Pattern Formation in Drosophila
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
Pattern Formation
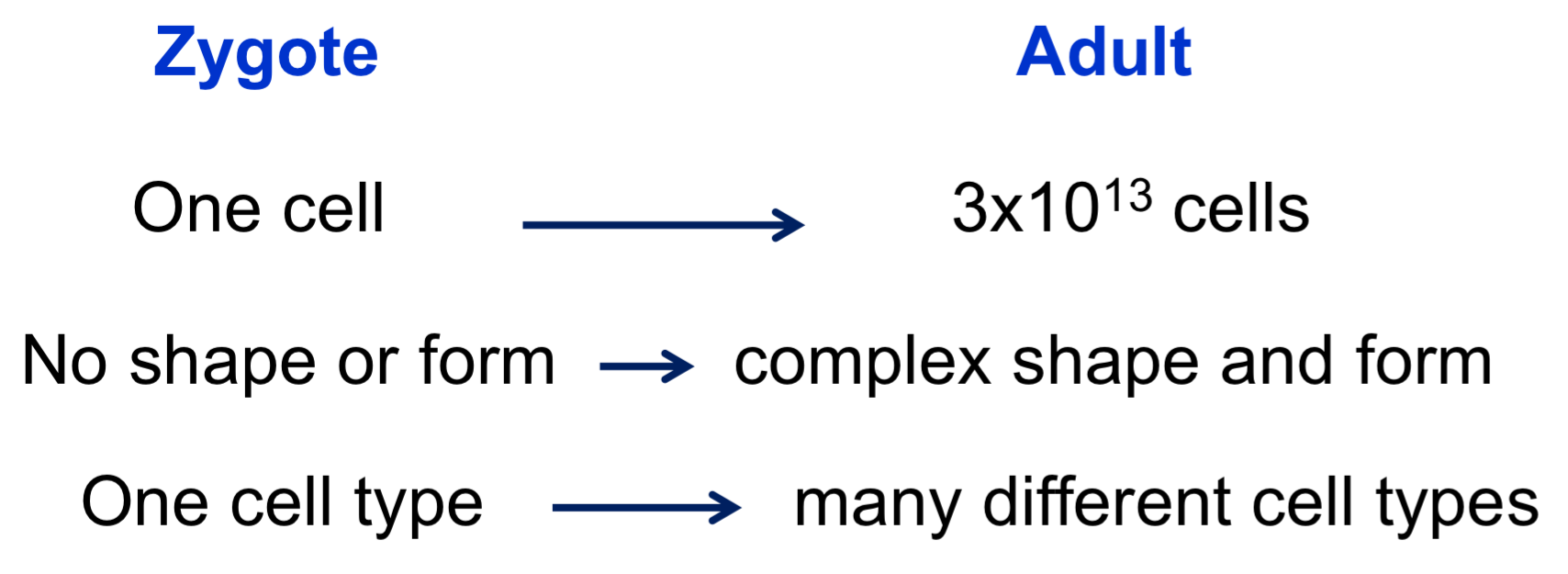
Principles of Pattern Formation
must establish positional information (spatial information) within the developing embryo (morphogens/environmental signals)
direct simple patterns early (anterior/posterior, dorsal/ventral), and use positional info to develop more complex patterns
use a cascade of determination events to coordinate timing (involves transcription factors and gene expression)
use master regulatory proteins (usually transcription proteins) each of which will be expressed in specific groups of cells marking them for a particular fate
Pattern Formation Axes
axes determined in the oocyte
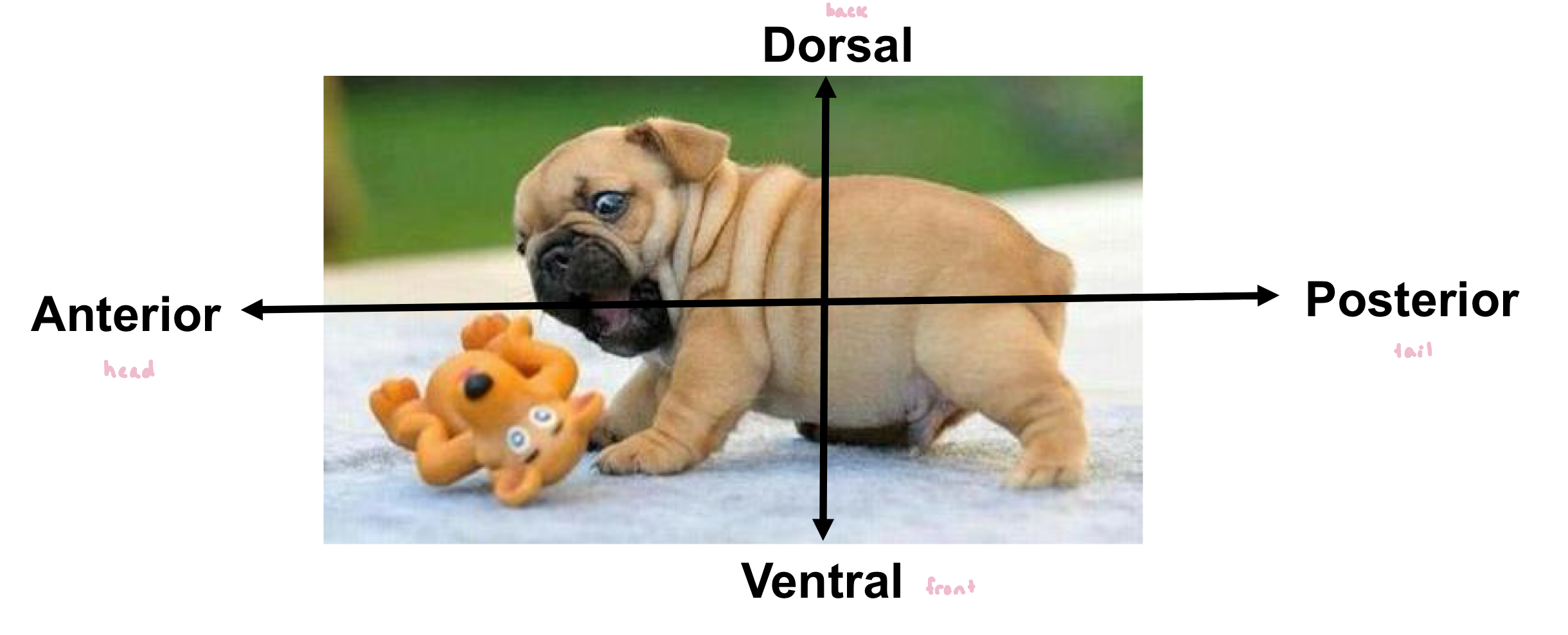
What are the key events during Drosophila embryogenesis?
oocyte cell is fertilized to generate the zygote (still single-celled embryo)
following fertilization, the zygotic nucleus (and all progenitor nuclei) undergoes 13 rounds of mitotic divisions without any cytokinesis, forming a syncytial blastoderm (giant multi-nucleated cell)
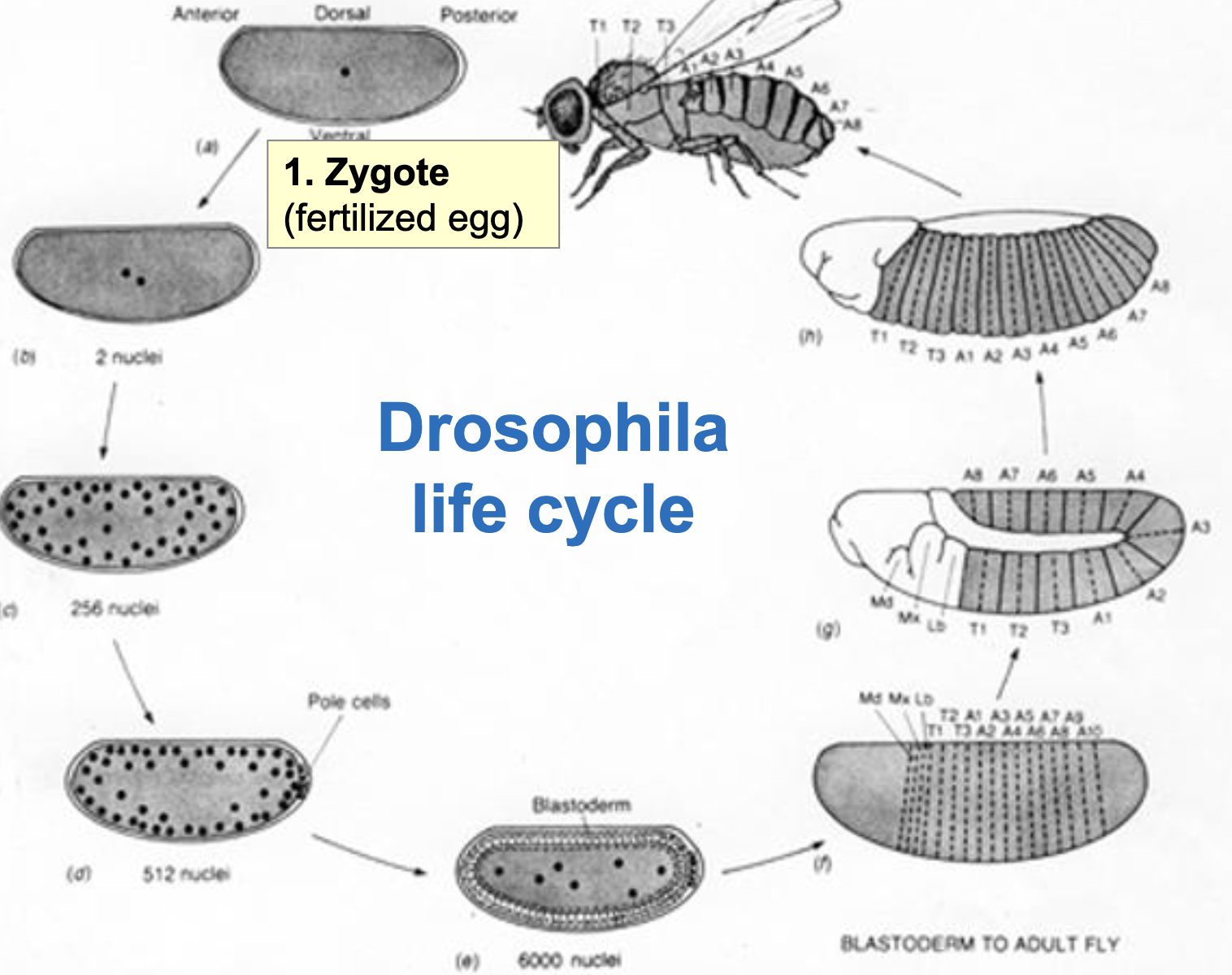
Drosophila Life Cycle
Zygote (fertilized egg)
Nuclear division w/o cytokinesis to form syncytial blastoderm
by the 9th or 10th round of divisions, the nuclei migrate to the periphery and the pole cells (germ line precursors) are formed
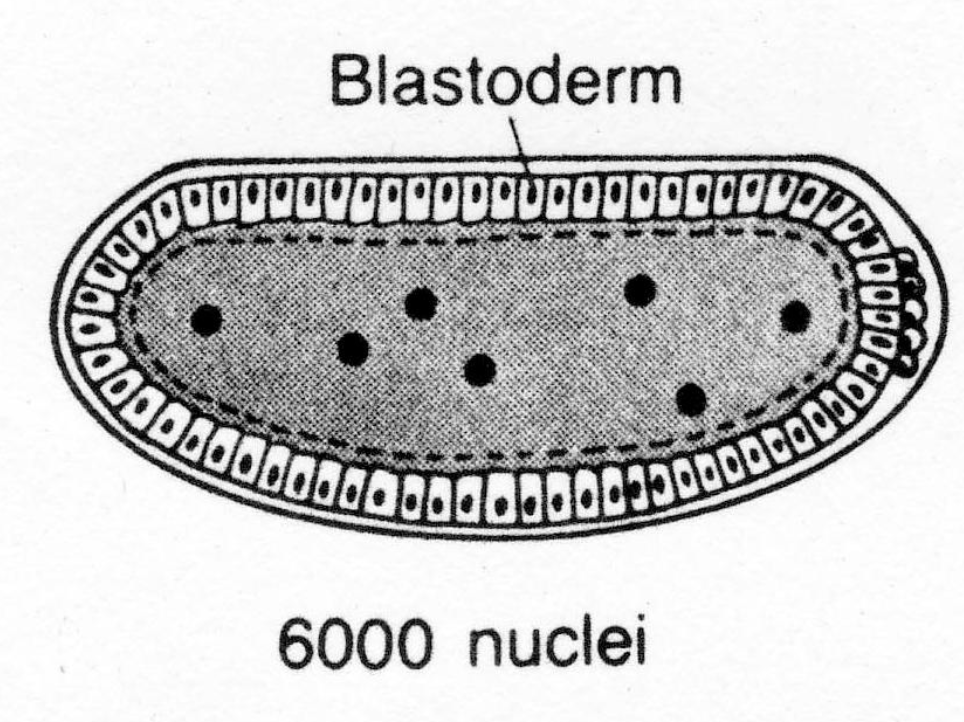
at the end of the 13th round of division (abt 6000 nuclei), cellularization (cytokinesis) occurs producing the cellular blastoderm (hollow ball of cells)
many cell membranes forming around diff nuclei
cell division and migration occurs during gastrulation, producing a segmented larva w/ shape and form
the “hollow ball” of cells flattens and folds inwards to form more layers of cells → forms the gut
cells have started to take on diff identities to become diff segments of the insect
Drosophila Life Cycle Figure
Are there cytoplasmic determinants in the zygote?
