A Level CIE Biology: 19 Genetic Technology
1/107
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
108 Terms
what is meant by genetic code being universal
almost every org using the same four nitrogenous bases - A, T, C, G with a few exceeptions. same codons code for the same aas in all living orgs meaning genetic info is transferable. scientists can aritificially change orgs dna by combining lengths of nucleotides = recombinant dna.
transgenic org
org w nucleotide seq from other species
gmo
any org that has introduced genetic material is a genetically modified organism
genetic engineering
technique used delib to modify specific characteristic(s) of an org by removing gene(s) w the desired characteristic from one org and transferring the gene with a vector into another, meaning there will be recombinant DNA and a GMO
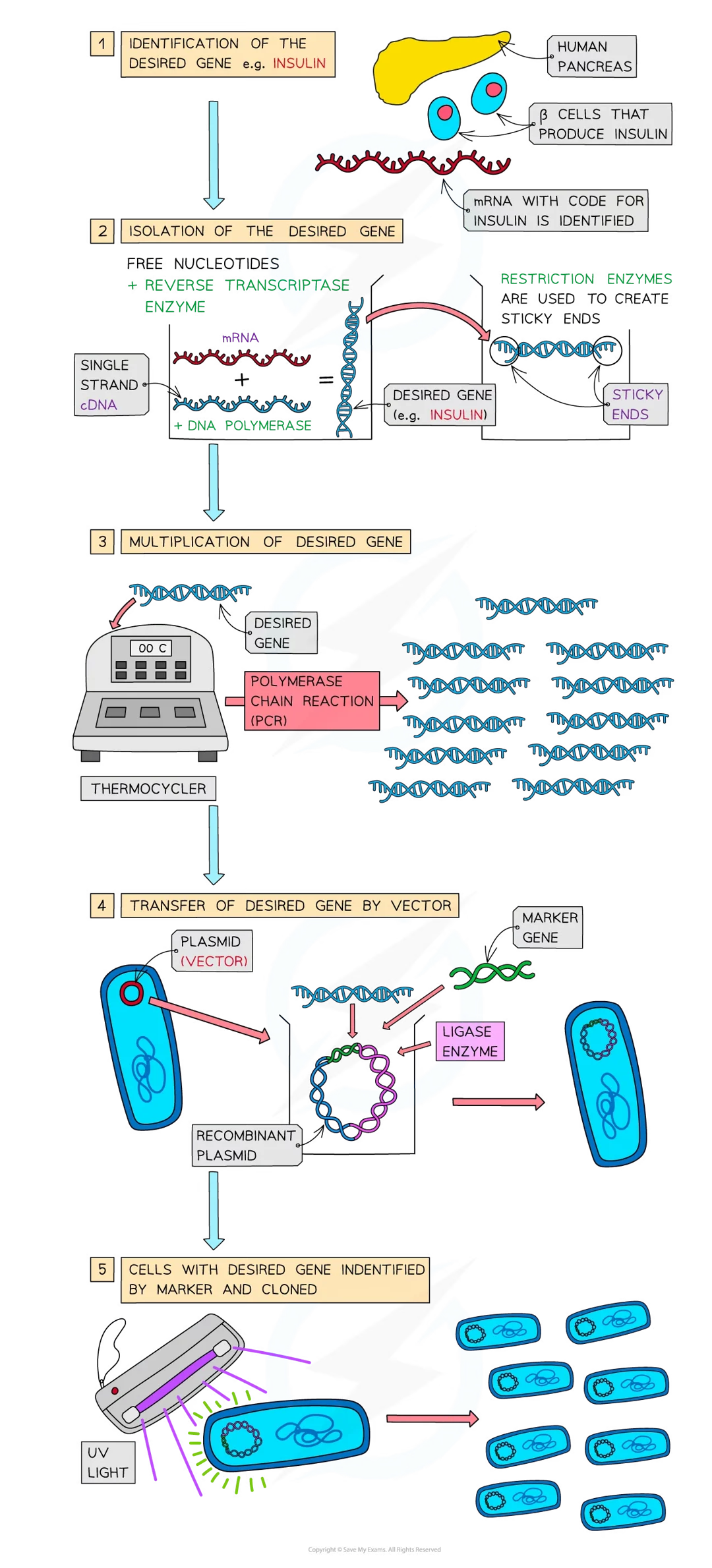
steps to genetic engineering: 5 (3)
identification of the desired gene
isolation of the desired gene by:
cutting from a chromosomes using enzymes (restriction endonucleases)
using reverse transcriptase to make a single strand of complementary DNA (cDNA) from mRNA
creating the gene artificially using nucleotides
multiplication of the gene (using polymerase chain reaction - PCR)
transfer into the organism using a vector (e.g. plasmids, viruses, liposomes)
identification of the cells with the new gene (by using a marker), which is then cloned.
what do genetic engineers need to modify organism 3
enzymes
vectors
markers
enz involved in genetic engineering
restriction endonucleases
ligase
reverse transcriptase
vectors
to deliver genes into cell e.g. plasmids, viruses, liposomes
markers
genes that code for identifiable substances that can be tracked (e.g. GFP - a green fluorescent protein which fluoresces under UV light or GUS - beta-glucuronidase enzyme which tarnasofrms colourless or non-fluorescent substrates into products that are coloured or fluorescent.)
synthetic biology
the field that ge is in in science, it designs construction of diff bio pathways, orgs, and devices as well as redesignign of exising natural bio systems.
what 3 ways can genes w specific characteristics be obtained
extracting the gene from the dna of a donor organism using enzymes (restriction endonucleases)
using reverse transcriptase to synthesise a single strand of complementary DNA (cDNA) from the mRNA of a donor organism
synthesising the gene artificially using nucleotides
what is required for extraction of genes
restriction endonucleases
what are restriction endonucleases
a class of enzymes found in bacteria that are used as a defence mechanism by bacteria agaisnt bacteriophages (viruses that infect bacteria, also known simply as phages)
what do restriction endonucleases do
restrict a viral infection by cutting the viral genetic material into smaller pieces at specific nucleotide sequences within the molecule (endo = within)
why are there many diff restriction endonucleases/enzymes
because each one binds to a specific restriction site (sepcific sequences of bases) on DNA e.g. Hindlll will always bind to AAGCTT
how are restriction enzymes named
according to bacteria they’re from and which numbered enz its from the source (e.g. Hindllll comes from Haemophilus influenzae and it is the third enz from that bac)
what do restriction endonucleases do to DNA
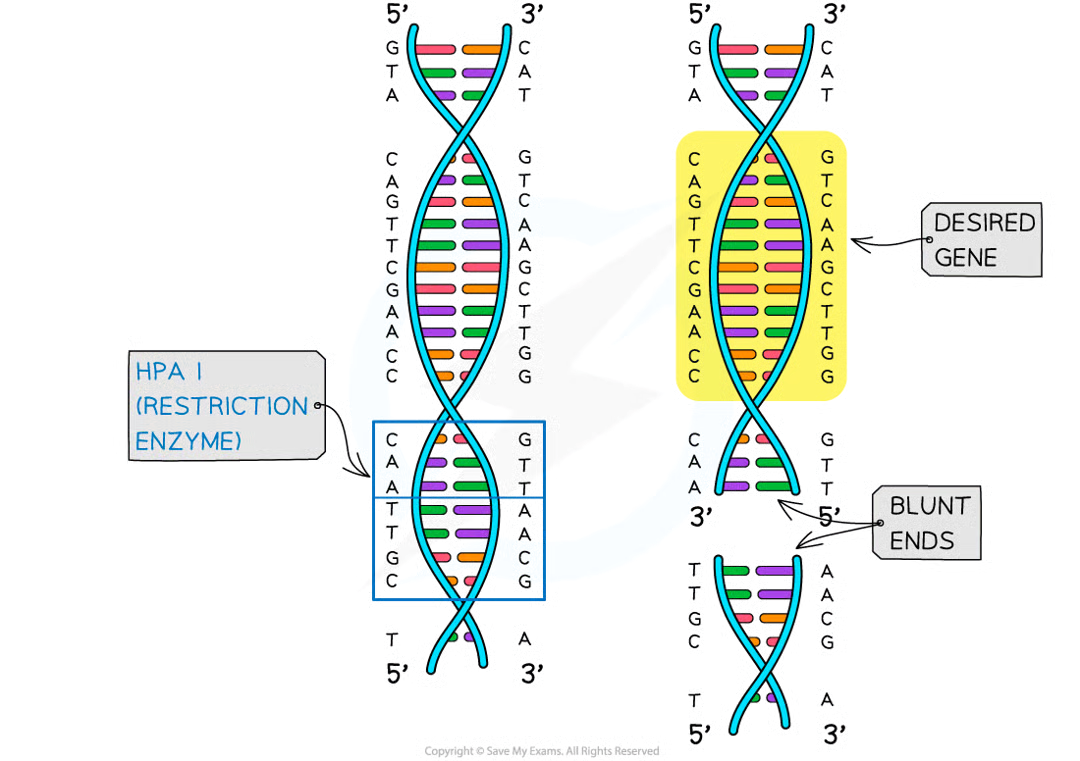
separate the two strands of dna at specific base sequence by cutting the sugar-phosphate backbone in an uneven way to give sticky ends or straight across to give blunt ends
sticky ends result in..
