrane)
Deoxyribose Nucleic Acid (DNA)
holds instructions for all living things
double helix with two strands of nucleotide strings
each nucleotide contains a sugar, a phosphate, and a base
DNA Bases
Adenine (A) = Thymine (T)
Guanine (G) = Cytosine (C)
hydrogen-bonding
Eukaryotic DNA is Organized into Chromosomes
DNA is organized into chromosomes in which the molecule is wound around histones (proteins)
the winding and unwinding of DNA around histones can expose or hide genes
regulation of gene expression
Ploidy
number of copies of unique chromosomes in a cell
haploid (n)
1 copy of each chromosome
diploid (2n)
2 copies of each chromosome
triploid (3n)
3 copies of each chromosome
tetaploid (4n)
4 copies of each chromosome
Sex Chromosome
a chromosome that pairs during meiosis but differs in copy number between males and females (X and Y)
Autosome
a chromosome that does not differ between sexes
Gene
segment of DNA whose nucleotide sequences code for proteins/RNA or regulates the expression of other genes
Gene Expression
process by which information of a gene is transformed into a product
RNA Polymerase
the enzyme that builds the single-stranded RNA molecule from the DNA template during transcription
Transcription
the process that takes place where RNA polymerase reads a coding sequence of DNA to the ribosome where it can be translated into protein
Translation
the process that takes place when a strand of mRNA is decoded by a ribosome to produce a protein
Ribosomes Translate mRNA into Protein
each mRNA acts as a template for building a protein
the ribosome reads 3 bases at a time (codon)
tRNA adds the correct amino acid
Hormones
molecular signals that flow through the body that can alter the expression of genes
Upstream and Downstream
upstream: → 5’ end of RNA/DNA
downstream: → 3’ end of RNA/DNA
Gene Control Region
an upstream section of DNA that includes the promoter region and other regulatory sequences that influence transcription of DNA
Repressor
protein that binds to a sequence of DNA or RNA and inhibits the expression of one or more genes
Transcription Factor
protein that regulates the expression of a gene by binding to a specific DNA sequence in association with the gene sequence
Enhancer
short sequence of DNA within the gene control region where activator proteins bind to initiate gene expression
MicroRNA
group of RNAs that act as post-transcriptional regulators of gene expression
bind to complementary sequences on specific mRNAs and can enhance or silence gene translation
Introns
non-coding sequences
longer than exons
removed during RNA splicing
Exons
coding sequences
RNA Splicing
spliceosome: group of proteins that removes introns from transcripts
occurs after transcription
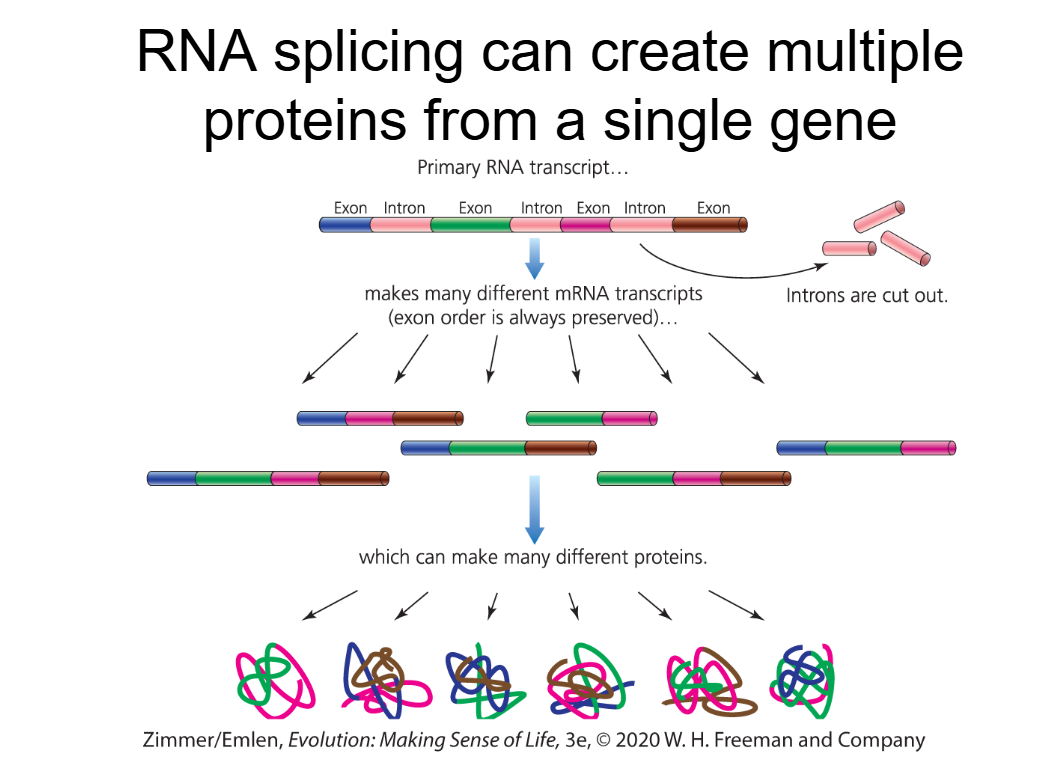
Alternative Splicing
creating multiple proteins from a single gene
Prokaryotic Gene Expression
primarily controlled at the level of transcription
translation and transcription occur simultaneously in the cytoplasm
Eukaryotic Gene Expression
controlled at the levels of epigenetics, transcription, post-transcription, translation, and post-translation
regulated during transcription and RNA processing (nucleus), protein translation (cytoplasm)