Removing Cytoplasm experiment:
removing cytoplasm from anterior → allow to develop to gastrulation → embryo develops with no head or thorax but a normal abdomen
remove cytoplasm from posterior → allow to develop to gastrulation → embryo develops w/ no abdomen but a normal head and thorax
Cytoplasm Transfer Exp:
transfer cytoplasm from posterior to anterior → allow to develop to gastrulation → embryo develops w/ posterior segments on both ends
based on these results, the zygote has specified commitment: we reversed the decision of which end is becoming anterior and which is becoming posterior (responded to changes in whatever was in the posterior cytoplasm)
Conclusions from these Experiments
cytoplasmic determinants are localized in the anterior and posterior regions of the zygote
the developmental decisions affected by the cytoplasmic transfer exps are broad ones; these early decisions are also reversible
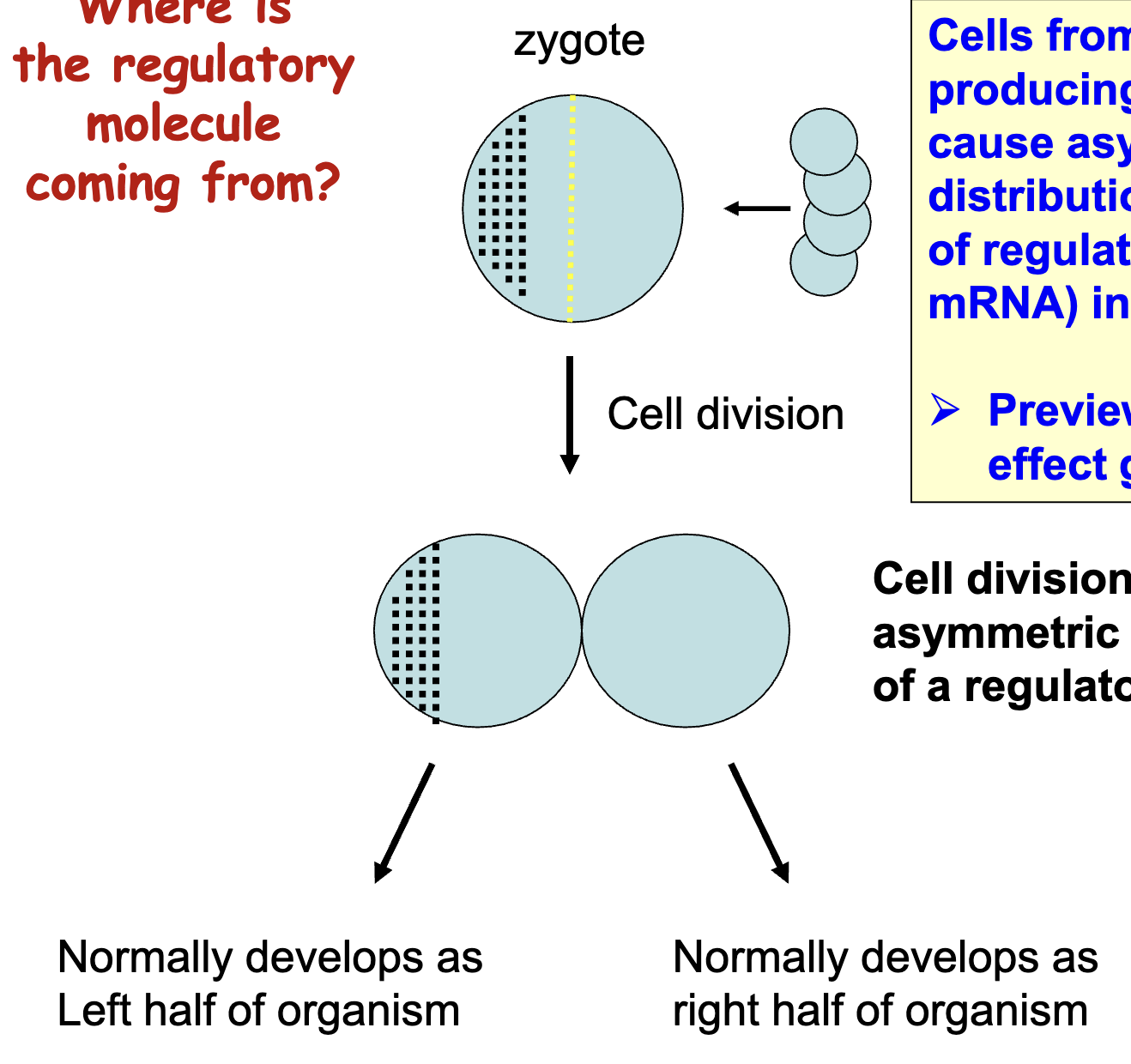
Where is the regulatory molecule coming from (Preview of maternal effect genes)
cell division results in an asymmetric distribution of a regulatory molecule
egg (zygote) cell is surrounded by other cells that deposit mRNA into egg (in specific locations)
asymmetric distribution gives positional info in zygote, which is maintained when cells start dividing
cells from female (egg-producing) parent cause asymmetric distributions of regulators (maternal mRNA in the egg), even before fertilization
Are Developmental Fates of Blastoderm Cells Fixed?
Cellular blastoderm exp: transfer blastoderm cells to larval abdomen
result: anterior cells develop as head and thoracic structures in the abdomen of the developing larva
we can’t tell if these cells are specified, determined or differenciated b/c of autonomous specification
autonomous specification: cell determinants already exist inside cells and doesn’t respond to outside signals (looks like determination b/c the cells will not respond to changes in position)
continuing to develop in the way they were supposed to
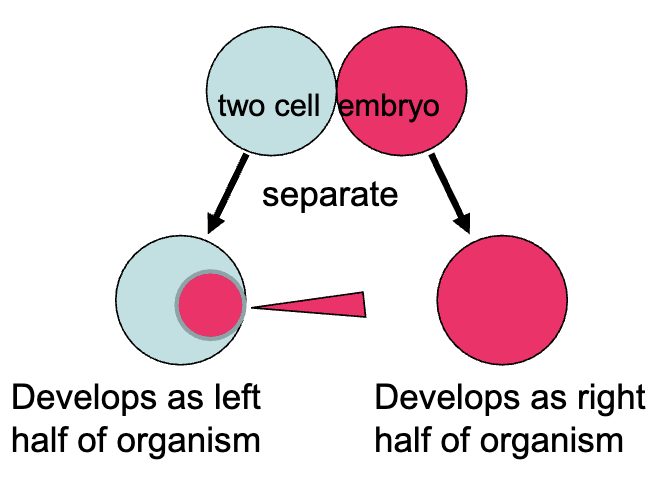
How could one tell if cells are autonomously specified or determined?
What if we were to inject cytoplasm from the right cell into left cell?
What would you predict if the cell was still specified?
the injected cell would develop as the right half of the organism
commitment to cell fate can still be reverse in a specified cell (it may not respond to outside signals but it would respond to changes in internal signals