one strand of DNa fragment being longer than the other
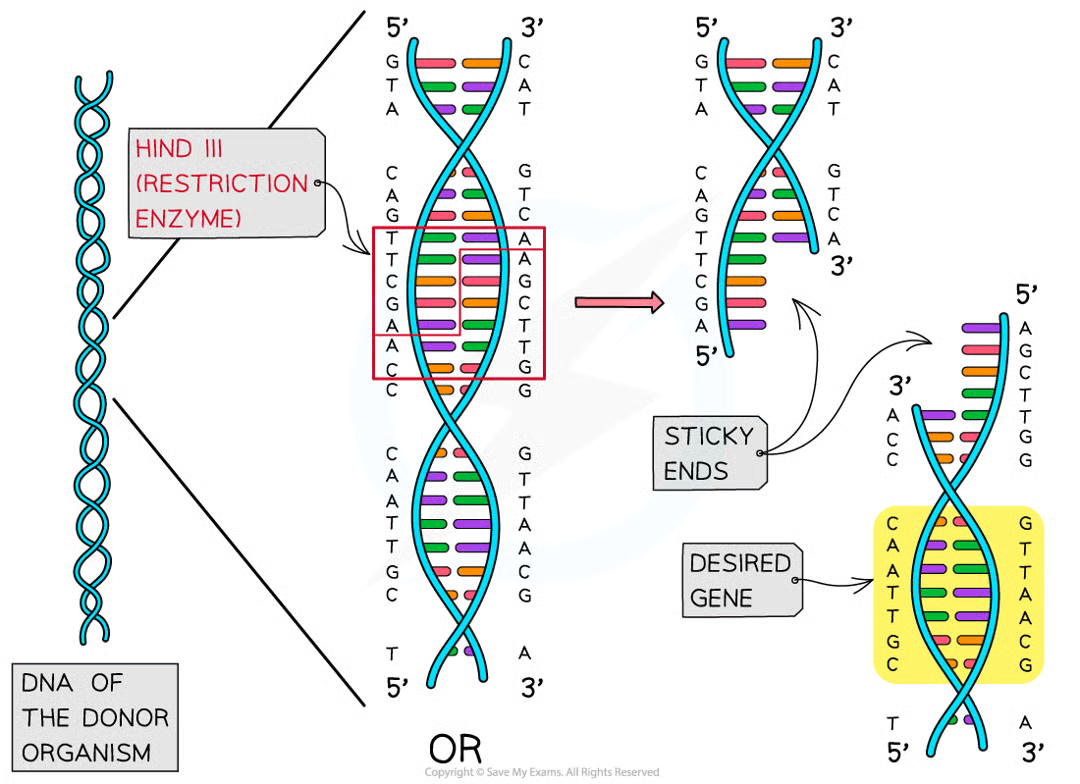
why are sticky ends useful
make it easier to insert desired gene into another organisms DNA as they can easily form H bonds w complementary base sequences on other pieces of DNA that have been cut with the same restriction enz
what if restriction enzymes give blunt ends
nucleotides can be added to make stickye nds
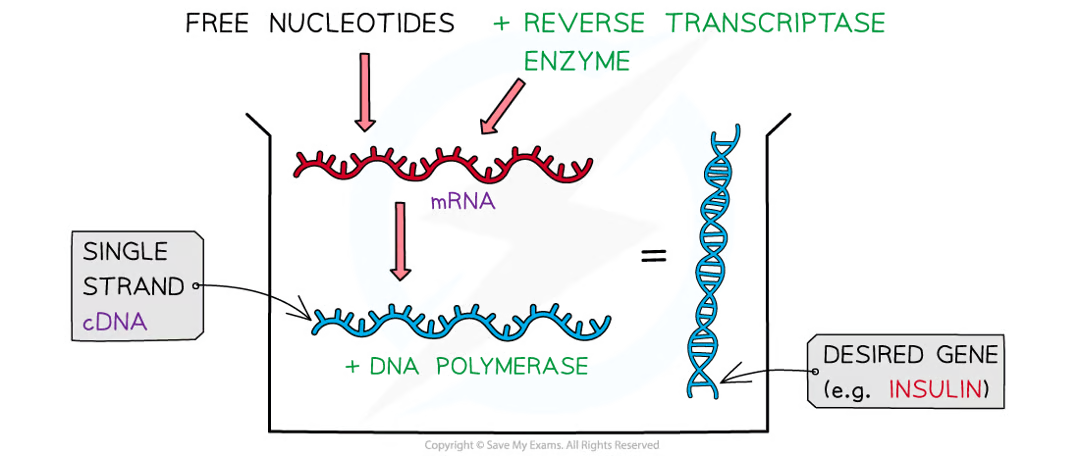
how are mRNA and reverse transcriptase used for isolating desired genes
mRna is combined w reverse transcriptase enzyme and nucleotides to create single strand of complementary DNA (cDNA)
where are reverse transciprtase enzymes sourced
retroviruses and they catalyse the reaction taht reverse transcription. mRNa used as template for cDNA
what does dna polymerase do
used to convert single strand of cDNA into double stranded DNa mol that contains desired code for gene
whys using mrna,r everse transcriptase and dna polymerase considered advantageouss
easier for scientists to find the gene bc of specialisied cells making very specific types of mRNA (e.g. beta cells of pancrease = insulin mRNA) and the mRNA (therefore cDNA) has no introns
how does artificial synthesis work
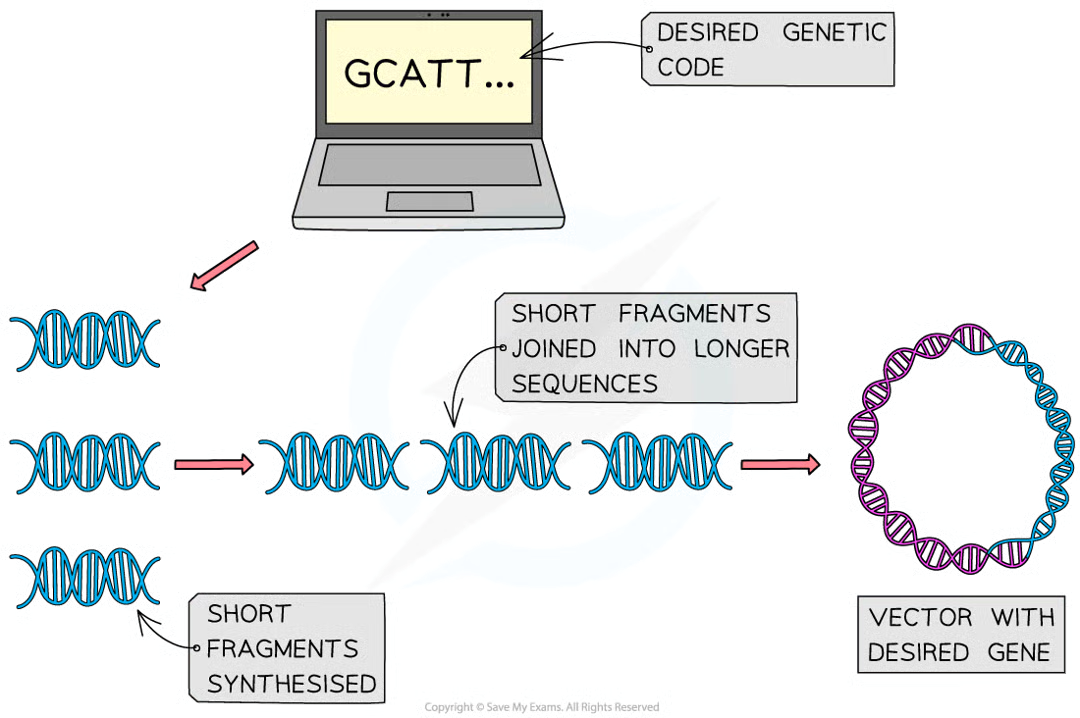
with the knwoledge of genetic code (aas required), scientists use computers to generate nucleotide sequence (rather than mRNA template) to produce the gene
short fragments of dna are first produce that are joined to make longer sequences of nucleotides thne inserted into vectors (plasmids)
artificial synthessis main use and advantage
used to create novel genes used to make vaccines and even to synthesise new bacterial genomes
method has adv that synthesised gene doesnt have to be cut out of existing genome and willl not contain introns or other unecessary lengths of non coding dna
4 enzymes reuqired in Genetic engineering
restriction endonucleases (enzymes) - cut dna strands so that desired gene can be isolated or spliced (inserted) into vector
reverse transcriptase - reverses transcription to produce single strand complementary DNA (cDNA) from an mRNA strand w the code for the desired gene
dna pollymerase - used to convert single-stranded cDNA into double stranded DNA mol of desired gene
DNA ligase - used to splice (insert) gene into vector