further regulation can occur through post-translational modifications of proteins
Mobile Genetic Element
type of DNA that can move around in the genome and plasmids
Plasmid
a molecule of DNA found most often in bacteria that can replicate independently of chromosomal DNA
Vertical Gene Transfer
receiving genetic material from an ancestor
Horizontal Gene Transfer
transfer of genetic material between organisms without reproduction
can be inherited once added to genome
Pseudogenes
nonfunctional
often form after a gene has been duplicated and one or more of the redundant copies lose their function
Types of Mutation
point mutation
insertion
deletion
frameshift
duplication
inversion
chromosome fusion
aneuploidy
genome duplication
Point Mutation
a single base changes from one nucleotide to another (substitution)
Insertion
a segment of DNA is inserted into the middle of an existing sequence
Deletion
a segment of DNA is deleted
Frameshift Mutation
insertion of 1 or 2 bases changes the codon, modifying all amino acids coded downstream
Duplication
a segment of DNA is copied a second time
Inversion
a segment of DNA is flipped around and inserted backwards into its original position
Chromosome Fusion
two chromosomes are joined together
Aneuploidy
chromosomes are duplicated or lost
Genome Duplication
leads to increased ploidy
Cis-Acting Element
a stretch of DNA located near a gene that influences the expression of that gene
Trans-Acting Element
sequence of DNA located away from a gene (e.g. on another chromosome) that codes for a protein, microRNA, or other diffusible molecules that they influence gene expression
Somatic Mutation
mutation that affects cells in the body of an organism
not passed down to offspring in animals
Germline Mutation
mutation that affects the gametes of an individual and can be transmitted from parents to offspring
results in heritable genetic variation
Coding Region
Type of Mutation
substitution, insertion, deletion, duplication
Consequences for Gene Action
alter the product of the gene and thus its function or activity
Cis-Regulatory Regions
Type of Mutation
substitution, insertion, deletion, duplication that alters the binding affinity of promoters, activators, repressors, etc.
Consequences for Gene Action
alters the timing, location, or level of expression of the gene. Alters the developmental or environmental context in which the gene is expressed
Trans-Regulatory Regions
Type of Mutation
mutation to coding regions of trans-acting factor
mutatiion to cis- or trans-regulatory regions of trans-acting factors
Consequences for Gene Action
alters the binding affinity and thus the activity of a promoter, activator, repressor, etc.
alters where, when, or to what extent inhibitory, activating, or other trans-acting regulatory factors are expressed
Physiological Pathways (ex. Hormones)
Type of Mutation
mutations altering where, when, or how much an endocrine signal is produced
Consequences for Gene Action
alters the timing, location, or level of expression of the gene.
alters the developmental or environmental context in which the gene is expressed
Albinism is an example of what mutation?
