Gap Genes
1/50
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
51 Terms
Zygotic Embryo-Lethal Patterning Genes
these genes are encoded by the zyote
4 general classes of lof mutant A-P patterning phenotypes that segregated zygotically
mutants follow Mendelian inheritance
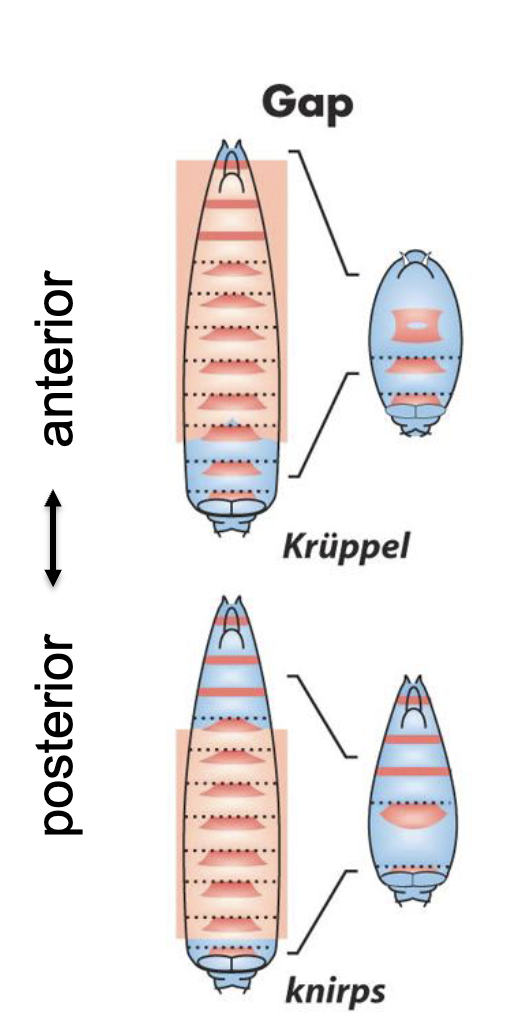
Gap Mutants
phenotype involves large gaps in the array of segments
used earliest in development
broad decisions made earlier in development, so if the genes controlling these decisions aren’t working then a broader part of the body pattern would be affected.
most likely to be the direct targets of the maternal effect transcription factors
These are the earliest zygotic genes to be expressed, so the only other TFs around to induce their expression would be from maternal effect genes.
bcd-, hb-, cad-, nos- are likely to be gap mutants based on their mutant phenotypes
Most similar because there are large gaps of missing segments ie. Head/thorax segments or abdominal segments
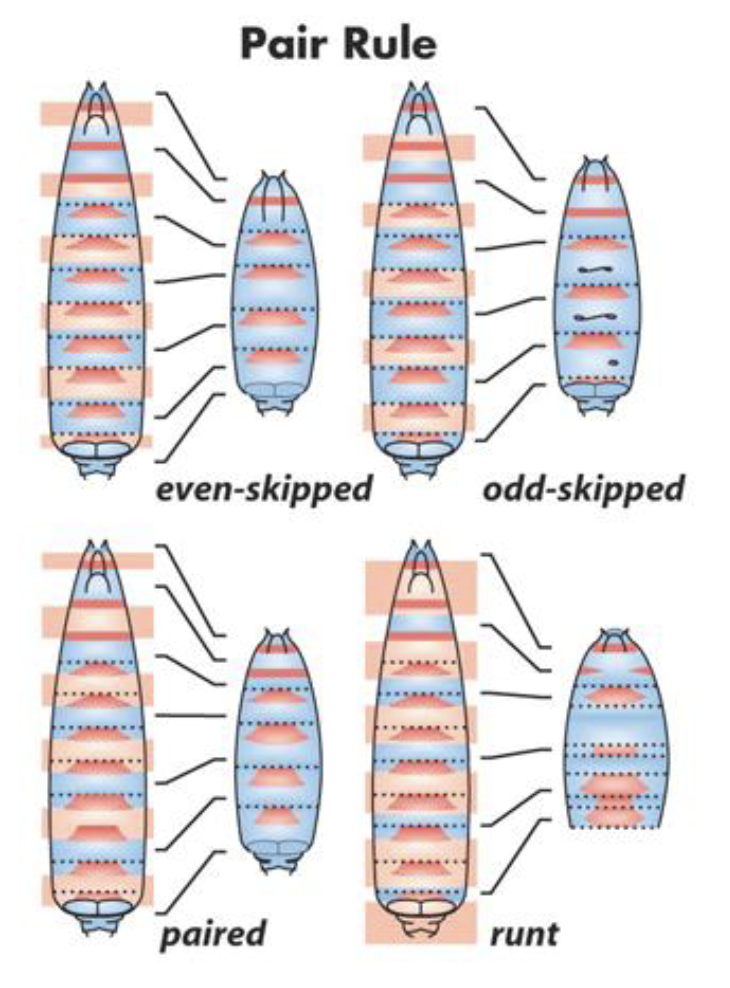
Pair-Rule Mutants
phenotype involves loss of every other segment
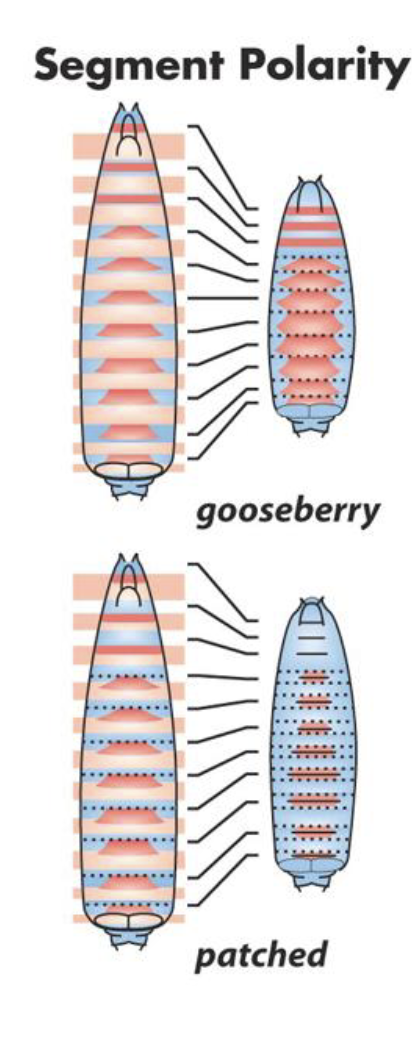
Segment Polarity Mutants
phenotype involves polarity defects of every segment
Each segment is present but internally disorganized.
e.g. missing posterior segments only
Mirror-image duplications within each segment
Loss of proper anterior/posterior compartment identity
Homeotic Selector Mutants
phenotype involves the change in identity of several adjacent segments
One segment develops as if it were another segment
The number of segments is normal, the segmentation pattern is normal, but the identity is wrong
What do the bcd, hb and cad transcription factor gradients do in the syncytial blastoderm?
directly activate the transcription of several zygotic embryo lethal patterning genes (gap genes) in nuclei of the syncytial blastoderm
i.e regulate expression of gap genes —> control which nuclei transcribe gap genes
these genes are expressed in specific regions of the embryo such that the gap gene proteins form several localized gradients
Gap Genes
developmental transcription factors that define the segmented embryo's body plan along the anterior-posterior axis
earliest transcribed: hunchback (hb-Z, not the maternal hb), Krüppel (Kr), knirps (kni)
all encode transcription factors
hb is a maternal effect gene that is needed at the earliest stage of development, but expression of zygotic hb is also induced for the next stages of development
How is the pattern of gap gene transcript established?
we have an incomplete understanding because the system is complex and we don’t understand all the interactions
in general, specific concentrations of maternal Bcd, Hb and Cad protein activates or represses the expression of specific gap genes
Establishing the Bicoid gradient experiment
a) reporter gene construct expressed in Drosophila embryo
b) reporter gene under the control of Hb regulatory sequences that contain Bicoid binding sties
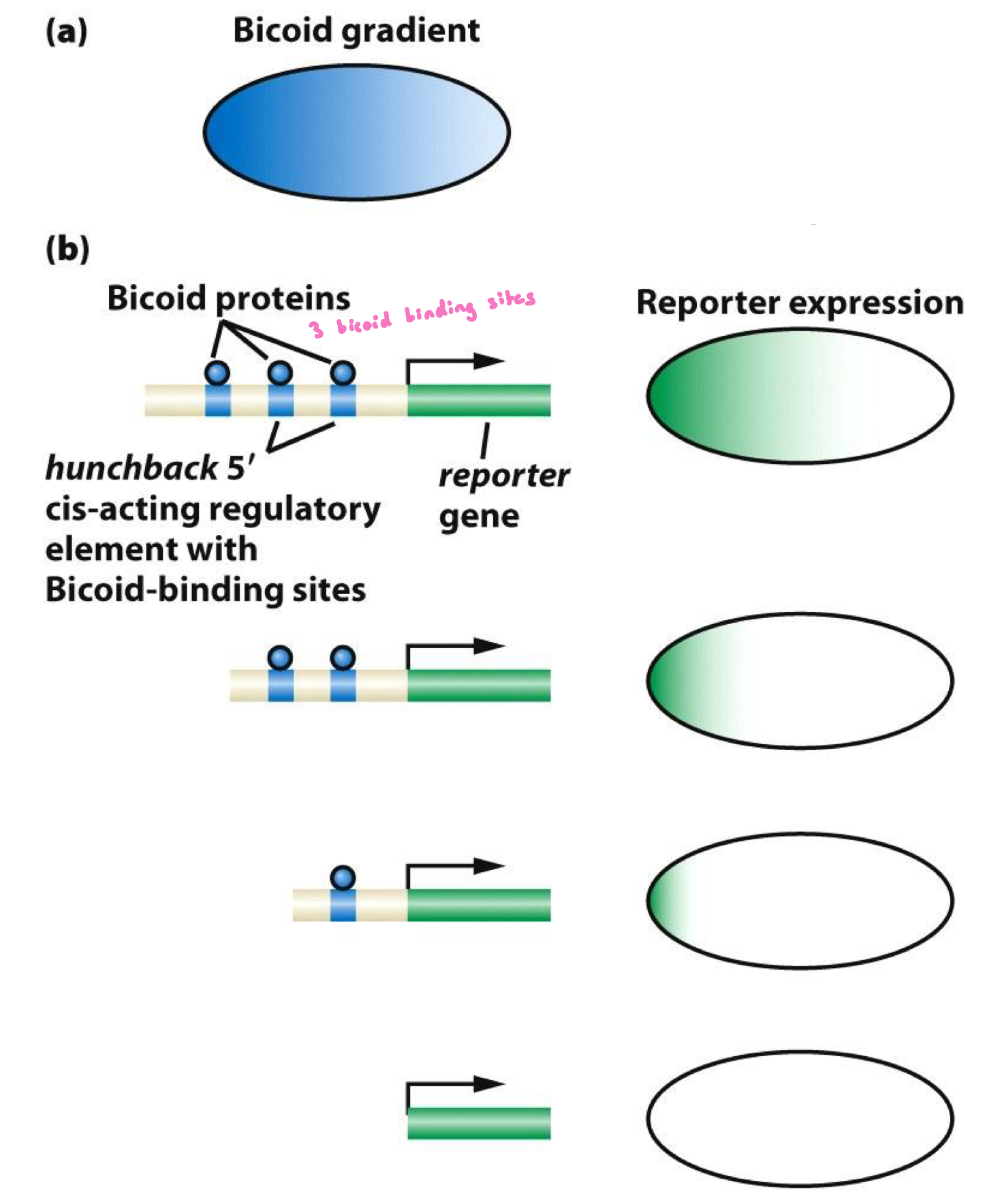
Establishing the Bicoid gradient experiment RESULTS
Results:
Mutation/deletion of the bcd binding sites of the hb enhancer/promoter controlling expression of a reporter gene alters zygotic expression of the reporter gene.
Deletion of some of the bcd binding sites decreases the size of the expression domain and the level of expression.
Deletion of all binding sites eliminates expression entirely
What does Maternal Bcd Protein Activate?
maternal Bcd protein activates zygotic hb+ transcription
Bicoid protein bidns to specific cis-acting sequences (9 bps in length) in the enhancer region of the hb gene
Hb-Z gene is transcribed in the syncytial blastoderm only by nuclei having Bcd transcription factor in high concentrations
higher Bicoid conc = all binding sites are occupied
lower Bicoid conc = less likely that all binding sites are occupied
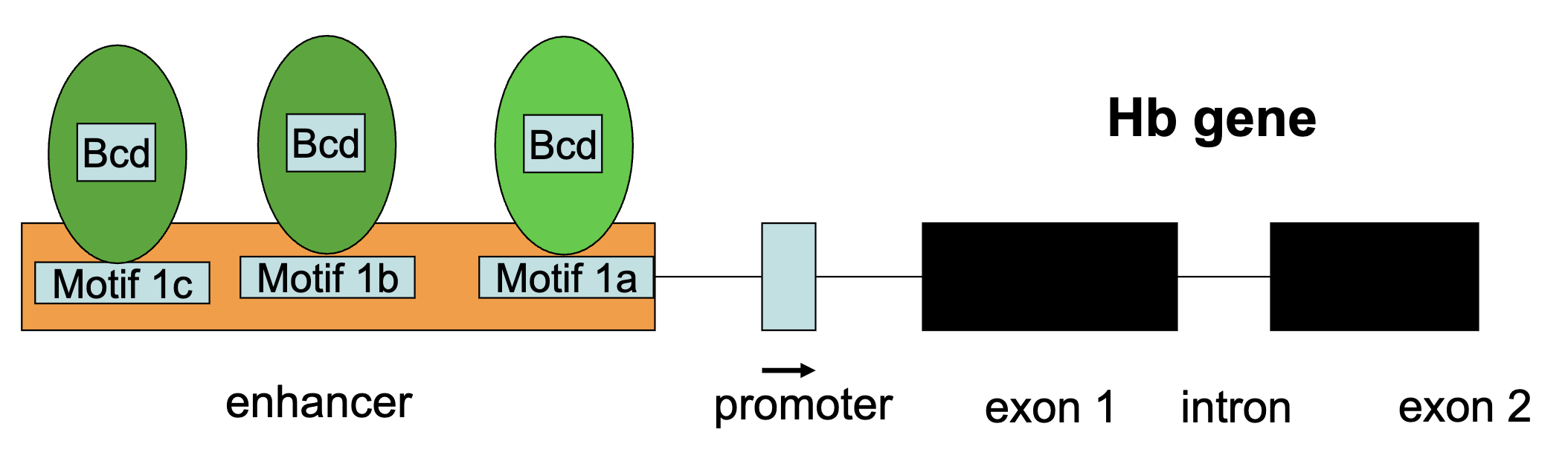
Bcd Protein Gradient Figure
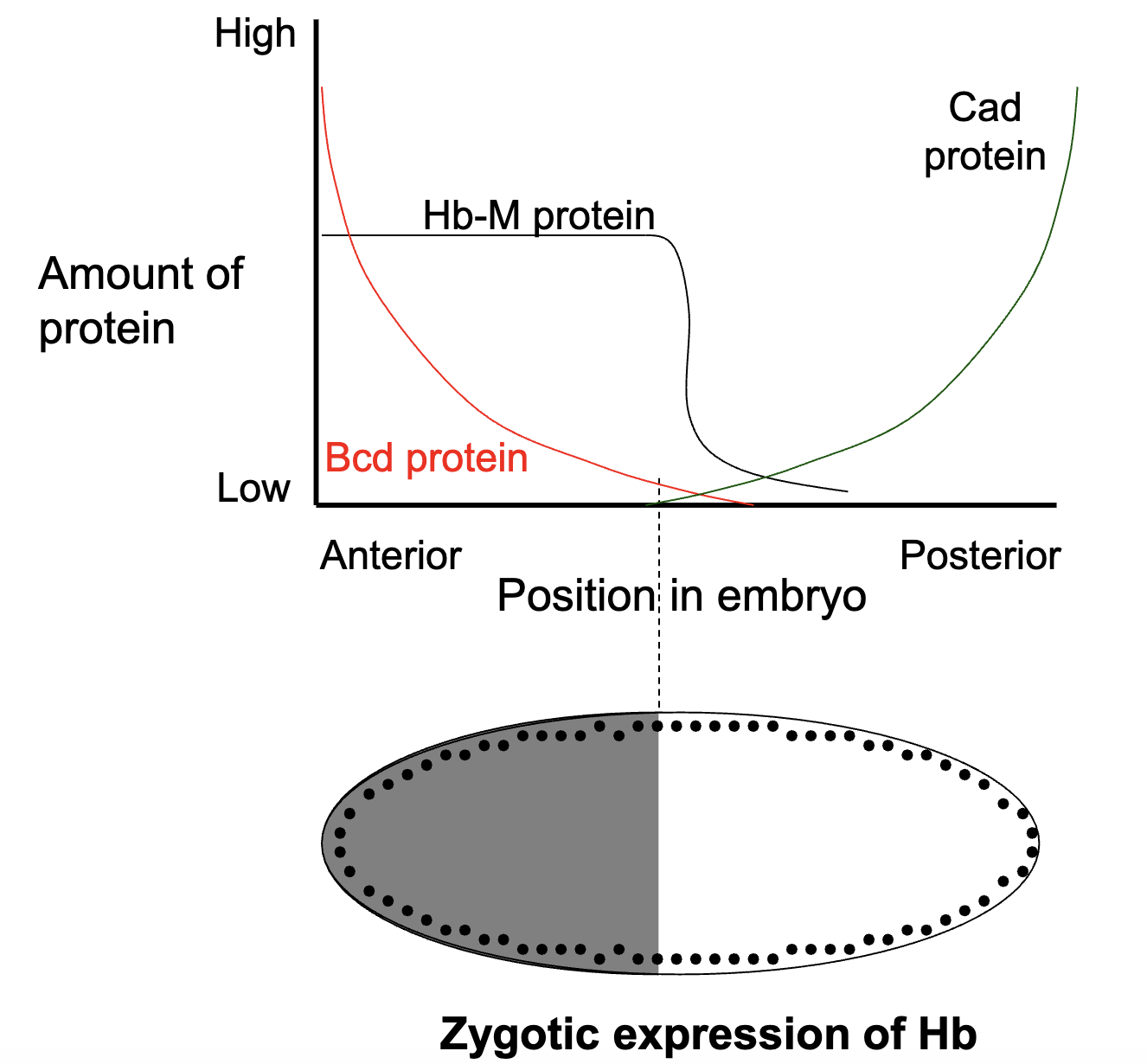
How is the pattern of Kr+ gene expression established?
