Lecture 1 of Exam 4 Content
1/27
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
28 Terms
What are fatty acids?
Carboxylic acids with long alkyl (hydrocarbon) chains as their R groups.
What are the 3 overall stages of fatty acid synthesis?
When the cell has a sufficient amount of ATP, Acetyl CoA (the building blocks of fatty acid synthesis) is transferred from the mitochondrial matrix to the cytosol in the form of citrate. Oxaloacetate is also returned to the mitochondrial matrix.
Recall that Acetyl CoA is made after pyruvate oxidation, which occurs in the mitochondrial matrix.
It can either continue on and be used to make ATP or be moved to the cytosol to create fatty acid chains.
Once in the cytosol, some of the Acetyl CoA is converted to malonyl CoA. Both Acetyl CoA and malonyl CoA are then attached to acyl carrier proteins to yield Acetyl ACP and Malonyl ACP.
Then, fatty acid elongation occurs 2 carbons at a time. 2 carbons are added after each complete cycle.
Describe how Acetyl CoA is moved from the mitochondrial matrix to the cytosol (step 1)
Acetyl CoA (2 carbons) and oxaloacetate (4 carbons) combine to form citrate (6 carbons) in the mitochondrial matrix. This reaction is catalyzed by citrate synthase.
Citrate then passes the inner mitochondrial membrane via a citrate transporter and then easily passes the outer mitochondrial membrane to now enter the cytosol!
Once in the cytosol, citrate is cleaved by ATP-Citrate Lyase to yield Acetyl CoA and oxaloacetate once again. This is essentially the reverse of the reaction catalyzed by citrate synthase.
Now we have Acetyl CoA in the cytosol. We also have oxaloacetate in the cytosol. However, for this process to occur again, oxaloacetate must be in the mitochondrial matrix so that it can be combined with acetyl CoA to form citrate that can easily enter the cytosol:
Oxaloacetate is reduced to malate via cytoplasmic malate dehydrogenase. This is coupled with the oxidation of NADH to NAD+.
Malate (4 carbons) is then oxidatively decarboxylated by the Malic Enzyme to yield pyruvate (3 carbons).
So, malate is oxidized, which is coupled with the reduction of NADP+ to NADPH.
Malate also loses CO2 (is decarboxylated).
Thus, pyruvate is an oxidized (because one of the carbons— the carbonyl— has a double bond with oxygen) and decarboxylated version of malate.
The newly formed pyruvate can now easily return to the mitochondrial matrix through a pyruvate transporter. Once there, it is carboxylated by pyruvate carboxylase to yield oxaloacetate.
Oxaloacetate has now returned to the mitochondrial matrix where it can be used to transport another Acetyl CoA to the cytosol for more fatty acid synthesis.
Note that fatty acid synthesis uses NADPH rather than NADH. This NADPH is generated during the oxidative decarboxylation of malate to yield pyruvate by the malic enzyme. It can also be generated by the pentose phosphate pathway.
Describe the structure of pyruvate, oxaloacetate, and malate
Pyruvate is a 3 carbon molecule with the 3 carbon functional groups— Carboxyl, carbonyl, and then methyl.
Oxaloacetate is a 4 carbon molecule. It has the same overall structure as pyruvate but the methyl group at the end has one if its hydrogens replaced with a carboxyl group (so its no longer actually a methyl group).
Malate is also a 4 carbon molecule, but is the reduced version of oxaloacetate. Its carbonyl group on C2 is replaced with a carbon bound to hydrogen and an OH.
To get from malate to pyruvate, you remove a carbon and oxidize— hence oxidative decarboxylation.
Describe what occurs during step 2 of fatty acid synthesis
Acetyl CoA carboxylase converts some of the acetyl CoA (2 carbons) to malonyl CoA (3 carbons) in 2 steps:
1: A biotin-enzyme, ATP, and a bicarbonate ion combine to yield carboxybiotin (intermediate), Pi, and H+.
2: Then, carboxybiotin transfers its CO2 to Acetyl CoA to yield Malonyl CoA and regenerate the biotin-enzyme. Thus, malonyl CoA is acetyl CoA but with another COO- added to its left side.