if the injected cell was already determined, then it would continue to develop as the left half of the organism
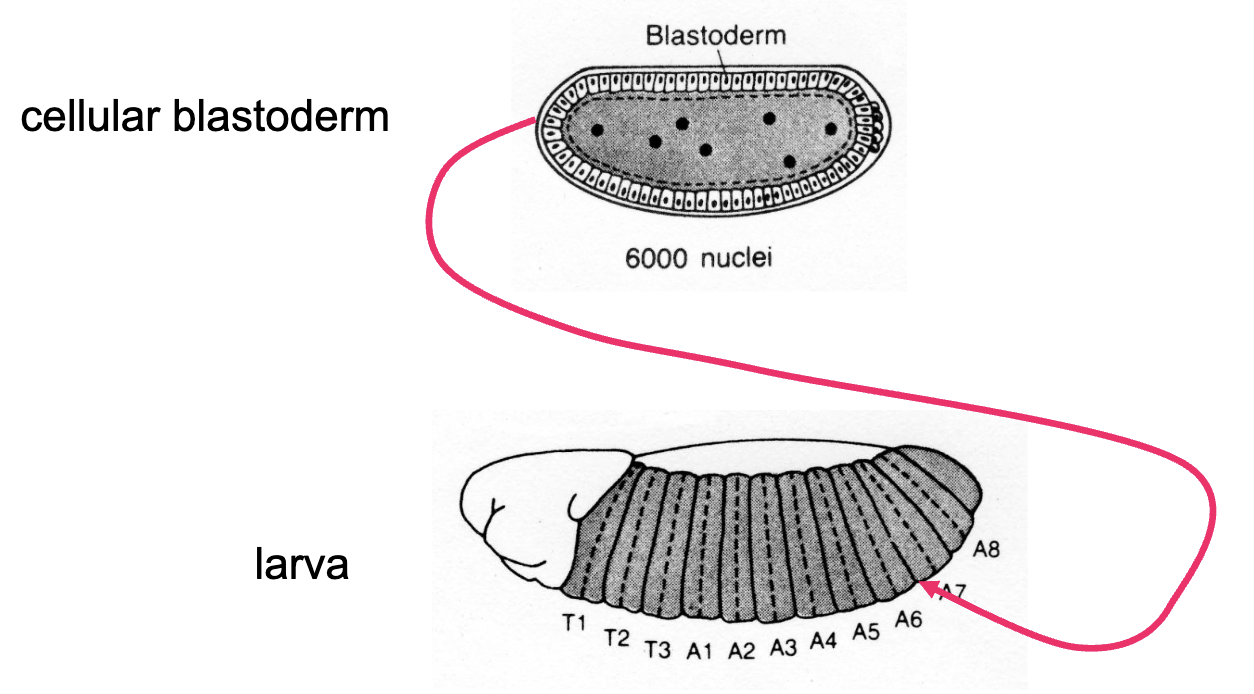
Blastoderm Cell Conclusion
researchers concluded that blastoderm cells are determined
if these cells are determined, if we were to inject cytoplasm from an anterior cell into a posterior cell, we would see:
that injected cells continue developing into posterior structures
posterior cells no longer responding to signals in anterior cytoplasm, no longer changing fate
the anterior-posterior and dorsal-ventral axes are already established
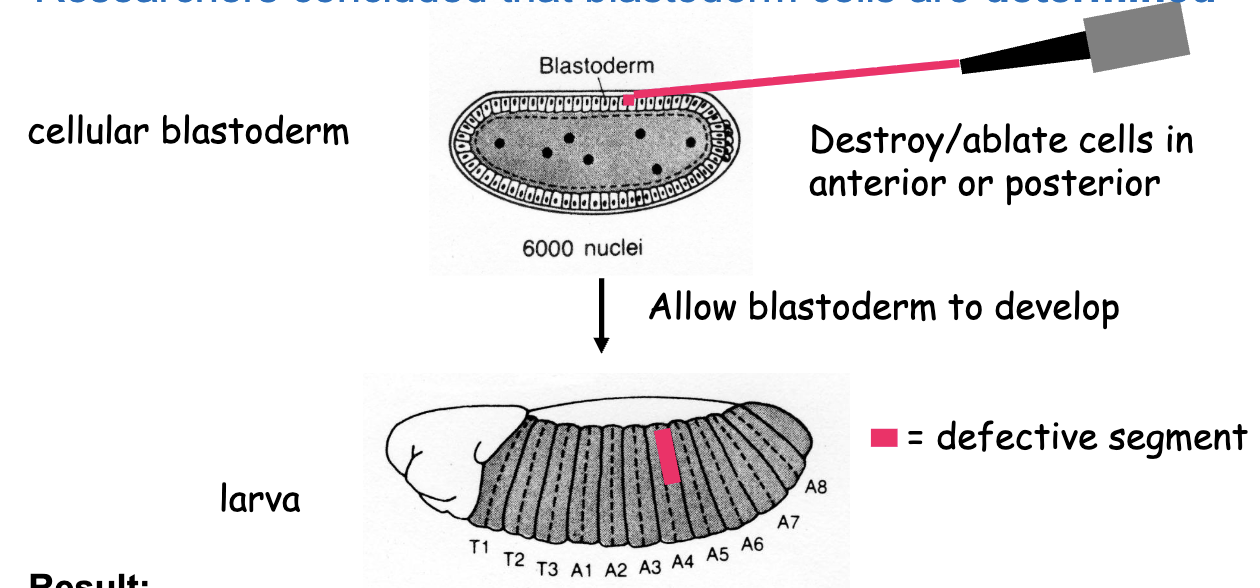
Ablation Experiment
destroy/ablate cells in anterior or posterior of cellular blastoderm →allow blastoderm to develop
results: anterior cell ablation results in a partly defective anterior segment
posterior cell ablation results in a partly defective posterior segment
a stripe of 4 blastoderm cells in width gives rise to 1 segment in the larva
conclusion: segmentation is already determined in the cellular blastoderm
Summary (so far)
during development, multicellular organisms must establish spatial and temporal cues using positional information and a cascade of master regulatory proteins
following fertilization in Drosophila, a series of embryonic nuclear divisions and migrations produce a syncytial blastoderm w/ nuclei lying close to the membrane
subsequently, nuclei are enclosed by membranes such that a blastoderm of 6000 cells results
morphogenesis is not apparent until much later during gastrulation
anterior-posterior fates and segmentation pattern are determined by the cellular blastoderm stage
What is the best approach to identifying the morphological determinants? Biochemical or Genetic
Genetic Approach: identify the genes required for morphogenesis
isolate mutants that are defective in morphogenesis
use the mutant phenotype to clone the gene required for morphogenesis
use the cloned gene to work out the function of the product, time and place of expression
Kruppel Gene Expected Phenotype Ratios
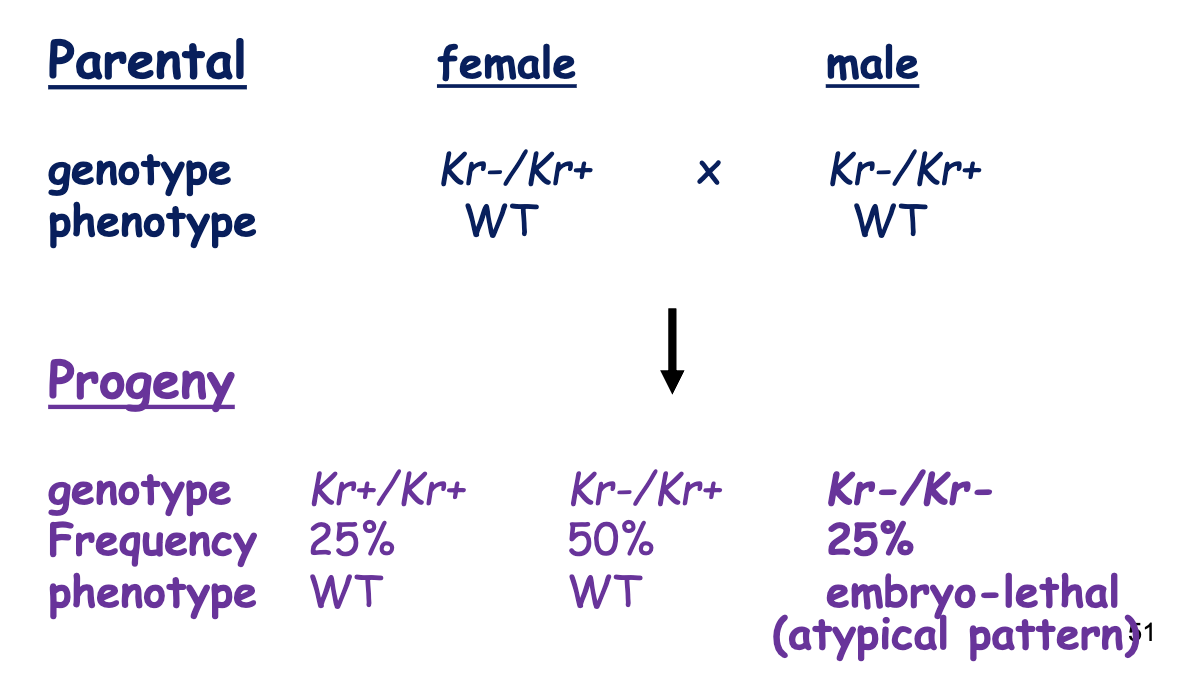
bcd Gene Phenotype Ratios: Heterozygous Female Parent
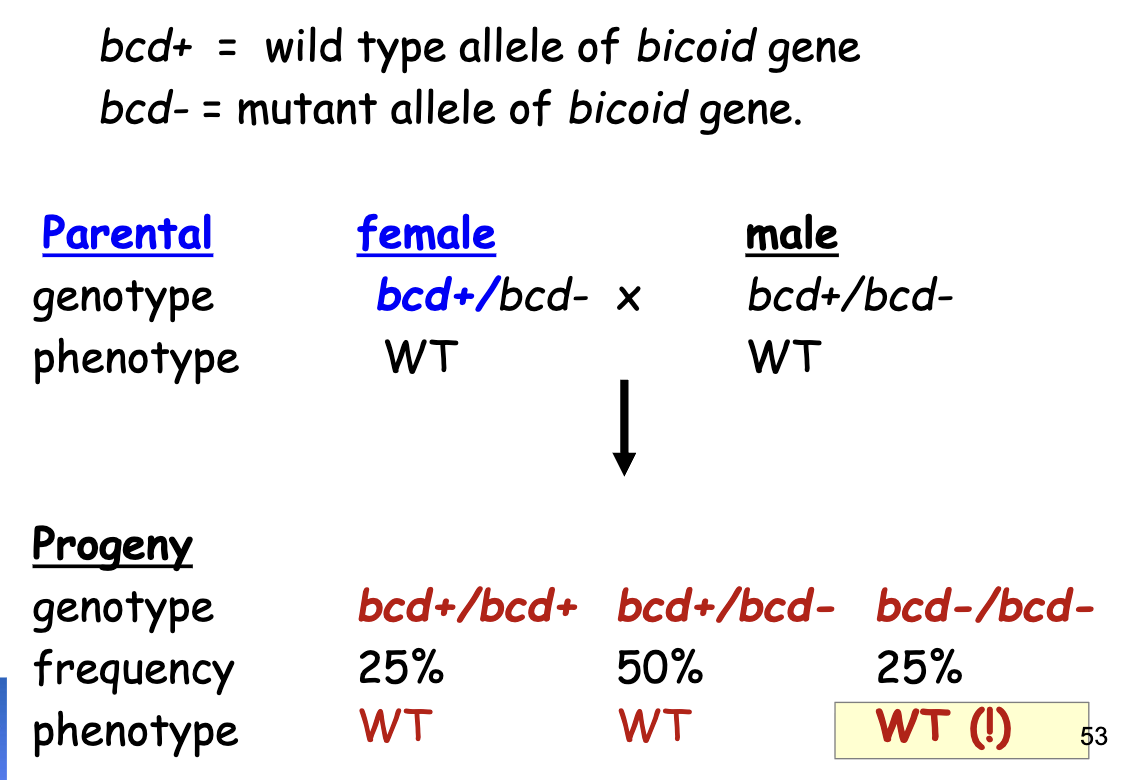
bcd Gene Phenotype Ratios: Homozygous Recessive (lf) female parent
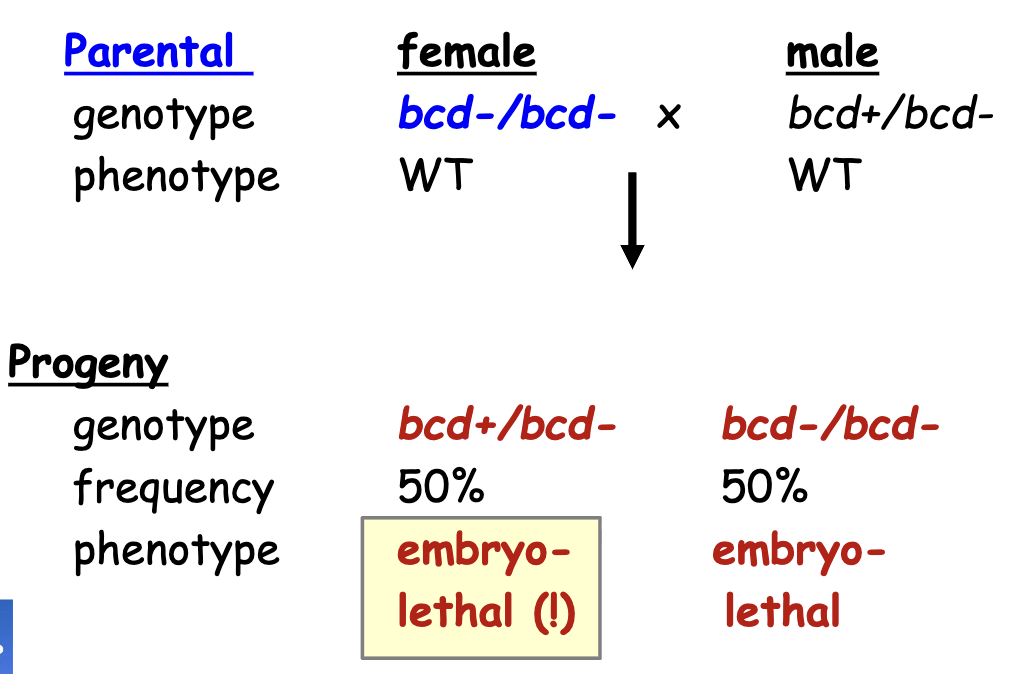
bcd Gene Phenotype Ratios: Heterozygous Female w/ homozygous recessive male
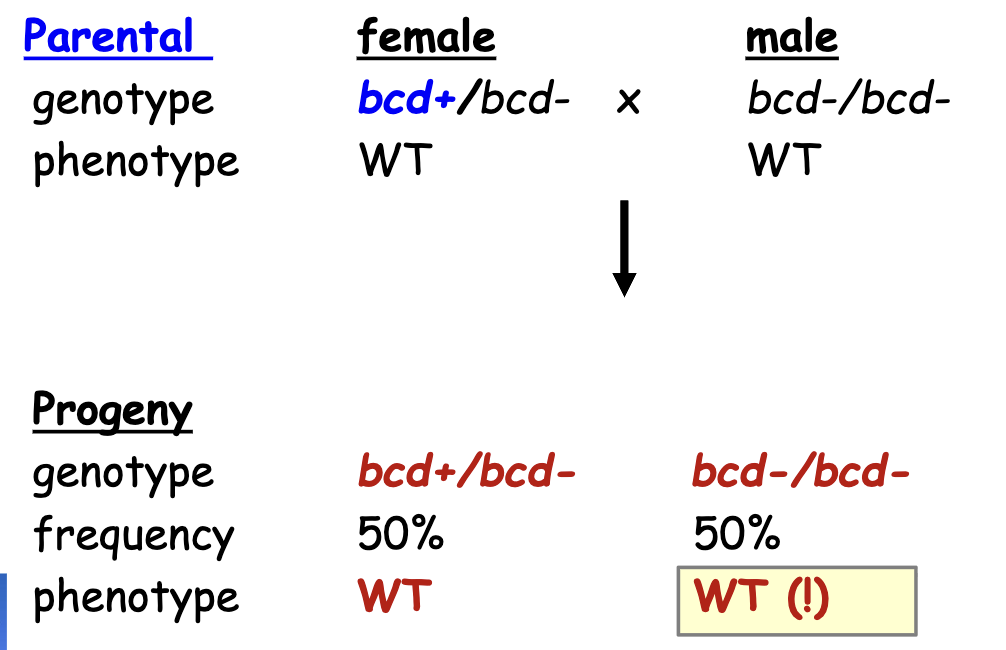
Explanation for Discrepancies in Genotype
Maternal Inheritance: as long as the female parent had a working copy of bcd, the progeny survived (even if progeny doesn’t have working copy of bcd)