2 roles of restriction endonucleases in transfer of gene into org
isolate desired gene
separate dna strands at same base sequence in vector so desired gene can be inserted
how do restriction enzymes/endonucleases work
bind to specific restriction site (therefore many diff)
separate the 2 strands of dna at the specific base sequence by cutting the sugar-phosphate backbone in an uneven way to give sticky ends or straight across to give blunt ends
sticky ends result in 1 strand of dna fragment being longer than other
sticky ends make easier to isnert desired gene into another orgs dna or into vector as they can easily form H bonds w comp base sequences on other pieces of dna that have been cut w same restriction enz
reverse transcriptase role
produce single strand complemetary DNA mol (cDNA) that contains code for desired characteristic, this will then be inserted into vector (after being converted into double stranded dna mol)
dna polym role
convert single strand of cdna into double stranded dna mol which contains desired code for gene and builds second strand by pairing free nucleotides w the comp bases on cdna strand exactly like in normal dna rep
dna ligase role
catalyses formotion of phosphodiester bonds in dna sugar phosphate backbone and enables isolated desired gene to be spliced into vector (plasimd) so can be transferred to new org
what are vectors used for
transfer desired genes into a foreign cells. commonly plasmids but viruses and liposomes (small vesicle w phospholip layer) can
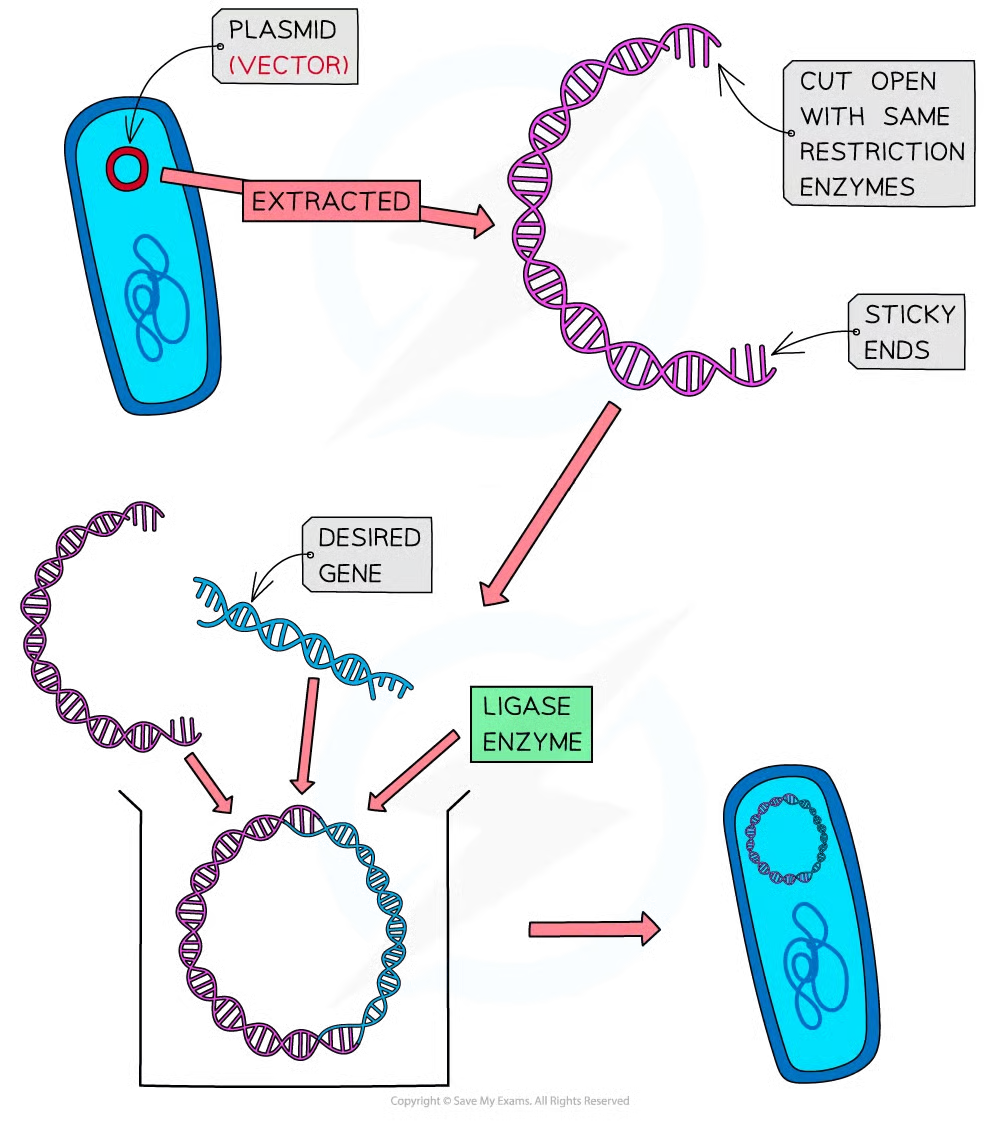
plasmid features 3
small circular rings of double stranded dna
occur naturally in abcteria, but can be found in archaea and euk orgs (yeast and fungi) and can contain genes for antibioitc res
can self replicate
how are plasmids used in transfering desired gene to new org 4
cricular dna of plasmid is cut open
same restriction endonuclease used to isolate desired gene is used to cut open plasmids
= plasmid complementary sticky ends to sticky ends on desired gene fragment
dna ligase forms phosphodiester bodns between sugar phosphate backbone of dna frag and plasmid to form recombinant plasmid (closed circle of double stranded dna containign desired gene
benefit of plasmids