specific concentrations of maternal Bcd and Hb protein activates or represses the expression of specific gap genes
high levels of Bcd and Hb protein INHIBIT Kr+ transcription
low levels of Bcd or Hb protein ACTIVATE Kr+ transcription
Kni protein also inhibits Kr+ transcription
figure = where we expect Kr+ to be expressed
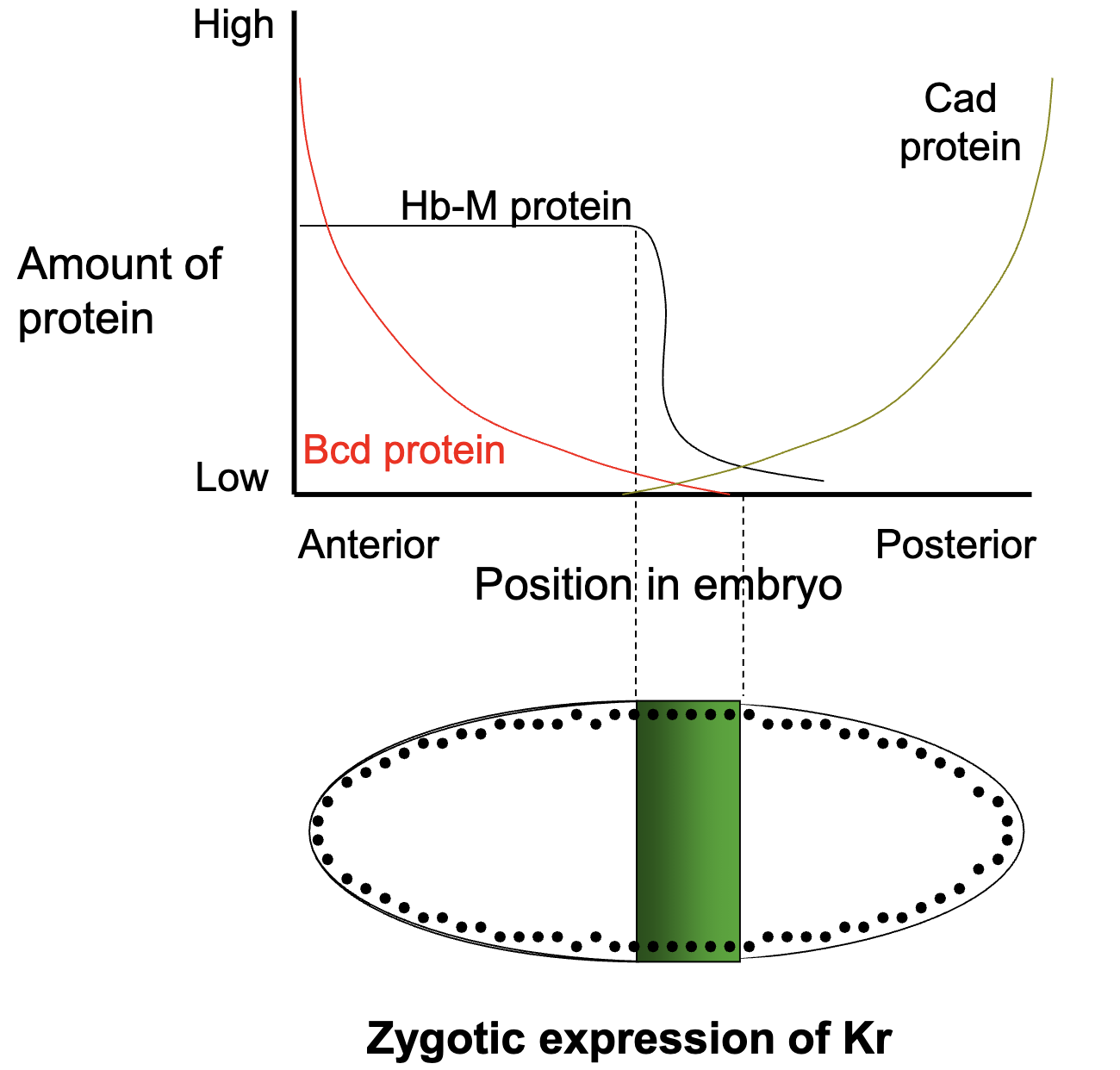
Kr Gene
Kr gene is transcribed only when either Hb+ transcription factor or bcd+ transcription are present in the correct concentrations (and when Kni protein is absent)
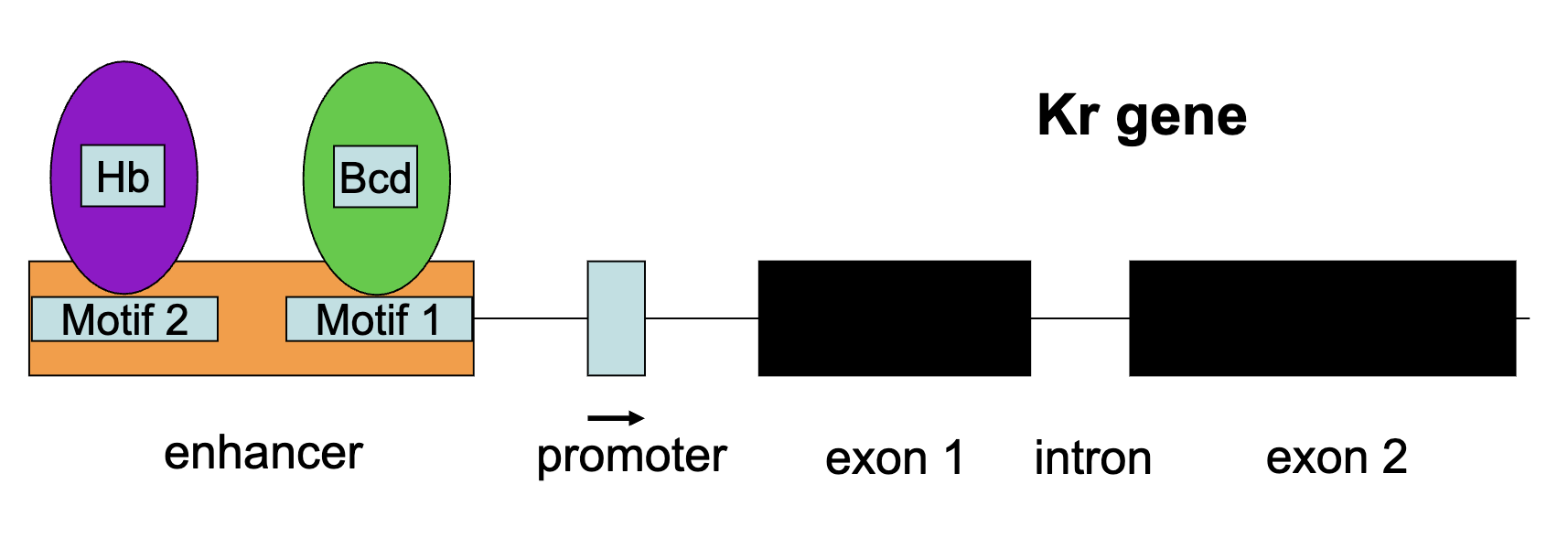
How is the pattern of Kni+ gene expression established?
specific concentrations of maternal Bcd, Hb and Cad protein activates or represses the expression of specific gap genes
Bcd protein INHIBITS kni+ expression
low levels of Cad and Hb ACTIVATES kni+ expression
Figure: where we expect Kni+ to be expression (where Bcd is absent but Cad and Hb are present)
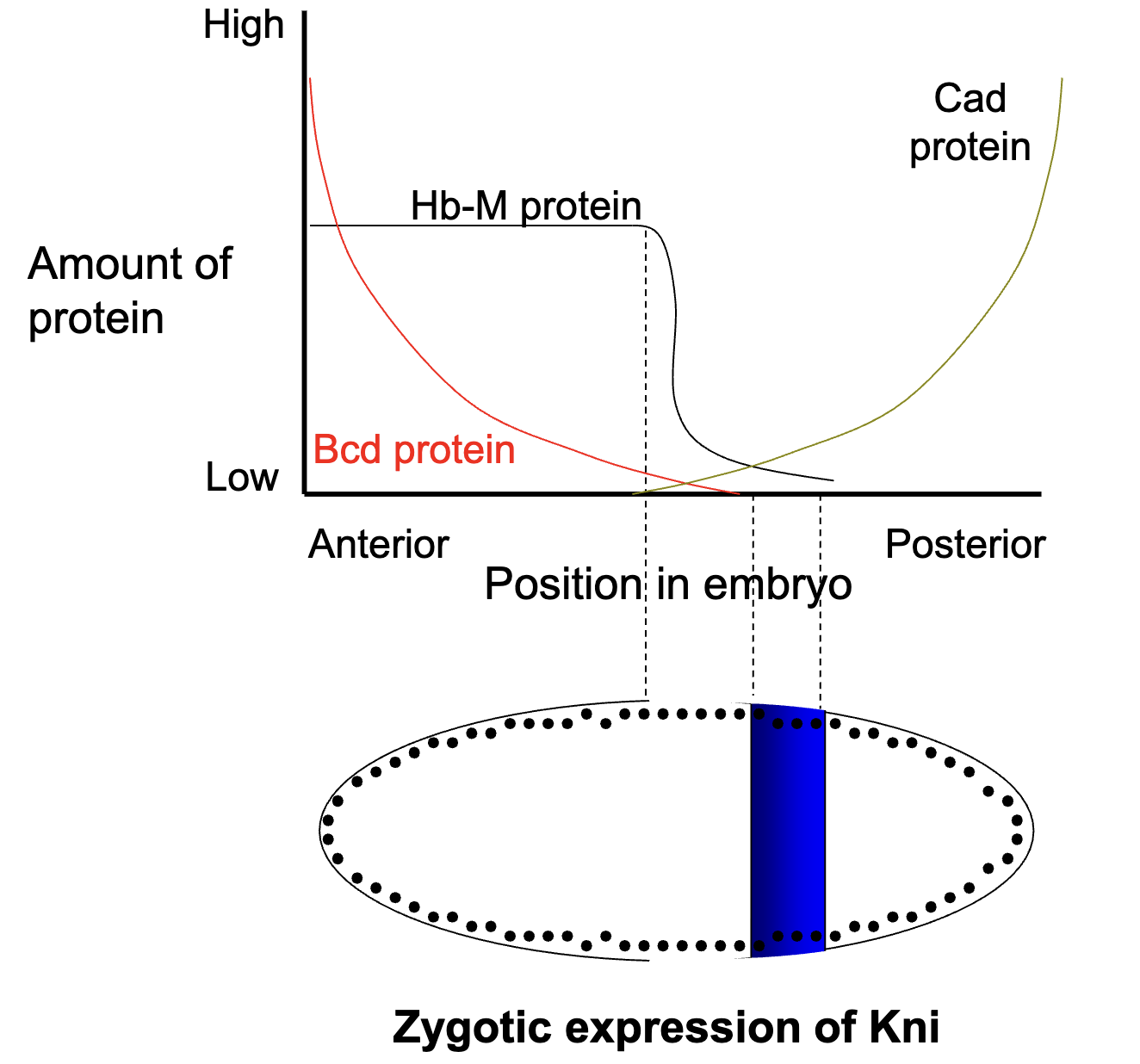
Pattern of gap gene transcript in the syncytial blastoderm
hb, Kr and kni are transcribed by contiguous nuclei in specific regions of the embryo, with hb more anterior to Kr and Kr more anterior to kni
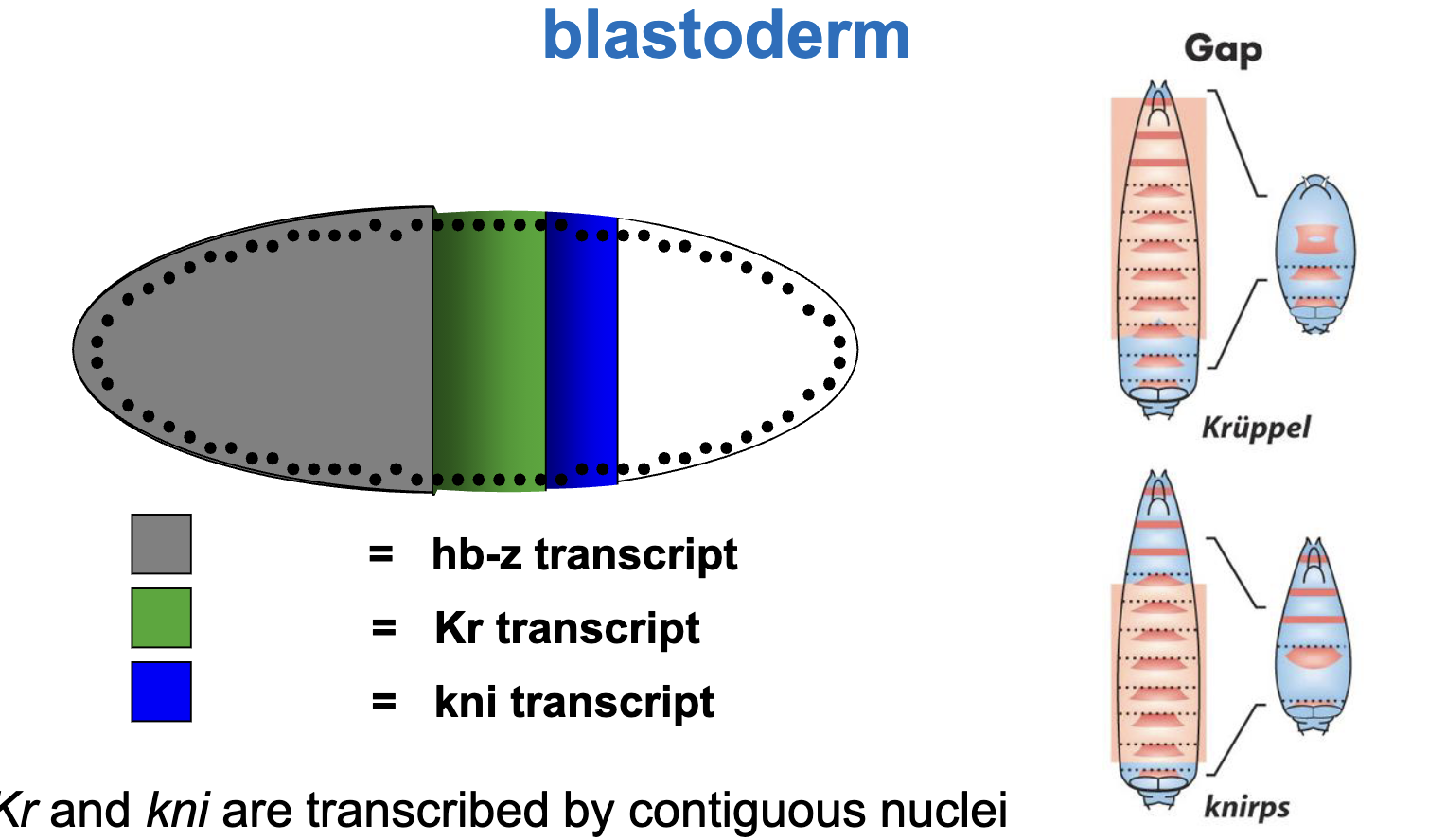
Why are the regions deleted in the mutants broader than the regions where the mRNA is localized?