All 3 carbons in malonyl CoA are destined for fatty acid synthesis. Thus, this is the 1st committed step towards fatty acid synthesis and is a point of regulation.
Also note that not all Acetyl CoA is converted to Malonyl CoA. So now, there is a population of acetyl CoA and malonyl CoA.
Acetyl CoA and Malonyl CoA get attached to acyl carrier protein (ACP) to yield Acetyl ACP and Malonyl ACP.
The ACP is made up of 77 residues and has the same business end as coenzyme A (the reactive SH group at the left end). Thus, Acetyl and Malonyl attach/get acylated to the same place on the ACP as they would on Coenzyme A.
They specifically get acylated onto the SH on the left end of the ACP by their respective enzymes.
Acetyl transacylase acylates Acetyl CoA onto ACP to yield Acetyl ACP and CoA.
Malonyl transacylase acylates Malonyl onto ACP to yield Malonyl ACP and CoA.
ACP functions as the foundation and scaffolding for fatty acid synthesis.
Note that it is transACYLase not transACETYLase.
Describe what occurs during step 3 of fatty acid synthesis
Fatty acid elongation occurs 2 carbons at a time. Each cycle has 4 steps. These 4 steps are repeated each cycle.
Condensation— acetyl ACP and malonyl ACP are combined by β-ketoacyl synthase to yield Acetoacetyl ACP, which is a β-ketoacyl ACP.
The carbon that was added to create malonyl CoA is now removed as CO2 (malonyl gets decarboxylated) so that acetyl ACP can bind to malonyl ACP in its place. The carbons from acetyl ACP are carbons 3 and 4 on the left.
The product now has 4 carbons.
Numbering of carbons begins at the carbon bound to the sulfur of ACP (this is carbon 1). The alpha carbon is carbon 2. The β carbon is a ketone which is acylated onto ACP, hence the name of the product.
Reduction— β-ketoacyl ACP is reduced by β-ketoacyl reductase to yield D-3-hydroxybutyryl ACP.
The carbonyl of the β carbon/C3 gets reduced and C3 gains an OH (hence the name of the product).
This is coupled with the oxidation of NADPH to NADP+.
Dehydration— D-3-hydroxylbutyryl loses H2O to yield Crotonyl ACP. This is catalyzed by 3-hydroxyacyl dehydratase.
The OH from carbon 3 and an H from C1 are lost. This allows for the formation of a double bond between C2 and C3, which creates an enol.
Reduction— crotonyl ACP is reduced by Enoyl reductase to yield butyryl ACP.
The double bond between C2 and C3 is broken because each carbon gains an electron.
This is coupled with the oxidation of NADPH to NADP+. NADPH supplies the electrons.
This yields a saturated, 4 carbon butyryl fatty acid that is attached to ACP. So we started with 2 carbons on the chain and now we have 4 🙂.
Fun fact: Enoyl reductase is inhibited by a common antibacterial— Triclosan.
Describe continuation of fatty acid elongation
The cycle with those 4 steps occurs again, adding two carbons to the chain each time. All of the same enzymes and substrates are used.
However, during the condensation step, instead of combining Acetyl ACP and Malonyl ACP, Malonyl ACP is decarboxylated and then combined with the end product of the previous cycle. So in our example, the two newly synthesized carbons from butryl ACP are added on to malonyl ACP to yield C6-β-ketoacyl ACP.
Describe the regulation of fatty acid synthesis
Regulation of fatty acid synthesis focuses on regulating Acetyl CoA carboxylase— the enzyme that catalyzes the committed step. This is the step in which some of the Acetyl CoA is converted to malonyl CoA by that enzyme. All of the carbons in the newly formed Malonyl CoA are destined for fatty acid synthesis.
Allosteric regulation of acetyl CoA carboxylase:
Citrate, which is formed during the CAC and eventually becomes Acetyl CoA, stimulates the activity of this enzyme because it will eventually form this enzyme’s substrate.
Palmitoyl (a fatty acid with16 carbons) inhibits the activity of this enzyme. This fatty acid is very long, so since this long fatty acid chain has already been formed, it inhibits the activity of the enzyme so that it doesn’t form more fatty acids.
Acetyl CoA carboxylase is also covalently regulated via phosphorylation. This is continued on the next card.