scientists can modify bacterial plasmids or produce artificially. one benefit is that plasmids can have one or more marker genes so cells that have recomb plasmids can be identified
what happens after plasmids become recomb
transfered to host cells (bacteria often) thru transformation but only a small prop of bacteria will be transformed nad markers used to identiy transofrmed cells
3 ways transformation can occur
bathing plasmids and bacteria in ice cold calcium chloride solution and then briefly incubating at 40 degrees c
this makes bacterial memb permeable
electroporation - where bacteria are given small electrical shock, making membranes porous (technique aslo can be used to get dna frags intoe uk cells)
promotor
region of dna that determines which gene will be expressed because its the site where rna polym binds in order to begin transcription (promoter =noncoding dna w specific function)
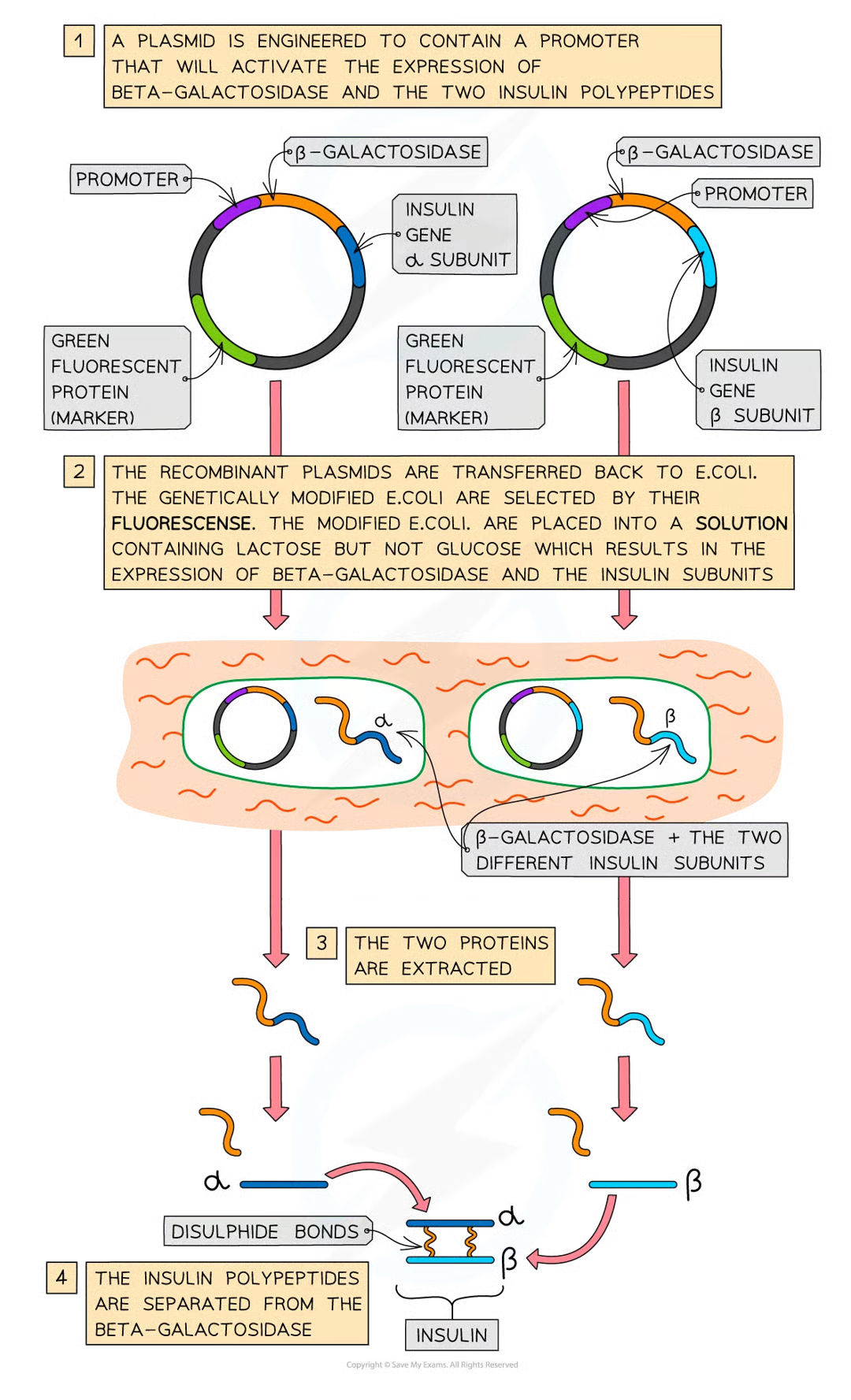
promoters role
regulate gene expression bc only if present will transcription and therefore gene expression occur
ensures rna polym can recognise which strand is dna template strand, rna polym recognizes template strand as promoter contains transcription start point (first nucleotide of gene to be transcribed) which is where enz will bind

marker
gene that is transferred w desired gene to enable scientists to identify which cells have been successfully altered and now contain recombinant dna. antibiotic resistant genes often used
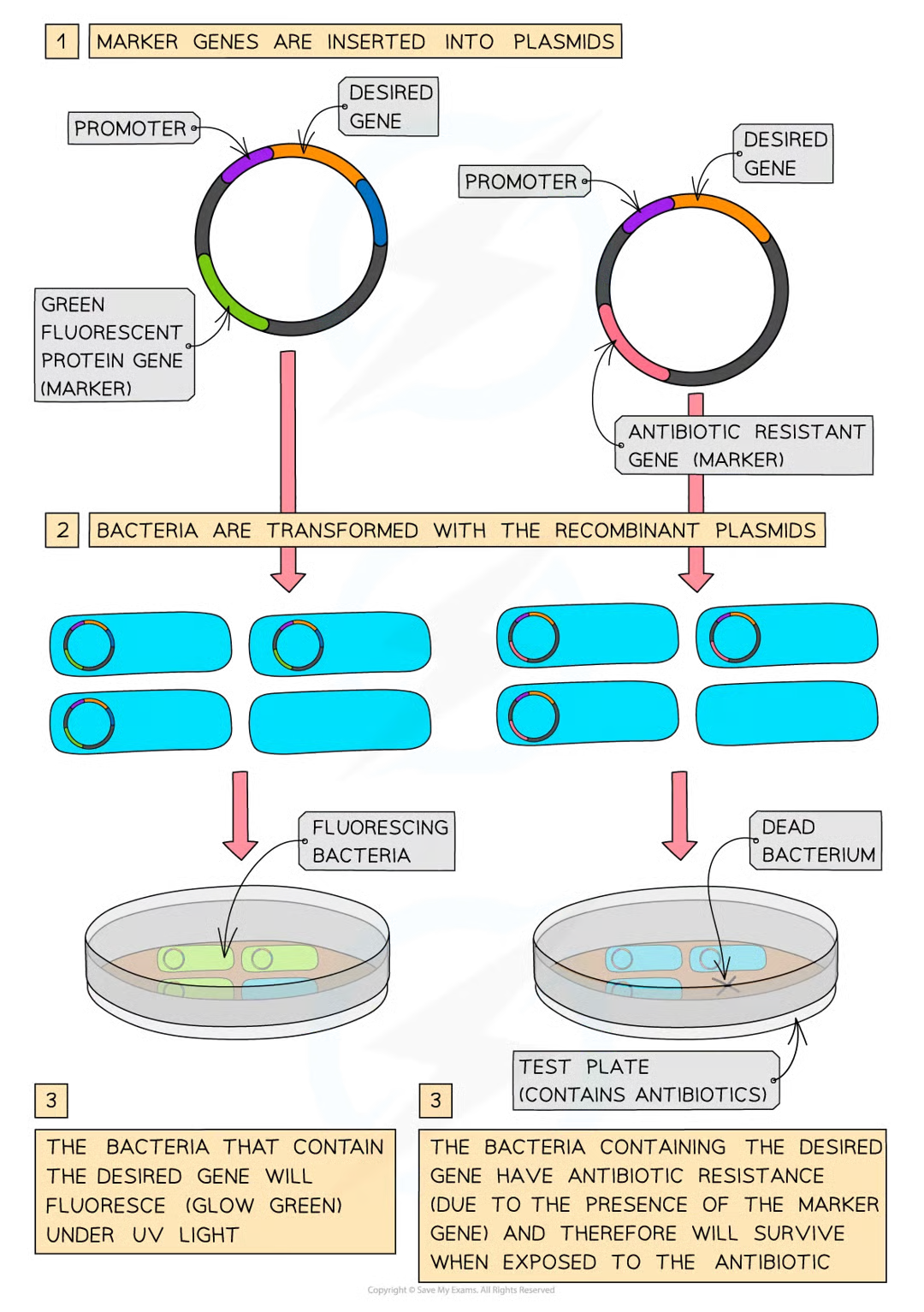
why were antibiotic resistant genes commoon markers
scientsits gmed the bacteria so taht the plasmid contained desired gene along w specific antibioitc ersistant gene (and promoter) and then grew the bacteria on agar plates w the anibioitic
bac w recombinant plasmids could be idenitified as they grew successuflly
why did antibioitic resistant genes as markers raise concerns 2
risk of antibiotic resistant genees transferring to other bacteria including pathogenic strains creating pathogenic antibiotic res bac making less effective antibioitics if res spreads
2 reasons antibioitic resistnat genes can occur
conjugation (transfer of genetic material fromone bacaterium to another)
transduction (transfer of gen material from one bacteria to another via a virus)
what are used as markers now
genes that express fluorescent proteins bc of presence of GFP (green fluorescent protein).