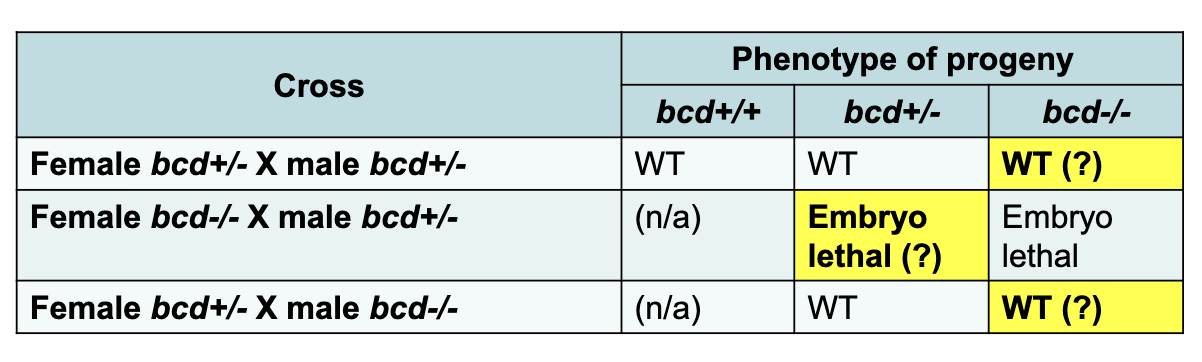
2 Classes of Embryo Lethal Patterning Mutants
zygotic: expected correlation of genotype and phenotype (e.g. Kruppel)
this pattern of inheritance implies that the product of the gene of interest is made in the developing embryo, to specify patterning for the developing embryo
the progeny has to have a copy of the working gene to be WT
the mutant gastrula is homozygous for a recessive lof allele of a gene
Maternal effect: phenotype of zygote influenced only by the female parent’s genotype, not the zygote’s genotype
female parent is depositing something in the egg that will be expressed in the zygote
this pattern of inheritance implies that the maternal tissue makes a product that is required by the embryo for patterning
eg. bicoid
Maternal Embryo-Lethal Patterning Mutants (More Detail)
all gastrula produced from a female heterozygous for a recessive, lof mutant allele of a maternal effect gene → develop normally regardless of the genotype of embryo
all gastrula produced by a female homozygous for a recessive lof mutant allele of a maternal effect gene have developmental defects in patterning regardless of the genotype of embryo
i.e the female parent needs to have one working allele for the embryo to develop normally. Genotype of the male parent and embryo does not matter for maternal effect genes
the female parent is expressing smth that is made available to the progeny
Gastrula
later stage embryo where the tissues have started to fold inwards
can start seeing what’s going to be the head and what’s going to be the abdomen
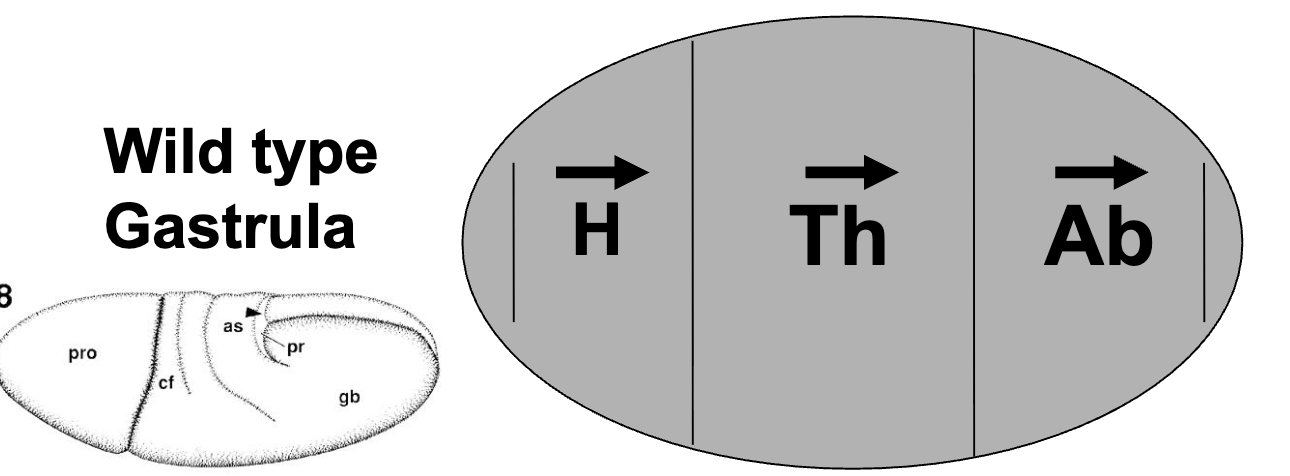
Maternal Effect Genes on Pattern Formation: Background
mutations in 30 different Drosophila genes result in maternal-effect patterning defects
of these mutants, 18 affect anterior-posterior patterning (the rest affect dorsal-ventral patterning)
most of the 18 mutant phenotypes fell into one of 2 broad classes: lacking anterior segments or lacking posterior segments
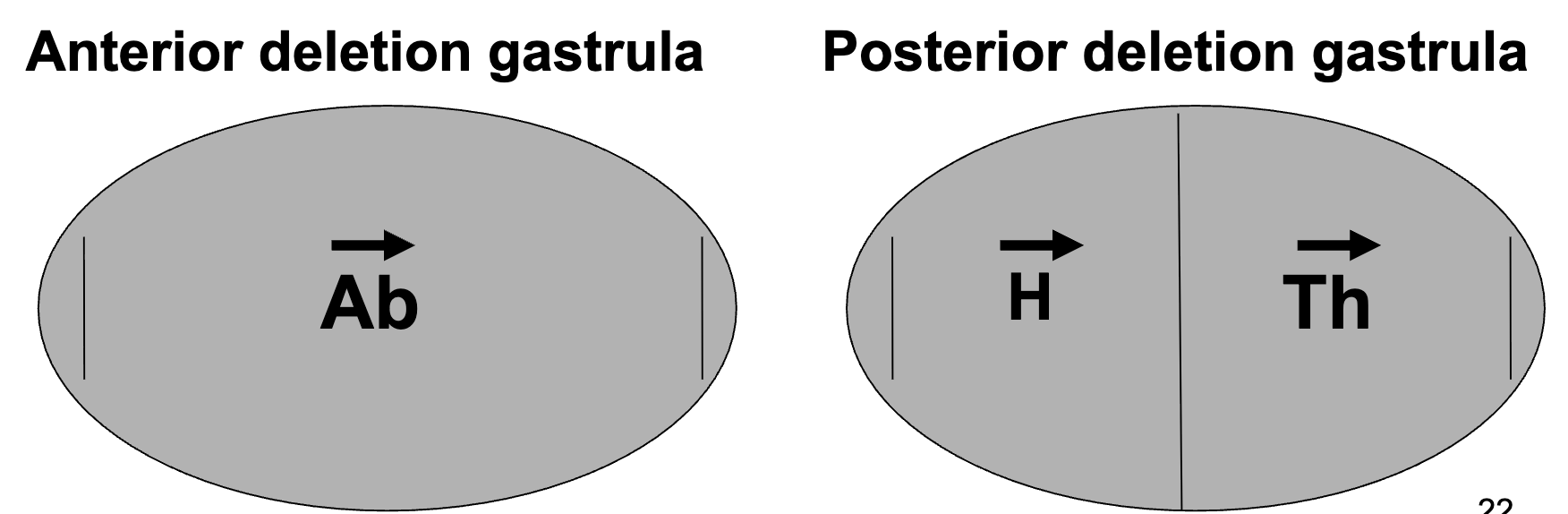
Maternal Effect Genes on Pattern Formation: Gene Localization
Anterior Deletion Phenotype: bicoid (bcd-), hunchback (hb-)
these proteins are typically found in anterior end → lose functions → lose anterior
Posterior Deletion Phenotype: nanos (nos-), caudal (cad-)
these proteins are typically found in posterior end
Maternal Effect Genes: Morphogens
maternal effect genes must code for morphogens
Morphogens:
molecules that can induce the acquisition of diff cell fates, on the basis of the concentration of molecule to which a cell is exposed
