Genetic Analysis Exam 2
1/110
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
111 Terms
When two genes interact…
Mendelian ratios change!!!
Epistasis
Ability of one mutant to mask another.
Epistasis Auxotrophic Yeast Example
Example = auxotrophic yeast mutants ade2 and ade3 (fail to grow on media lacking Adenine). ade2 = red. ade3 = white. Double mutant = white. These mutants are on different genes in an enzymatic pathway. ade3 is epistatic to ade2 (stands upon) - epistatic gene is first in the pathway (upstream).
ADE3 = enzyme converts to red pigment. ADE2 = enzyme converts to white pigment (after ADE3). ade2 mutant = no conversion to white… remains red. ade3 mutant = no conversion… remains white. Both mutants = stays white (no conversion at all).
Epistasis Radiation-Sensitive Mutants
Increased radiation = decreased survival. WT strain = can withstand radiation. rfa1 mutant = steeper decline with more radiation. rad52 mutant = even STEEPER decline. Double mutant = same survival curve as rad52 (masks effect of rfa1). rad52 is epistatic to rfa1. Two mutants in same pathway OR two different, albeit redundant, pathways.
Epistasis Mice Pigment Pathway
WT = agouti (brown). Requires two enzymes to move from white (no pigment; aabb) —> dark hair —> agouti (AABB). Cross agouti and white mouse —> F1 = all heterozygotes. Self-fertilize to create F2 —> 9:3:4 Mendelian ratios! Why? If you have “aa,” masks the impact of whether BB, Bb, or bb (ALL still white) because it occurs FIRST in the pathway (must convert from white to black before you can progress down enzymatic pathway). aa is epistatic over the B locus!
Inhibitors
Suppress a phenotype when NOT part of the pathway itself. These enzymes impact the functioning of enzymes in biosynthetic pathways.
Inhibitors - Chicken Feather Pigmentation
Inhibition of an enzyme results in lack of colored feathers (need enzyme for coloration) or colorless chickens (leghorn chickens). Inhibitor gene codes for inhibitor enzyme. Genes independently segregate. Perform a dihybrid cross (ccii x CCII = CcIi) and self-fertilize the F1 —> F2 = anytime an I is expressed means colorless no matter what “C” is OR anytime cc is expressed means colorless (leads to a 13:3 white:colored feather ratio).

Genetic Model System
Living organism that scientists use to study genetic phenomena by studying genes/functions. Chosen because 1) it has a well-understood genetic makeup, 2) easy to manipulate in lab, and 3) can be used to gain insight into broader biological processes (e.g., human genetics).
Examples of Genetic Model Systems
Yeast (Saccharomyces cerevisiae), fruit fly (Drosophila melanogaster), nematode worm (C. elegans), and mice (Mus musculus).
Why is it good to study genetics in haploids (and tricky in diploids)?
WHAT YOU SEE IS WHAT YOU GET (if you get a genotype, you see the phenotype in haploids… no dom-rec relationships).
Saccharomyces cerevisiae (Yeast)
16 chromosomes as haploid. 60% genes conserved in humans (can perform cross-species complementation = insert human gene in yeast, makes up for function, and can thus study human genes). Great for APOG = genetic differences manifest in phenotypic differences.
Yeast Life Cycle
Reproduce asexually via budding. Have haploid and diploid sexual life stages → a or alpha mating types (MATa, MATalpha). Two spores of opposing mating type fuse to form diploid (MATa/alpha). Meiosis/sporulation can form spores from the diploid (tetrad).
Yeast Double Mutants
Combine two separate mutations (one in each haploid) in a diploid to see how they interact (epistasis, complementation, synthetic lethality, inhibition, suppressors). GENETIC REDUNDANCY!
Synthetic Lethality
When a double mutant (mutants A and B) results in loss of function (e.g., two proteins serve as redundant backup mechanisms and the loss of BOTH = loss of function). Mutant in one = still function but slightly reduced (genetic redundancy). Indicated by ABSENCE OF DOUBLE MUTANT!