This is happening in the syncytial blastoderm, so the proteins diffuse out from where they are translated
What kinds of evidence do you need to prove that gene Y is a target gene of transcription factor X?
a) mutating X affects the expression of Y
done by examining expression of Gap genes in embryos where a maternal gene was mutated or missing
b) X can bind to cis elements in the regulatory sequence of Y in vitro (EMSA)
c) X binds to the regulatory sequence of Y in vivo (ChIP-PCR)
b) and c): molecular genetic analysis of enhancer regions of the gap genes
analysis of binding between the maternal effect TFs to the cis-acting sequences of the gap genes
mutation analysis of the binding sites to determine whether they are required for controlling the expression of a reporter gene
a) How would Kr expression change in the absence of Bcd activity (embryos from bcd -/- mothers)
No Bcd protein gradient
No zygotic expression of hunchback–lower Hb protein gradient.
Therefore Kr expression occurs more to the anterior
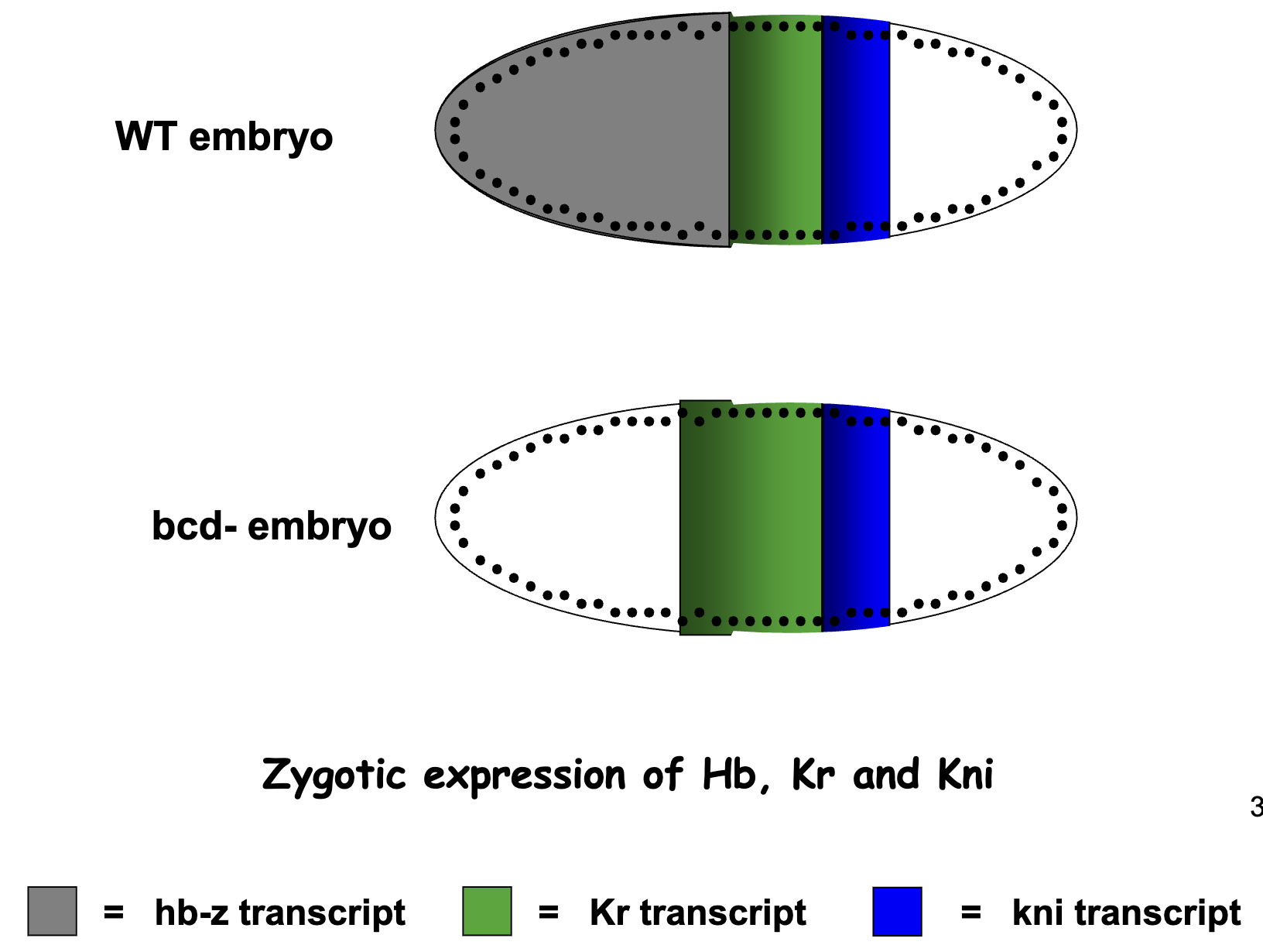
b) + c) Kni expression Experiment
delete the Bcd binding site in the kni regulatory region.
normally, Bcd represses Kni in the anterior, so Kni is expressed more posteriorly
in mutant: Kni expression expands more towards anterior
effect on Kr expression: if Kni expands, it represses Kr in more regions
figure: solid blue = WT zygotic expression of Kni
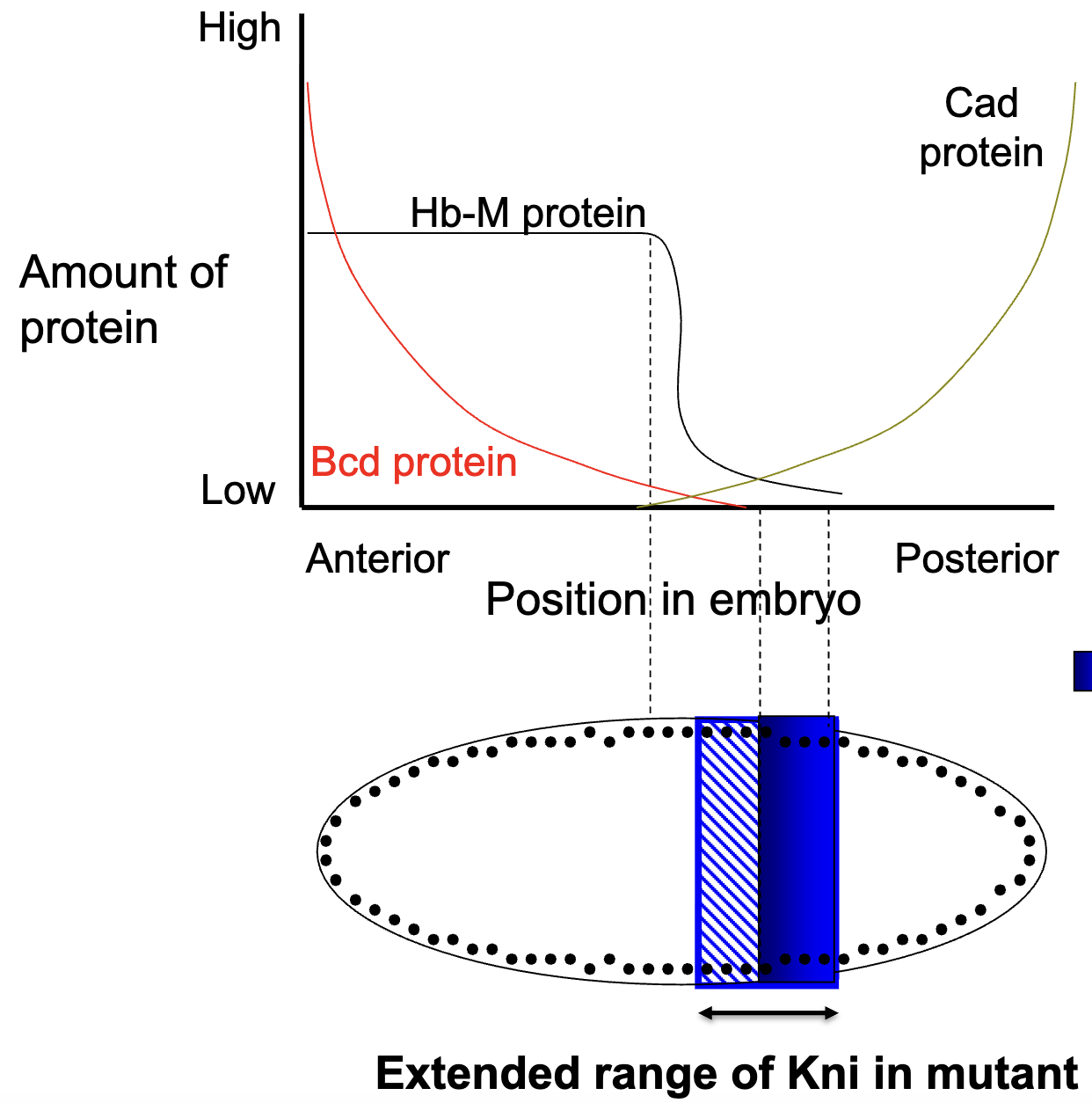
Summary: Gap Gene Expression Patterns
gap gene is transcribed in nuclei of a certain region within the syncytial blastoderm
the transcription patterns of the gap genes in the syncytial blastoderm are initially determined by activaition and/or repression by the maternal morphogens (transcription factors) bcd, hb-M, and cad
translation of the localized gap gene mRNA molecules results in a series of localized gap protein gradients
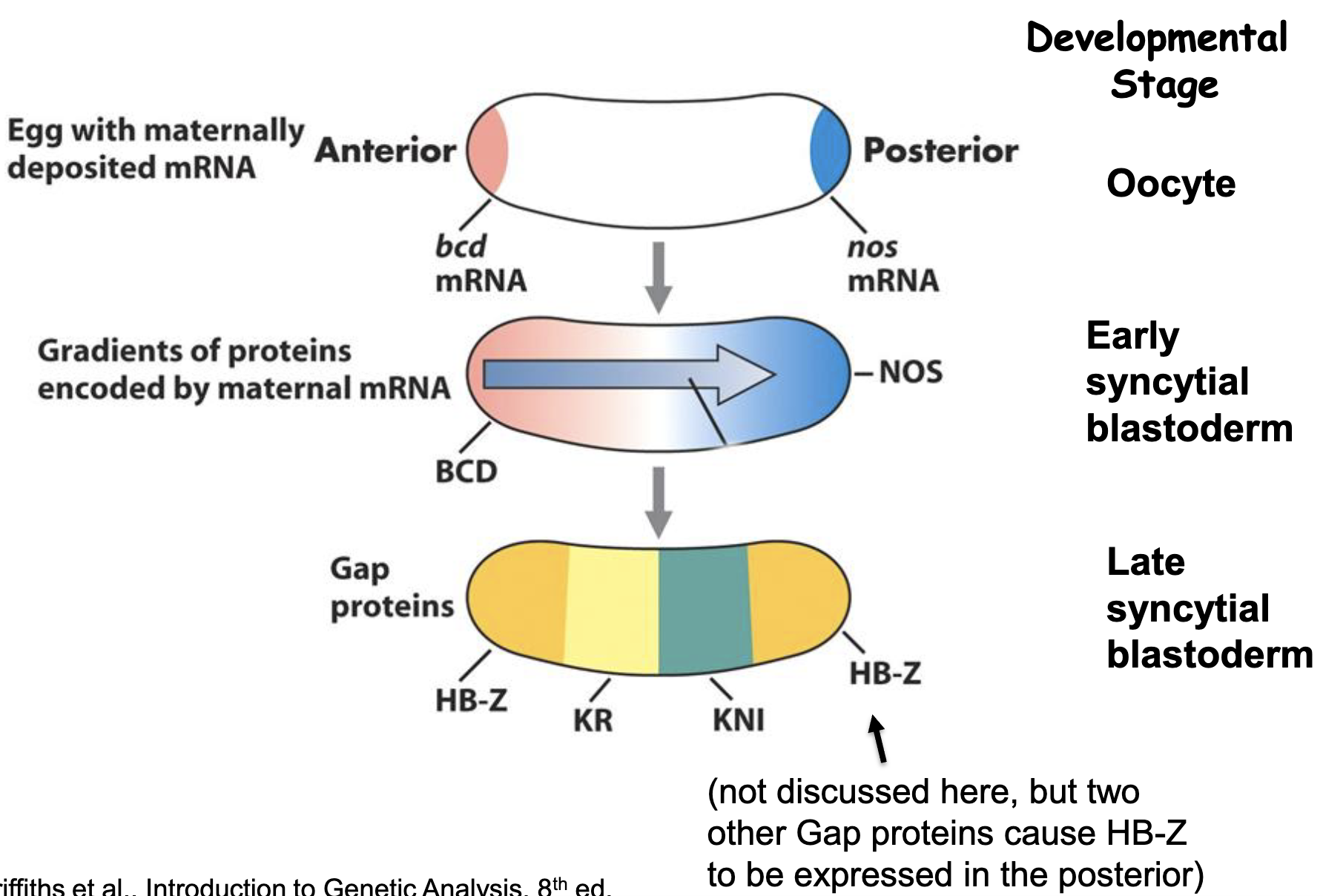
Pair Rule Genes
an embryo homologous for a loss-of-function mutation in a pair rule gene lacks every other segment
looking at the mutant phenotype: we would expect pair rule genes to be normally expressed in the missing areas
the role of the gene is to promote segment formation
several pair rule genes have been cloned and all encode transcription factors
pair rule genes are expressed in seven stripes, 3-4 cells wide and perpendicular to the anterior-posterior axis
stripes of expression are separated by a 3-4 cell wide stripe of no expression; the total number of stripes (expression + no expression) correspond to the number of segments (14)
transcribed in a pattern of seven stripes around the circumference of the embryo in cellular blastoderm
Impact of GAP proteins on Pair Rule Genes and Homeotic Selector Genes
pair rule genes and homeotic selector genes are directly activated by GAP proteins (which are expressed in syncytial blastoderm)
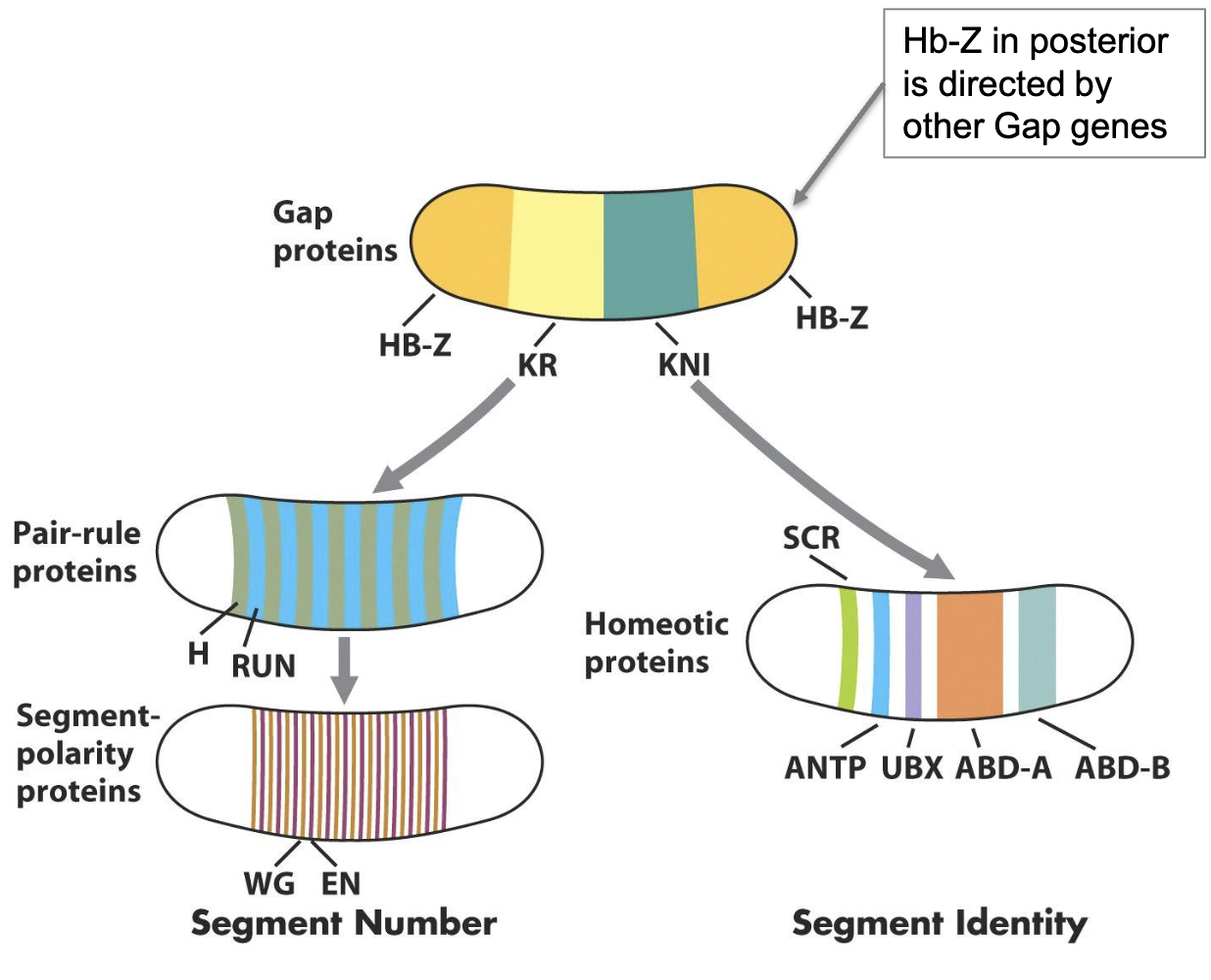
How is the expression of primary pair rule genes established by gap gene transcription factors 1A)
1) pair rule genes are activated by a combination of the gap gene morphogens
pair rule genes are expressed after blastoderm cellularization
during the syncytial blastoderm stage, nuclei share the same cytoplasm, gap genes (e.g. hb, Kr, kni) are transcribed in broad domains along the anterior-posterior axis.