a gradient of morphogen across an embryo or developing tissue provides positional information for the cells along the gradient → cells develop accordingly
usually produced at some distance from the targets
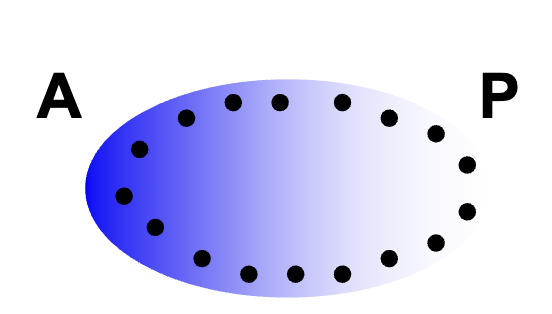
Maternal Effect Gemes: lof phenotypes
e.g. bicoid (bcd), hunchback (hb), nanos (nos), caudal (cad)
all code for proteins w/ morphogen function
likely were the cytoplasmic protein determinants in the cytoplasm exchange exp
several lines of evidence suggested that these 4 genes identified by mutation (2 anterior, 2 posterior), encode morphogens and could be the cytoplasmic elements present in the zygote identified by developmental experiments
Bicoid+ gene product
the bicoid+ gene product acts like a morphogen
the bicoid- embryo lacks head structures (all abdomen); didn’t have bcd at the critical time
cytoplasmic transplant of bicoid+ to bicoid- anterior partially rescues the phenotype
a cytoplasmic transplant to the middle would result in head and thoracic segments in the middle
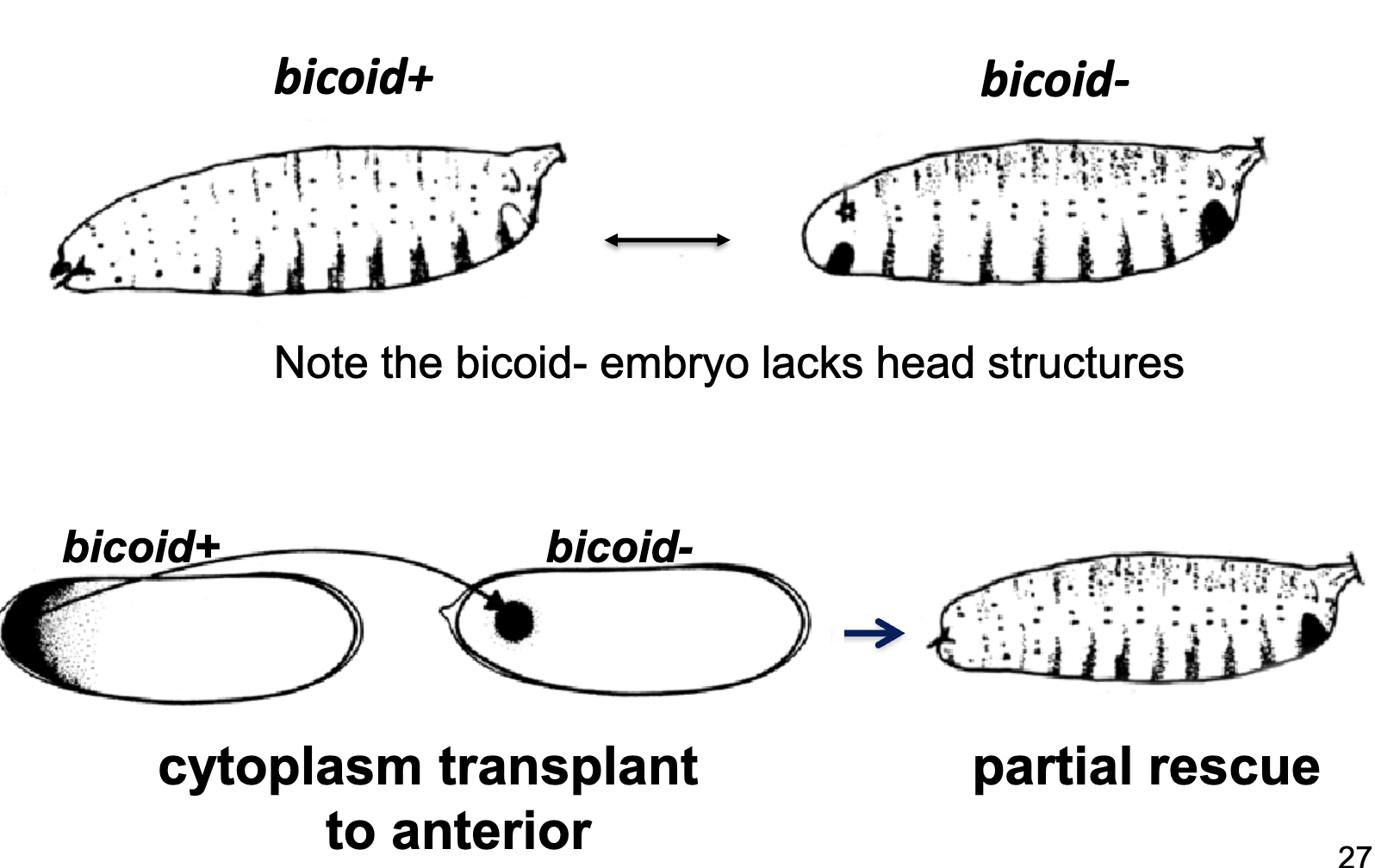
Transplantation Experiment Summary
transplanting cytoplasm from the anterior of a bicoid+ embryo (but not bicoid- embryo) to a bicoid- embryo can partially rescue the bicoid- phenotype
therefore, cytoplasmic determinants are missing in a bicoid- embryo
if the bicoid+ cytoplasm is delivered to the centre of a bicoid- embryo, the head/thoracic segments are induced in both directions from the site of injection, suggesting a morphogen-like effect
Gene Dosage Experiments
gene dosage corresponds to amount of mRNA deposited and amount of Bicoid protein produced in zygote
changing the number of copies of the bicoid gene in the female parent changes the relative position at which the head furrow is formed in the gastrula
an increase in copy number moves the furrow towards the posterior, a decrease towards the anterior
more copies = more translation of bcd
therefore the amount of bicoid gene product influences the exact positioning of anterior elements
the more bcd copies the female parent has, the more mRNA gets made → the more mRNA gets deposited in oocyte before fertilization
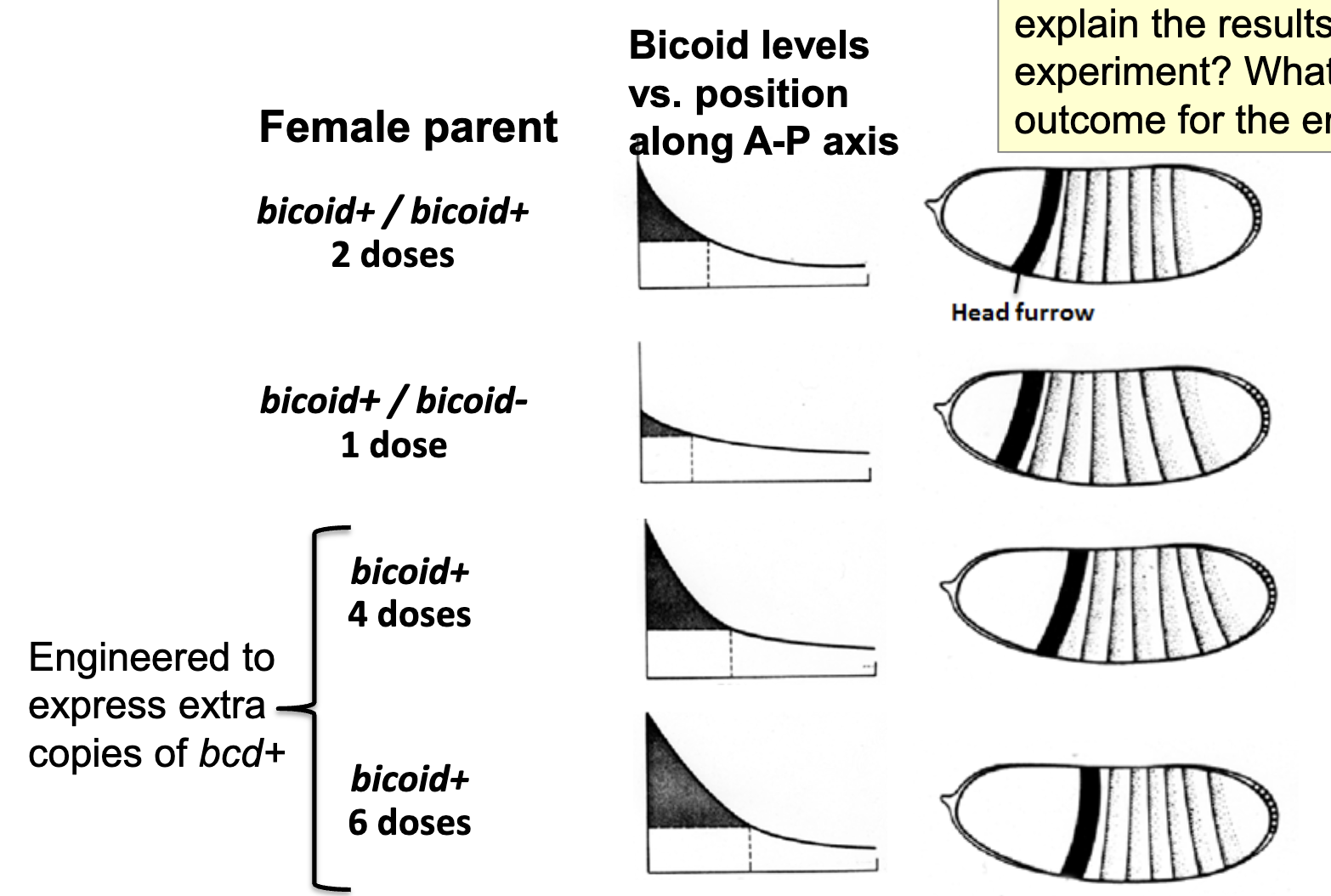
Gene Dosage Experiments: Another Figure
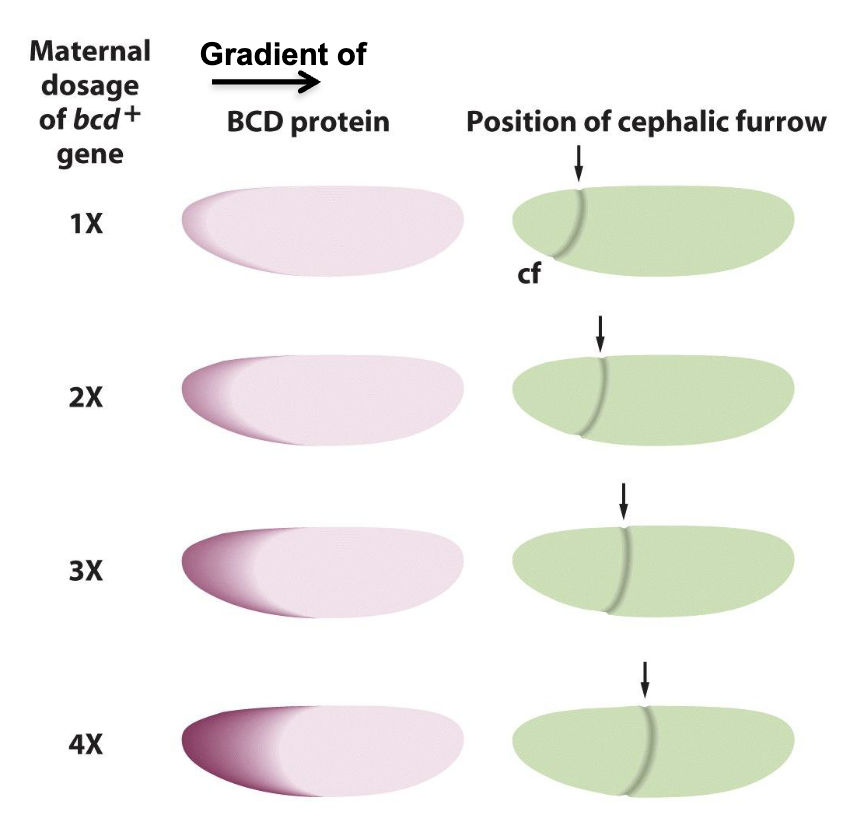
Can Transcription Factors function as Morphogens
Yes! it acts in a gradient and is a molecule that directs cell fate
transcription factors could act as morphogens by activating target genes in diff nuclei in a concentration-dependent manner