How can you study synthetic lethality?
Used to test whether a second gene is involved in redundant backup mechanisms (when only knowing one gene). Cross two haploid spores, each one with a mutant in a diff gene —> diploid yeast (MmBb) —> after sporulation, if synthetic lethality occurred, we would see NO double mutant (e.g., only 3 spores live). In diploids… 9:3:3 ratio (mmbb = dead).
Seen in yeast: from a heterozygous diploid (MmBb), we would see only three spores survive after sporulation (mb = synthetic lethality = dead).
Why is studying synthetic lethality important?
1) Assigns function to uncharacterized genes, 2) identifies genetic relationships between biological processes, and 3) increase understanding of genetic underpinnings of complex inheritable diseases.
Essential Genes
Essential for functioning (mutant knockout = lethal to cell).
Non-essential Genes
When knocked out, no phenotype! Usually where redundant backup mechanisms are located/have been involved (suggests EXTREME importance in functioning).
How could you study essential genes (normally mess with it = death)?
Create a conditional temperature-sensitive mutant (alive/functions at room T but DIES at higher T (37 degrees C… loss of function)). Aka conditional lethal!
High Copy Suppressor Gene
A gene that codes for EXTRA substrate of a protein produced by a mutated temperature sensitive mutant. Extra copies of substrate = mutation protein interacts with substrate more = functions/cell is viable = NEGATES MUTANT.
Suppressors
A gene that, when expressed, negates the effects of a mutant (usually temperature-sensitive) by producing extra substrate for the protein of the mutant to interact with (more likely to interact). HELPS a mutant.
Genomic Library
Chopped-up pieces of DNA of a WT strain (collection of DNA that represents an entire WT genome). Mutants subjected to one piece of DNA each. Which pieces of added DNA assist in the survival of the mutant?
Screening for a High-Copy Suppressor
Use a temp-sensitive conditional mutant, subject the strain to a genomic library (one piece added to each mutant colony), mutant grown at 30 degrees C. Then, heat up colony to 37 degrees C —> look at the colonies that do not die. Two explanations: 1) received WT copy of mutant gene (complementation) or 2) extra copy of a suppressor gene (more substrate-producing gene). Gene fragment assists survival! Allows for identification of suppressors!
Note - if diploid, make homozygous for mutant!
Two Major Methods of Genetic Analysis
Screen
Assay
Screen
A fishing expedition! Uses transformation of mutant with genomic library and selecting survivors under high T (isolate and study genes that permit growth at high T).
Assay
Cherry picking! Transform a mutant with a single gene and test if it supports growth at high T. Study the gene if yes!
Complementation
Taking a mutant and inserting the WT gene, making up for the loss of function of the mutant. The WT complements a conditional mutant!
Complementation Analysis in Yeast
First step is always mutagenesis. Often done in yeast —> mutagenize each spore with a different mutant (a and alpha) and cross. In the diploid, does each mutant complement each other (his1/HIS3 x HIS1/his3 - diploid can grow on media lacking histidine)?
Only complements if mutants (his1, his3) are on separate genes (e.g., different enzymes in a biosynthetic pathway). Same gene = would be a double mutant with the cross (NO function).
Output of Complementation Analysis
Complementation = if mutants are on DIFFERENT genes
NO Comp = if mutants are on SAME gene
Intragenic Complementation = when one mutation is on one end of a gene and can complement a mutation on the other end (RARE). Functions in tetramers (protein with subunits) when each mutation codes for different domains/subunits (each mutant can fulfill each other’s roles).
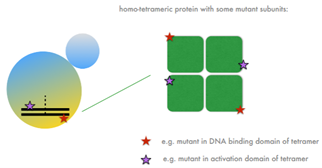
Complex Complementation Analysis
Cross MANY mutated haploids with a variety of mutants (ex. his-) to test for number of genes involved in pathway. Streak each mutant on rich media plates (separate). Combine streaks with OVERLAP (of diff mating types) in a cross format on a rich media —> move to plate lacking the necessary nutrient —> mating occurs only where streaks overlap AND where complementation occurs. Growth = complementation = diff genes! No growth at crosses = N/A = same gene.