GFP and desired gene linked to specific promotor, so what happens when promo activated
protein expressed, recomb bac detected bc glow green under UV light
4 reasons for GFP>antibioitic
easier to idenityf (uv light)
more economical (dont nede to grow bac on plates of agar w antibioitics
no riks of antbibiotic res being passed to other bac
antibioitics no longer effecftive therefore wont stop bac growth
gene(genome) editing
allows genetic engineers to alter dna of org by inserting, deleting or replacing dna at specific sites in genome known to cause disease. foreign dna is not introduced to genome.
inaccurate methods using vectors used in past
modifying viruses to insert dna into gene causing disease = dna being inserted into other genes causing unforeseen consequences
liposomes (smal spheres of lipid mols) containing normal gene was sprayed into noses = short term sol as epithelial lining nasal passageway were short lived.
modern gene editing techniques
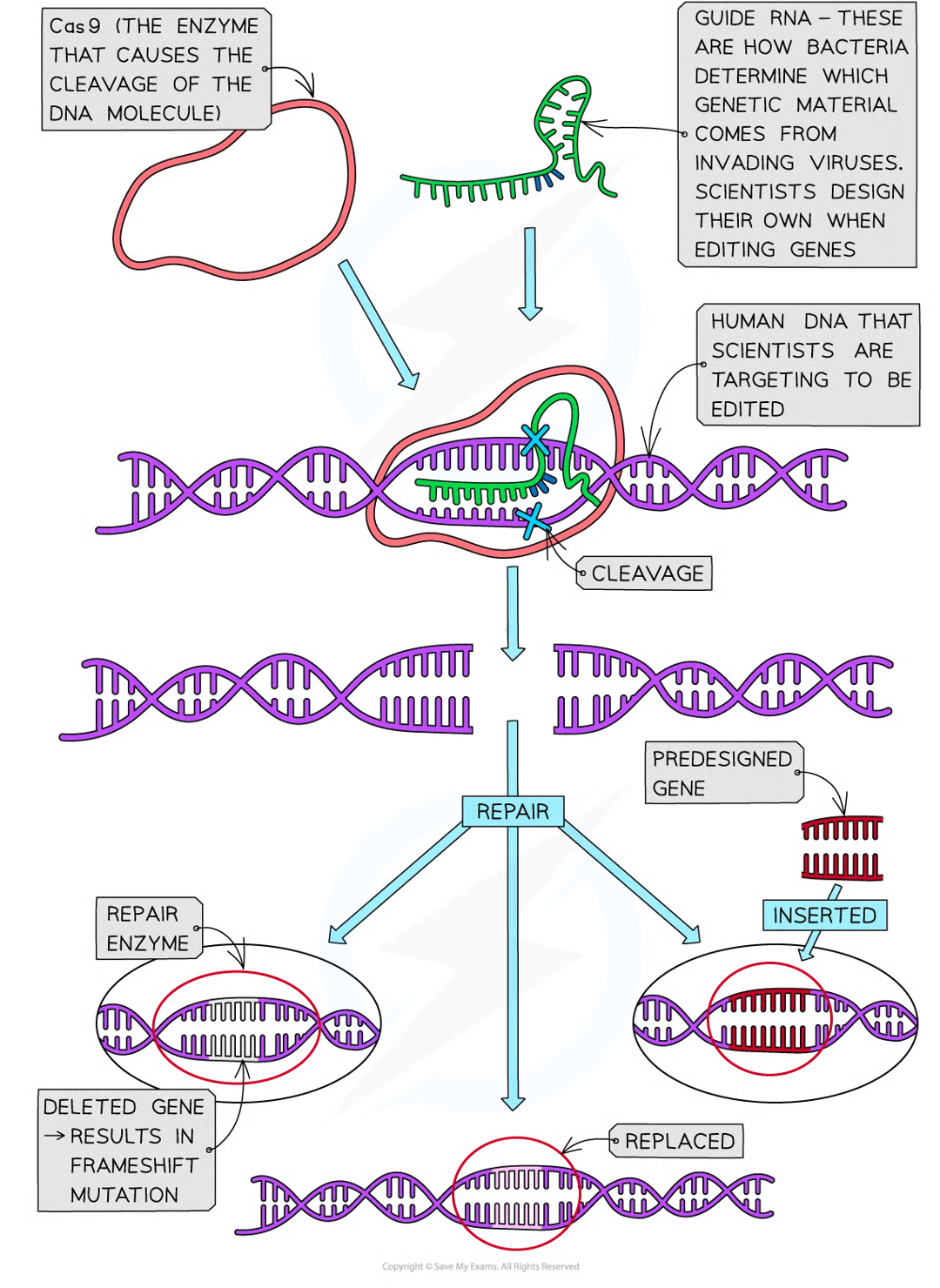
CRISPR (clustered regularly interspaced short palindromic repeats)
invovles using natural defence mech that bacteria (and archaea) have evolved to cut dna strands at sepcific point as determined by guide rna attached to enz (cas9)
one cut scientists can insert, delete, or replace faulty dna w normal
what is gene editing currently involved in
gene therapies e.g. cystic fibrosis, sicke cell anemia, treats people by altering genotype.