the bicoid+ gene encodes a transcription factor; conc. of bcd transcription factor influences whether target gene is expressed and to what level
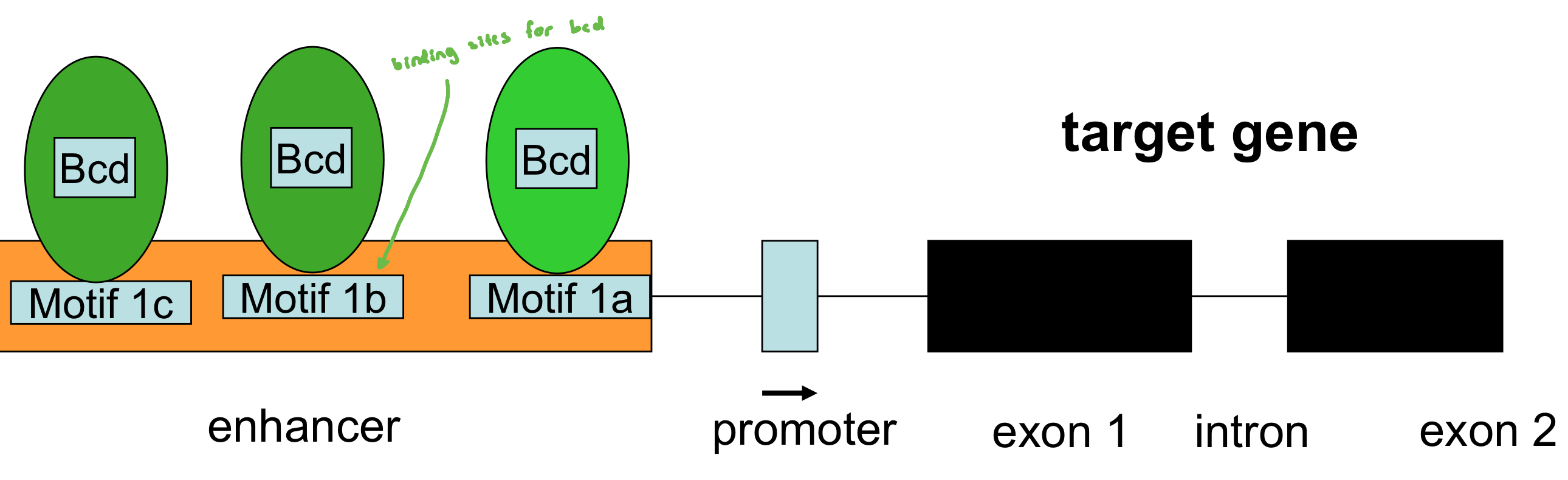
bcd, hb, cad, nos: Transcription Factor or RNA Binding protein
bcd = transcription factor
hb = transcription factor
cad = transcription factor
nos = RNA binding protein
Location of Protein in Syncytial Embryo
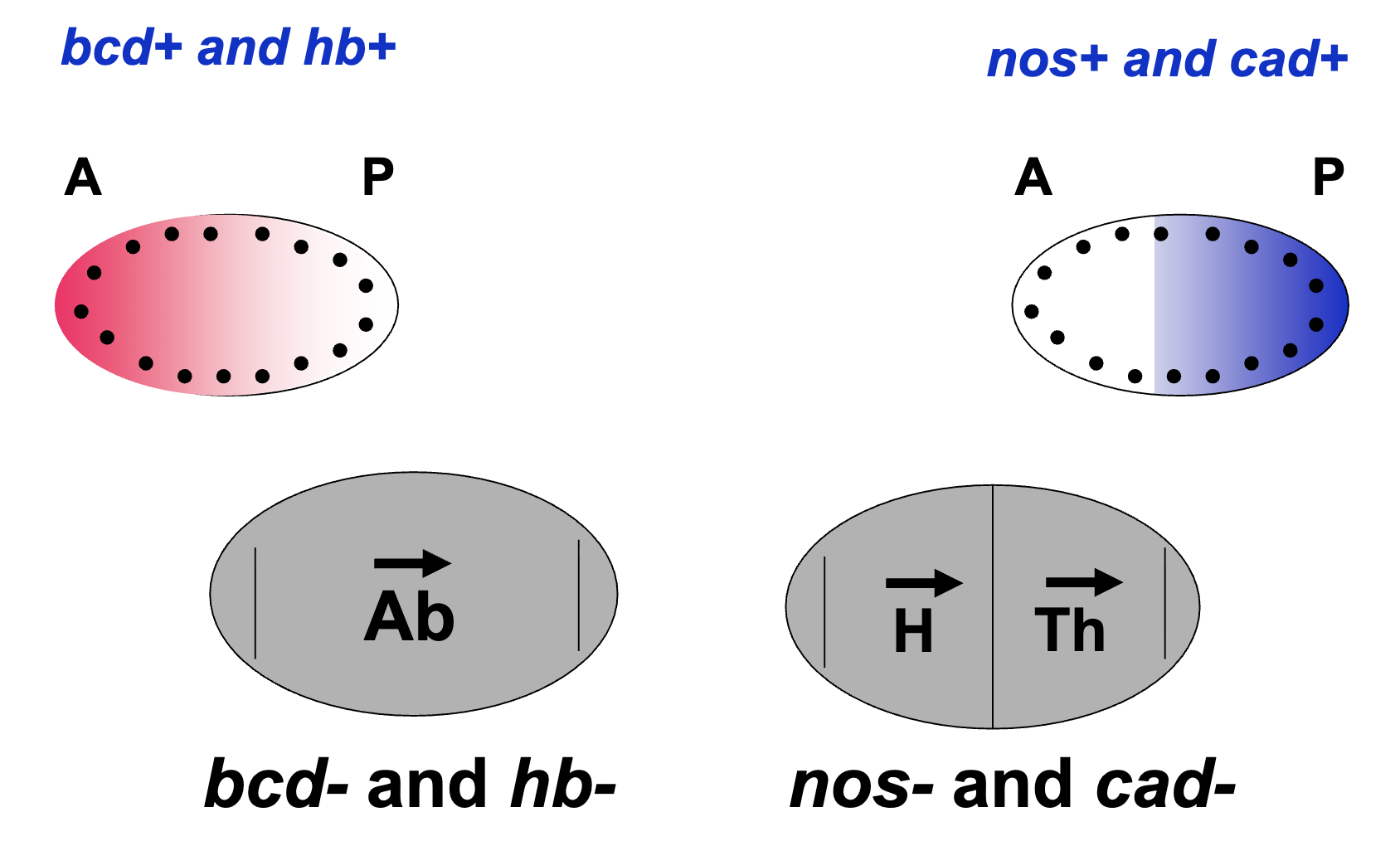
gradient of bcd and hb protein is highest at the anterior
bcd- and hb- embryos lack head and thorax structures
gradient of nos and cad protein is highest at the posterior pole
nos- and cad- embryos lack abdominal structures
the localization of these 4 proteins correlate with the phenotypes of the mutant
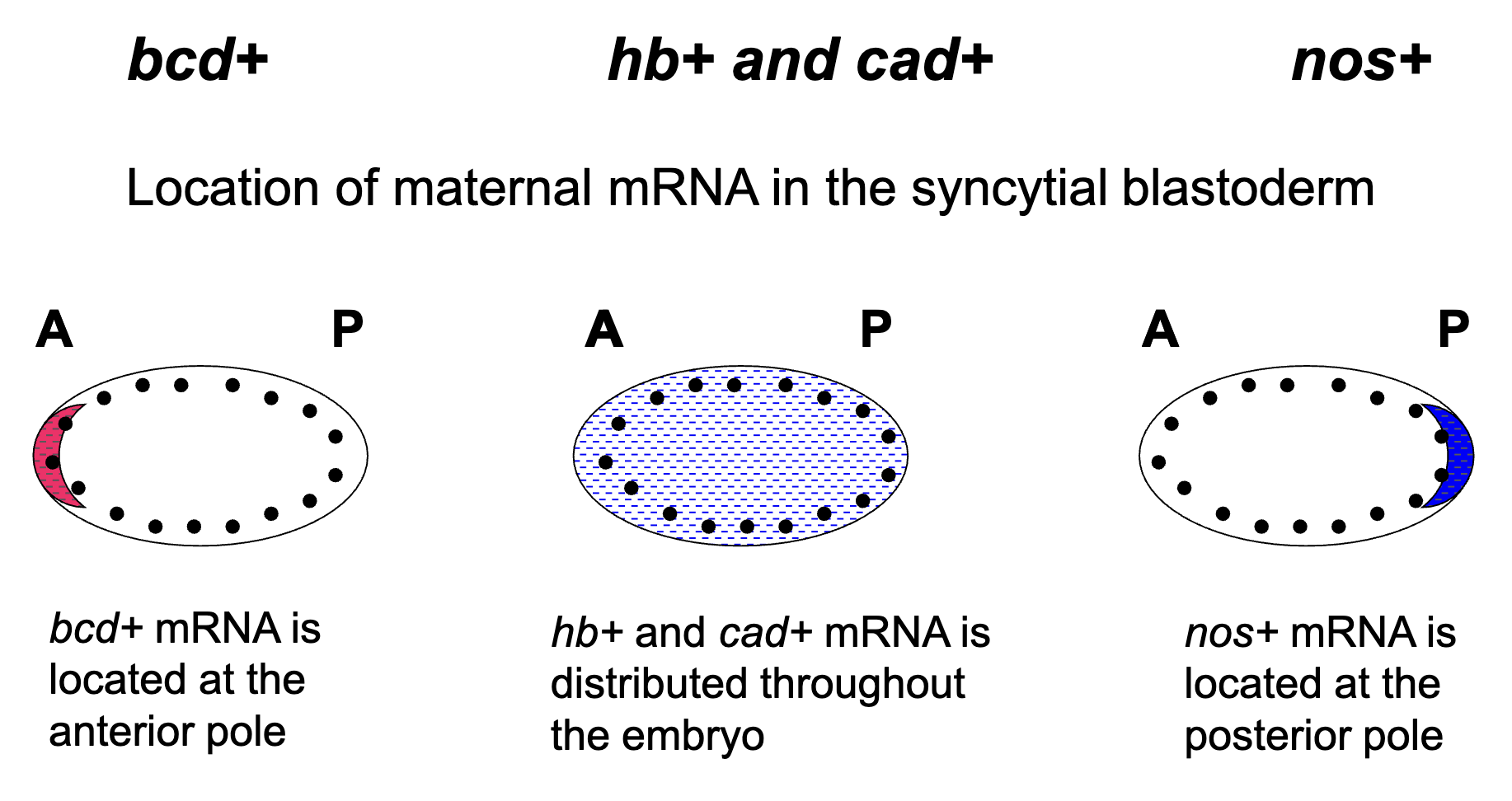
Location of the Maternally Synthesized mRNA
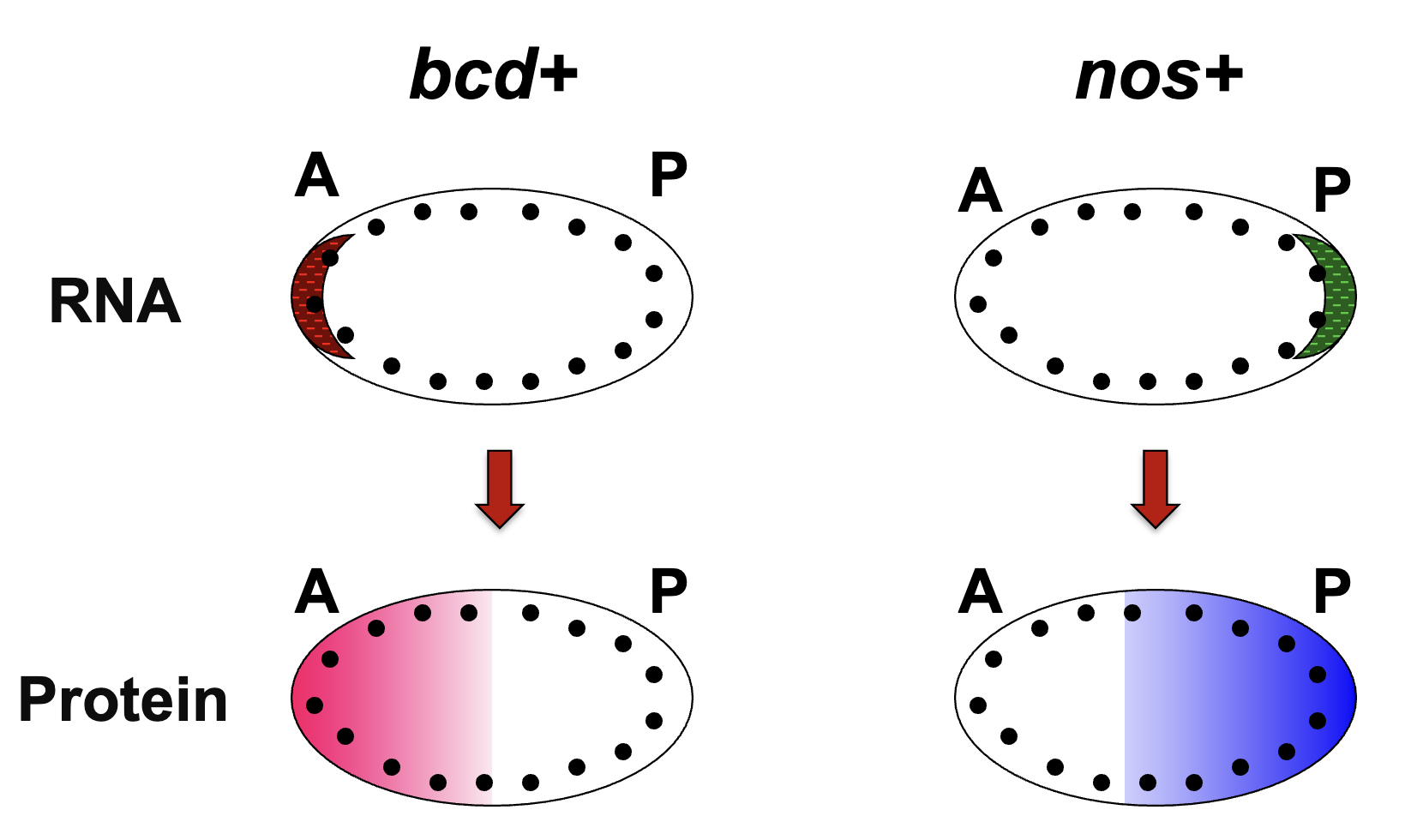
How are the protein gradients established?
bcd+ and nos+ protein gradients are established thru diffusion from site of translation
the syncytial blastoderm stage of Drosophila embryogenesis allows gradients to form (stage of many nuclear divisions w/o cytokinesis)
nuclei are being exposed to diff concs of bcd and nos
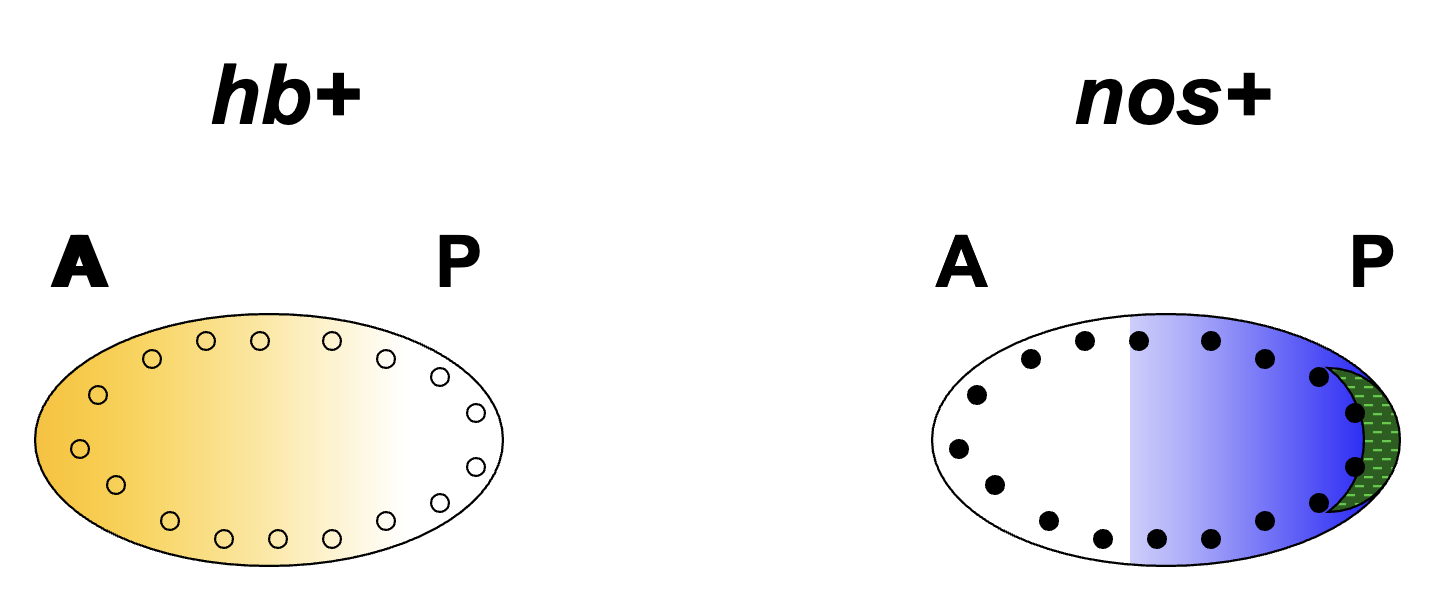
hb+ and nos+ mRNA Protein Gradients
location of hb+ protein in the syncytial blastoderm is regulated by nos+ protein
hb+ mRNA is located throughout the embryo
nos+ protein binds and inhibits translation of hb+ mRNA (hb+ is a target of nos+)
thus hb+ mRNA is translated only in the anterior blastoderm and repressed in the posterior
nos+ protein is located in the posterior part of the blastoderm
bcd+ and cad+ protein gradients
bcd+ protein is located in the anterior part of the blastoderm (dual functions, transcription factor and binds Cad+)
cad+ mRNA is locaed thoghout the embryo
bcd+ protein binds and inhibits translation of cad+ mRNA
Summary of mRNA and protein gradients in syncytial blastoderm
bcd+ and nos+ protein gradients are established through diffusion from that site of translation
establishment of the hb+ and cad+ protein gradients are more complex:
the nos RNA binding protein binds to the hb+ RNA and inhibits its translation in a conc-dependent manner (also inhibits bcd+ mRNA translation)
the bcd+ protein can bind cad+ mRNA and inhibits its translation in a conc. dependent mannger
the gradients of bcd+ and nos+ proteins establish the gradients of hb+ and cad+ proteins
How does bcd+ and nos+ mRNA get localized to the anterior/posterior poles of the zygote?