at the cellular blastoderm stage, membranes form around nuclei and gradients become fixed concs in each cell (each cell now contains a specific combination of gap gene transcription factors)
combinations activate primary rule gene
How is the expression of primary pair rule genes established by gap gene transcription factors: Figure of Gap Gene Transcript Pattern
hb, Kr and kni are transcribed by contiguous nuclei in specific regions of the embryo with hb more anterior to Kr and Kr more anterior to kni
gap genes are first expressed 11-13th cycles of nuclear division
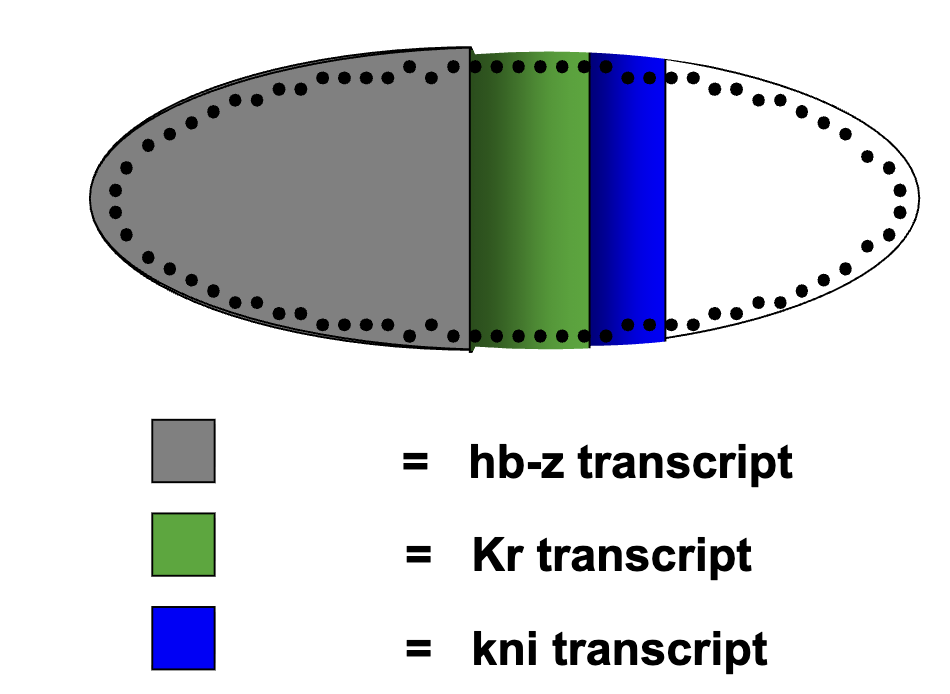
How is the expression of primary pair rule genes established by gap gene transcription factors: Figure of Distribution of Gap Proteins
Cellular blastoderm: membrane forms around the nucleus and Gap proteins are trapped inside → protein gradients get fixed
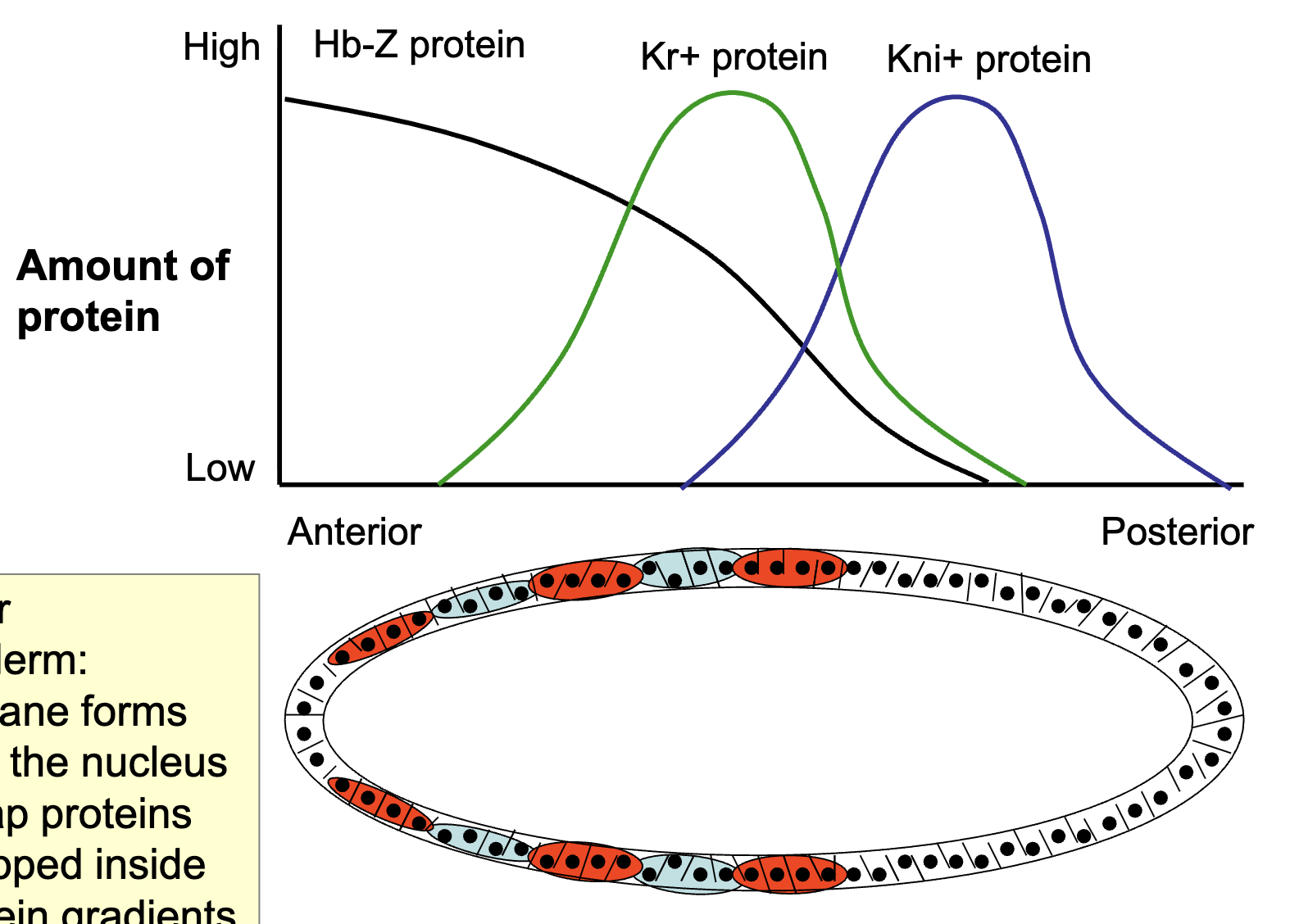
Pair rule genes are activated by the Gap proteins FIGURE
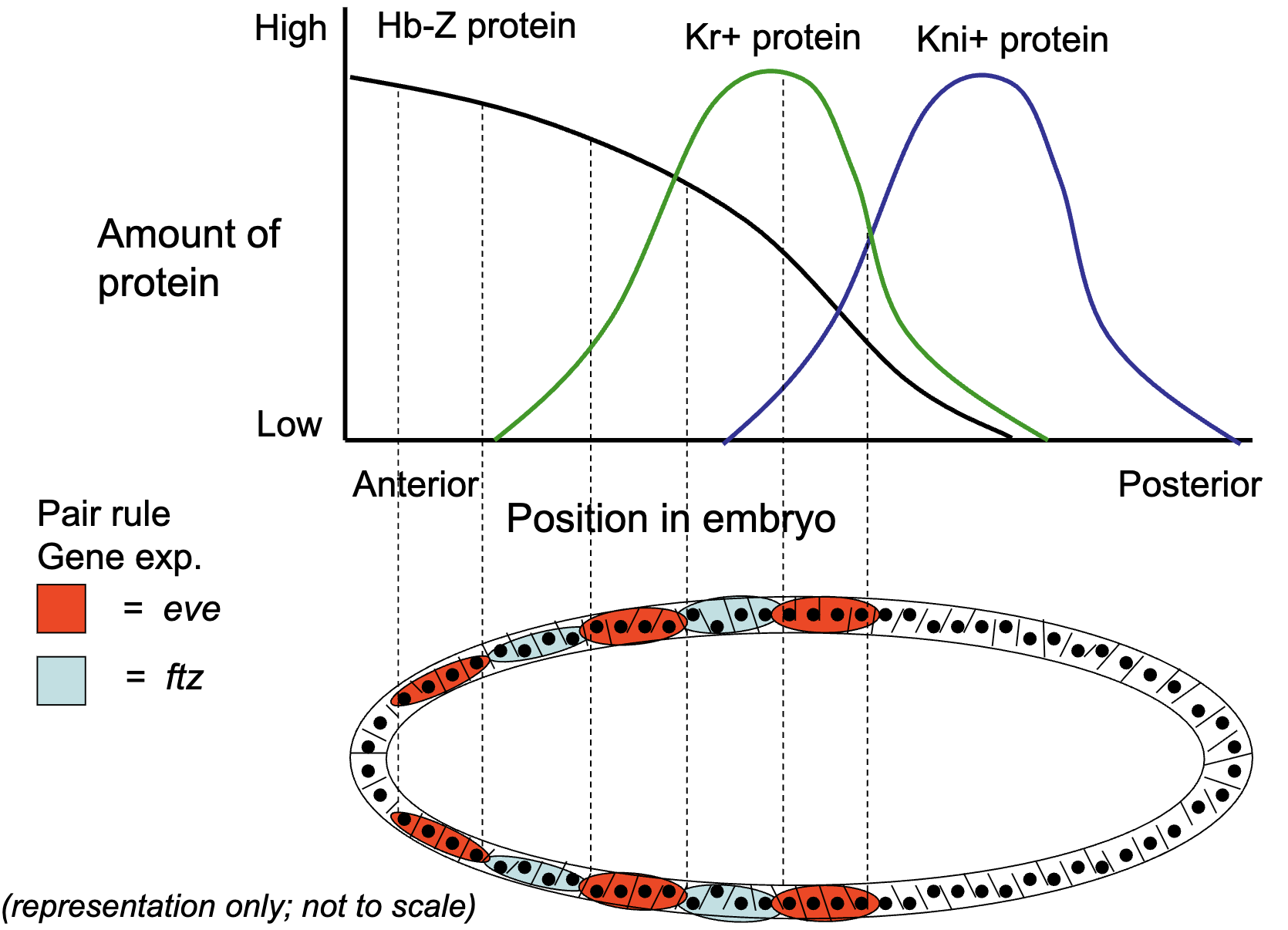
How is the expression of primary pair rule genes established by gap gene transcription factors 1B)
pair rule genes are activated by a combination of the gap gene morphogens
in gap gene mutants, the pair rule genes are not expressed correctly, some of the stripes of expression do not form
if gap gene loses function → all striped of pair-rule genes are not transcribed
Figure: e.g. in Hb-Z LOF mutants
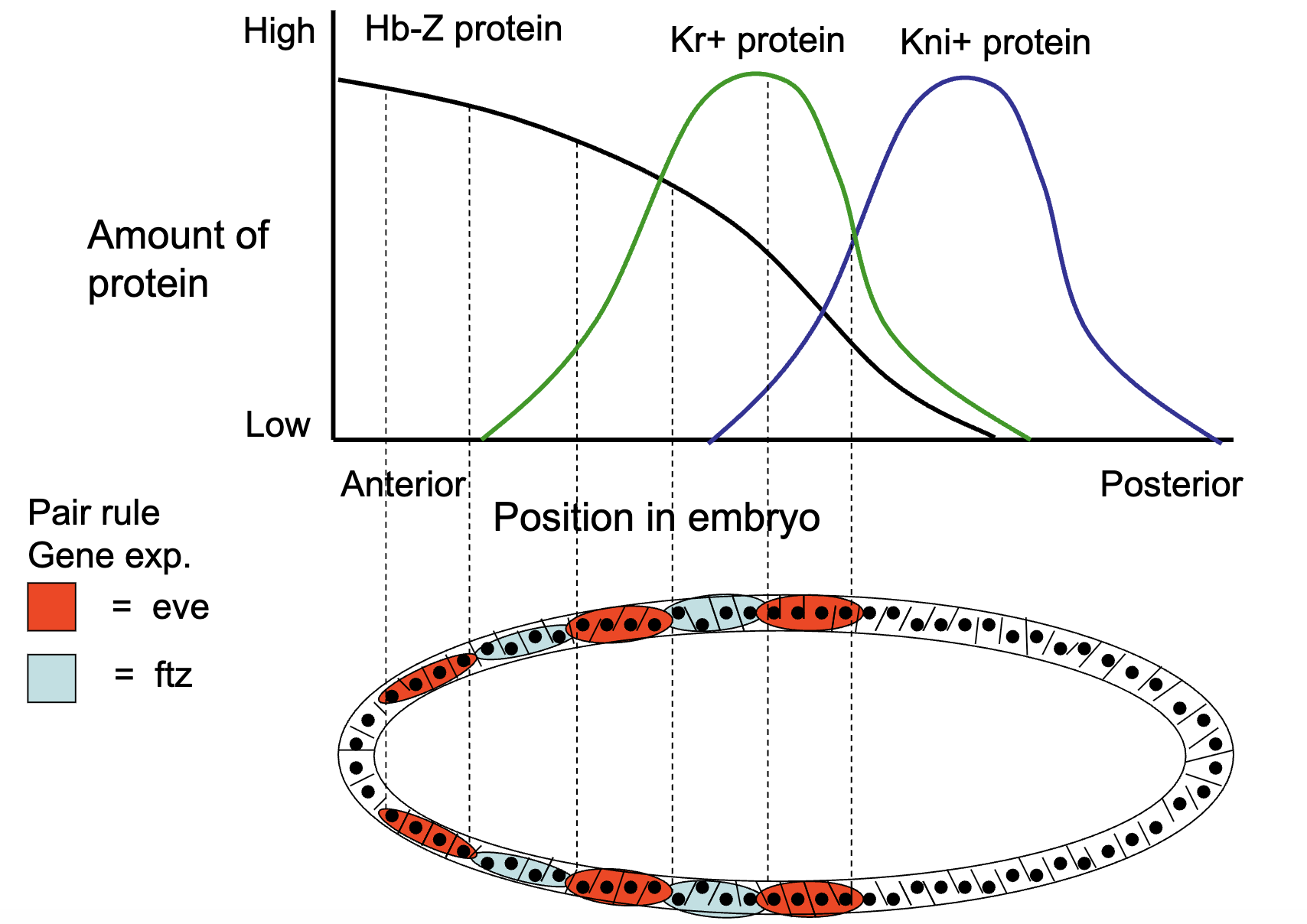
A Drosophila mutant is discovered that lacks one thoracic segment of the developing embryo and is found to be missing one stripe of odd-skipped gene expression from the exact region of the missing segment
In which of the following genes is the mutation causing this defect most likely to be?