point mutation
Independent Assortment
ensures novel combinations of alleles
genes are inherited independently of each other
Genetic Recombination
generates variation
during the production of gametes, each pair of chromosomes crosses over and exchanges segments of DNA
Genotype
the genetic makeup of an individual
Phenotype
an observable, measurable characteristic as the manifestation of the genotype of an organism
Polyphenic Trait
single genotype produces multiple phenotypes depending on environment
Quantitative Traits
have a continuous distribution of phenotypic variation
influenced by multiple genes
normal distribution
Quantitative Trait Locus (QTL)
the analysis of such can help discover genes influencing quantitative traits
Environmental Influences on Gene Expression
during development, cells respond to a multitude of signals from their environment
morphogen
phenotypic plasticity
Morphogen
signaling molecule that flows between nearby cells
alters the expression of target genes
Phenotypic Plasticity
changes in phenotype produced by a single genotype in different environments
tailors organism to environment
Population Genetics
the study of the distribution and frequencies of alleles in populations
how and why allele frequencies change
Genetic Locus
location of a specific gene or sequence of DNA on a chromosome
Homozygous
individual carries two copies of the same allele at a locus
Heterozygous
individual carries different alleles at a locus
Hardy-Weinburg Conditions
no mutations
mating is random
no selection (equal survival)
very large population size
no gene flow in or out
Hardy-Weinberg Equation
Alleles:
p = frequency of dominant allele
q = frequency of dominant allele
p + q = 1
Genotypes:
p² + 2pq + q² = 1
p² = frequency of homozygous dominant genotype
2pq = frequency of heterozygous genotype
q² = frequency of homozygous recessive genotype
Microevolution
an evolving population is one that is showing genetic change over generations
Hardy-Weinberg lets us detect microevolution
Hardy-Weinberg Equilibrium Assumptions
allele frequencies of a population will not change if:
population is infinitely large
genotypes do not confer differences in fitness
there is no mutation
mating is random
there is no migration
Are the Hardy-Weinberg Equilibrium assumptions reasonable?
population is infinitely large
populations are always finite, but some are large enough to function nearly as though they are infinite
genotypes do not confer differences in fitness
natural selection imposes differential survival and reproduction
there is no mutation
mutation rates have been studied and are known
mating is random
mating is assumed to be random at specific loci of interest
there is no migration
this may occasionally be true but not often
What results in genetic drift?
random sampling error
higher in a smaller sample
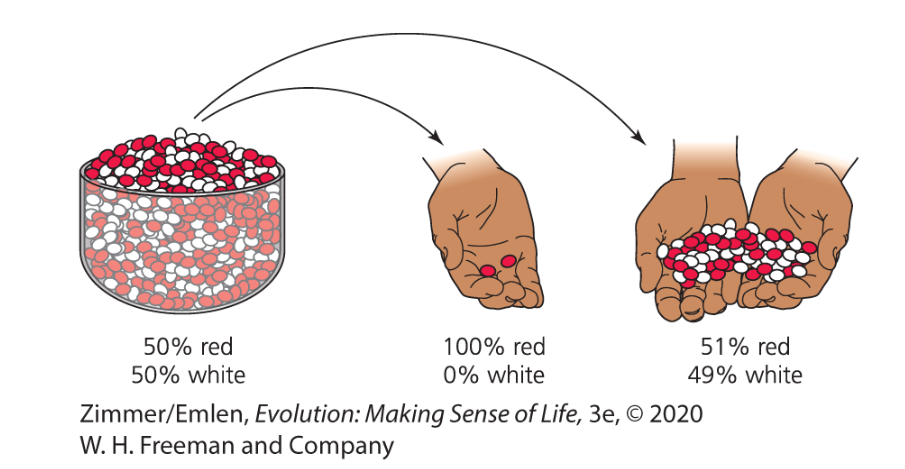
Genetic Drift Reduces Genetic Variation
small populations experience strong drift
some alleles become fixed in the population
some alleles disappear
Bottlenecks
reduce genetic variation
results in nonrepresentative set of alleles for subsequent populations
even after population size rebounds
rare alleles more likely to be lost during bottleneck event
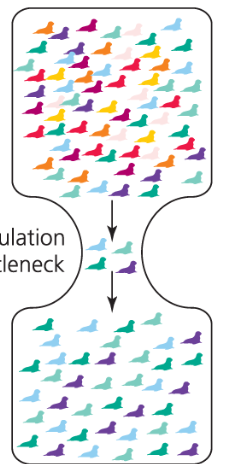
Founder Effect
a type of bottleneck resulting from a small number of individuals colonizing a new, isolated habitat
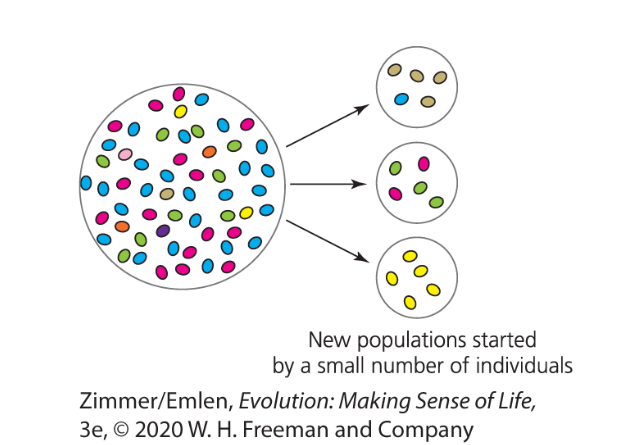
Fitness
the survival and reproductive success of an individual with a particular
components:
survival to reproductive age
mating success
fecundity
Relative Fitness (w)
contribution of individuals with one genotype compared with the average contribution of all individuals in the population
Contribution of Alleles to Fitness
fitness is a product of an organism’s entire phenotype, but this is difficult to assess
Average Excess Fitness
difference between the relative contribution of individuals with one genotype and the average fitness of the population as a whole
Δp = p x (aA1/ϖ)
Δp = change in allele frequency due to selection
p = frequency of the A1 allele
ϖ = average fitness of the population
aA1 = average excess of fitness for the A1 allele
In what kind of population is natural selection more effective in?