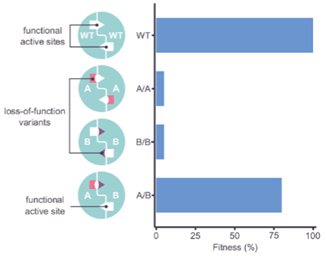
Clinical Correlation on Complementation
Looked at gene ASL involved in urea cycle. Discovered harmful mutants. Single mutants died (much LOWER fitness). When mutants are combined, recovery (pairing deleterious single mutants = GROWTH). Occurs in 4% of human genes. This is an example of intragenic complementation (two harmful variants = restore protein function). This find is interesting from a clinical perspective - this could be important for gene therapy (e.g., mutagenize an already-mutated gene). CRISPR gene editing.
Why do kids grow but adults do not? Why do sick people lose weight? Why is there a max to growth?
Tissue homeostasis where rate of cell death = rate of cell generation (reach at specific ages). Sickness = imbalance (death > generation). Based on mitosis or division of cells
Cyclical Division of Cells
Mitosis, meiosis, etc. Some cells divide more often than others (ex. muscle cells are slow to divide). Differential growth of cells can be seen in growth rates between organisms (ex. trees vs bamboo).
Cell Cycle of Non-germ line Cells
Gap 1 (G1) = growth. S phase = replication of DNA and prep for segregation (2n —> 4n… 46 to 92 in humans). Gap 2 (G2) = prep for mitosis. Mitosis!!!
Mitosis
Production of two identical daughter cells compared to the parent cell (e.g., 4n —> 2× 2n). Occurs in a ______ manner.
SYNCHRONIZED
Every chromosome replicates, condensates, aligns, and segregates at the SAME TIME. If not synchronized, could lose or gain chromosomes (aneuploidy = cause BIG problems).
Mitosis Steps
Interphase (G1, S, G2) —> Mitosis —> next interphase.
Mitosis: prophase, metaphase, anaphase, and telophase/cytokinesis.
Interphase
Cell growth and DNA replication
Early Prophase
Cohesion of replicated DNA into chromosomes. Condensed in nucleus. Produces structures that are more easily moved (b/c coiled and condensed by histones).
Late Prophase
Chromosomes appear double-stranded with each chromosome being composed of two sister chromatids joined at the centromere. Formation of spindle that attaches to kinetochores. The nuclear envelope (NE) starts to dissolve.
Spindle
Array of microtubules (parallel fibers) that extend from the poles of the cell to attach to each chromatid (1 per chromatid) at the kinetochore.
Kinetochore
Site of attachment at the centromere on chromosomes.
Metaphase
NE is gone. Microtubules exert tension on replicated chromosomes (pull), causing chromosomes to align at the metaphase plate (balance of tension from either direction). Can have mechanical issues —> only one microtubule attaches (replicated pair pulled to one side). What happens then…
Spindle Checkpoint
Cell checks whether microtubules are attached to each kinetochore and whether there is the generation of equivalent TENSION. No tension = cell is arrested in metaphase (before anaphase entry… no further mitosis).
Metaphase Plate
Imaginary line that is equidistant from the two spindle poles. MOST IMPORTANT MOMENT FOR SYNCHRONIZATION and INDEPENDENT ASSORTMENT.
Anaphase
Alignment is complete! Cohesins release. Individual chromatids move to the poles (sister chromatids pulled APART). Forms V-shapes as two arms of each chromatid follow the pulled centromere.
How do these microtubules pull on each chromatid?
At the kinetochore on each chromatid, the microtubules slowly depolymerize at each kinetochore end (e.g., slowly cut off portions of the microtubule… shortening distance of kinetochore to pole).
Telophase
NE reforms. Spindle dissolves. Cytokinesis is in progress (division of cytoplasm in two). Chromosomes decondense/uncoil).
