Post-Transcriptional Control
1/50
Earn XP
Description and Tags
lecture 9
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
51 Terms
What does post-transcriptional control regulate?
mRNA processing (premRNA → mature mRNA)
pre-mRNA 5’ end capping
pre-mRNA splicing
pre-mRNA 3’ Poly A tail
mRNA transport from nucleus to cytoplasm
mRNA stability (how long mRNA persists before degradation)
mRNA degradation
What is the main point of mRNA decay?
All cellular mRNAs are eventually degraded; mRNA decay is a regulated post-transcriptional process controlling gene expression and mRNA half life.
What is mRNA half life?
The time it takes to degrade half of existing mRNA molecules.
What determines mRNA half life?
Specific interactions between cis-acting mRNA elements and the proteins that bind these elements.
What are the three pathways of cytoplasmic mRNA decay in eukaryotes?
Deadentlation-dependent mRNA decay
Deadenylation-independent mRNA decay
Endonuclease-mediated mRNA decay
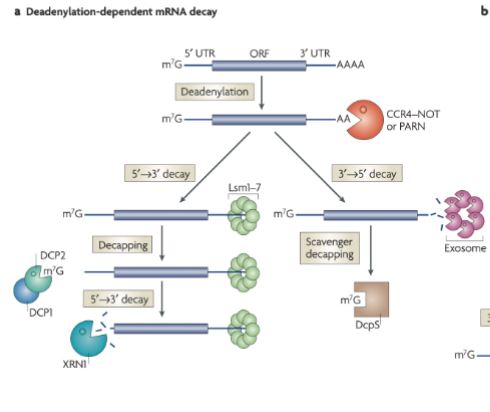
What happens in Deadenylation-dependent mRNA decay?
Removal of poly A tail at the 3’ end (deadenylation by deadenylation protein)
Path splits → either decay 5’ → 3’ or 3’ → 5’ direction
Path A: 5’ → 3’ decay
3’ tail is stabilized with a protein
Cap removed (decapping from decapping enzyme)
RNA is then degraded 5’ → 3’ direction (exonuclease)
Path B: 3’ → 5’ decay
Exosomes chew up mRNA 3’ → 5’ direction
Cap removed at the end (scavenger decapping)
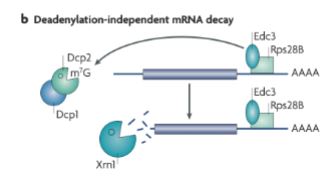
What happens in Deadenylation-independent mRNA decay?
Poly A tail is not removed first.
Proteins bind mRNA (usually 3’ end) for stabilization.
decapping enzymes are recruited.
Cap is removed.
RNA is degraded 5' → 3’ (exonuclease)
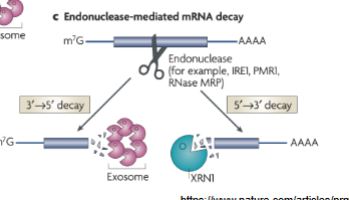
What happens in Endonuclease-mediated mRNA decay?
mRNA is cut in the middle (with endonuclease), it goes for shapes + structure (like sequences and stem loops)
it can then go through either pathway = 5’ → 3’ or 3’ → 5’
What are deadenylases?
They:
remove poly A tail
initiate mRNA decay
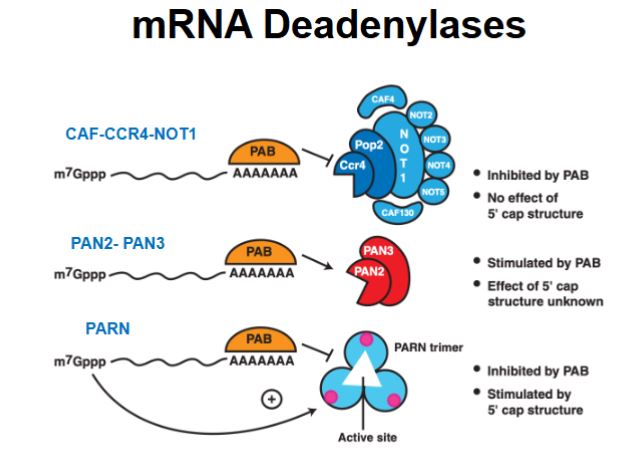
What is the role of Poly A binding protein (PAB)?