Describe covalent regulation of Acetyl CoA carboxylase
Acetyl CoA carboxylase is also covalently regulated via phosphorylation:
Phosphorylation inactivates the enzyme, thereby reducing its activity
Dephosphorylation activates the enzyme, thereby stimulating its activity.
Specific covalent regulators:
Insulin stimulates the enzyme’s activity by activating Protein Phosphatase 2A, which removes the phosphate group from acetyl CoA carboxylase, thereby activating it.
Glucagon and epinephrine inhibit the enzyme’s activity by activating AMP-activated Protein Kinase (not PKA), which adds a phosphate group to acetyl CoA carboxylase, thereby reducing its activity. There are two ways to get AMP:
When cAMP levels rise, it gets hydrolyzed to AMP, which activates AMP-activated Protein Kinase which then inhibits the enzyme.
ATP can be broken down to yield AMP
Citrate can also partially activate a phopshorylated Acetyl CoA carboxylase. The phosphate group remains on the enzyme, but it regains some of its function. Its level of activity would be between its level when phosphorylated and when unphosphorylated.
Note that citrate’s ability to be an allosteric regulator and covalent regulator is NOT mutually exclusive. If the enzyme is phosphorylated and citrate levels increase from the CAC, citrate will allosterically stimulate the enzyme’s activity.
Insulin is released when blood glucose levels are high. Once it’s released, sugars get metabolized to acetyl CoA, which can then be used for fatty acid synthesis. Thus, insulin helps us store energy in 2 ways:
By stimulating glycogen synthesis
By stimulating fatty acid synthesis
^^Both of which store energy in our bodies.
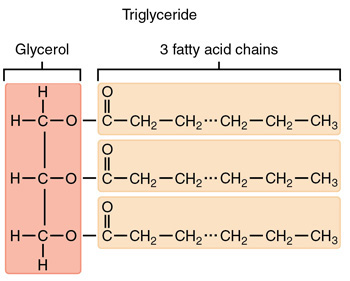
What are triacylglycerols?
They are the storage form of fatty acids and are formed through esterification. They are made up of:
A glycerol backbone
3 fatty acid chains acylated onto the glycerol backbone
How are triacylglycerols broken down/metabolized?
They are first acted on by lipases, which hydrolyze the fatty acid chains/remove them from the glycerol backbone. Lipases remove fatty acid chains one at a time to yield 3 free fatty acids. This is called lipolysis.
So, the triacylglycerol becomes a diacylglycerol and then a monoacylglycerol and then that last fatty acid can be removed to yield 3 free fatty acids and a glycerol backbone. These 3 free fatty acid chains and the glycerol backbone are all used to make energy.
Recall that fatty acids are chains of hydrocarbons that terminate with a carboxylic acid group. The first carbon after the carbonyl (C1) is the alpha carbon (C2).
Describe the different types of lipases found inside cells and their regulation
Adipocyte Triglyceride Lipase (ATGL) hydrolyzes triacylglycerols to diacylglycerols and releases a free fatty acid.
CGI-58 regulates its activity.
Hormone sensitive lipase (HSL) hydrolyzes diacylglycerols to monoacylglycerols and releases a free fatty acid.
Perilipin A regulates HSL’s activity
Both CGI-58 and Perilipin A are regulated by phosphorylation.
When they are phosphorylated, they are active, and activate ATGL and HSL, and thus triacylglycerols are broken down.
They are both phosphorylated by PKA.
Phosphorylated CGI-58 binds to ATGL, and then PKA phosphorylates ATGL and now it’s active and lipolysis occurs.
Phosphorylated Perilipin A binds to HSL, and then PKA phosphorylates HSL and now it’s active and lipolysis occurs.
When they are unphosphorylated, they are unstimulated and are bound together, inhibited. When they are inhibited, triacylglycerols are NOT broken down.
Lastly, after removal of the first 2 fatty acids, Monoacylglycerol Lipase (MGL) easily removes the last fatty acid to yield another fatty acid and the glycerol bacbone. This enzyme is unregulated because it only removes the last fatty acid chain after the first 2 have been removed.
Describe glucagon/epinephrine and insulin’s role in the regulation of lipases
Glucagon/epinephrine stimulate the cAMP/PKA pathway via the 7TM receptor that they bind to and thus promote lipolysis and fatty acid metabolism.
