BIO130
1/136
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
137 Terms
Genome
entirety of an organism’s genetic info (all the DNA in a cell)
sequencing a genome
determining the order of all the bases in the DNA
human genome has ~__ base pairs per genome
3 billion
in a standard cell, you have _ genomes
2 (one from each parent)
genome size is or isn’t always correlated with complexity?
isn’t
_% of your genome is repetitive DNA
50
less than _% of your genome encodes protein
1
nonrepetitive DNA (~50%)
includes sequences that tell the cell how much RNA to make and what to transcribe
nonrepetitive DNA includes __ and __
introns (transcribed then spliced out) and exons (transcribed and translated)
mobile genetic elements
sequences that are sometimes spliced out of the DNA, sometimes copied, sometimes pasted back in
LINEs
Long Interspersed Nuclear Elements
A type of mobile genetic element
>= 500 base pairs
SINEs
Short Interspersed Nuclear Elements
noncoding
Type of mobile genetic element
< 500 base pairs
retrotransposons
transcribed (made into RNA)
DNA-only transposons
not transcribed into RNA
Genome occupies a large portion of the cell volume
false
DNA packaging in prokaryotes
have no nucleus
but DNA is in compact structure called the nucleoid
Chromosome Painting Hybridization
a technique that uses many fluorescent probes, one for each chromosome, so you can visualize which are present in a sample. (FISH with a lot of probes)
Fluorescence in Situ Hybridization (FISH)
A diagnostic technique for testing the presence of a particular sequence
probe DNA binds to the sequence you want to detect (bc it’s antiparallel complementary)
use fluorscent dye that attaches to the probe
separate sample DNA strands so that they’re available for the probes to attach
once it cools back down look at the sample. If fluorescence, then probe sequence had been able to bond to the target sequence confirming the target’s presence in the sample.
__ pairs of chromosomes in humans
23
chromatin
single, long, linear DNA molecule and associated proteins
“beads on a string”
chromatid vs chromosome
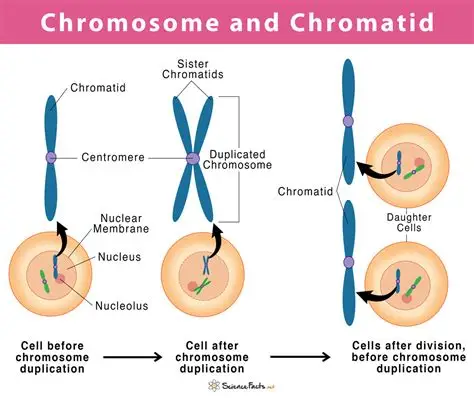
30 nm fiber
name of the structure where chromatin fiber is packed into nucleosomes
histones
small proteins rich in lysine and guanine
have positive charges that neutralize DNA’s negative charge
is an octamer core with a pair of each 4 core histone proteins (H2A, H2B, H3, H4)
there’s also H1, the linker histone
What does H1, the linker histone do?