Odd-skipped
What kind of mutation within a Pair Rule gene could result in an embryo that is missing alternative segments (corresponding to the pair-rule)?
deletion within the protein coding sequence
mutation of the core promoter
deletion that removes the entire enhancer
mutation affecting splice sites
The mutation likely affects function of protein and its expression
Where in a Pair Rule Gene would you expect to find a mutation if a single segment was missing from the embryo
Deletion that removes part of the enhancer
How is the expression of primary pair rule genes established by gap gene transcription factors 2A)
the pair rule enhancer regions are very complex, involving multiple binding sites for all of the gap gene transcription factors
a) the binding of diff amount and combinations of gap gene proteins to the pair rule regulatory region results in its activation or repression
specific enhancer regions regulate expression in specific tissues (e.g. eve stripe-specific enhancers)
2A) Hypothetical Pair-Rule Gene
specific enhancer elements are recognized by diff combinations of gap morphogens to produce specific stripes
deletion of one element can result in loss of one stripe
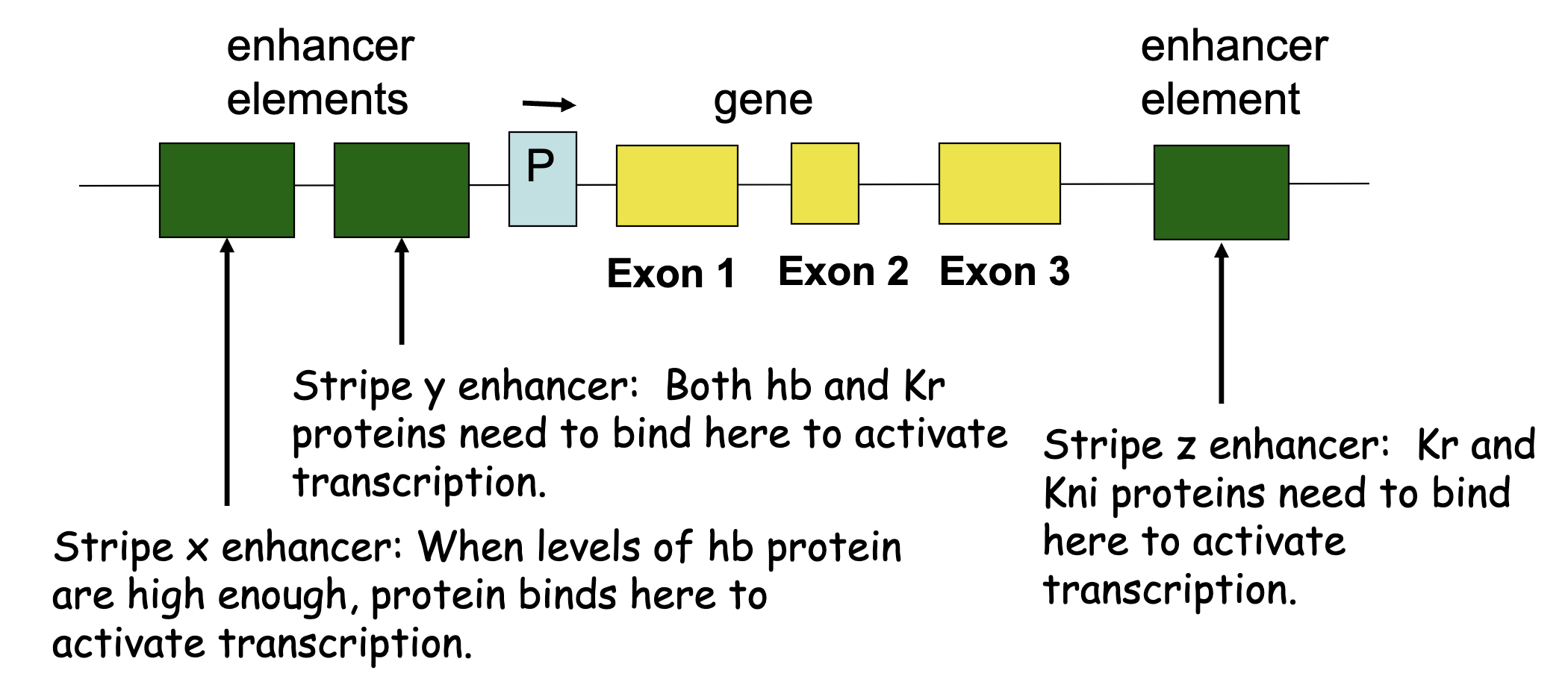
Impact of maternal effect and gap proteins in combination on pair rule stripe formation
combinations of maternal effect and gap proteins control individual pair rule stripe formation
figure: eve regulatory region
e.g. at high [Bcd] and [Hb], and low [Kr] and [Gt] we’d see stripe 2 enhancer to be active, driving eve expression
![<ul><li><p>combinations of maternal effect and gap proteins control individual pair rule stripe formation </p></li><li><p>figure: <em>eve</em> regulatory region</p><ul><li><p>e.g. at high [Bcd] and [Hb], and low [Kr] and [Gt] we’d see stripe 2 enhancer to be active, driving <em>eve </em>expression</p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/e41f3e4c-89c2-4876-bc3d-21aef1a0f4d3.png)