Glucagon/epinephrine bind to the 7TM receptor and after downstream signaling, produce cAMP that activates PKA. PKA then phosphorylates and activates CGI-58 and Perilipin A, which can then go bind to HSL and AGTL and once they are again phosphorylated by PKA, lipolysis occurs.
Insulin, via the P13K/Protein Kinase B (Akt) pathway, activates phosphodiesterase (PDE).
Insulin binds to the recetptor
Stimulates the breakdown of cAMP to yield 5’AMP.
This turns off cAMP’s activation of PKA, which thus turns off lipolysis since nothing can get phosphorylated.
Insulin and glucagon are released under opposite conditions and do opposing things.
Insulin is released when blood glucose is high and the cell is about to metabolize all of that glucose and get energy, so fatty acid synthesis is stimulated and lipolysis is inhibited since we do not need any more energy so we do not need to break down triacylglycerols to make energy. Thus, insulin stimulates fatty acid synthesis and inhibits lipolysis.
Glucagon is released when blood glucose is low and the cell needs energy. So, it stimulates fatty acid degradation/lipolysis so that we can get energy from those triacylglycerols. It also stimulates the breakdown of glycogen so that we can get energy from those stored sugars.
What is Chanarin-Dorfman Syndrome?
Is a rare disease in which the CGI-58 gene is mutated, which affects the breakdown of lipids. Is characterized by the accumulation of dead skin cells on the face that are brown.
Because of the mutation, CGI-58 cannot get phosphorylated, so it cannot bind to ATGL, so the first fatty acid cannot be removed from triacylglycerol, so the other 2 cannot be removed either. This causes triacylglycerols to accumulate, which increases the amount of lipids inside cells and leads to necrosis of cells due to lipotoxicity.
What are the fates of free fatty acids and glycerol?
Glycerol is processed in the liver. There, it can either go through the glycolytic or gluconeogenic pathway depending on the cell’s energy needs.
Fatty acids are taken up by cells and undergo fatty acid oxidation to be used as fuel. They are also packaged with serum albumin and transported to other tissues.
Describe the processing of glycerol
Glycerol goes to the liver. There, it can either by used for glycolysis OR gluconeogenesis.
Glycerol is phosphorylated by glycerol kinase to yield glycerol-3-phosphate. Glycerol kinase uses ATP as its source of energy and phosphate group.
Glycerol-3-phosphate is oxidized by glycerol phosphate dehydrogenase to yield DHAP. This oxidation is coupled with the reduction of NAD+ to NADH.
The fate of DHAP depends on the cell’s energy state:
If the cell needs ATP, DHAP will be isomerized to glyceraldehyde 3-phosphate (G3P) via triose phosphate isomerase, which can then be metabolized to pyruvate to eventually yield ATP.
If the cell has enough ATP, some DHAP is isomerized to G3P and then they are combined to yield fructose-1,6-bisphosphate which can enter gluconeogenesis and yield glucose.
Describe the first part of the fate of fatty acids
Fatty acids will undergo oxidation, which occurs in the mitochondrial matrix. But lipolysis occurs in the cytosol.
To bring fatty acids from the cytosol into the mitochondrial matrix, we must form acyl carnitine:
Fatty acids attach onto coenzyme A to yield acyl-CoA via acyl CoA synthetase. This occurs in 2 steps:
The fatty acid reacts with ATP to yield acyl adenylate. This also releases pyrophosphate— the 2 terminal phosphate groups of ATP.
The released pyrophosphate is unstable under aqueous conditions and is thus quickly becomes 2 inorganic phosphates. This makes the reaction pretty much irreversible.
The release of this pyrophosphate is equivalent to losing 2 high-energy phosphate bonds, which is equivalent to losing 2 ATP molecules.
Then, the SH of Coenzyme A attacks acyl adenylate to yield acyl-CoA (fatty acid attached to Coenzyme A) and AMP.
Carnitine acyltransferase 1 transfers the acyl group from acyl CoA to the OH group of carnitine. This yields acyl-carnitine— fatty acid chain attached to the oxygen of carnitine.
Acyl carnitine is then shuttled across the inner mitochondrial membrane by translocase, which binds to carnitine.