clips the DNA onto the histone octamer
nucleosome core particle vs nucleosome
nucleosome core particle = core histone + DNA around it
nucleosome = nucleosome core particle + H1 + linker DNA
sequence-specific clamp proteins and cohesins
involved in forming chromatin loops
condensins
as cells enter mitosis, condensins replace most cohesins to form double loops of chromatin to generate compact chromosome
chromatin remodeling complexes/histone modifying enzymes
proteins that can make changes in chromatin structure and alter access to DNA for replication or transcription
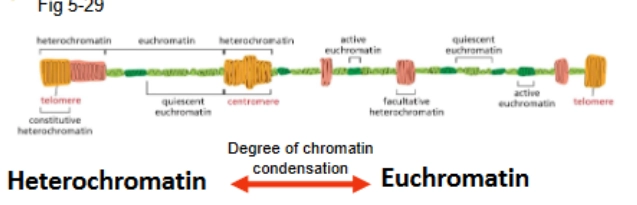
heterochromatin vs euchromatin
h: highly condensed → gene expression suppresses
meiotic/mitotic chromosomes
centromeres and telomeres
time spent highly condensed varies (constitutive vs facultative (more temporary))
e: relatively non-condensed → genes tend to be expressed
degree of condensation varies
level of activity varies (quiescent vs active)
active = doing something w the DNA
DNA synthesis is always __conservative
semi
DNA polymerase
catalyzes DNA synthesis (adding on base pairs to template strand)
3 rules for DNA synthesis:
DNA is antiparallel and complimentary
new DNA is synthesized from 5’ - 3’
the template is read 3’ - 5’
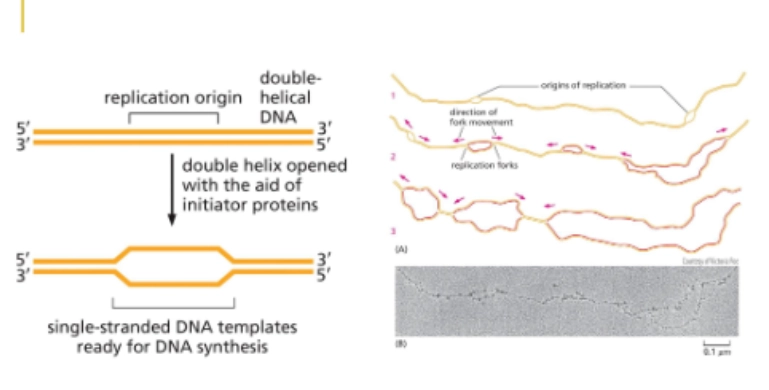
origin of replication
starting point for bidirectional growth
easy to open, A-T rich (only 2 H bonds)
recognized by initiator proteins that bind to the DNA
Bacteria have 1, Eukaryotes have multiple
prokaryote vs eukaryotic cell
P:
no nuclei
single-celled
no membrane-bound organelles
smaller
less DNA
(bacteria and archaea)
E:
nuclei
single OR multi-celled
membrane-bound organelles
larger
more complex
(plants, fungi, animals)
mitochondria
uses oxygen to generate ATP through aerobic respiration
endosymbiont, started as ectosymbiont
3 lines of evidence that supports endosymbiont hypothesis for origin of mitochondria and chloroplasts
they have their own cell walls that resemble that of prokaryotes
they have remnants their own genomes and genetic systems that resemble that of prokaryotes
they have their own protein and DNA synthesis components
general attributes of a model organism
readily available
tractability - ease of maniipulation/modification
we understand its genetics
rapid development with short life cycles
small adult (reproductive) size
(like a rat!)
3 types of RNA that eukaryotes and prokaryotes have in common
Messenger RNA (mRNA) - blueprint
Transfer RNA (tRNA) - brings amino acids
Ribosomal RNA (rRNA) - structural for ribosome, break + form proteins