It can either protect the poly-A tail (inhibiting decay) or recruit deadenylases (promoting decay).
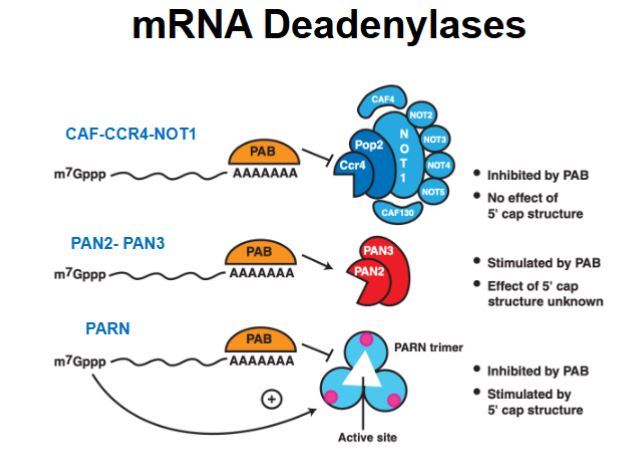
If the poly A tail is protected (deadenylase is inhibited), what happens?
Deadenylation cannot occur; other decay pathways must be used:
Deadenylation independent decay pathway
Endonuclease-mediated decay pathway
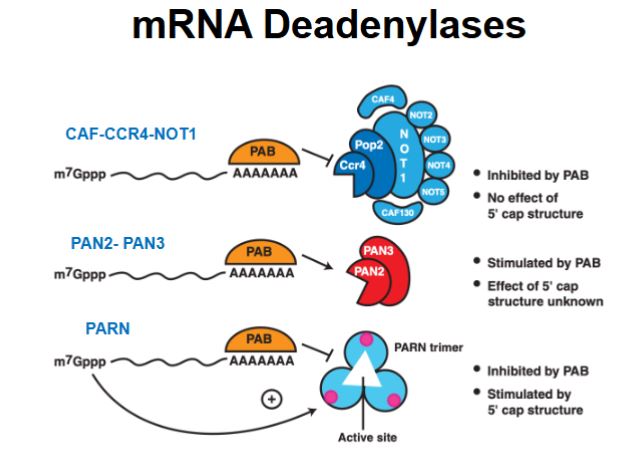
What determines which mRNA decay pathway occurs?
The proteins bound to the mRNA and what they recruit or inhibit.
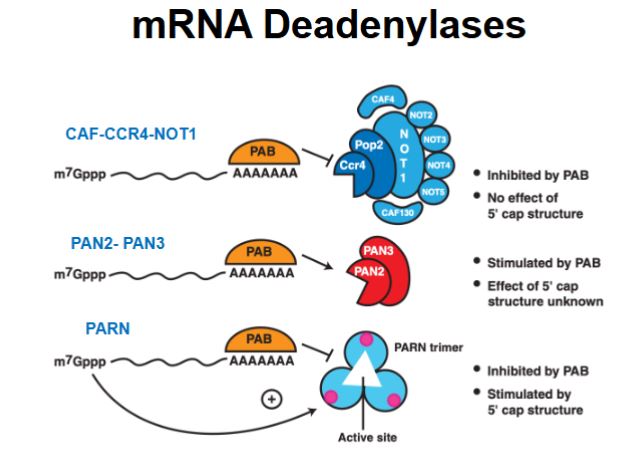
What is an exosome and what is it’s function?
A large protein complex with 3’ → 5’ exonuclease activity that degrades and processes RNA.
Where does the exosome function?
Nucleus: processes RNA (rRNA, snRNA, snoRNA) and degrades improperly processed transcripts ( improper 5’ cap, 3’ tail, and/or splicing(introns)).
Cytoplasm: degrades normal and defective mRNAs
What determines how the exosome is recruited?
The state of the mRNA and the proteins bound to it.
“Depending on whats happening with the mRNA → different proteins will be used.”
What is nonstop decay?
Decay of mRNA lacking a stop codon; ribosome stalls and recruties exosome via ski proteins.