What is important about mitosis?
INPUT = OUTPUT for haploids and diploids. What goes in is replicated, and each daughter cell gets one copy of the original genes (exception = mutations OR something went wrong during mitosis). How does this happen…
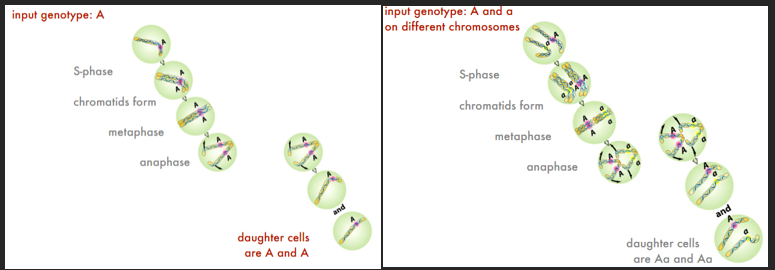
Metaphase Plate Alignment/Magic
Ensures that microtubules from each pole attach to ONE replicate (other replicate = other pole). Thus, NO REPEAT of replicates in one daughter cell. Metaphase is critical to the process.
Asexual Budding/Mitosis in Yeast
First budding = S phase. Closed cell cycle (NE never dissolves, but the nucleus positions itself at the neck of the bud… anaphase entry signal = can see spindle). Chromosomes are being segregated (only fibers/nucleus visible). After, two nuclei separate and bud pops off.
How did researchers discover the “pulley” mechanism?
Identify/clone proteins that connect microtubules/spindles to chromosome separation (e.g., KINETOCHORE PROTEIN CLONING).
Irradiate WT yeast cells —> collect T-sensitive yeast mutants (die at warm T) —> test hundreds of mutants for microtubule staining at non-permissive temp (e.g., what mutants have improper attachment; see phenotype via staining) —> identify mutants that fail to form attachments at high T (ex. ndc-10-1) —> grow mutant at permissive low T —> use screen of genomic library (see suppressor card) —> grow mutants at permissive low T —> raise T until only some survivors (received WT gene for mutated one (complementation)… or suppressor; what makes up function of mutation) —> grow/clone the gene (on plasmid).
Kinetochore Proteins
Bind centromeres on chromosomes (specialized DNA region). Each chromosome has only one centromere, but there are many different types and copies of kinetochore proteins. Bind to centromeres AND microtubules (made up of many proteins). Chromosomes can be lost with defective spindle attachment.
Mitotic Spindle Checkpoint
If improper attachment (due to mutation), cells will arrest mitosis and try to capture lost chromosomes.
Nocodazole
Microtubule-destabilizing chemical. Induces unattached chromosomes to cause the prevention of anaphase entry/mitotic arrest. Microtubules can reform after washing out nocodazole (cells can recover).
What genes control mitotic arrest (e.g., prevent anaphase entry)?
Create a mutant that undergoes mitosis even when treated with nocodazole (e.g., mutated cells did not survive nocodazole (WT do… recover) because they tried to divide in the presence of destabilized MTs (NO arrest)). Results in mitotic catastrophe (death). Cancer cells also ignore the mitotic spindle checkpoint (NO stopping cell division… divide and die).
Can identify mutated gene using same screening/cloning process above. Mitotic arrest deficient (MAD) genes = sensor for MT integrity. When mutated, cannot sense that MTs are not present (continues to divide).
Key for Mitosis
Kinetochore proteins and mitotic checkpoint checkpoints! Highly conserved!!
Clinical Correlation of Mitotic Checkpoint Gene Mutations
Assessed whether cancerous and non-cancerous cell lines had active checkpoints. Cancerous = were they responsive to nocodazole chemicals (e.g., do they survive by arresting mitosis) —> many cell lines responded to Noc (arrested in mitosis; high mitotic index = measure of how many cells are in mitosis (arrested)), and others did not (NO arrest; low mitotic index).