nucleic acids
genetic material in a cell
DNA = deoxyribonucleic acid
RNA = ribonucleic acid
made up of nucleotides
nucleotide structure
phosphate group (backbone) - attached to 5’ C of sugar
nitrogenous base - attached to 3’ C of sugar
pentose sugar (scaffold for base)
two types of bases
pyrimidines - “U C The pyramids” → Uracil, Cystine, Thymine
purines - adenine, guanine
(side chains make bases different from each other)
H-bond together when base-pairing
nucleoside vs nucleotide
nucleotide if has at least one P, nucleoside if not (just base + sugar)
3 forces that keep DNA strands together
Hydrogen bonding between bases
hydrophobic interactions (bases want to get away from water, press in closer)
can der Waals attractions (molecules closely packed)
structure of amino acid
carboxyl group, R-group, amine group, hydrogen → connected to central C
initiator proteins
binds to replication origin to help DNA helicase bind (it destabilizes the AT-rich sequence)
helicase-loading protein
helps helicase load onto the DNA then leaves
helicase
unwinds and separates strands
predominant helicase moves 5’-3’ along the lagging strand template
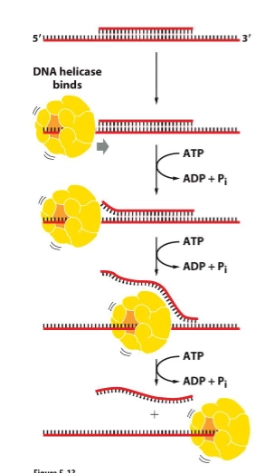
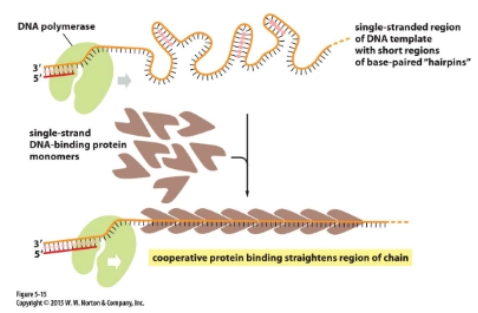
single-strand binding proteins
keep DNA strands separated after they are separated by helicase
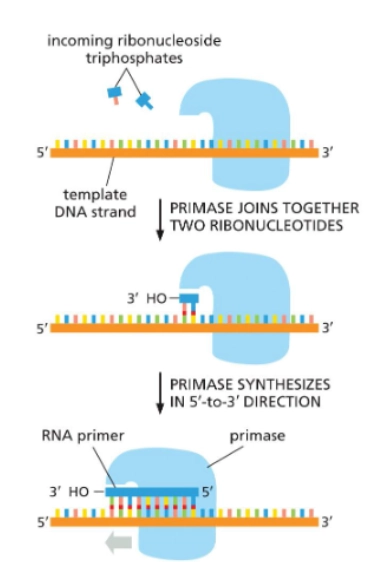
RNA Primers and DNA Primase
Primase makes primers, which is a short sequence of nucleotides with a free 3’OH. DNA polymerase requires a “bound primer” to begin adding nucleotides to the template strand.
Primase proceeds in the 3’→5’ direction
Primosome
DNA Polymerase + DNA Primase
Sliding clamp
holds polymerase onto the DNA so it doesn’t float away
DNA ligase
removes RNA primer and replaces it with a DNA sequence
is a “DNA repair system”
This is important for linking the Okazaki fragments together
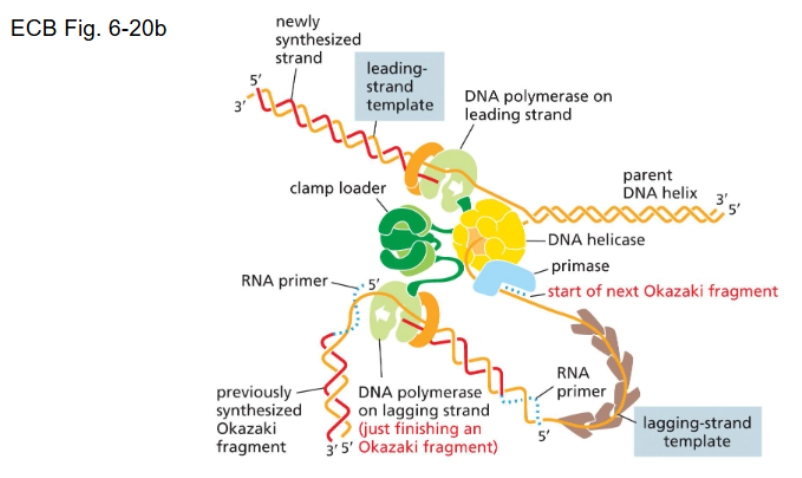
Replisome
A type of “molecular machine” - all the proteins that work together to replicate DNA
okay this is a hard one…recount the process of DNA replication
initiator protein → helicase-loading protein → helicase unwinds strands (single-strand binding proteins needed to keep them separated) → DNA polymerase + DNA primase = primosome (sliding clamp needed to keep polymerase from floating away) → leading strand and lagging strand synthesis are simultaneous (2 replication forks moving in opposite directions)
topoisomerase
solves the problem of supercoils that form when helicase unwinds DNA. It makes a cut to release tensions then seals the cut.