What are the two types of decapping enzymes?
DCP1/2 (standard) and scavenger decapping enzyme.
When does DCP1/2 act?
Early, initiating 5’ → 3’.
When does scavenger decapping enzyme act?
After 3’ → 5’ decay when only the cap remains.
What is Nonsense-mediate mRNA decay (NMD)?
A translation-dependent mRNA quality control (or mRNA surveillance) mechanism that degrades transcripts with premature stop codons.
Full NMD mechanism
mRNA made
Exported to cytoplasm
First (pioneer) round of translation
Ribosome encounters premature stop codon
Cell recognizes defect
NMD triggered
mRNA degraded (via decay pathways)
Why is NMD necessary?
To prevent production of truncated, potentially harmful proteins.
When does NMD detect the faulty mRNA/ What is required before NMD can occur?
During the pioneer (first) round of translation.
Why doesn’t the cell just degrade faulty proteins?
It is more energy-efficient to degrade the mRNA before protein synthesis.
Why make something we need to degrade anyway?
What mutation commonly triggers NMD?
Frameshifts often incorporates an early stop codon.
What is the 50-55 nucleotide rule?
A premature stop codon triggers NMD only if it is ≥50–55 nucleotides upstream of an exon–exon junction (EJC marks where the exon-exon junctions are).
What role do exon junction complexes (EJC) play in NMD?
They mark exon junctions and allow detection of premature stop codons if they remain downstream during translation.
Why must translation occur before NMD?
Because the cell must detect a premature stop codon during translation.
Besides quality control, what is another role of NMD?
Feedback regulation.
example: high amounts of protein → a protein can bind to own preRNA → introducing premature stop codon → triggers NMD pathway → mRNAs decay and inhibit making anymore protein.
What is the general principle behind NMD targeting?
mRNAs that appear abnormal during translation are recognized and degraded.
NMD mammalian targets (4 different classes that gets recognized for NMD)?
mRNAs that naturally contain an uORF (mRNAs with extra upstream start/stop regions)
Introns in 3’ UTR (untranslated region). Leaves exon junction signals downstream → triggers NMD
Selenoprotein mRNA: UGA can sometimes encode selenocysteine instead of functioning as a stop codon.
mRNAs of transposons and pseudogenes. Often defective → targeted by NMD
What are post-transcriptional modifications (PTMs)?
Chemical or structural changes made to proteins after translation.
Why are PTMs important?
They increase the diversity of the proteome (which increases functional capacity).
How do PTMs affect the proteome?
They greatly increase protein diversity beyond what genes alone produce.
What is co-translational modification?
Modificaiton of a protein while it is still attached to the ribosome.
What is post-translational modification?
Modification of a protein after it is released from the ribosome (no longer attached to the ribosome).
What is proteolytic cleavage (example of PTM)?
Cutting a protein to convert it from an inactive to an active form.
Example: procollagen (inactive) cut to collagen (active).
What can PTMs change in a protein?
Activity
Stability
localization
interactions
What does polyubiquitination signal?
Protein degradation.
What does monoubiquitination do?
Regulates protein function (not degradation).
What are writers in PTMs?
Enzymes that add tags (modification).
What are erasers in PTMs?
Enzymes that remove tags (modifications).
What are readers in PTMs?
Proteins that recognize the tag and carries out the function.
Phosphorylation → what enzyme adds and removes phosphate groups?
Kinase: adds phosphate groups
Phosphatase: removes phosphate groups
What is N-terminal acetylation?
Almost 80% of proteins have this added to the N-Terminus.
Addition of an acetyl group to the N-terminus that stabilizes proteins.
What does methylation do to proteins?
Can either activate or repress protein function.
Why was methylation and acetylation reffered to as yin and yang?
Acetylation generally activates proteins
Methylation can repress (but also activate) proteins
What is sumoylation?
Addition of SUMO protein tags that regulate protein function.
Glycosylation?
Addition of carbohydrate groups to proteins
glycoproteins are often involved in membrane localization and cell interactions
Lipoproteins
Addition of lipid groups to proteins.
Anchors proteins to membranes.
Metalloproteins?
Proteins that contain metal ion cofactors.
support structure or catalytic function.