Double checked —> did cell lines that respond to Noc (arrest, triggered checkpoint) stop replicating DNA? YES (no DNA replication in colon cancer cell lines). Other colon cancer cell lines did NOT stop replicating (e.g., no arrest) —> due to presence of a SNP in tumor cells compared to normal cells in checkpoint gene region.
Thus, checkpoint protein mutations = responsible for aggravation of some cancers.
Meiosis
Aka reductional cell division! Required for genetic diversity! Germ cells = NOT diploid in human body. More complicated than mitosis —> creations haploid germ cells that later combine to form zygotes (n + n = 2n; fertilization). Steps = replication (2n —> 4n) and then two rounds of division (4n —> 2× 2n —> 4× 1n). NO REPLICATION BETWEEN!
Types of Germ Cells
Spermatozoa (male)
Oocytes (female)
Spermatozoa
Meiosis/creation of male gametes —> occurs in seminiferous tubules of testes.
Oocytes
Develop in ovaries through a form of meiosis called oogenesis. Mammals = stored in ovary and arrested at prophase I of meiosis. During menstrual cycle = some oocytes proceed through rest of meiosis. Eggs NOT equal size = one large gamete and three small gametes (discarded).
Why two rounds of division?
Tetraploidy (4n) is lethal in humans (do not want 2n x 2n). Ensures sex cells are 1n and combine to form 2n. Toxic due to GENE BALANCING = too many gene products can imbalance cell (4x amount of gene products compared to haploid). Plants can handle tetraploid because they change cell size.
Synchrony in Meiosis
IMPORTANT! One difference - results in combination of maternal and paternal genetic material in zygotes that has NEVER been seen before. Unique combos due to independent assortment (metaphase I, II) + recombination (prophase I).
Steps of Meiosis
S-phase (replication before meiosis) —> prophase I (recombination) —> metaphase I (independent assortment) —> anaphase I —> telophase I —> interphase I —> prophase II —> metaphase II —> anaphase II —> telophase II
S-phase
Replicate DNA before meiosis (diploid —> 4n). Sister chromatids are attached (copes) and tight together at centromeres.
Prophase I
Chromosomes condense and contract. Active pairing of replicate chromosome sets (e.g., both pairs of sister chromatids together) (4n total) —> eventually fully-synapsed pairs where crossing-over occurs (eventually chiasmata (crossovers) are visible)). After final compaction, the synaptonemal complex disappears…
Synaptonemal Complex (SC)
Protein structure that forms between homologous chromosomes (e.g., between two pairs of sister chromatids). Tripartite structure - two parallel lateral regions and a central element. Functions as scaffold to facilitate completion of crossover activities (mediate chromosome pairing, synapsis, and helps recombination). NOT REQUIRED FOR RECOMBINATION! Holds onto…
Cohesins
Ring-like handcuffs that connect/pair replicated sister chromatids on either side of the SC. Crucial role in keeping sister chromatids together during meiosis I.
Metaphase I
NE is now gone. Spindle forms. Metaphase plate alignment of compacted chromatids (4n). Two microtubules for each dyad (BUT one is not active… each dyad pulled in one direction). Crossing over finished.
Metaphase I/Early Anaphase Transition
Alignment done (microtubules fully associated). Cohesins hold together crossover chromosome regions still.
Late Anaphase I
Cohesins hold sister chromatids together near the centromere. Movement to opposite poles of homologous chromosomes.
Telophase I
NE reforms. Chromosomes decondense somewhat. Spindle disappears. Cell readies to divide.
Interphase I
Division of cells. NO new DNA synthesis. Moves straight into meiosis II.
Prophase II
Chromosomes condense again
Metaphase II
Attached sister chromatids arrange on metaphase plate. Sister chromatids already partly separated. Held together at regions near centromeres (intact cohesins).
Anaphase II
Centromeres split, and sister chromatids separate.
Telophase II
NE reforms. Spindle disappears. Cells divide, creating 4 haploid cells.
How does germ cell formation produce variety?