Once in the mitochondrial matrix, the acyl group on acyl carnitine is transferred back to coenzyme A by carnitine acyltransferase II to reform acyl-CoA for fatty acid oxidation.
Describe the second half of fatty acid’s fate
Fatty acid oxidation has 4 steps in each cycle that are the opposite of the steps in fatty acid synthesis. These 4 steps repeat for each cycle.
First oxidation: Acyl CoA is oxidized by acyl CoA dehydrogenase to yield enoyl CoA with a trans double bond between C2 and C3 (it’s an enol). This is coupled with the reduction of FAD to FADH2.
Hydration: Enoyl CoA gains H2O, which breaks the double bond between C2 and C3 and forms 3-hydroxyacyl CoA (so the OH goes on C3). This is catalyzed by enoyl CoA hydratase.
Second oxidation: 3-hydroxyacyl CoA is oxidized by 3-hydroxyacyl CoA dehydrogenase to yield 3-ketoacyl CoA. This is coupled with the reduction of NAD+ to NADH. Thus, C3 loses its OH and instead becomes a carbonyl.
Thiolysis: the C2-C3 bond of 3-ketoacyl CoA is cleaved by β-ketothiolase to yield Acyl CoA but shorted by 2 carbons (so now it only has 2) and acetyl CoA. The SH group of another SH-CoA releases one acetyl CoA and one acyl CoA (it supplies the CoA).
Thus, the fatty acid chain has been shortened by 2 carbons and released an acetyl CoA.
The fatty acid can then undergo β-oxidation again to get rid of the last 2 carbons.
Explain how the the β-oxidation of unsaturated fatty acid differs from that of saturated fatty acids
Recall saturated vs unsaturated— saturated does not have a double bond, unsaturated has a double bond.
β-oxidation occurs normally until it reaches a point in which the double bond is between C3 and C4. If we continued, the OH after hydration would end up on C4 and so C4 would also get the carbonyl after the 2nd oxidation and thus β-oxidation cannot properly occur.
To fix this, cis-△3-Enoyl CoA isomerase moves the double bond from between C3 and C4 to where it should be— between C2 and C3. This yields trans-△2-Enoyl CoA, which can now continue on through β-oxidation.
Compare the amount of ATP that can be produced from saturated vs unsaturated fatty acids
Because unsaturated fatty acids already have a double bond, the first oxidation step is skipped and hydration occurs right away. Thus, no FADH2 is created.
The first oxidation step is skipped for every double bond. Thus, for each double bond, FADH2 is not made.
For every FADH2, 1.5 ATP is created. Thus, for every double bond in an unsaturated fatty acid, 1.5 ATP is missed out on.
Thus, unsaturated fatty acids produce less energy than saturated fatty acid.
How much ATP is produced from fatty acid oxidation?
For each cycle, an acyl CoA yields:
1 FADH2— 1.5 ATP
1 NADH— 2.5 ATP
1 Acetyl CoA— via the CAC, 10 ATP
Recall however that when bringing fatty acids into the mitochondrial matrix, 2 equivalents of ATP are lost. Thus, you must subtract 2 from the obtained number.
Each double bonds reduces the number of ATP produced by 1.5 since each double bond prevents the formation of 1 FADH2 which yields 1.5 ATP.
The ratio of ATP’s per carbon gets smaller as the fatty acid chain gets shorter. Thus, longer fatty acid chains give you more ATP per carbon.
Note that the number of cycles required to completely oxidize a fatty acid is equal to the # of Acetyl CoA groups - 1
What are nucleic acids?
They are a type of macromolecule. Examples of nucleic acids include DNA and RNA.
Their monomers are nucleotides that line up in a linear fashion to form the polymers. Nucleotides are made up of:
A sugar (ribose for RNA, deoxyribose for DNA)
At least 1 phosphate group bound to the sugar
The sugar and the phosphate group make up the backbone of the molecule. They are held together by phosphodiester linkages. Specifically phosphoesters— like a carbonyl but with the carbon replaced with a phosphoryl group to yield a phosphoester.
A nitrogenous base bound to the sugar
The nitrogenous base and the sugar are connected via a β- glycosidic linkage between C1 of the sugar and the N of the base.
