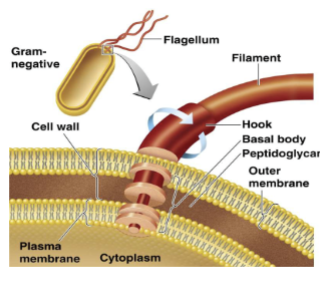
ch 3 prokaryote cell structure and function
1/45
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
46 Terms
bacterial cell envelope
contain outer membrane; possess cell wall; may consist of only cell/plasma membrane
inner cell membrane
phospholipids; proteins; maintains gradient
Cytoplasm
proteins, rna, macromolecules; water; inorganic ions; small organic molecules; nucleoid (DNA); ribosomes
cell wall
peptidoglycan - sugar peptide polymer; gram positive and negative bacteria
gram positive bacteria
thick cell wall
gram negative bacteria
thin cell wall; outer cell membrane; lipopolysaccharide layer (LPS)
external structure
pili/fimbriae; flagellum (motile); capsule - surrounds cell wall (virulence factor)
molecule abundancy
water is most abundant
rna, proteins, and dna make up 25%
transport across membranes
simple: o2, co2
facilitated: amino acids, sugars, transport proteins
osmosis: h2o
group translocation
modification of substrate during transport; source substrate moving down gradient
abc transporters
amino acid, carb transport; expends ATPs to transport
membrane permanent weal acids/bases
small neutral weak acid/weak basic diffuses into cell and causes pH to rise(basic) or fall(acidic); cell must counteract
prokaryote cell envelope
bacterial archaeal inner cell membrane protected by structural support layers: cell wall, s layer, outer membrane, complex mycobacteria species
cell envelopes of 2 major taxonomic groups distinguished by gram stain
bacterial cell wall
structural support, protection, and porous
composed of peptidoglycan (polymer of disaccharides, glycan, peptide tetramers)
peptidoglycan synthesis
synthesis complex extends chains of amino sugars
helical direction of synthesis by complect via MreB (cytoskeletal element)
MreB
polymerizes along arc beneath plasma membrane (inner)
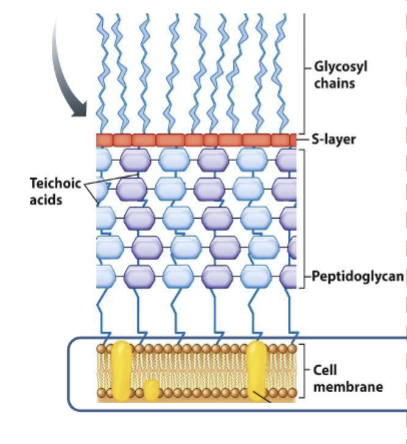
gram positive
thick cell wall; peptidoglycan layers w teichoic acids
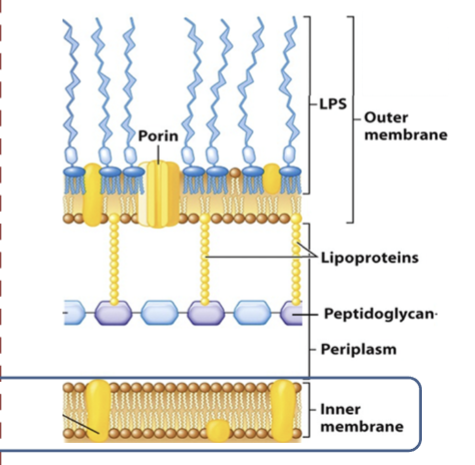
gram negative
porous to most ions and small organic compounds
outer membrane proteins contain nonspecific porins transporting sugars and peptides
LPS layer has endotoxin released when cell dies: lipid A - short chain fatty acids linked to glucosamine dimers, core polysaccharide- 5 sugars, O polysaccharide - linked to core polysaccharide
s layer
surface layer of crystalline array of protein or glycoprotein; porous
periplasm
between inner and outer membranes; contains enzymes and nutrient transporters
atypical cell envelops
mycoplasma: smallest, lack cell wall
archaea: comprised pf pseudomurien
mycobacteria: tuberculosis, leprosy; contain mycolic acids (waxy lipids)
external structures
capsule: polysaccharide layer outside envelope, protein in nature, can be virulence factor; organized and tightly associated to cell than slime layer; biofilms surrounded by exopolysaccharide matrix
bacterial cytoskeleton
provides shape and form; provide protection; FtsZ protein; MreB protein; CreS protein