METAPHASE PLATE MAGIC/REARRANGEMENT = orientation of maternal and paternal replicated copies for each chromosome is RANDOM (independently segregate). E.g., whether maternal/paternal goes to each pole/daughter cell in meiosis I is random between chromosomes. Dmany possible different combos in daughter cells!
Number of Combinations based on Metaphase Plate Magic
In a diploid, 2^n where n is the number of chromosomes (humans… 223 different combos).
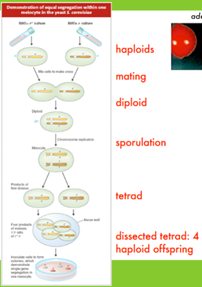
How can we prove that daughter haploid cells are receiving the same info from the parent cell (not missing genes)?
Markers/assays —> imagine ADE2 MATalpha yeast (white) and ade2 MATa yeast (red). Difference in phenotype color. With a cross of haploids to form diploid, we would expect a 1:1 phenotype ratio in a tetrad produced by the diploid (2 red, 2 white). TRUE!
Double mutants —> test for correct ratios if chromosomes were to independently segregate. Two mutants, one mutant per haploid, mutants on diff chromosomes. Cross to form diploid (would be heterozygous for both genes). The tetrad formed during sporulation would have one double mutant, two single mutants, and one WT (all differ in phenotype b/c haploid).
Cohesins
Chromosomal glue during mitosis and meiosis. Ring-like structures made of three proteins. Hold sister chromatids together in the metaphase plate until anaphase. Unglue rings by CUTTING THEM!
How to Unglue Cohesin Synchronously
Separase (Esp1) = enzyme that cleaves cohesin protein structure. Securin (Pds1) = protein that keeps separase inactive. Remove securin = allows separase to cleave cohesin (separation of sister chromatids). Spindle checkpoint proteins trigger the signal to breakdown securin.
Cohesin Cleavage in Mitosis
FULL!!
Cohesin Cleavage in Meiosis
INCOMPLETE CLEAVAGE IN MEIOSIS I!!!! Homologous dyads still attached via cohesins between crossed-over segments.
In metaphase I, cohesin at the edges of chomosomes (cross-over segments) is CLEAVED by separase (allows for separation of homologous dyads). Sister chromatids intact via cohesins near centromeres. Central cohesins remain intact by NOT being phosphorylated (must be to be cleaved). Shugoshin prevents cohesin phosphorylation around centromere.
In metaphase II, rest of cohesins is cleaved near centromere.
Cohesin-related Accidents
Can result in aneuploidy = incorrect number of chromosomes caused by an incomplete set (n) or extra chromosomes. Nondisjunction (lack of separation) at first or second divisions. Haploid products can be deprived (n-1; Turner’s) OR in excess of certain chromosomes (n+1; Down, Patau, Klinefelter’s).
Of 15% of conceptions that abort spontaneously, half show chromosomal abnormalities (UNDERESTIMATE). Among live births, 0.6% have chromosomal abnormalities (e.g., aneuploidy/chromosomal rearrangements are more likely to cause death).
Crossing-Over
Exchange of pieces of DNA/genes between non-sister chromatids. Generates LOTS of new genetic variation and combinations (1-2 crossovers per chromosome every meiosis). Occur during prophase I of meiosis. Leads to creation of parental genotypes (same as parent; no crossing over) or recombinant genotypes (crossing over).
Recombinant Sequences
Novel combinations of genetic info due to crossing-over.
Crossing over results in an _____
INCREASE in number of possible gametic combinations (recombinant). Each single reciprocal exchange = 2 new combinations at least.
Does crossing-over affect segregation of genes that reside on DIFF chromosomes?
Yes (ex. switching B and b on one chromosome), but this is indistinguishable from metaphase plate shuffling.
Does crossing over affect segregation of genes that reside on the SAME chromosome?
YES, it can impact the segregation of genes (a crossover between genes A/a and B/b can result in new combinations if parentals were AB/ab… aB or Ab). Would normally segregate together (e.g., A and B, a and b) because they are on the same chromosome —> can distinguish from metaphase plate shuffling.