the oocyte already has polarity (i.e. already has an established anterior-posterior axis)
the polarity of the oocyte is set up by the surrounding environment and is used to direct the orientation of the microtubules
the orientation of the microtubules is used to direct bcd+ and nos+ mRNA to one end or the other
Microtubule orientation in the oocyte is guided by cell-cell interactions between the oocyte and the asymmetric cellular environment of the follicle
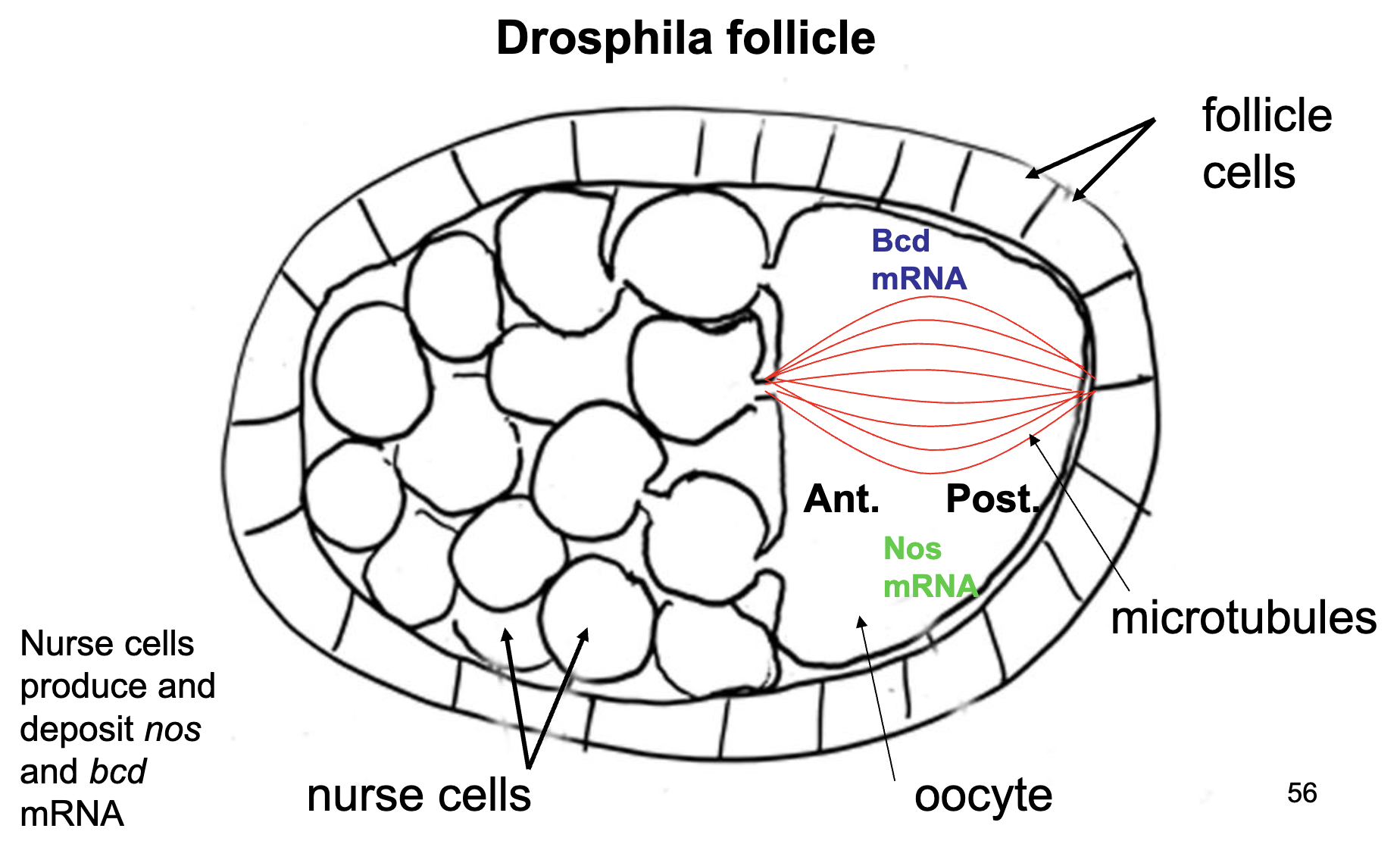
Nurse Cells
nurse cells deposit bcd+ and nos+ mRNA at the poles of the oocyte
bcd+ mRNA is anchored to the cytoskeleton at the anterior end of the oocyte
nos+ mRNA is anchored to the cytoskeleton at the posterior end of the oocyte
when they are translated, their respective proteins diffuse out from the mRNA
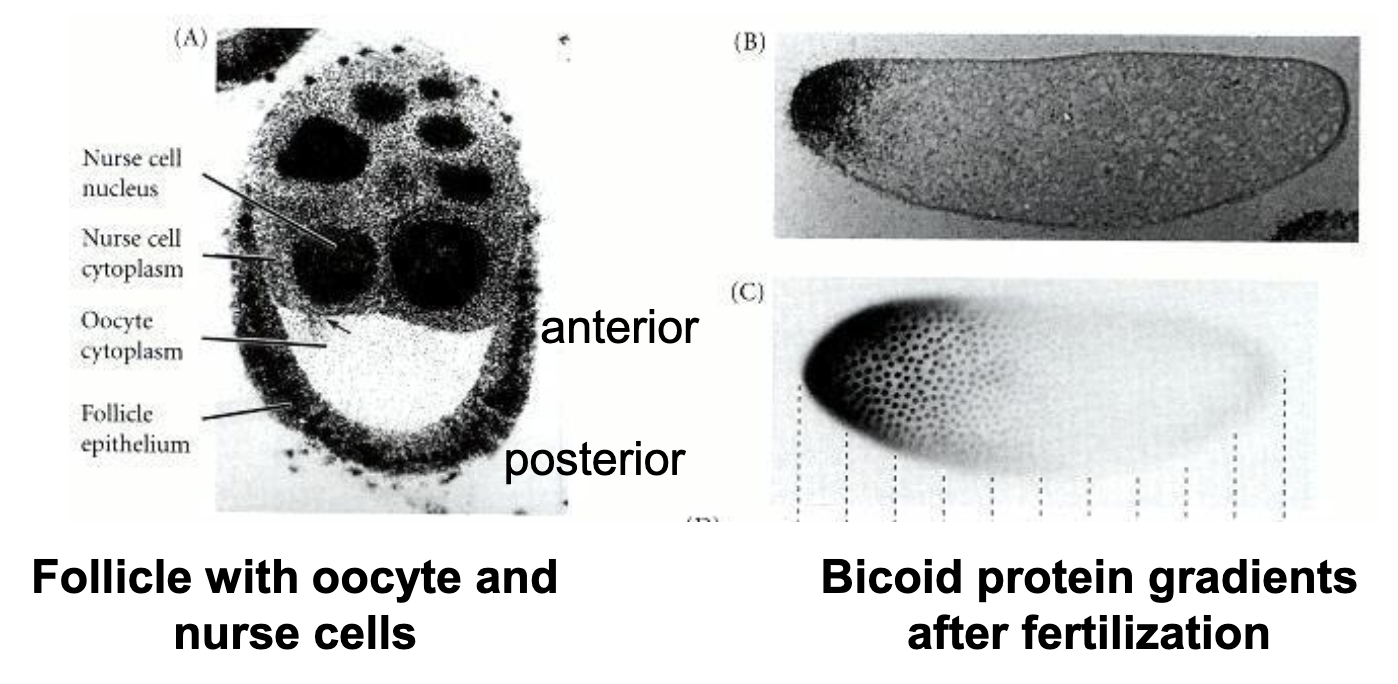
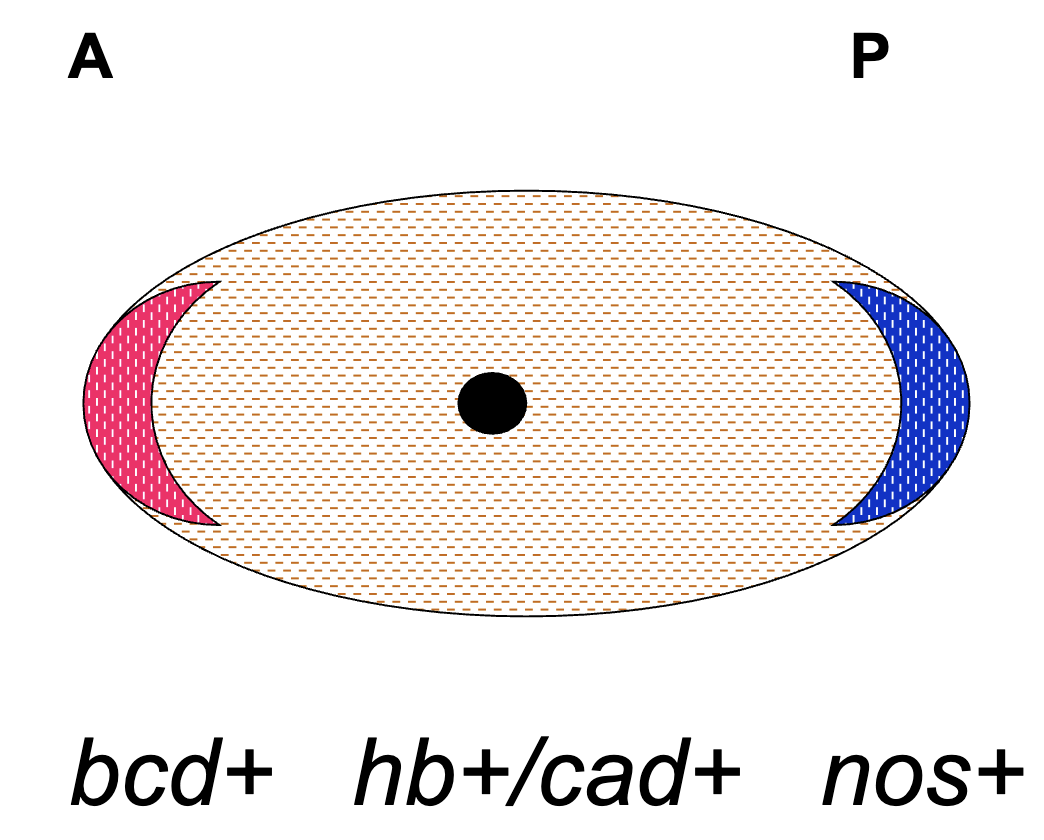
Zygote at Fertilization (egg)
bcd+ mRNA anchored to anterior end
nos+ mRNA anchored to posterior end
hb+ and cas+ mRNA present throughout
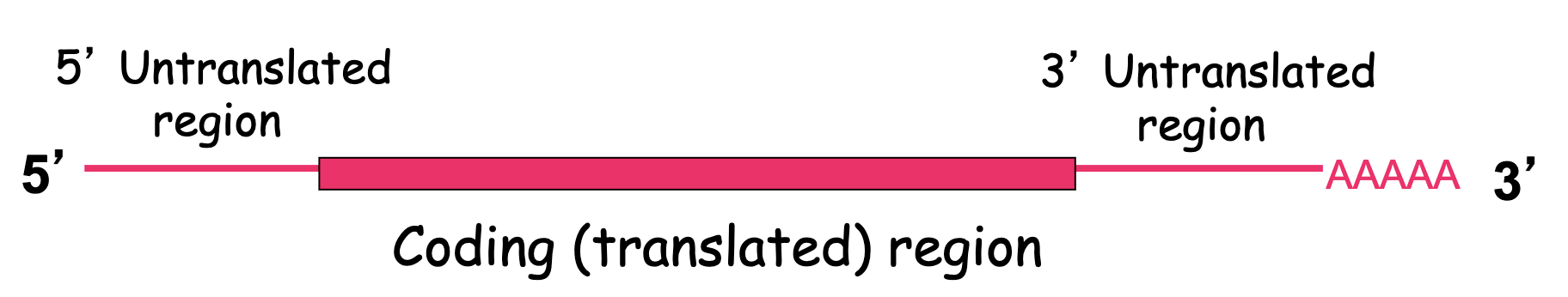
How do bcd mRNAs get deposited to the anterior end of the cytoskeleton?