Ftsz protein
determines cell diameter; forms z ring for cell division and septation
MreB protein
travels in helical arc under cell membrane; guides cell wall synthesis
CreS protein
crescentin; curves inner side of bacteria
division and septation
septum is partition splitting envelope; septum grows inward and constricts; rapid synthesis of envelope; managed by divisome; facilitated by FtsZ
polysome
can result in protein product from expressed gene
polar aging
related to cell division in rod shaped bacteria; asymmetrical
old poles age: cell wall degrades and susceptible to lysis; accumulation of non functional protein aggregates in stressed cells
differ in antibiotic resistance
bacillus: endospore forms at one pole
c bacteria: pole gens flagellum or stalk during division
replisome
one at each replication fork in DNA
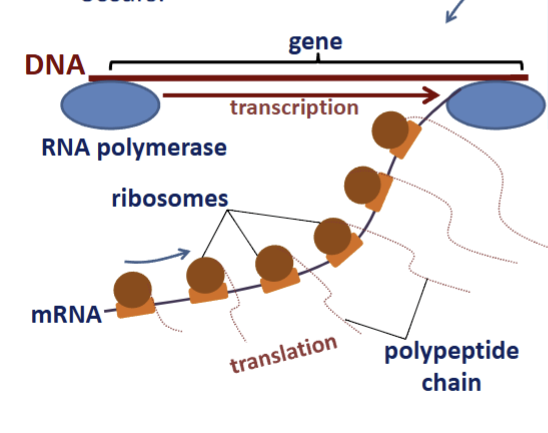
bacterial transcription and translation
occurs in nucleoid simultaneously
nucleoid
chromosome; singular, circular, double stranded DNA, loops/domains; ori is origin of replication; supercoiling compacts DNA (gyrase); DNA condensation via binding proteins
transcription/translation
tightly coupled; polysome/polyribosome formation
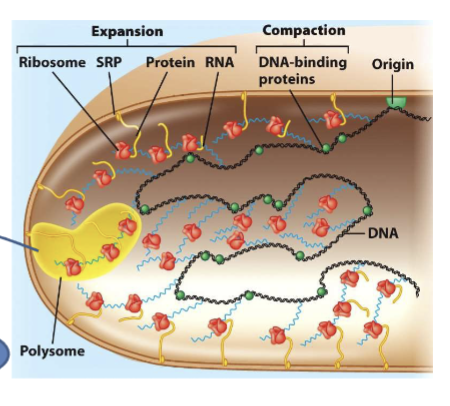
translated proteins
functions in cytosol; synthesized at membrane directed by signal recognition particles
cell division
fission; segregation and partitioning; clones; 20 min - 2 hrs
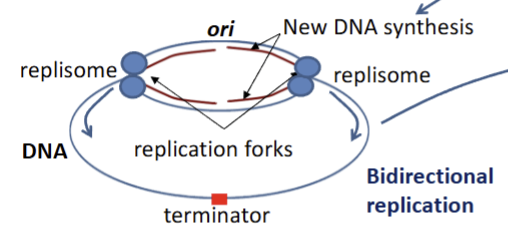
dna replication
begins at ori site, strands separate
dna polymerase complexes (repliosomes) synthesize dna: 2 dna polym per repliosome → leading and lagging strand synthesis
end of replication → z rinng formation via ftsz → septation and division
specialized structure
adapt to metabolic strategies and environments
phototrophs - thylakoids, gas vacuoles, carboxysomes
thylakoids
inner cell membrane foldings with photosynthetic components
carboxysomes
protein covered bodies with rubisco ( co2 fixation enzyme)gas
gas vacuole
maintain depthso
storage granules
metachromatic: inorganic phosphate
polysaccharide: glycogen, starch → glucose polymers
sulfur: insoluble, oxidation of h2s; h2s → s0
lipid inclusion: polyhydroxybutyrate
magnetosomes
magnetite crystals
orient along magnetic field
anaerobic or microaerophilic bacteria
move to lower oxygen depth
pili, fimbriae
adherence. protein monomer pilin
attach to mucous membrane
attach to surfaces → biofilm formation
twitching motility
sex pilus connects donor cell to recipient cell
stalk
extension of cytoplasm
attaches bacteria via secretion of adherence (holdfasts)
nanotubules
cell envelope extensions connecting cytoplasm between cells
share proteins and mrnaf
flagella
motility via rotary motion
filament of monomers of protein arranged in chains
h antigen: protein
filament attached to protein attached to hook attached to basal body anchored to wall and membrane
required ATP
clockwise → tumbles; counterclockwise → runs