Output of Crossover in One Chromosome in One Meiosis
NO crossovers = creation of two unique parental genotypes in four haploid gametes (input = output).
ONE crossover = creation of four unique genotypes (2 parental, 2 recombinant) in four haploid gametes (input NOT equal to output). Indicates crossing-over AFTER replication (if before… only two unique genotypes with replication; sister chromatids = COPIES).
Recombination Frequency
Recombinant genotypes / total genotypes!!!
Unlinked = 0.5. Genes are far apart (on diff chromosomes) - 50% maximum likelihood of being separated due to metaphase plate shuffling (thus 50% recombinants and 50% parental). 1:1:1:1 ratio genotypes.
Effectively Unlinked = ~0.5. Genes far apart (on SAME chromosome) - opposite ends of a centromere on a chromosome. Remember— 1-2 crossover events per chromosome per meiosis.
Linked = <0.5. Genes are close together (on SAME chromosome) - no metaphase plate shuffling BUT chance of crossovers (so some generation of recombinants). 10:10:1:1 ratio genotypes (parental > recombinant). Indicates parental allele combos are associated/linked (less recombinants).
Complete Linkage = 0 (no recombination). Genes are right next to each other - no metaphase plate shuffling NOR crossovers (always linked… no recombinants). 1:1 ratio genotypes (only parental/nonrecombinants). Indicates parental is linked! Crossing over never cuts between these genes.
If genes are far apart and 10 meiosis happen, 9 with recombination, what is the recombination frequency?
10 × 4 gametes = 40 total. 9 × 2 recombinant gametes = 18. 18 / 40 = 0.45 (pretty unlinked).
First Examination of Linkage in Diploids
Morgan/Sturtevant! Found that diff alleles on the same chromosome should always “travel” together but NOT ALWAYS TRUE. Found that markers can swap positions with respect to each other in X and Y sex chromosomes of white-eye flies! Why? Can see recombinants in the male (one X… recombinant genotypes not masked; gained from female; crossover during meiosis of female gametes = recombinant).
Morgan Fly Linkage Experiment
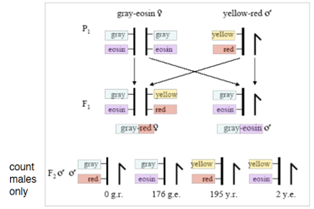
Used gray-eosin (linked) and yellow-red (linked) markers. Morgan crossed a homozygous gray-eosin female (both X) with a yellow-red X male. Examined the males of the F2 generation…
F1 generation = all heterozygous females and all grey-eosin males (X from female).
MALES of F2 generation = would theoretically be 50% gray-eosin X and 50% yellow-red X (b/c females heterozygous). BUT… another phenotype/genotype was seen: yellow-eosin (RECOMBINANT) at low numbers (2). Low # of recombinants suggests the genes are pretty close together/linked (but not completely).
How are all meiotic linkage analyses conducted?
Cross a HETEROZYGOTE for both genes with a homozygous recessive tester for both genes. AND must have LOTS of offspring.
Map Units (cM = centimorgans)
Genetic distance between 2 genes where 1% (0.01) recombination frequency = 1 map unit. A function of the likelihood that a crossover cuts between two genes (close together = linked = low RF = low map distance). 0 cM = full linkage. 50 cM = unlinked! RF * 100% = cM!
Summary of Linkage
Not all genes on the same chromosome are linked (crossovers). Far enough = unlinked. All genes on the same chromosome are SYNTENIC. Each pair of linked genes has a characteristic RF based on distance/likelihood of crossover between them.
Crossing-over in Detail
Exchange of segments of homologous chromosomes in prophase I. Involves chiasmata. The process of breaking and reattachment occurs in THREE dimensions (orientation determines directions/angles of cuts and rejoins). Known as the Chi structure or Holliday recombinant structure. Different cuts = genetic variation! See gene conversion cards for steps of crossing over.