What is the issue in DNA replication concerning the ends of linear chromosomes?
The lagging strand would miss information (if not for the solution that exists) because primase isn’t good at putting a primer at the very end, and once the 3’-most primer gets removed there’s no 5’ end to add onto—missing DNA!
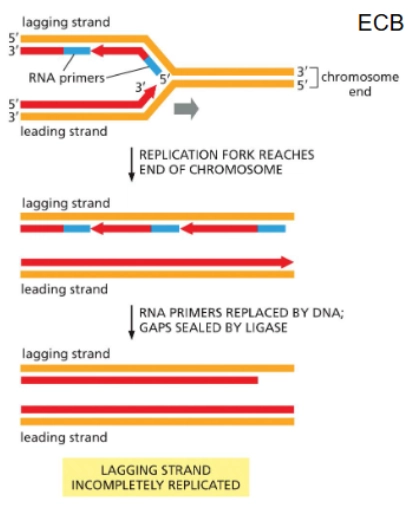
Telomerase
A solution for the lagging strand’s problem with missing DNA at its ends.
Telomerase uses an RNA template to make a DNA complementary copy
it adds a repetitive sequence to the 3’ end of the parental strand (lagging strand template) → it’s so long now that primase CAN be placed and polymerase can come in and fill the gap we had earlier
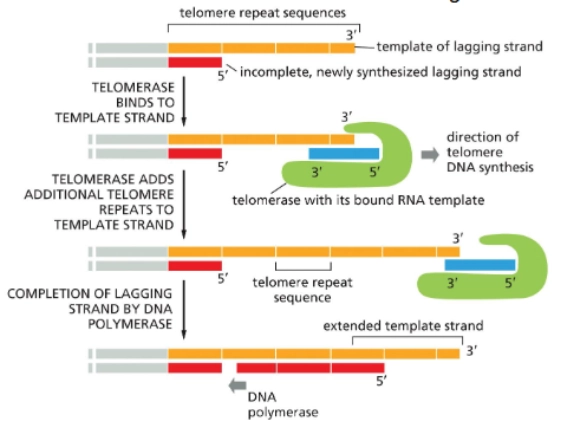
Telomerase is not abundant in __ cells
somatic (connects to aging!)
connect telomerase to aging/cancer
Telomerase is what adds telomeres to the ends of linear chromosomes. Telomerase is abundant in stem and germ-line cells, but NOT in somatic cells. So, loss of telomeres occurs normally during DNA replication, and limits the number of round of cell division.
on the other hand, cancer tells have high levels of telomerase and so replicate abundantly.
Fill in the blanks with numbers:
RNA polymerases typically have an error rate of about 1 in __. DNA polymerases are only about 1 in __. The human genome (3×10^9) is only changed by about _ nucleotides every time a cell divides!
10^4, 10^9, 3
What are the two mechanisms in place that ensure accurate DNA synthesis?
3’ to 5’ exonuclease
strand-directed mismatch repair
3’-5’ exonuclease
is on the DNA polymerase’s exonuclease (“editing”) site. Removes incorrectly placed nucleotides. “Proofreads” DNA.