polymerase chain reaction (pcr)
common molecular biology technique used in most qapplications of gene technology e.g. dna profiling (e.g. identification of criminals and determining paternity), or GE
can be described as in vitro method of dna amplification
can be used to produce large qts of specific fragments of dna or rna from v small qts even just 1 mol of dna or rna so scientists can have identical copies of dna or rna samples in short time
pcr process 2 details
in each cycle dna is doubled so in a standard run of 20 cycles a million dna mols are produced
3 stages undertaken in pcr instrument/thermal cycle which automatically provides optimal temp for each stage and control length of time spent at each stage
5 requirements of pcr reaction
target DNA or rna being amplified
primers (forward and reverse) - short sequences of single stranded dna that have base seq comp to 3’ end of dna or rna being copied, define region that is to be amplified by signalling to dna polym where to begin building new strands
dna polym - enz used to build new dna or rna strand (most commonly taq polym as from thermophiliic bacterium thermus aquaticus so doesnt denature at high temps involved during 1st stage of pcr reaction) and opt temp high enoguh to prevent annealing of uncopied dna strands
free nucleotides - used in construction of dna or rna strnads
buffer solutoin - to provide opt ph for reactions to occur
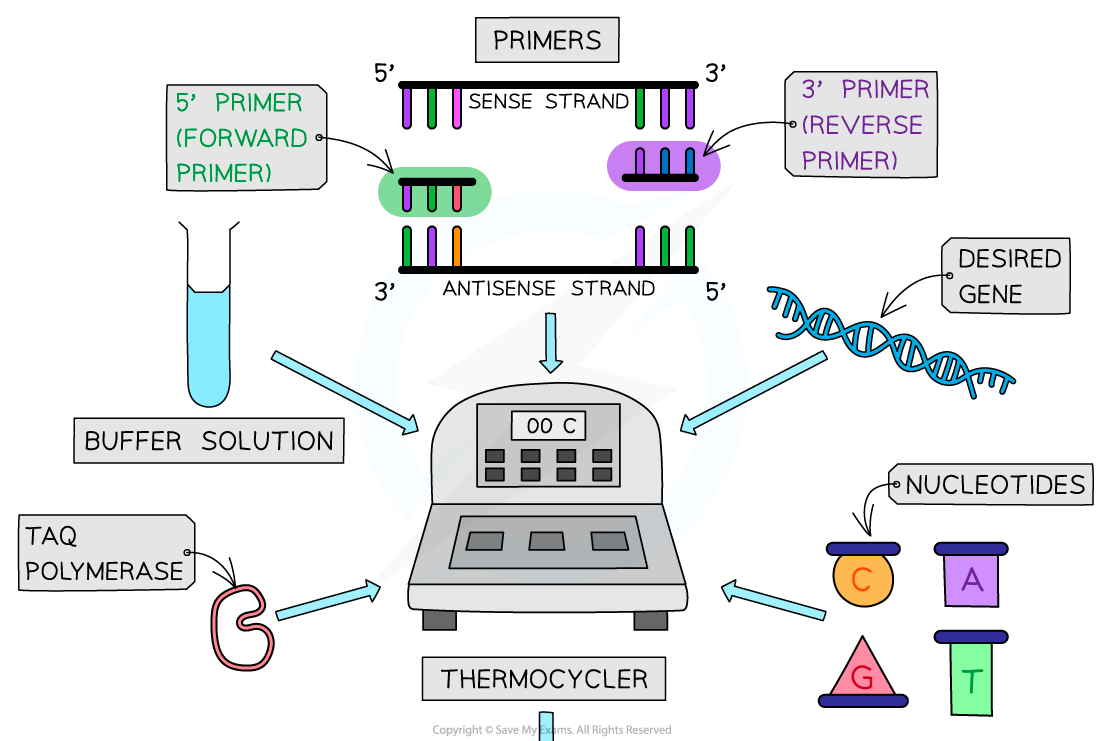
3 stages of pcr reaction
denaturation
annealing
elognation/extension
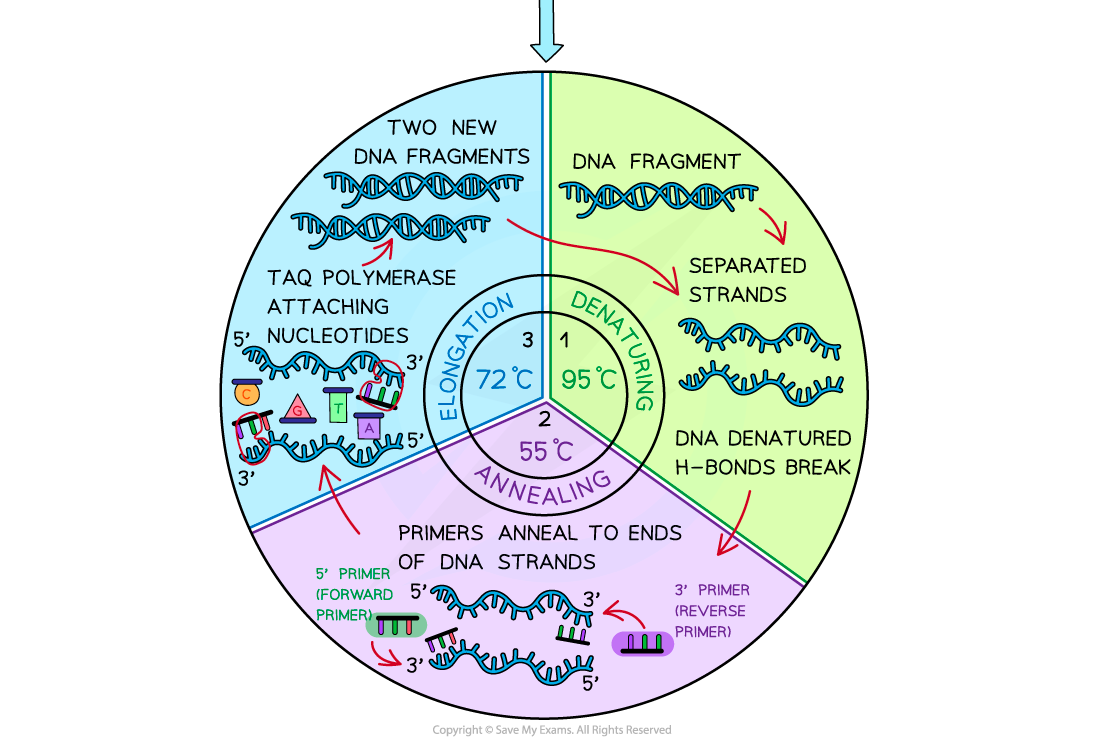
denaturation
double stranded dna is heated to 95C which rbeaks H bonds that bond 2 dna strands tg
annealing
temp decreased between 50-60 C so that primers (fwd and reverse) can anneal to ends of single strands of dna
elongation/ext
temp increased to 72C for at least a min, as this is opt temp for taq polym to build comp strands of dna to produce new identical doubel stranded dna mols
gel electrophoresis
widely used to analyse dna, rna and proteins. the molecules are separated according to their size/mass and their net change.
why does separation occur in gel electrophoresis 6
of the electrical charge that molecules carry - positively charged molecules will move towards the cathode (negative pole), whereas negatively charged molecules will move towards the anode (positive pole)
dna is neg charged due to phosphate grps and thus when placed in an electric field, the mols move towards anode
diff sized mols move thru gel (agarose for dna and polyacrylamide - pag for proteins) at diff rates
tiny pores in gel result in smaller mols moving quickly, whereas larger mols move slowly
diff types of gel have diff sized pores
proe size affects speed at which mols can move through them
dna separation process 9
dna can be collected from almost any body tissue e.g. saliva, hair root
after collection, dna must be prepped for gel ellectrophoresis so that dna can be sequenced or analsyed for genetic profiling (fingerprintg)
to prep the fragments, scientists must first increase/amplify no. dna mols by PCR
thenresstirction endonucleases enz are used to cut dna into frags
diff restriction enz cut dna at diff base sequences
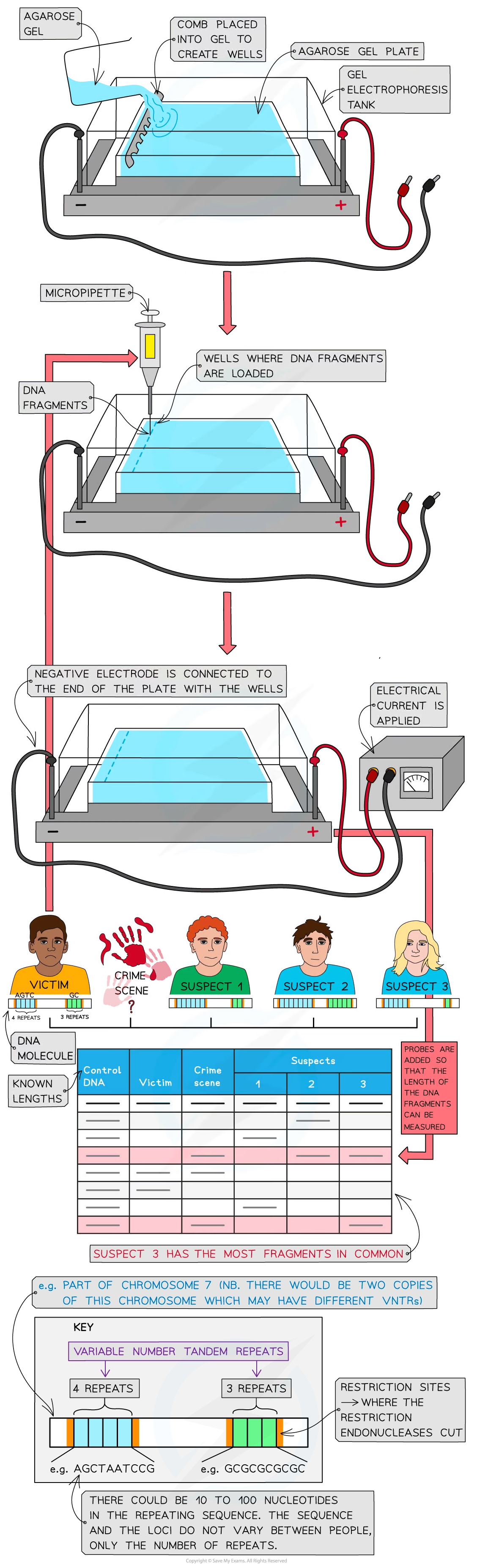
therefore scientists use enzymes that will cut close to variable number tandem repeat (VNTR) regions
variable no. tandem repeats (VNTRs) are regions found in non coding parts of dna
vntrs contian variable no. repeated dna sequences and are known to vary between diff ppl (except identical twins)
these vntrs may be referred to as satellite or microsatellite dna
how scientists separate dna fragments in gel electrophoresis 11
create agarose gel plate in tank by pouring liquid agarose and waitigng for it to set as a gel
mould wells (series of grooves) into gel at one end, using comb taht creates wells as the agaros gel solidifies.