3’UTR regions
they must contain some sort of regulatory sequence that is recognized by another factor that helps bind this region
3’UTR sequence recognized by RNA binding protein that attaches to the microtubule and moves the mRNA to anterior pole
3’UTR of bcd+ and nos+ mRNAs have cis-acting sequences
Chimeric nos gene w/ bcd 3’UTR Experiment
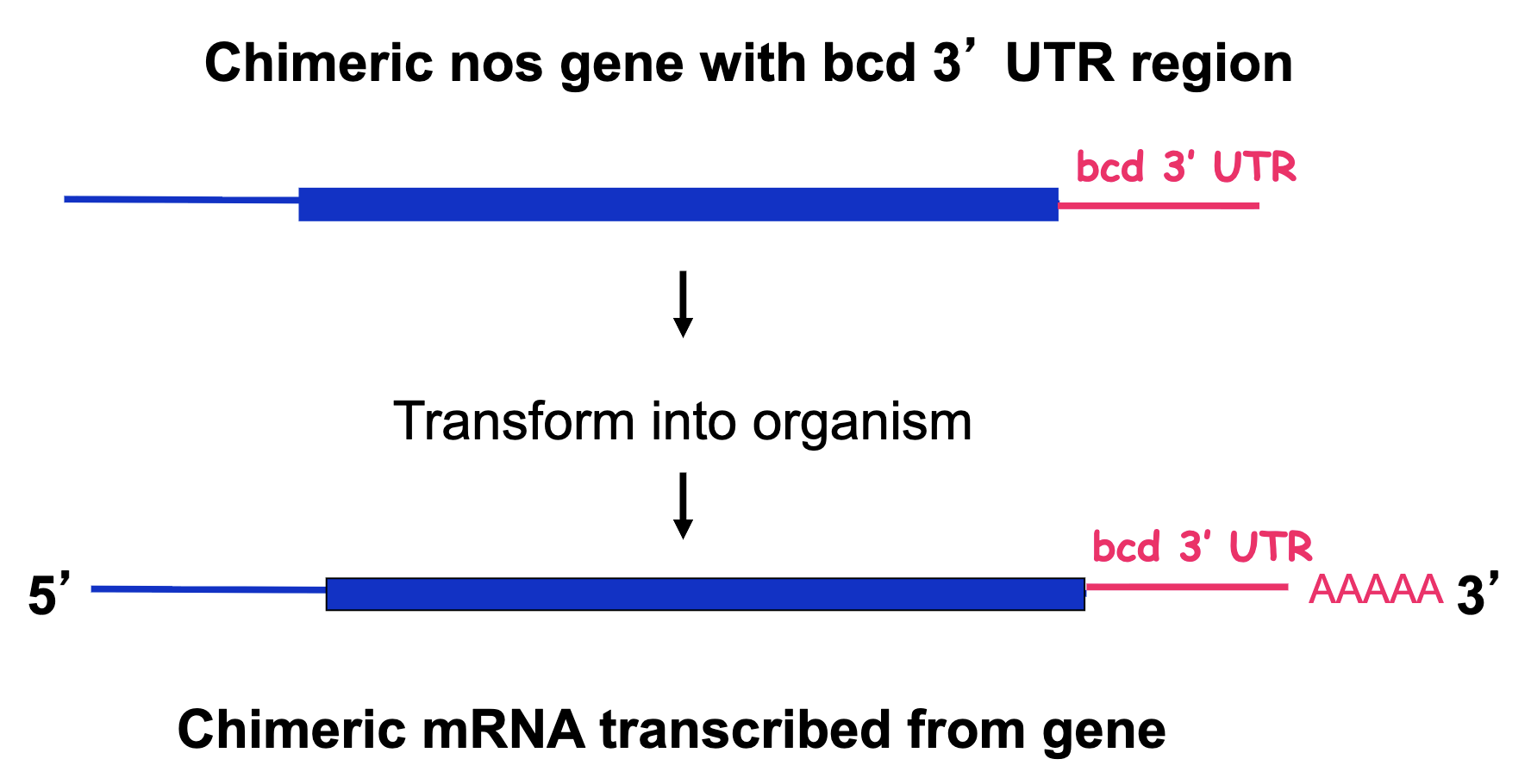
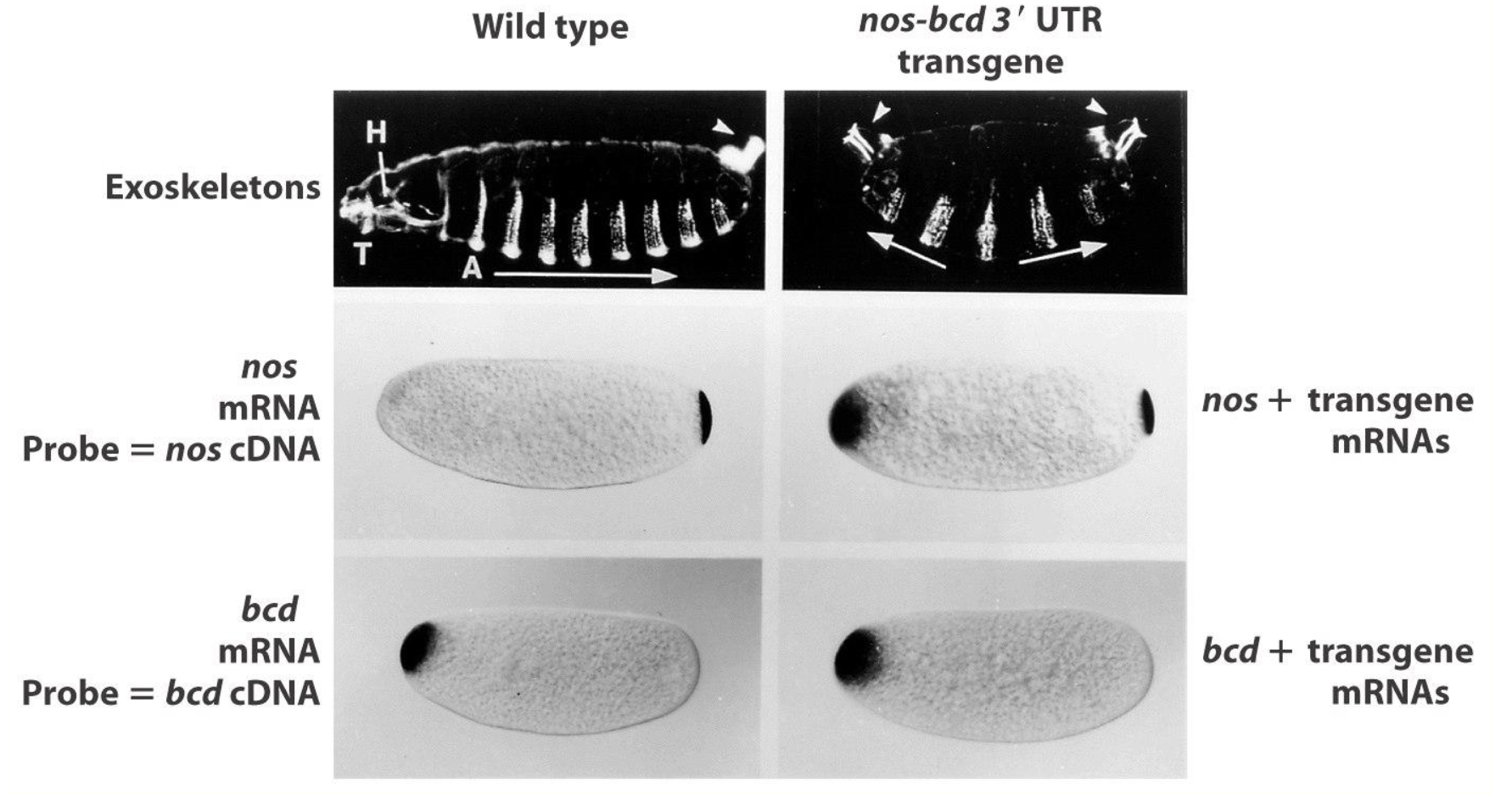
Chimeric nos gene w/ bcd 3’UTR Experiment: Results
we see nos localized to both ends
nos is suppressing the effect of bcd, resulting in posterior structures in the anterior → nos protein suppresses translation of bcd mRNA
Zygote Figure of nos/bcd chimeric mRNA
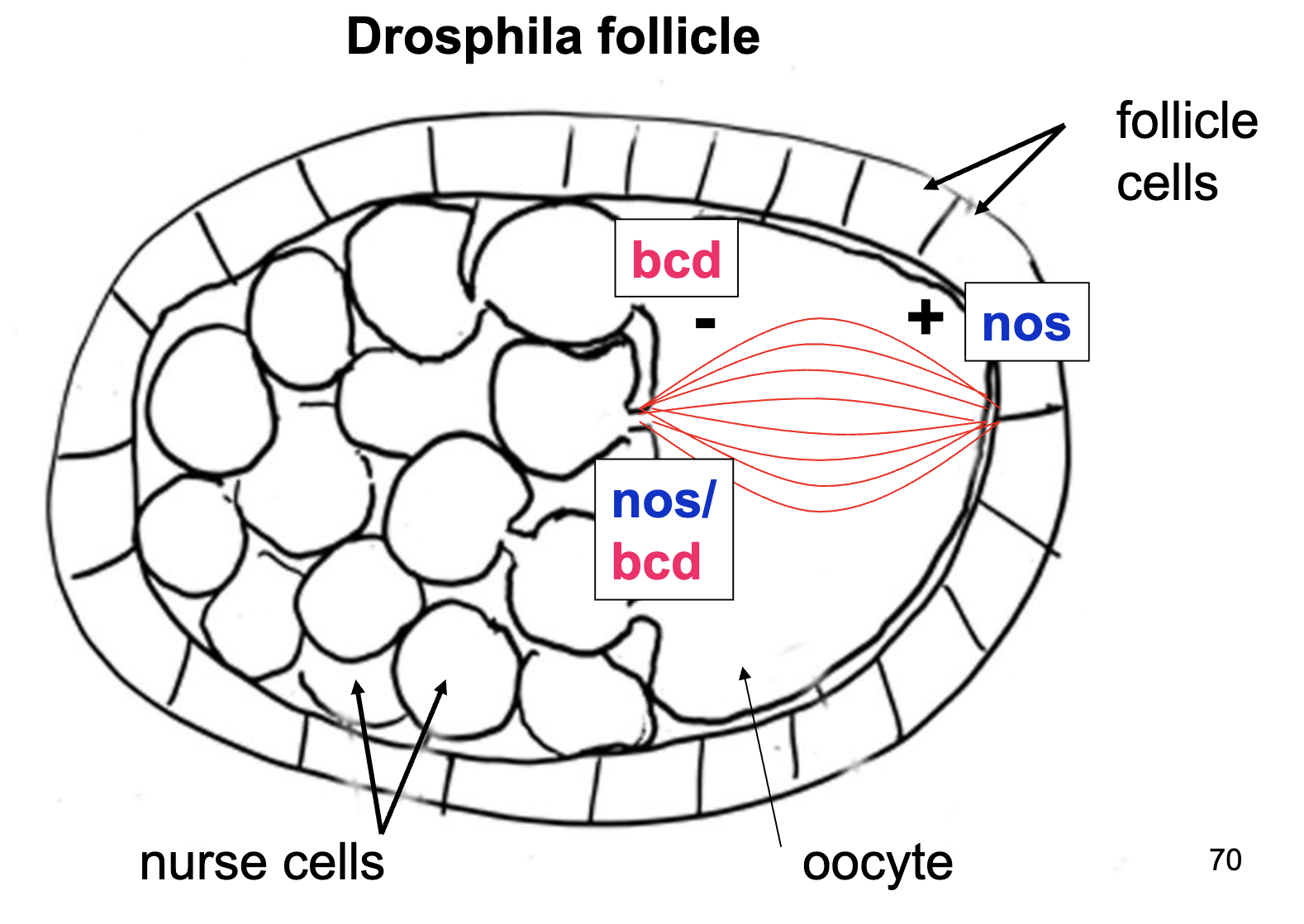
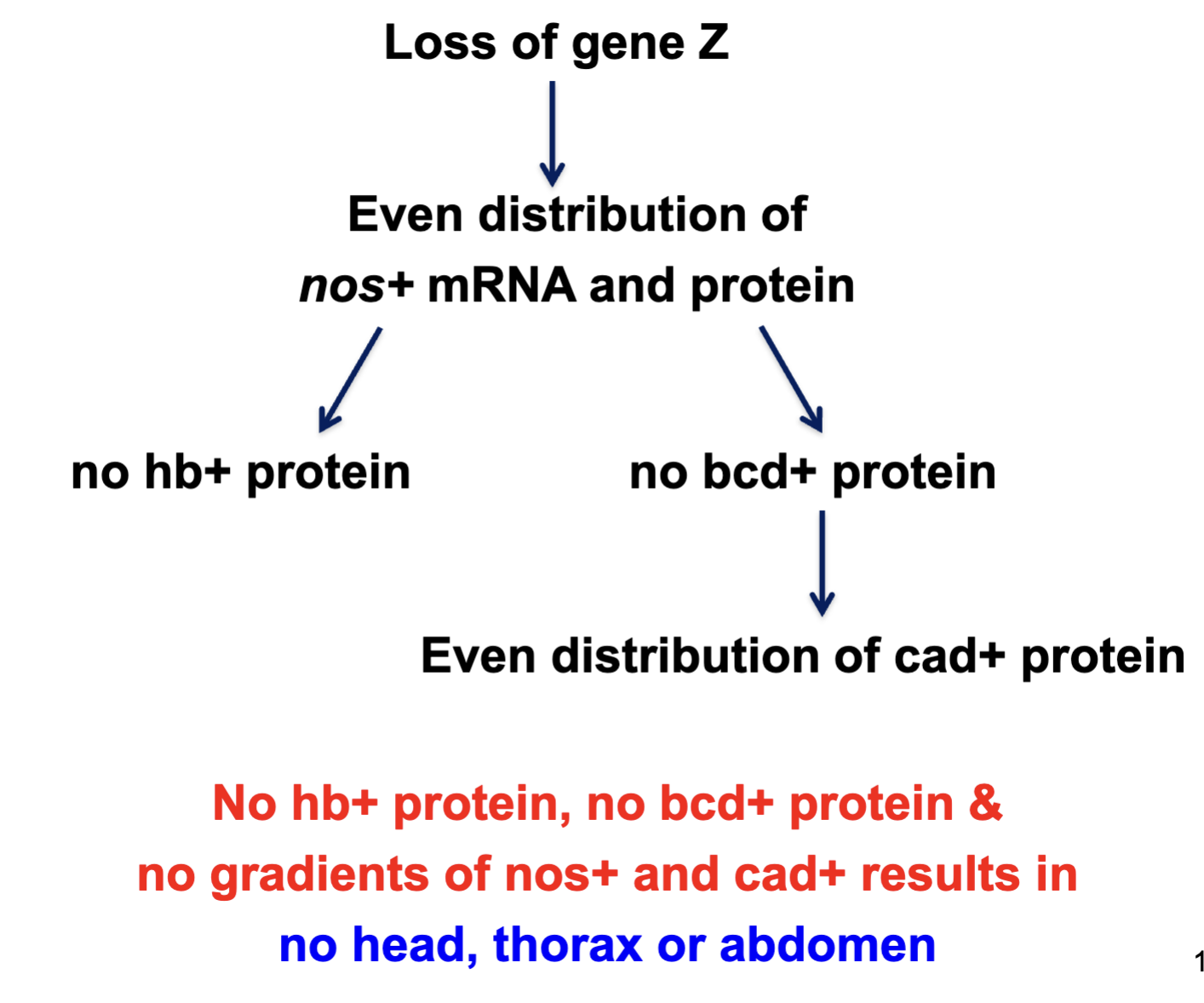
A specific protein (Z) is required to bind nos mRNA and localize it to the posterior pole. What kind of gene codes for Z?
Maternal effect gene
This protein is responsible for positioning nos mRNA in the oocyte, before fertilization. Thus it must be expressed by the female, egg-producing parent.
A mutant homozygous for a loss-of-function mutation in the gene encoding Z would be expected to what phenotype>
No head, thorax or abdomen