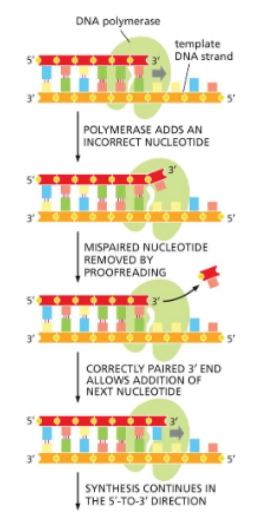
strand-directed mismatch repair (in eukaryotes)
A DNA replication error repair process (for if proofreading fails). It is initiated by the detection of the distortion in geometry of the double helix generated by mismatched base pairs
(for prokaryotes: detect unmethylated adenines, because parental strand have them but not new strands)
DNA can be damaged by:
oxidation
radiation
heat
chemicals
and more
Pyrimidine Dimers
when UV radiation causes covalent bonds when they’re not supposed to between adjacent bases
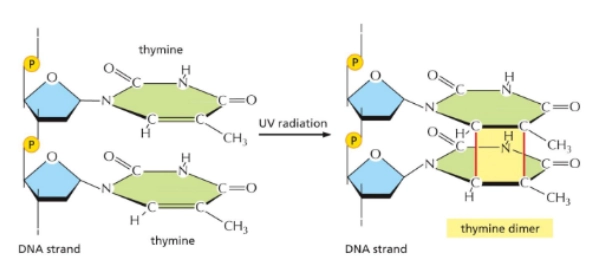
Depurination
a base is lost spontaneously because a water molecule hit the DNA strand at the wrong time/energy/space/spot
Deamination
An amine is spontaneously bumped (by a water molecule) from the cytosine and it becomes a uracil instead
The two general mechanisms of DNA repair of incorrect DNA:
Base Excision Repair (BER)
Nucleotide Excision Repair (NER)
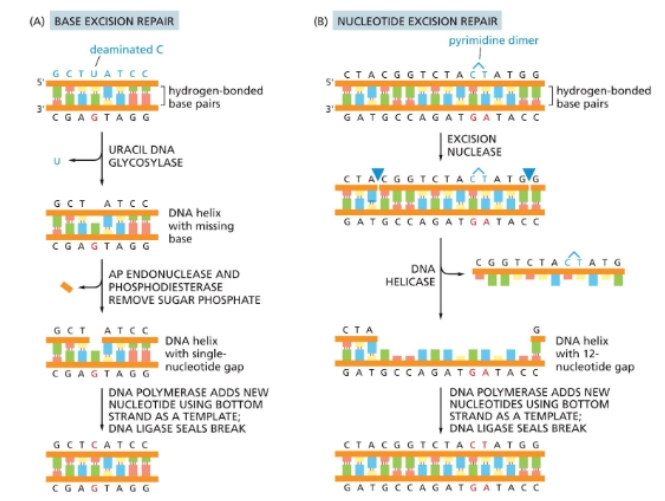
Base excision repair (BER)
Fixes one nucleotide at a time. Following the example of the image:
Uracil DNA glycosylase comes in a removes the incorrectly placed uracil
two enzymes (AP endonuclease and phospodiesterase) come in and remove the sugar phosphate backbone
so then polymerase can come in and repair
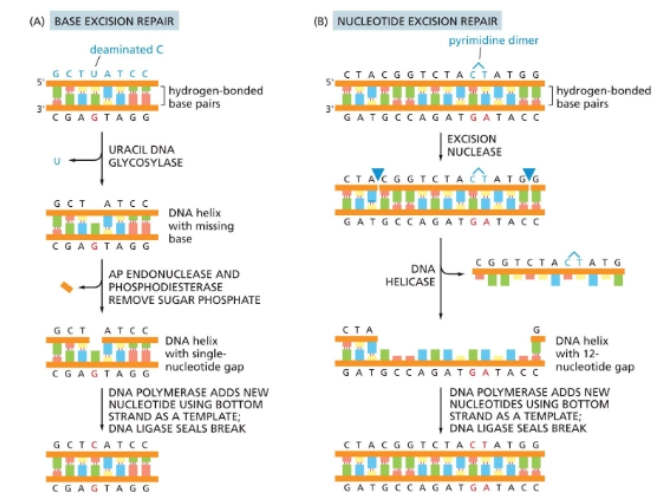
Nucleotide Excision Repair
Fixes a couple incorrectly placed nucleotides at a time.