submerge gel in an electrolyte solution (salt solution that conducts electricity)( in tank
load insert) fragments into wells using a micropippette
apply an electrical current to the tank
the engative electrode must be connected to the ned of th eplate w the wells as the dna frags will then move towards the anode (positive pole) due to the attraction between neg charged phosphates of dna and the anode
smaller mass/shorter dna frags move faster and further from wells than larger frags
frags not visible so muct be trasnferred onto absorbent paper or nitrocellulose which is then heated to separate the two dna strands
probes then added after whihc an x ray image is taken or uv light is shone onto the paper, producing a pattern of bands which is generally compared to a control fragment of dna
probes are single stranded dna sequences that are comp to vntr regions sought by scientists
probes also contain a means by which to be identified whihc is either:
radioactive label e.g. phosphorus isotope which causes probes to emit radiation that makes x ray film go dark creating pattern of dark bands
fluorescent stain/dye (eg ethidium bromide) which fluoresces (shines) when exposed to uv light creating pattern of coloured bands
how does protein separation work 3
aa comp of protein determines charge on each protein because each indiviudal aa r group diff charge
charge of r group depends on pH so buffer solutions used during sep of proteins to keep constant
gel electrophoresis used to separated ppt chains produced by diff allleles e.g. haemoglobin variants (a globin, b gloin and sickle cell anemia variant.)
microarrays
lab tools used to detect the expression of thousands of genes at the same time and to identify the genes present in an organisms genome
microarrays 3 diff uses
pathology - comparison between healthy cells and diseased cells to find disease characteristics
biotech (e.g. agri to identify insect pests)
crime (e.g. foresic analysts)
why are microarrays valuable to science
large numbers of genes can be studied in a short period ot ime since a microarray consists of a small (usually 2cm²) piece of glass/plastic/silicon (chips) that have probes attached to a spot (gene spot) in a grid pattern with more than 10k spots per cm²
probes
short lengths of single stranded dna (oligonucleotides) or rna which are synthesised to be comp for a specific base sequence (depends on purpose of microarray
how to use microarrays to analyse genomes
dna is collected from the species going to be compared
restirction enzymes are used to cut the dna into fragments
these frags denatured to created single stranded dna mols
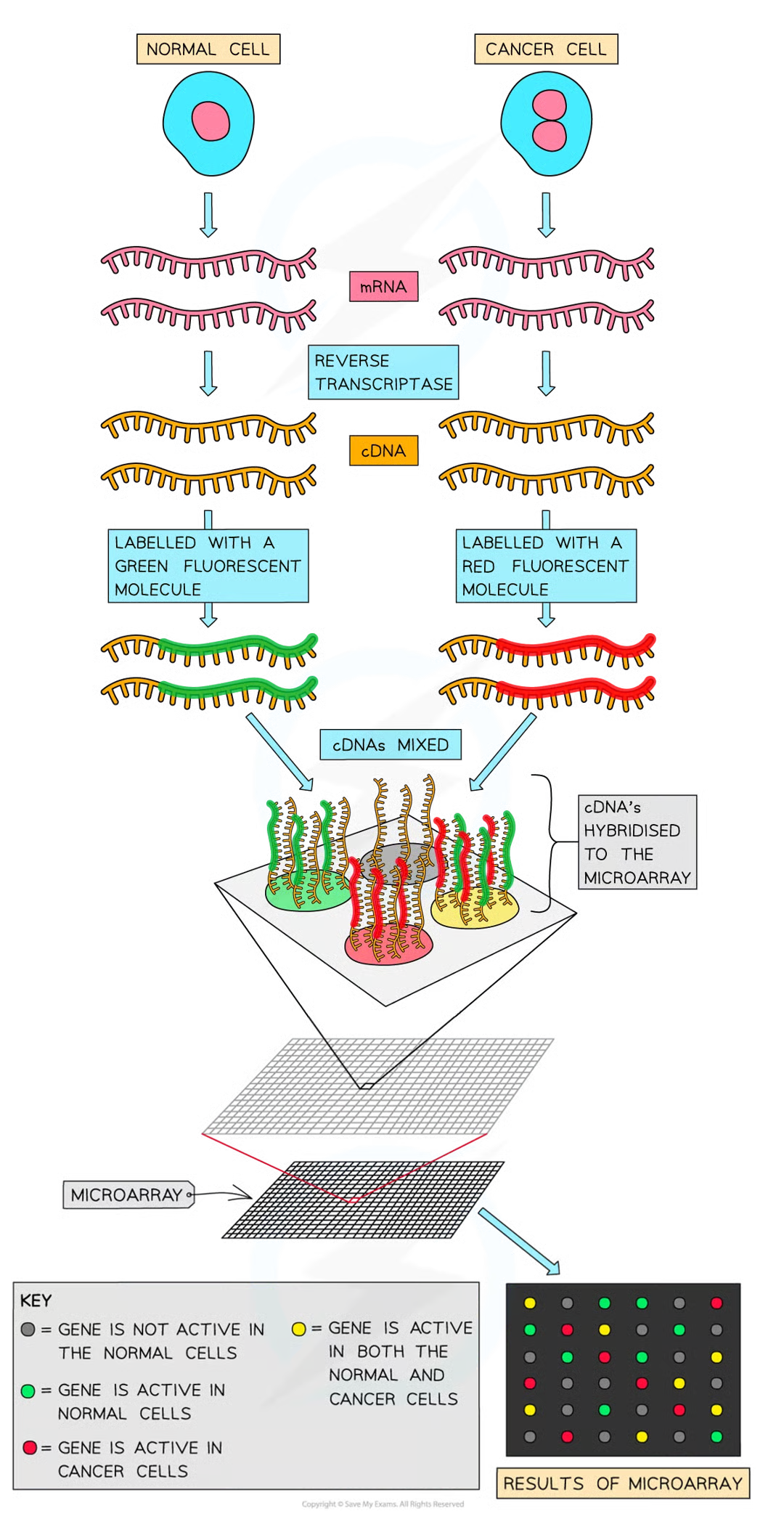
DNA frags labelled using fluorescent tags (frags form diff sources tagged diff colours - usualy red and green)
once these frags mixed tg they’re allowed to hybridise w probes on microarray
after set period, any dna that didn’t hybridise w the probes is washed off
microarray then examined w uv light (causes tags to fluoresce) or scanned (colours detected by comp and info analysed and stored)
presence of colour indicates where hybridisation occured as the dna frag is comp to probe
if red green fluorescent spots appear then only 1 species of dna has hybridised but if its yellow then both hybridised w dna frag = both species have gene in common
if spot lacks colour = gene not present in either species
why can scientists indirectly determine which geenes are being expressed in cells by assessing qt of mRNAS
bc when genes r being expressed or in active state, many copies of mRNA rpoduced by transcription. corresponding protein then produced from these during translation
steps to compare which genes are being expressed using microarrays: 7
mRNA collected from both types of cells and reverse transcriptase is used to convert mRNa into cDNA
pcr may b used to increase qt of cDNA (occurs for all samples to remain proportional so a comparison can be made when anal coccurs)
fluorescent tags added to cDNA
cDNA denatured to produce single stranded DNA
single strande ddna mols allowed to hybridise w probes on microarray
when uv light shone on microarray the spots that fluoresce indicate that gene was transcribed (expressed) and intensity of light emitting from spot indicates qt of mrna produced (how active gene is)
if light emitted is of high intensity, then many mrna were present while low intensity emission indicates few mrna present
data ocllected in todays techniques (3)
the sequences of genomes
when genes are express during orgs life
structure aa seq and functions of proteins
bioinformatics
interdisciplinary science (incorporating bio w computer tech and stats) where bio data is collected, organised, manipulated, analysed and stored. large databases are craeted containing info ranging from gene sequences to aa seq of prroteins which are available online and can performa nalaysis of data selected.