Nucleotides in a DNA strand are connected via phosphodiester linkages.
List the different nitrogenous bases and their categories and characteristics
Purines: Angels are pure— Adenine and Guanine
Guanine has an oxygen from a carbonyl sticking out of the ring (GO Guanine)
Adenine has an amino group sticking out of the ring (AC)
Pyrimidines: pyramids cut: Cytosine, Uracil, Thymine
Thymine has a methyl group sticking out of the ring (methyl)
Uracil looks like thymine but without the methyl group (unmethylated)
Cytosine has an amino group sticking out of the ring (AC)
Purines have 2 rings, pyrimidines have 1 ring (inverse correlation)
Describe the structure of the DNA double helix
Each DNA molecule is made up of two helical and linear strands that wrap around a common axis to form a double helix.
These strands are antiparallel to each other, meaning that the 5’ end of one lines up with the 3’ end of the other and vice versa.
The 5’ end is indicated by the phosphate group.
The 3’ end is indicated by the 3’ OH on the sugar
Each strand has its own sugar-phosphate backbone held together by phosphodiester linkages and nitrogenous bases bound to C1 of the sugar via glycosidic linkages.
The two strands are held together by hydrogen bonds between complementary base pairs. This base pairing is what makes the DNA double helix stable.
A and T (U if RNA) pair and form H bonds
C and G pair and form H bonds
The sugar-phosphate backbone is on the “outside” of the helix. The interior of the helix is made up of the nitrogenous bases of each strand held together by H bonds.
The bases are nearly perpendicular to the helix axis.
The diameter of the helix is about 20 Angstroms. A 360 degree rotation takes about 10.4 bases— about 34 Angstroms. Thus, each base pair is separated by about 3.4 Angstroms.
Major and minor groove:
Between two bases with hydrogen bonds, one side has more space between the two glycosidic bonds than the other side does due to the angles/orientation of the glycosidic bonds.
Because of this difference in space, when you stack the nitrogenous bases on top of each other and then twist in a helical manner to form the DNA double helix, the uneven spacing creates a major and minor groove.
Major groove: has a lot for space between the backbones on either side
Minor groove: has less space between the backbones on either side and is more narrow
Some proteins only interact with DNA at specific seuquences. These proteins find their sequencesby interacting with DNA in the major groove because the major groove exposes a larger surface of base pairs for this sequence-specific recognition.
Describe the conservation of DNA Replication
DNA replication is semi-conservative, meaning that each of the new DNA molecules created after replication have 1 strand that is newly synthesized and 1 strand from the parent DNA molecule. Thus, the daughter molecules are 50:50 old and new.
If this process wasn’t conservative, there wouldn’t be an entire strand from the parent DNA molecule. There would only be bits and pieces embedded here and there into the DNA molecule.
Describe the Meselson and Stahl Experiment
This was the first experiment to determine that DNA replication is semi-conservative.
Details:
They labeled E.coli DNA strands with 14N and 15N radioactive isotopes. The 15N is heavier than the 14N.
They used a CsCl gradient to separate DNA with different densities based on the radioactive isotope bound to the DNA.
They then looked at a piece of film with bands that each correspond to one of the radioactively-labeled DNA molecules.
Steps:
They initially just fed the bacteria the heavy nitrogen (15N) . Consequently, all of the DNA was labeled with it.
Then the cells were transferred to a medium that only had the light nitrogen. They then took samples, isolated the DNA, and put it in the CsCl gradient.
After 1 new generation of DNA replication, they observed the CsCl gradient and the band pattern on the film. They saw that the band on the film didn’t line up completely with the heavy nor the light isotope. Rather, it was in between both of them. Thus, this told them that the DNA molecule after replication was a hybrid of both original and newly synthesized DNA— thus semi-conservative replication.
Technically however, even if replication was unconserved, you could still end up with a 50:50 OG and newly synthesized just mixed in each strand rather than each having its own strand. Thus, it is good that they took it further and did 1 more step because this first result could be caused by non-conservative replication.
They then allowed the DNA to replicate for another generation. In this 2nd generation, they saw:
2 of the bands lining up in the middle of the bands for the heavy and light isotopes— so 2 hybrid DNA molecules.