excision nuclease comes in, makes two cuts
a helicase removes a strand
polymerase comes in and fixes the strand
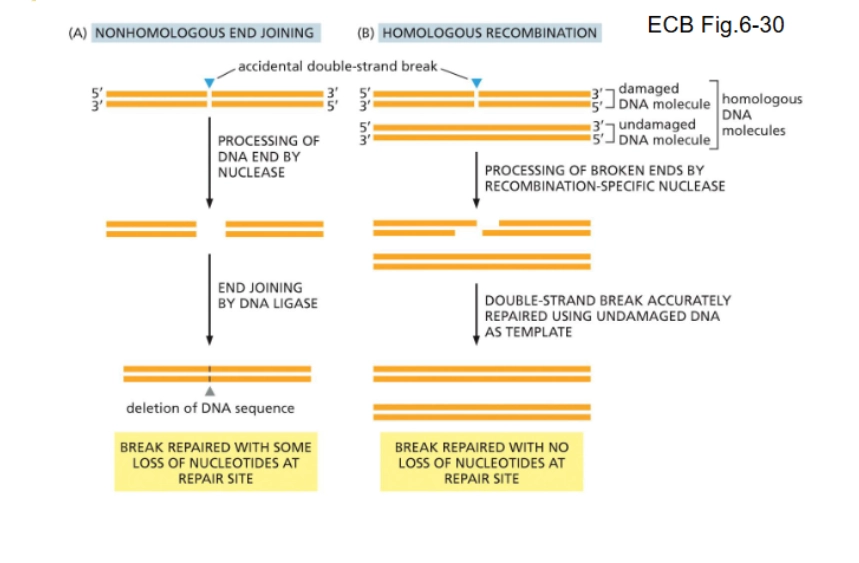
The two methods of DNA repair of double-stranded breaks:
Nonhomologous end joining (NHJ)
Homologous recombination (HR)
Nonhomologous end joining (NHJ)
A method of repair for double stranded breaks in DNA. Nuclease trims the strand then joins it together with a special DNA ligase. “quick and dirty”, used in emergency in cell division to make sure daughter cells have the right number of chromosomes.
Homologous recombination (HR)
The more accurate method (but takes longer) of DNA repair of double stranded breaks. Uses info from the homologous chromosome.
Gene
The entire nucleic acid sequence necessary for the synthesis of a protieins or RNA. “segments of DNA that are transcribed into RNA”
ssDNA
single stranded DNA (the template for RNA)
phosphodiester bonds
bonds between nucleotides (in DNA and RNA)
sigma factor
A protein required in transcription for bacteria. It binds to RNA Polymerase core enzyme, searches for the promoter sequence on the DNA and binds to it. Once transcription begins, it’s released.
RNAP Holoenzyme
RNAP core enzyme + sigma factor
Promoter
The sequence of DNA where transcription starts for bacteria
where sigma factor binds
commonly at ~ -10 and ~ -35
Describe how terminator sequences work in bacteria
the end of a gene contains a lot of Gs and Cs, followed by As and Ts. The strong base pairing of the Gs and Cs gets in the way of the RNA Polymerase (“hairpin” structure).
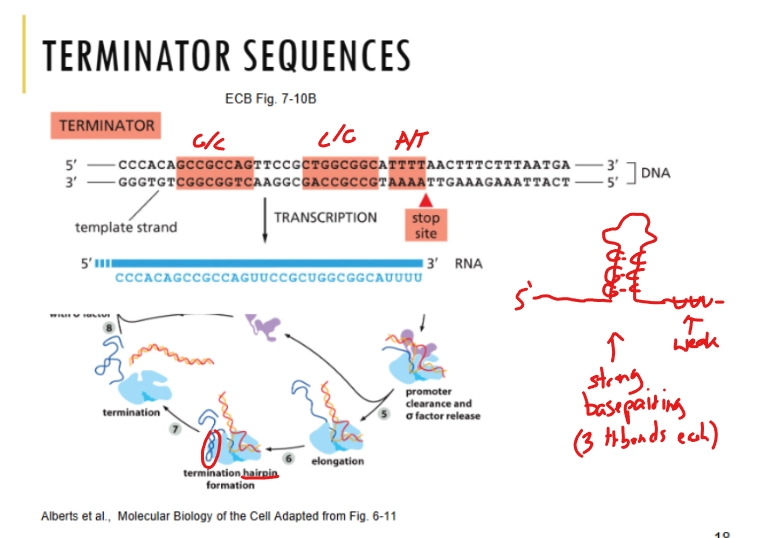
primary or pre- mRNA transcript
the full mRNA transcript, including introns. but doesn’t have a 5’ cap or 3’ poly-A tail yet.