once genome sequenced, bioinformatics allows scientists to make comparisons w genomes of other orgs using many databases available whihch can help to find degree of similarity between orgs which then gives indication of how closely related orgs are and whether there are orgs that cld be used in experiments as modelf or humans (e.g. furit fly drosophila)
bio informatics 4 databases
the european molecular biology laboratory - nucleotide seq database
arrayexpress - microarray databse w level and types of mrna expressed in diff cells
protein data bank at europe - protein seq searches
blast (basic local alignment search tool) - used by researches to find similarities between sequecnes theyre studying w those already in data base
recomb dna
dna that has been altered by introducing nucleotides from another source. dna from one org has been recombined w dna from another
transgenic organism
if an organism contains nucleotides from a diff species
gmo
any org that has received genetic material from another source
recombinant proteins and their use
produced from recomb dna and are mainpulated forms of og protein. generated using m/os e.g. bacteria, yeast, animal cells in culture for reseaarch and treatments (diabetes, cancer, infectious diseases, haemophilia)
how are rp (human) produced
using euk cells (e.g. yeast, animal cells in culture) rather than prok bc euk carry out post-translational modification bc presence of golgi apparatus (enzymes) that is required to produce a suitable human protein.
advantages of g engineering orgs to produce recomb human proteins 6
more cost effective to produce large vols (unlimited availability)
simpler (w regards to using prokaryotic cells)
faster to produce many proteins
reliable supply available
proteins engineered to be identical to human proteins or have modifications that are beneficial
can solve the issue for ppl who have moral, ethical, religious concerns against using cow or pork produced proteins
how was recomb insulin made 4
made to treat diabetes by modifiying bacterial plasmids to include the human insulin gene
restriction endonucleases are used to cut open plasmids and dna ligase is used to splice the plasmid and human dna together
these plasmids then reinserted into bacterium escherichia coli by trnasformation (bath of ca2+ and then heat or electric shock
once transgenic bacteria identified by markers, theyre isolated, purified and placed into fermenters that provide optimal conditions
transgenic bacteria multiply by binary fission an d express human protein insuling which is eventually extracted and purified
advantages for scientists to use recombinant insulin 6
identical to human insulin, unless modified to have different properties (e.g. act faster which is useful for taking immediately after a meal or to act more slowly)
there’s a reliable suplpy avialble to meet demand (no dependence on availability of meat stock)
fewer ethical, moral or religious concerns (proteins not extracted from cows/pigs
fewer rejection problems, side effects or allergic reactions
cheaper to produce in large volumes
useful for people who have animal insulin tolerance
how was recomb factor viii created
done to help haemophiliacs who cant naturally produce this blood clotting protein from gm kidney and ovary hamster cells
recomb cells placed into fermenter and cultlured once modified
due to opt conditions in fermenter, hamster cells constantly express factor viiii which is extracted, purified and used in injectable teratment for heamophilia
3 adv for scientists using recomb factor viii
fewer ethical, moral, religious concerned (proteins not extracted form human blood
less risk of transmitting infection (hiv) or disease
greater prod rate
how was recomb adenosine deaminase made 2
made to treat inherited condiitoin adenosine deaminase deficiency as it causes severe combined immunodeficiency (scid) bc immune system damaged
larva of cabbage looper moth trichoplusia ni has been gm (using virus vector) to produce enz adenosine deaminase so that it can be used as treatment while patient waits for gene therpay or when gene therapy isnt possible
4 adv for using recomb adenosine deaminase
fewer ethical, moral, religious concerns (proteins not extr from cows)
less risk fo transmitting infection or disease (from cows)
most reliable prod of enz
fsater to produce mnay protiens
genetic screening
can help identify individuals hwo are caryring an allele at a gene locus for a particular disorder. the testing of an embryo, foetus or adult to analyse the dna
how is genetic screening dna sample obtained 2
taking tissue samples from adults or embryos made IVF
chorionic villus sampling or amniocentesis of embryso and fetuses in uterus
genetic counsellors role and 6 discussions
reads results and explains to them
chances of couple having child w certain disease
termination of pregnancy
therapeutic treatments possible for child
financial implications of having the child
effect on existing siblings
ethical issues
brca1 and brca2
genes that produce tumour suppresor proteins and thus they play an important role in regulating cell growth
how is breast cancer brough about
faulty alleles of brca1 and 2 increase risk of indi dev breast and ovarian cancers during lifetime and can be inherited from either parent+
advantages of gennetic screening for those w history of brca1 and 2 mutations: 3
person may decide to take preventative measures (e..g elective mastectomy - breasst removal - to reduce risk of dev cancer
screening for breast cancer may begin from earlier age or more frequently, and inidivudla (if female) will haev more frequent clinical examinations of ovaries
enables person to participate in research and clinical trials
huntingtons
progressive disease that affects brain at 40+, shown by uncontrolled movement, lower cognitive thinkingability and emotional problems without a cure, only treatment. it is autosomal dominant.
3 adv of gen screening for huntingtons
ppl to plan for future (how they will live and b cared for)
couples make informed reproductive decisions (as risk is 50%)
ppl participate in research and clinical trials
cystic fibrosis
autosomal recessive genetic disorder caused by mutation of gene that codes for transported protein called cftr. progressive and causes mucus in various organs (lungs, pancreas) to become thick and sticky. no cure but treatments and can cause death from bac infection in lungs
what happens if cftr protein is faulty
no longer transports chloride ions across cell plasma memb and therefore water doesnt move by osmosis across memb either (presence of water wolud normally make mucus thinner enabling cilia to remove it)
2 adv of genetic screening for cyst fibro
enables couples to make informed decs aas both may be carriers w/o symptoms
ppl can participate in research and clinical tries.
gene therpay
involves using various mechanisms to alter a persons’ genetic material to treat or cure diseases. possibilities of replacing faulty gene, inactivating it or inserting a new gene. experimental technqiues being used to treat and research treatments for scid, leber congenital amaurosis (blindness), beta thalassaemia and haemophilia b. most in clinical trial stage as hard to find delivery systems that can transfer normal alleles to persons cells and ensure expression
why is finding an appropriate delivery system one of the problems in gene therapy
vectors are currently used, with viruses being most common but non-viral vectors also being researched e.g. liposomes and naked dna
viruses (retroviruses and lentiviruses) most commonly used vectors as have mechanisms needed to recognise cells and deliver genetic material into them
why is effects of changing somatic cells short lived
changes in gen mat are targeted to specific cells and so will not be inherited by future generations (as somatic gene therpay doesnt target gametes)
2 types of somatic gene therapy
ex vivo - new gene inserted via virus vector into cell outside body
blood or bone marrow clels extracted and exposed to virus which inserts gene into cells
these cells grown in lab and returned to person by injection into vein
in vivo - new gene inserted via a vector into cells inside the body