2 of the bands lining up completely with the light isotopes band. These are both newly synthesized strands or were conserved from the creation of the previous newly-synthesized strand.
Because these hybrids are still present in addition to only light strands, they were able to conclude that DNA replication is semi-conservative.
If replication was non-conservative, there wouldn’t be any more hybrids left and we wouldn’t see 2 distinct bands on the film because the remnants from the OG parent molecule would be so small. Thus, they were able to conclude that replication must be semi-conservative.
Describe DNA Replication in Prokaryotes
Replication begins at the origin of replication, oriC. Note that replicarion in prokaryotes is bi-directional— it goes both ways after this oriC.
This is 245 BP region with 3 13 BP sites that have a lot of A and T.
OriC also has 5 binding sites that DnaA recognizes and binds to.
Once it binds it recruits DnaB to bind to the A-T rich sites. DnaB is a helicase that breaks the H bonds between nitrogenous bases to open/unwind the DNA at these sites. This creates 2 single-stranded DNA molecules. DnaB requires ATP to do this.
Note that DnaB binds to and breaks the H bonds between A and T because A and T are only held together by 2 h bonds so they are easier to separate than C and G which are held together by 3 H bonds.
Single-strand binding proteins (SSB) then bind to each of the single strands to prevent them from reforming the double helix.
A replication bubble has now been created. Replication can proceed both left and right. Each direction has its own fork, so the 2 forks are going in opposite directions.
Primase, aka RNA Polymerase, synthesizes a 10 BP primer/starting block that is complementary to the template strand.This primer provides a 3’OH that nucleotides can be added on to. Thus, the primers are made of RNA.
The part of the DNA that has not yet been reached by helicase and the rest of the DNA can start to supercoil, which can stop the replication fork from moving ahead. Topoisomerases move ahead of the replication fork and relieve this supercoil. Two types:
Type II: require ATP and breaks a phosphodiester linkage on both of the strands (double stranded) so that the supercoil unwinds and they rejoin.
Type I: break a phosphodiester linkage on 1 strand (single stranded) so that the other strand can unwind around it and the 2 strands can rejoin.
Describe DNA replication in eukaryotes
Replication in eukaryotes is catalyzed by 2 types of DNA Polymerases. They use dNTPs— dATP, dCTP, dGTP, and dTTP— a primer, the template strand, and Mg2+ as a cofactor.
Replication is uni-directional— the new DNA strand is always synthesized in the 5’ to 3’ direction.
The newly synthesized strand is obviously anti-parallel and complementary to the template strand.
A primer is put in, and then a base is added complementary to the base on the template strand. This base is always added to the 3’ end. Then a DNA polymerase forms a phosphodiester linkage to create the backbone.
The two DNA polymerases:
DNA Polymerase III: does most of the work. Has 5’ to 3’ polymerase function for synthesis and 3’ to 5’ exonuclease function to go backwards and proofread and repair if needed.
DNA Polymerase I: has the same two functions as III but also has an additional function— 5’ to 3’ exonuclease function that allows it to chew away at the RNA primers that are at the 5’ end of the Okazaki fragments.
In eukaryotes, the replication fork only moves in 1 direction because DNA Polymerase only adds nucleotides 5’ to 3.
The strand synthesized in the same direction as the replication machinery is the leading strand and is always intact. Bases are always added to the 3’ end. Thus, the strand synthesized complementary to the 3’ to 5’ strand will be the leading strand.
The strand synthesized in the opposite direction as the movement of the replication fork is known as the lagging strand. This strand is made up of many short fragments called Okazaki fragments.Thus, the strand synthesized complementary to the 5’ to 3’ strand will be the lagging strand because DNA Polymerase must attach and unattach constantly to synthesize the strand. But it is nonetheless synthesized 5’ to 3’.
This works via a “trombone slide”. The lagging strand, while being synthesized, forms a loop. As bases are added to the 3’ end, the loop gets larger until it gets too large and then the loop is released, which forms the short, Okazaki fragments. This loop then forms again and again until all of the DNA is replicated. This loop ensures that the leading and lagging strand are still synthesized at the same time.
DNA Ligase: seals the gaps between Okazaki fragments and thus connects all of the DNA fragments to yield a continuous strand.