How is the expression of primary pair rule genes established by gap gene transcription factors 2B)
The pair rule enhancer regions are very complex involving multiple binding sites for all of the gap genes transcription factors
b) deletions of parts of the enhancer region of pair rule genes can result in the loss of only one or two of the stripes
2B) Each enhancer region controls expression of eve within a diff stripe
Each enhancer region controls expression of eve within a diff stripe
lacZ reporter genes fused to diff enhancers found within eve; eve protein stained with orange, lacZ reporter activity detected with brown
stripe 5 has a very broad expression of the reporter gene in the giant-deficient embryo, this means Giant is a repressor of stripe 5 eve
Next flashcard continues this one
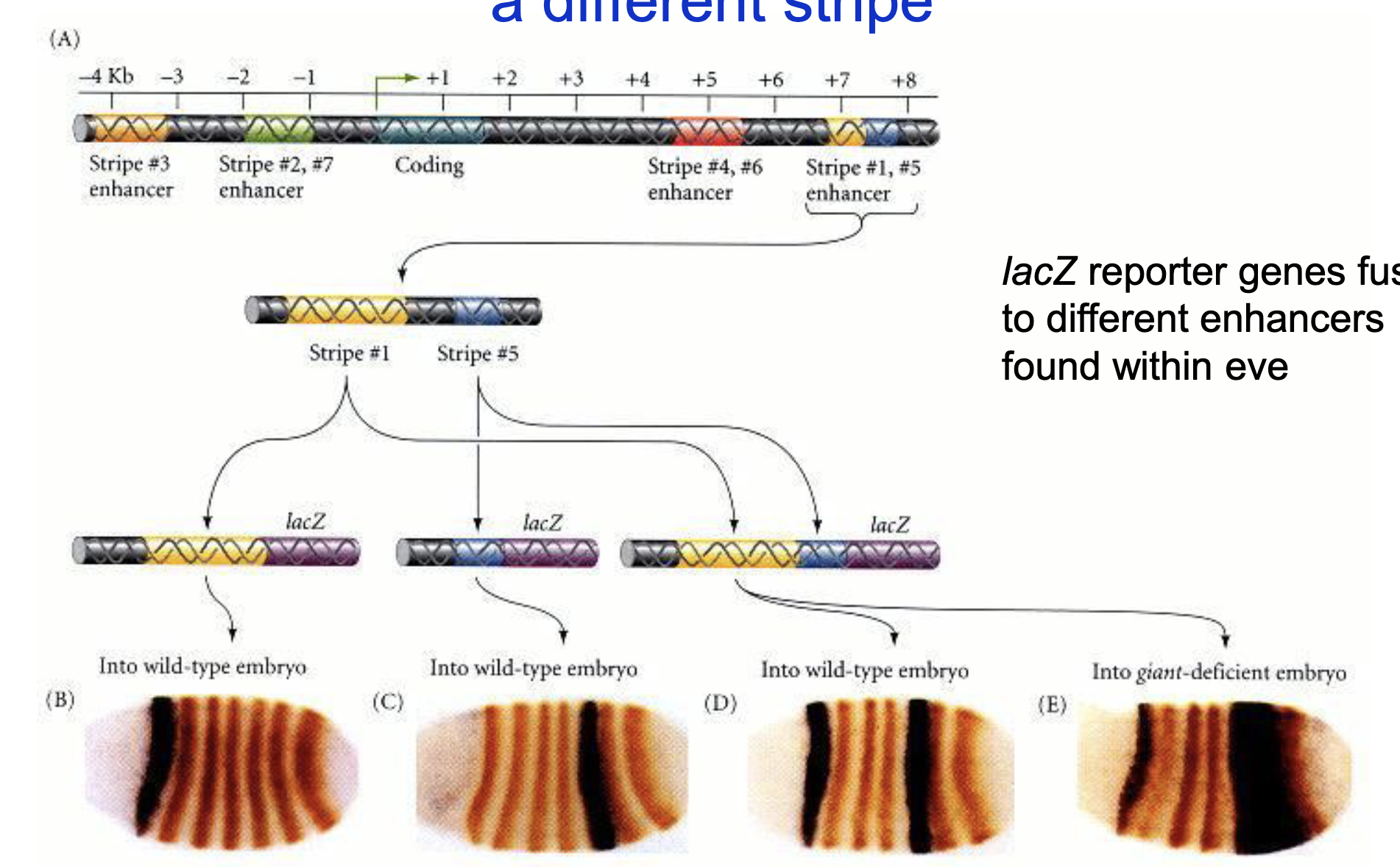
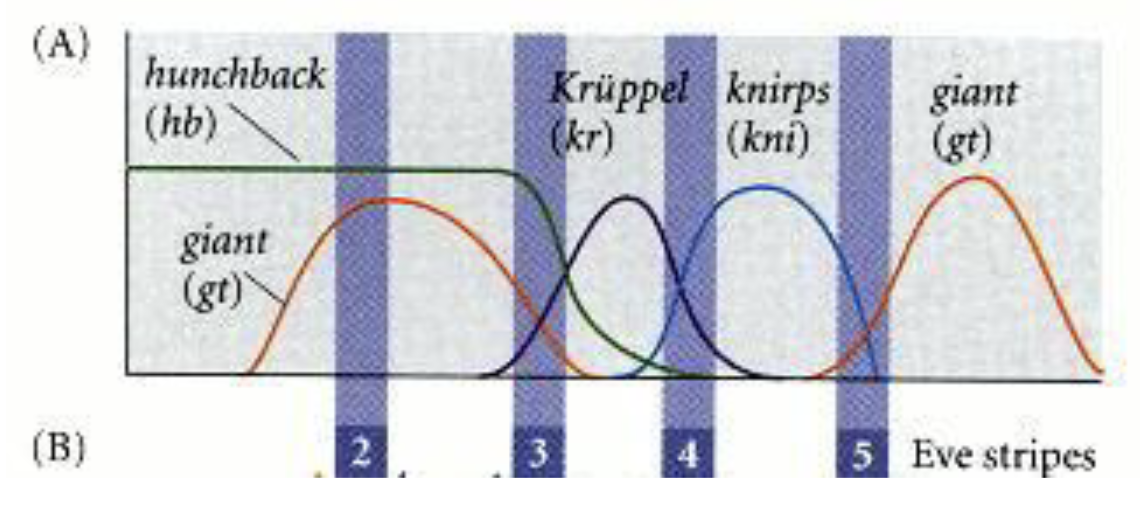
normally high knirps (anterior) and high giant (posterior) define stripe 5, but now with not gt, the posterior boundary is gone
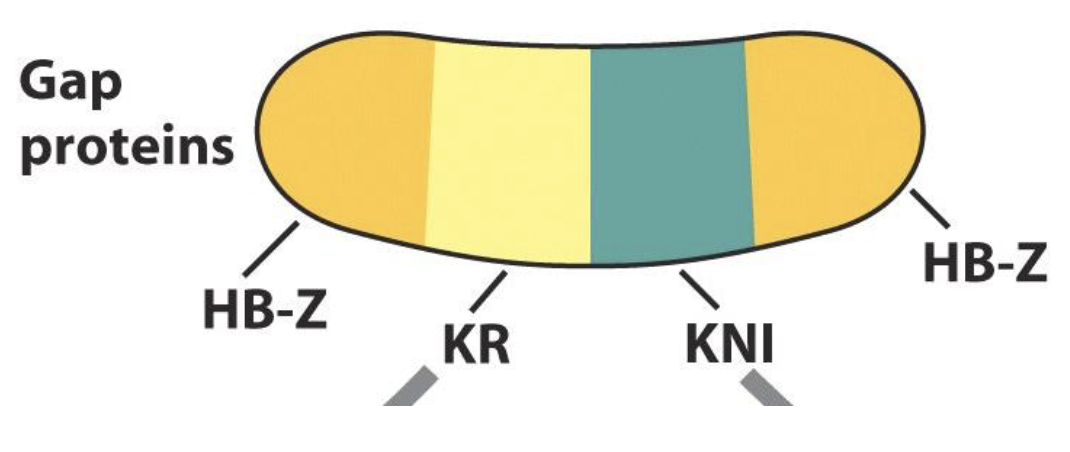
Zygotic Hunchback (HB-Z)
zygotic Hunchback protein is expressed at both the anterior and posterior ends of the early Drosophila embryo
HB-Z expression at the anterior end is regulated by Bicoid, where as HB-Z expression at the posterior end is regulated by Gap proteins
the female parent also expersses and deposits HB mRNA in the oocyte before it is fertilized to become the zygote
we would expect the regulatory region for HB to look like 3 regulatory regions
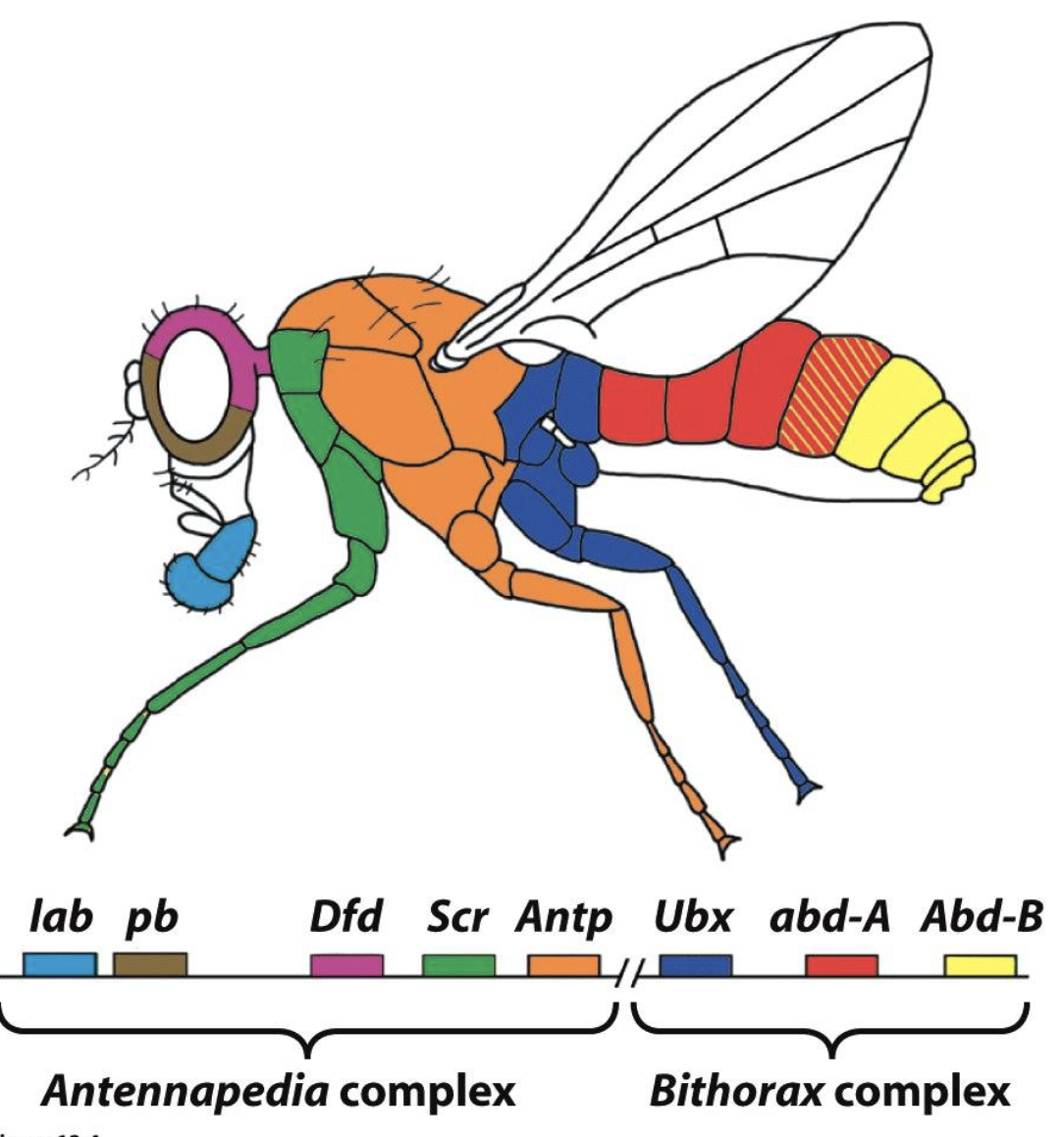
Homeotic Selector Genes (Hox Genes)
Hox genes regulate the identity of body parts (regulate segment identity)
code for transcription factors
expression is regulated by gap proteins thru mechanisms similar to pair rule genes
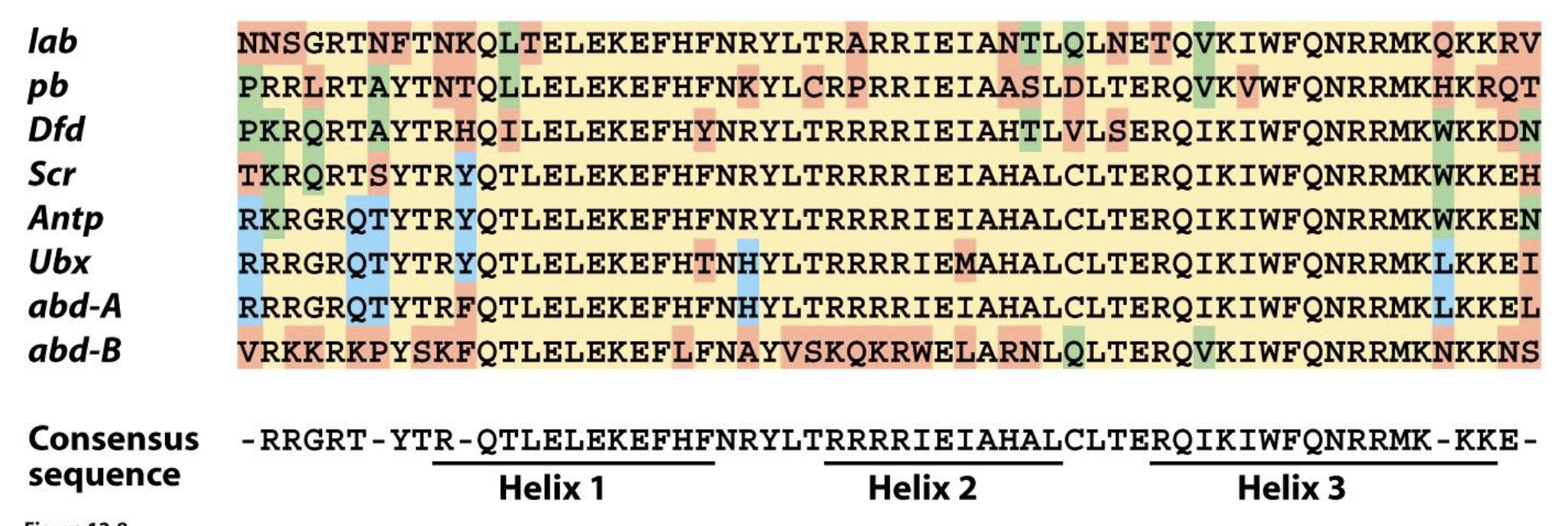