UTR
Untranslated regions (at the end of mRNA). They do not code for proteins.
translation and transcription are coupled in bacteria or eukaryotes?
bacteria
What is the start codon in mRNA for eukaryotic translation?
AUG
name what the three different RNA polymerases are mainly used for transcribing
RNAP I: most rRNA genes
RNAP II: all protein-coding genes
RNAP III: tRNA genes
carboxyl terminal domain tail (CDT)
special tail of the largest subunit of RNAP II. Tandem repeats of 7 amino acids. Allows transcription and RNA processing to be coupled by undergoing dynamic phosphorylation and recruiting proteins involved with capping, polyadenylation, and splicing.
transcription factors
Proteins required by RNAP to help position them at the promoter (similar to the sigma factor in bacteria)
TATA box
A common element (promoter sequence), bound ~ 30 bps upstream from the start site for transcription. It helps position RNAP II and general transcription factors.
TFIIH
A transcription factor that helps separate DNA strands and phosphorylates CTD
(can remember that “H” is for “Helicase”)
The 4 steps in eukaryotic transcription initiation (this is pretty detailed, but something we have to know)
TBP, a subunit of TFIID, binds to the TATA box promoter in the minor groove, bending and distorting the DNA
this attracts other transcription factors, which help orient and bind RNAP II to the DNA
the helicase activity of TFIIH uses ATP to pry apart DNA strands of transcription start site
TFIIH also phosphorylates the CTD of RNAP II, activating it so that transcription can begin
exonucleases
proteins that chop off nucleotides one at a time from nucleic acids
protected against by poly-A tail and 5’ capping
describe how a intron is removed
Branch point A attacks the 5’ splice site
3’ end of one exon connects with 5’ end of the other
exon junction complex added (like glue)
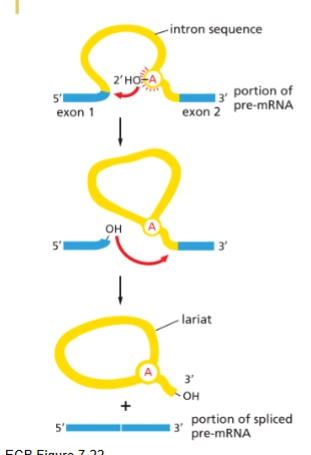
spliceosome
enzyme that catalyzes splicing
How it works:
takes the 2’ oxygen off the ribose and forms a phosphodiester bond with the 5’ end of the intron
that frees up a 3’ OH at the end of exon 1, and so a bond is formed with the 5’ end of exon 2
poly-A binding protein
Proteins added to the poly-A tail that make it hard for 3’ exonucleases to chop of the Adenines.
poly-A polymerase
a protein involved in RNA processing that adds an poly-A tail to mRNA
does NOT bind to CTD
does NOT require a template
snRNPs
proteins that make up the spliceosome. Recognizes splice-sites of mRNA, helps create the lariat. Made of snRNAs bound to a protein.
alternative splicing
pre-mRNAs can be spliced in multiple ways to create different versions of a protein from one gene (eukaryotes)
redundancy
most amino acids are encoded by multiple codons
3 types of substitution mutations:
silent: codon still refers to same amino acid
missense: codon refers to different amino acid
nonsense: codon changed to a premature STOP codon
other than substitution, what are the other two types of mutations?
1 nucleotide-pair insertion or deletion
changes reading frame (really bad!)
3 nucleotide-pair deletion
causes a missing amino acid