Large populations
Pleiotropy
mutation in a single gene affects more than one phenotypic trait
may constrain evolution
Antagonist Pleiotropy
beneficial effects for one trait but detrimental effects for other traits
net effect on fitness determines outcome of selection
Pesticide Resistance and Pleiotropy
the frequency of the Ester1 (resistance to pesticides) gene increased in response to the use of pesticides in coastal areas
mosquitoes with this gene are more susceptible to predation by spiders
Pesticide Resistance and Antagonistic Pleiotropy
Ester1 raised the fitness of carriers in coastal areas because of insecticide use
carrying this allele farther inland proved detrimental in escaping predation
Negative Selection
alleles that lower fitness experience negative selection
Positive Selection
alleles that increase fitness experience positive selection
What model is useful for running evolution experiments and why?
bacteria
haploid genetics are easier to study
diploid genetics are more complex due to interactions between two alleles
Additive Alleles
homozygous condition yields twice the phenotypic effect for the gene as compared with heterozygotes
Dominance
dominant allele masks presence of recessive in heterozygote
Mutations Generate Variations in Populations
mutation rates for any given gene are low
per genome and population, many new mutations arise each generation
source of variation upon which selection and drift act
Mutation-Selection Balance
equilibrium frequency reached through “tug-of-war” between negative selection on deleterious alleles and new mutations
explains persistence of deleterious mutations in populations
Balancing Selection
selection that favors more than one allele
maintains genetic diversity in a population by keeping alleles at frequencies higher than would be expected by chance or mutation alone
Negative Frequency-Dependent Selection
common phenotypes are selected against
rare phenotypes are favored
heterozygote advantage
malaria and sickle-cell anemia
Inbreeding Coefficient (F)
F = probability that two alleles at any locus of an individual are identical by descent
Inbreeding Depression
results in reduced fitness
rare recessive alleles are expressed in homozygous state
high inbreeding associated with low infant survival rates
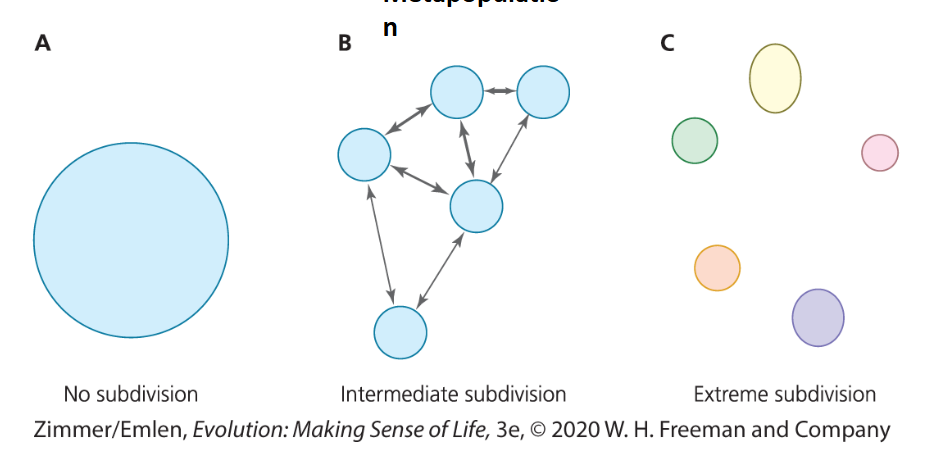
Population Subdivision
depends on landscape features and the relative degree of motility of individuals in the population
subdivided populations show distinct genetic structure (heterogeneity in allelic frequencies)