Homeotic (Hox) proteins have a sequence in common
highly conserved protein domain (homeodomain) that encodes 3 ⍺-helices
helices 2 and 3 form a DNA binding motif (helix-turn-helix) found in many DNA binding proteins that regulate gene expression
Hox proteins are sequence specific DNA binding proteins that regulate gene expression
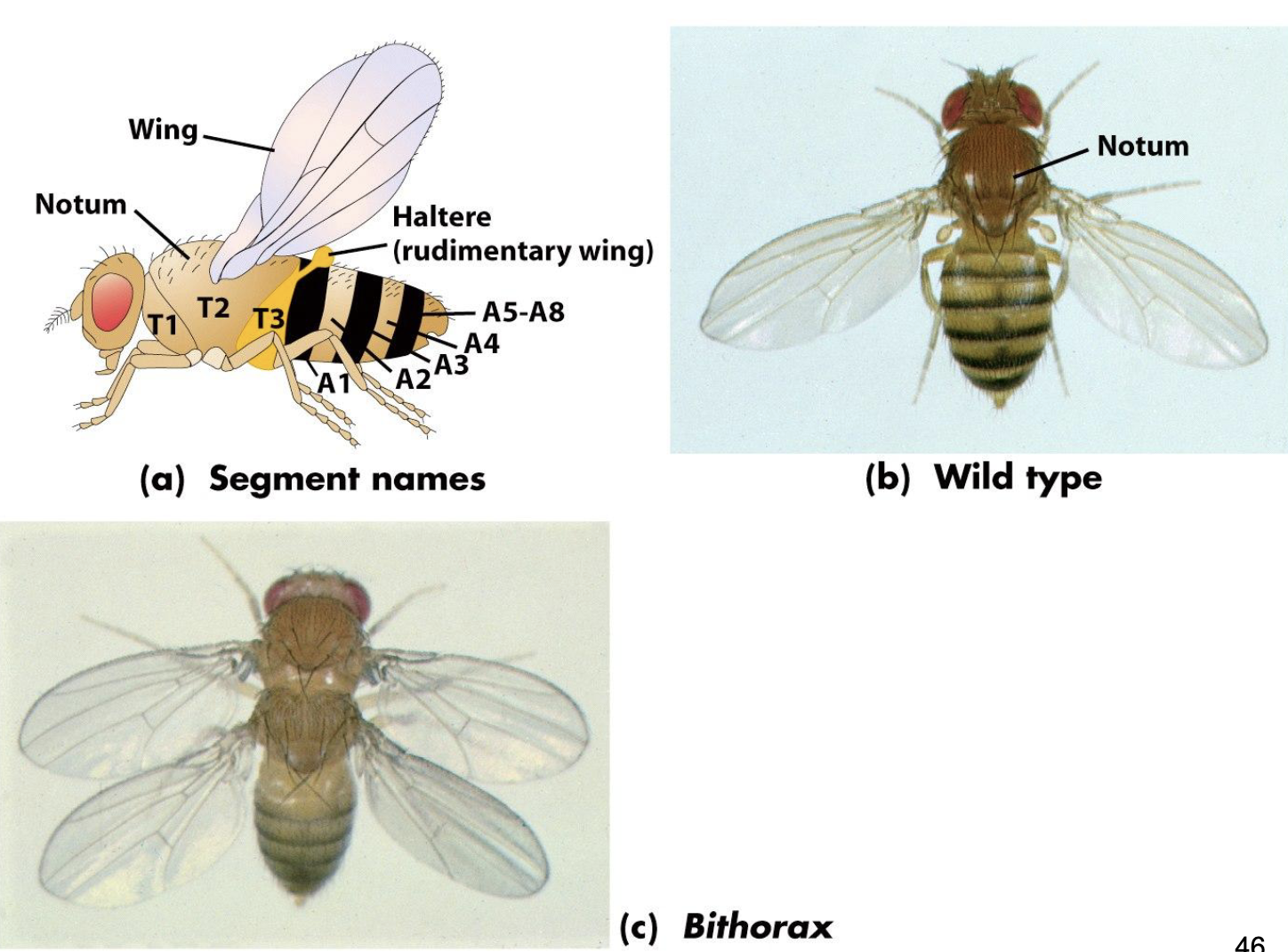
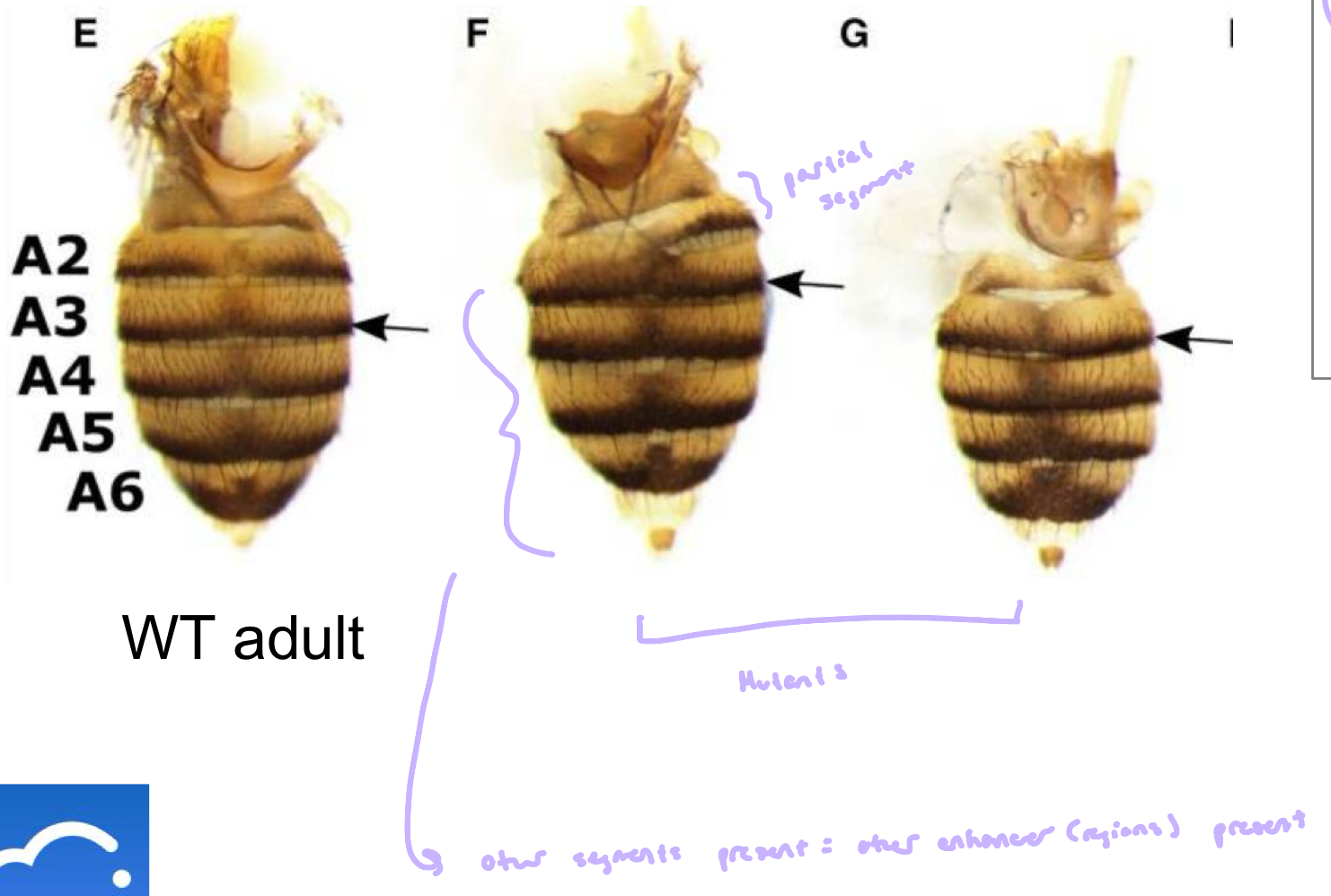
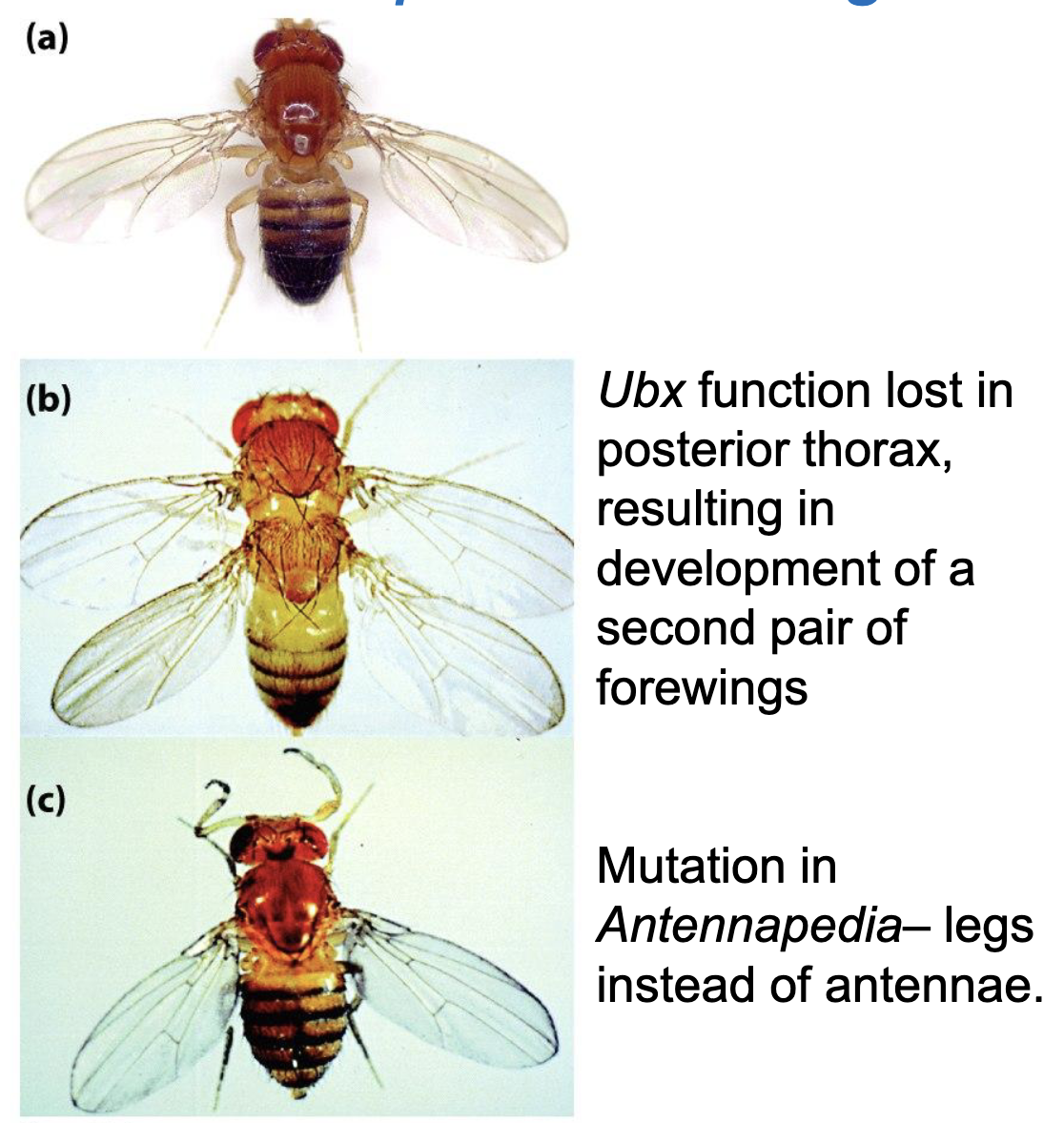
Examples of Homeotic Mutants of Drosophila
a) WT
b) Ubx function lost in posterior thorax, resulting in development of a second pair of forewings
c) Mutation in Antennapedia, legs instead of antenna
Hox Genes: Distal-less (DII)
Distal-less is a target of Hox homeotic selector genes
gene directs development of appendages (antenna, mouth parts, legs)
expression is restricted to anterior segments (so no appendages in the abdomen)
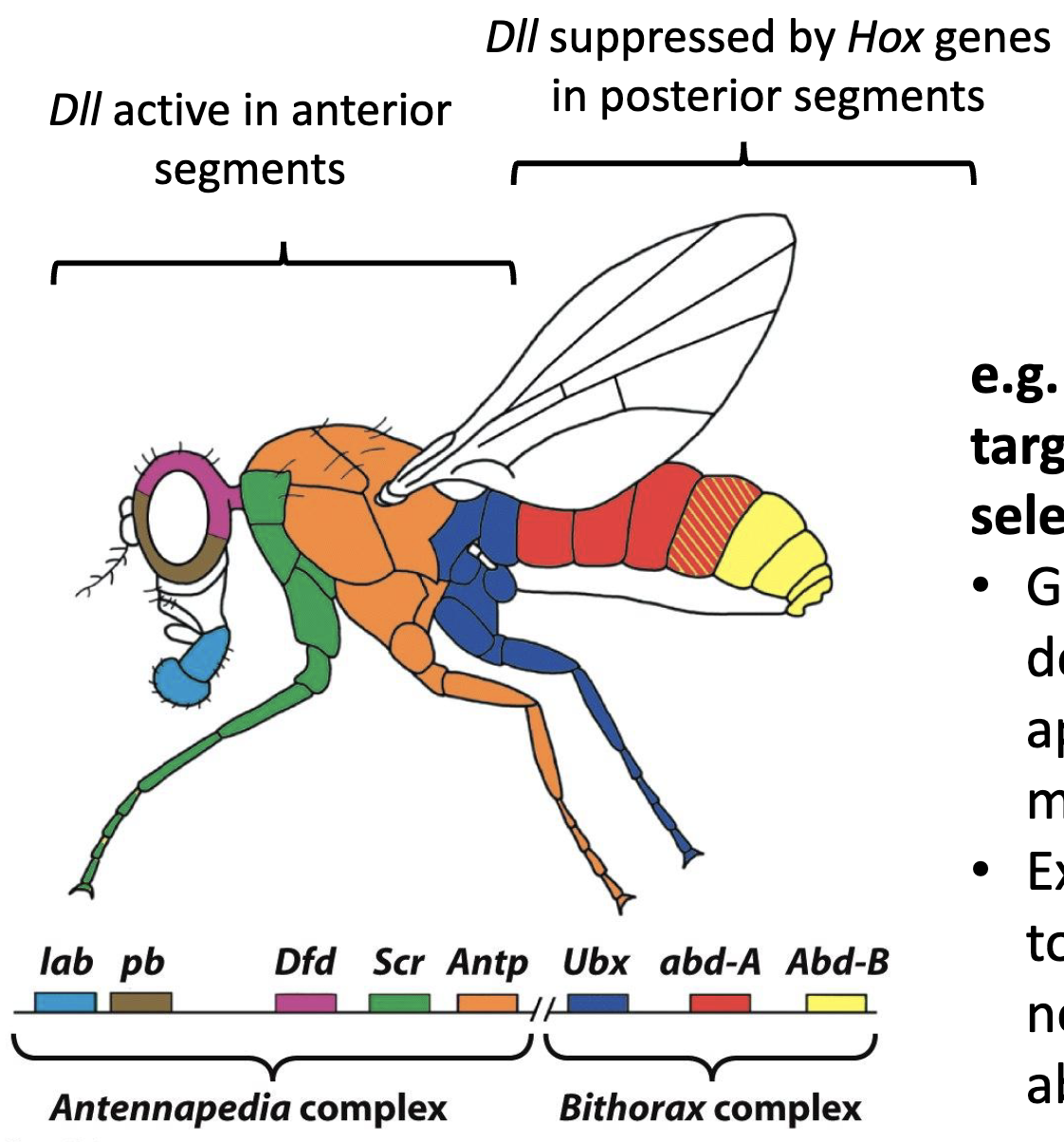
What is DII repressed by?
DII is repressed by Hox proteins Ultrabithorax (Utx)
DII is absent in segments where Ubx is expressed
when DII is derepressed in abdominal segment A1, the segment develops appendages (wings) and fly has two sets of wings (change in A1 identity)
Figure: WT expression
Blue: Engrailed (expressed in posterior of each segment, used here to visualize segments)
Purple: Ubx expression
Red: DII expression, this marks position of future appendages
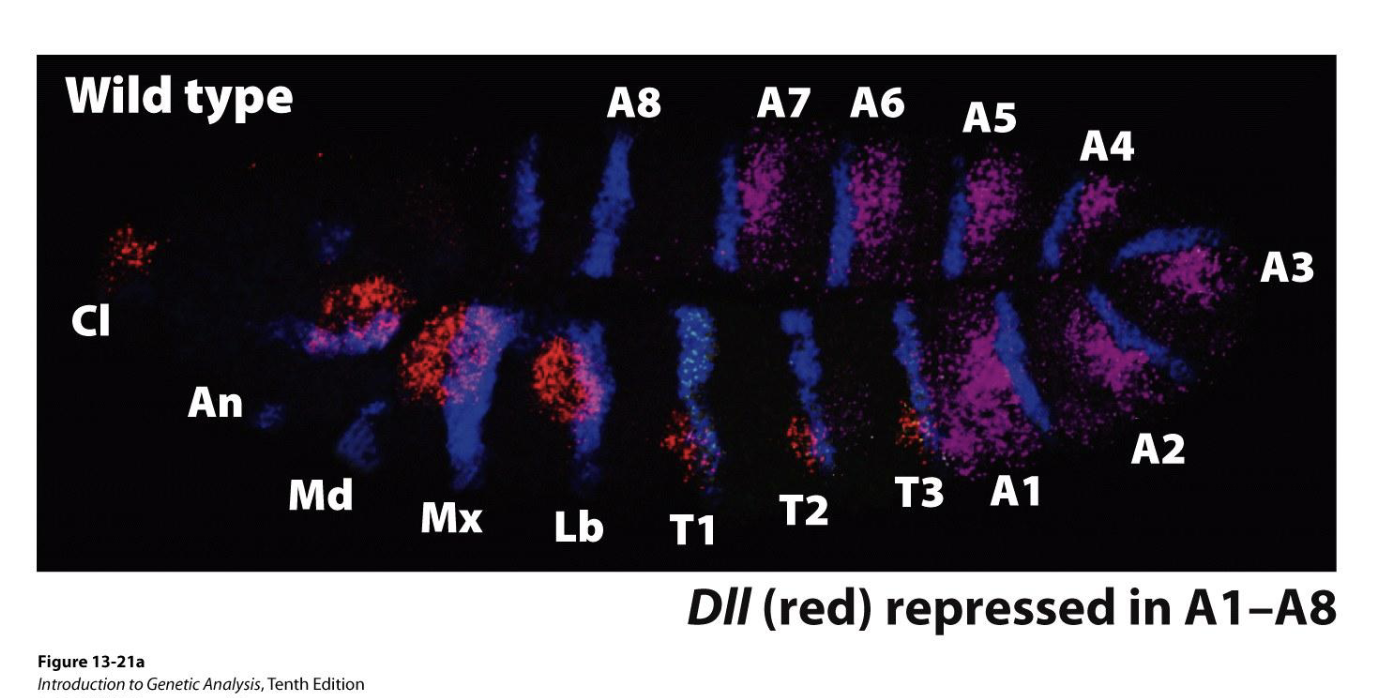
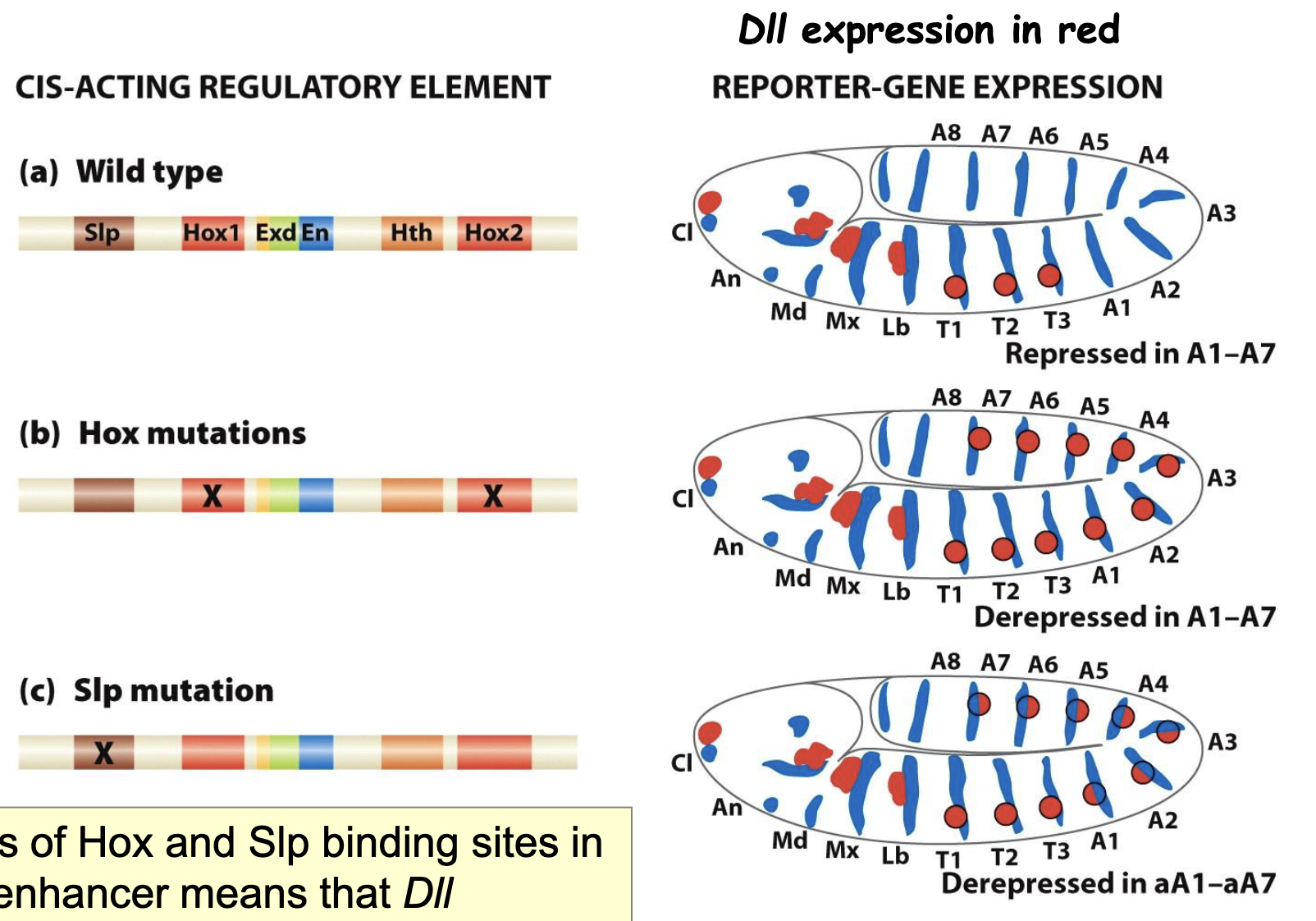
Regulation of DII by multiple Hox and segmentation proteins
Cis-acting regulatory elements: binding sites for the different TFs that regulate DII expression
Hox mutations
Slp mutations
loss of Hox and Slp binding sites in DII enhancer means that DII expresseion cannot be suppressed in segments A1-A7
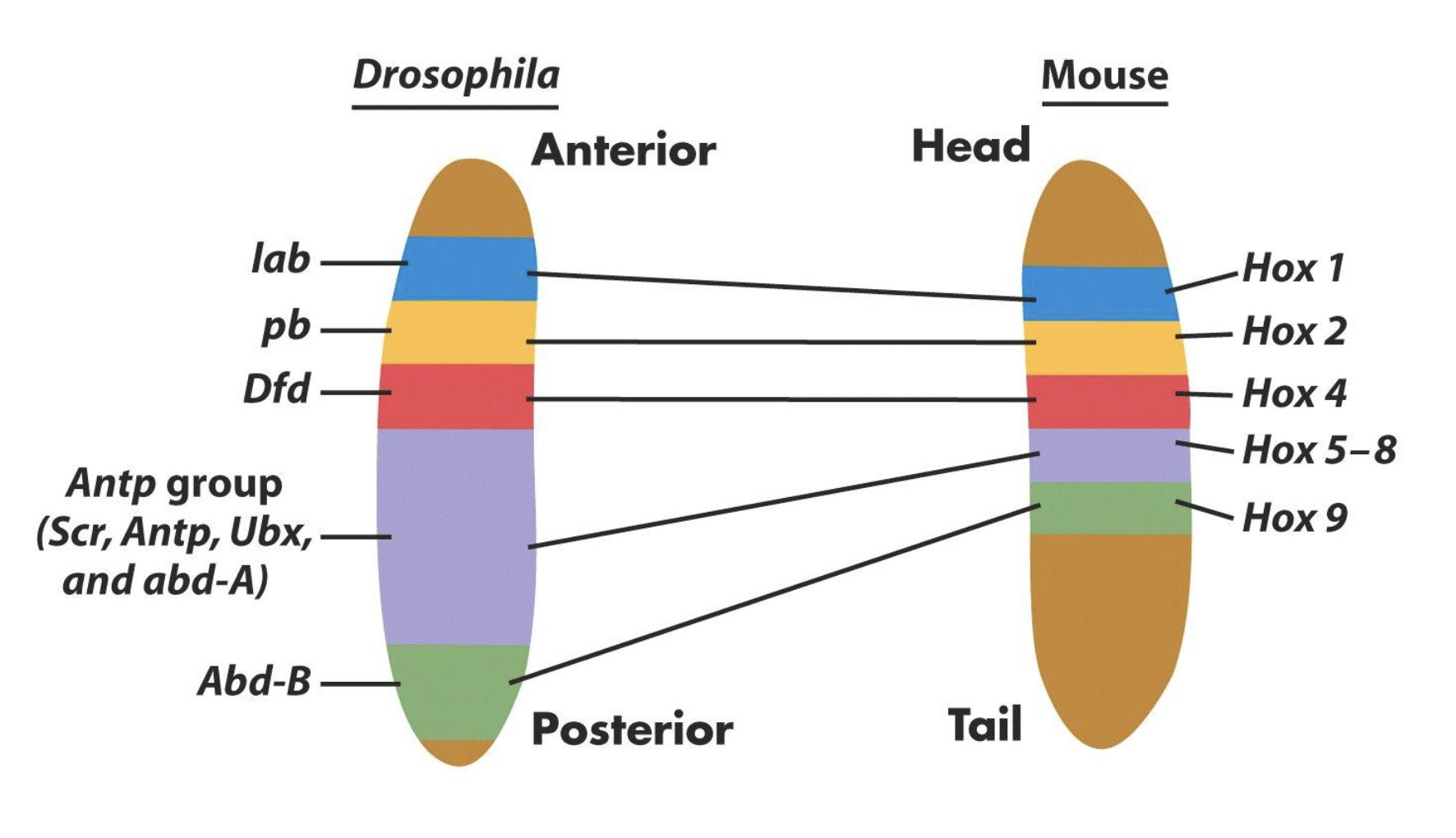
Similarities b/w Drosophila and Vertebrates
Drosophila and vertebrate Hox protein show striking similarities (sequence similarity)
Hox genes regulate segment identities in both Drosophila and vertebrates

Hox genes regulate the identity of serially repeated structures in vertebrates
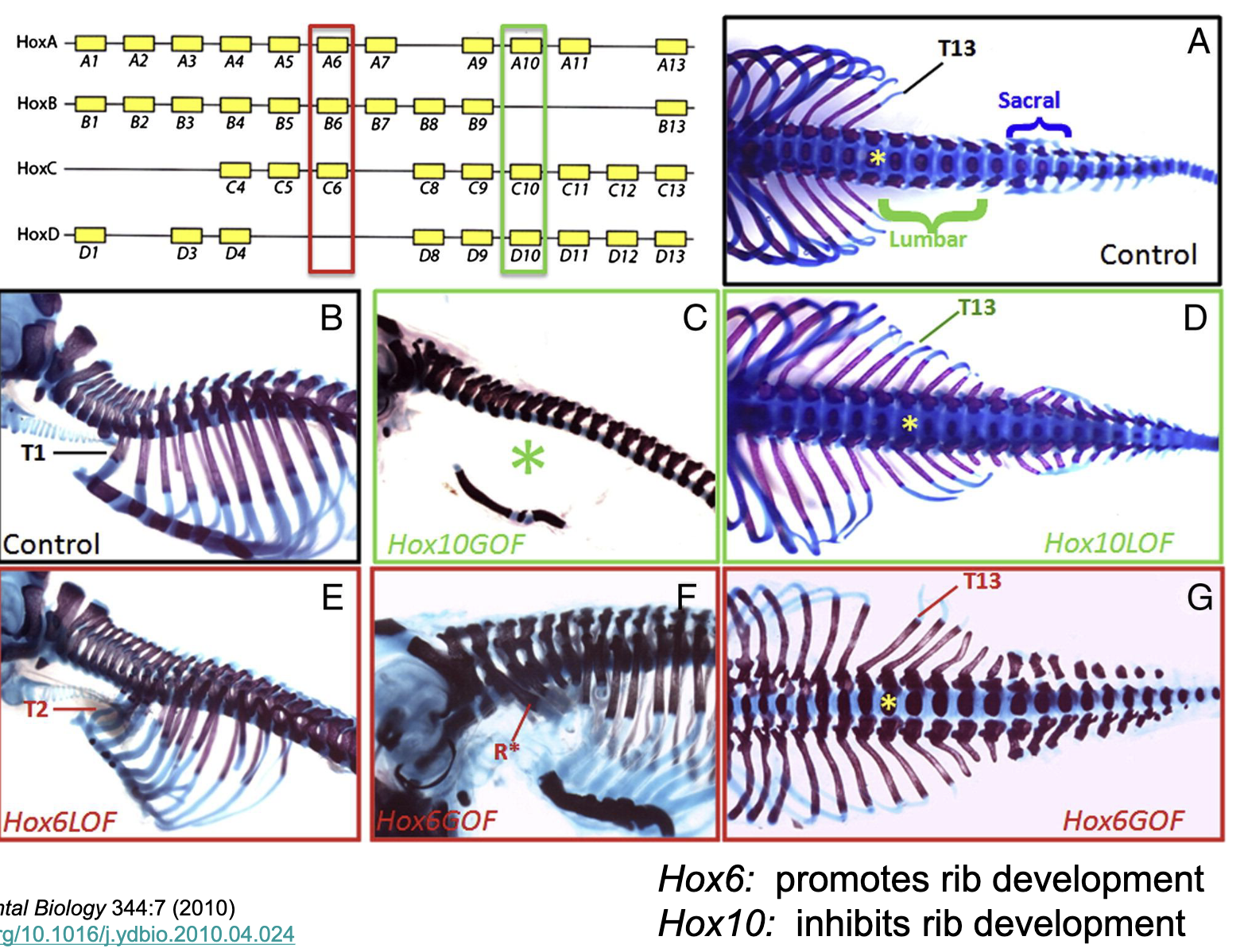
Control of TF in time and space
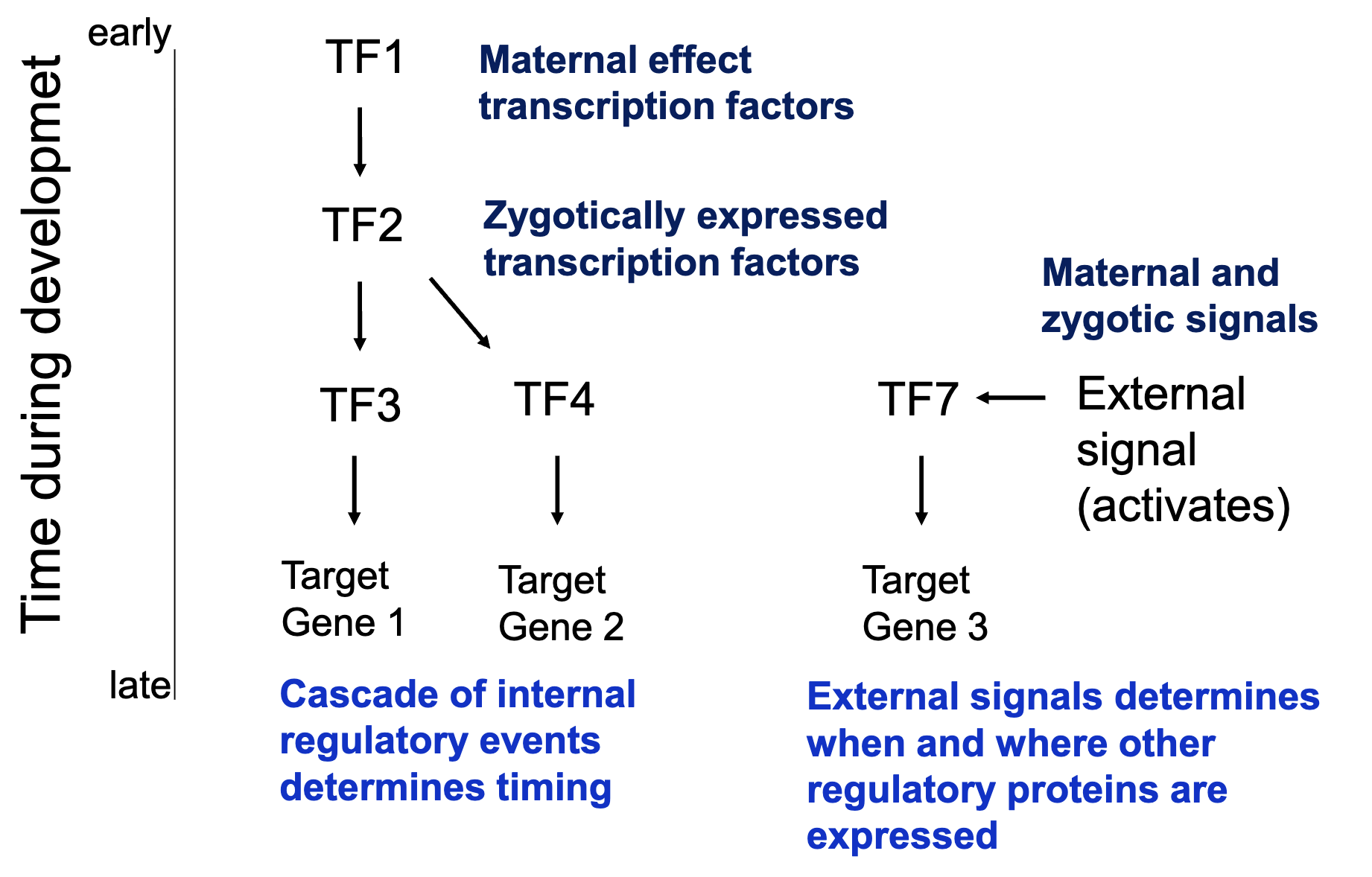
Control of patterning in time and space
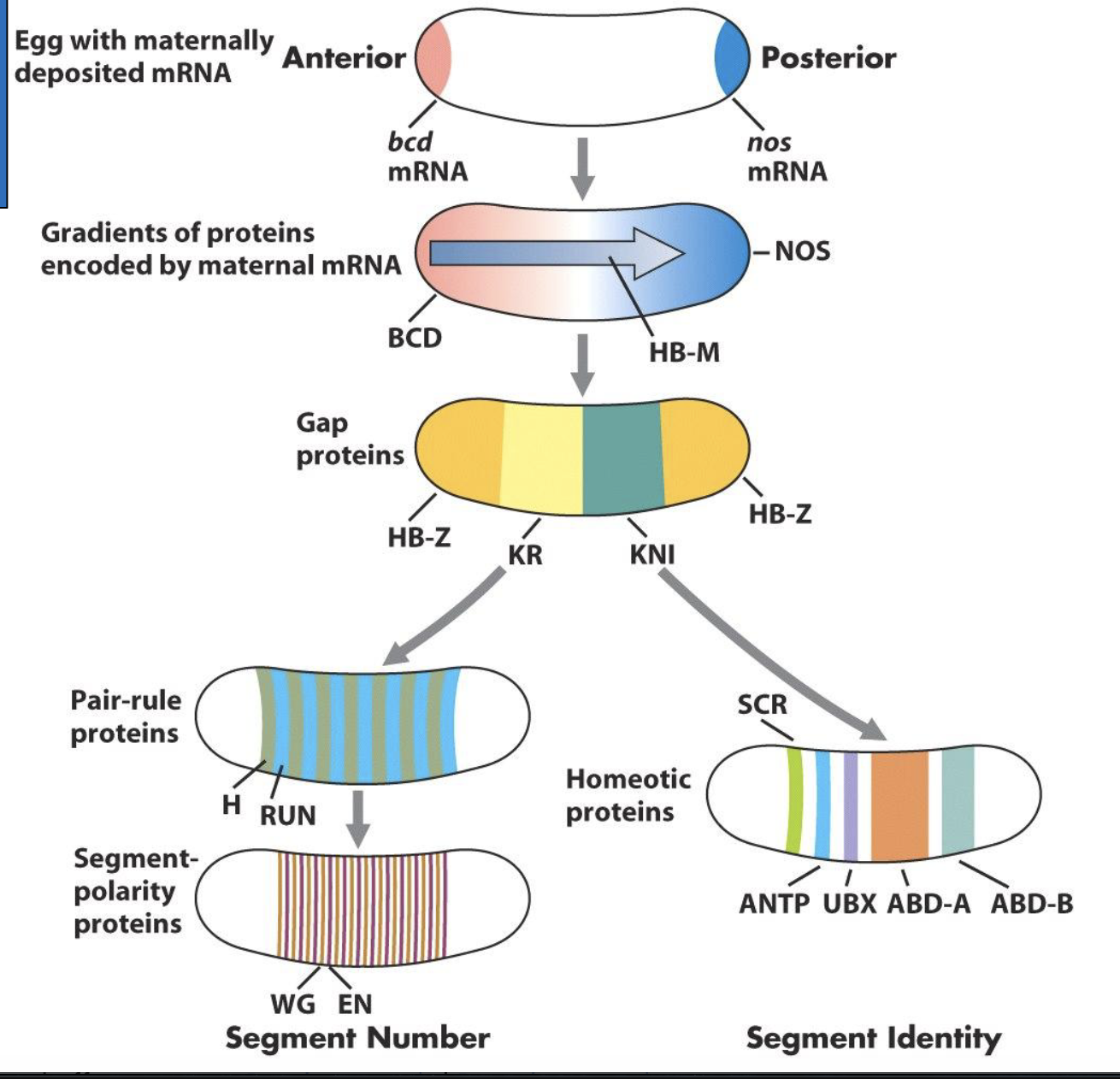
Summary!
gap gene proteins are TFs that act as morphogens, activating and/or repressing expression of pair-rule and homeotic selector genes in diff cells of the cellular blastoderm
pair-rule genes encode TFs and are expressed in a pattern of 7 stripes. The role of the pair-rule genes is to determine the position and number of segments in the Drosophila embryo
the homeotic selector genes are first transcribed in broad overlapping domains of contiguous cells in the cellular blastoderm and function tgt to specify the identity of each segment
What have we learned concerning pattern formation?
asymmetry in the female is used to generate asymmetry in the oocyte
asymmetry in the oocyte is used to generate maternal morphogens in the early embryo
maternal morphogens establish a specific pattern of zygotically expressed morphogens
zygotic morphogens activate a complex set of genes required to determine the differential fates of cells in the blastoderm thus establishing a spatial pattern of morphological structures
Principles of pattern formation
must establish positional information (spatial information) within the developing embryo (morphogen? environmental signals?)
direct simple patterns (anterior–posterior, dorsal–ventral) early and use to elaborate more complex patterns
use a cascade of determination events to coordinate timing
use master regulatory proteins each of which will be expressed in specific groups of cells marking them for a particular fate