McSharry Intro to Bio Test #3
1/90
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
91 Terms
Universal Steps of Cell Division (4)
Cell Division Signals
DNA Replication
DNA Segregation
Cytokinesis
Binary Fission
Form of cell division specific to prokaryotes, usually caused by external factors.
Prokaryote Cell Division Causes
Favorable environmental conditions like nutrient availability, proper temperature, and pH that lead to certain organisms dividing.
These organisms will most likely divide when conditions are met.
Eukaryote Cell Division Causes
Favorable environmental and internal conditions (division signals) that lead to certain organisms dividing.
Even with correct conditions met, organisms may still not divide.
Phases of Eukaryotic Cell Cycle
In order:
Gap/Growth 1 Phase (G1)
Either
Gap/Growth 0 Phase (G0)
Synthesis Phase (S)
Gap/Growth 2 Phase (G2)
Mitosis Phase (M)
Repeat
G0 Phase
Stage where cells that don’t commit to division rest, perform functions, and wait for signals to divide.
Cells may enter this phase from G1.
G1 Phase
Initial, active growth stage of interphase in cell cycle where cell prepares for DNA synthesis.
Cells can enter G0 from this stage.
After mitosis (M) and before DNA synthesis (S).
S Phase (DNA Synthesis)
Stage of interphase in the cell cycle where cell replicates its genetic information.
After G1 and Before G2.
Theta Replication
Process of chromosome duplication in prokaryotes.
Division of chromosome starts at the point of origin (Ori).
Named for the resemblance to the greek letter during the process.
Point of Origin (Ori)
The place on a prokaryotic chromosome where theta replication starts.
G2 Phase
Final, crucial growth stage of interphase in eukaryotic cell cycle.
Focus on rapid cell growth, protein synthesis, and organelle replication before division.
Also checks for DNA damage
M Phase (Mitosis)
Stage of the cell cycle where cell’s nucleus divides once, resulting in two genetically identical nuclei.
Characteristic of somatic cell division.
After telophase and before cytokinesis.
Cell Cycle
Process that only happens in eukaryote kingdom of organisms where they grow, replicate DNA, and divide.
Interphase
Crucial preparatory period between cell divisions consisting of G1, S, and G2 phases (NOT MITOSIS).
Cyclin Dependent Kinase (CDK)
Proteins responsible for controlling progress of the cell cycle.
Phosphorylates target proteins to activate or deactivate them.
Different kinds for different parts of cell cycle, each one is activated by specific cyclin.
Cyclin
Proteins responsible for activating CDKs.
Each one binds to a specific CDK to activate it.
Degrade over time and must be replaced.
Cell Cycle Checkpoint
Surveillance mechanisms that pause the cycle at G1, G2, and M to monitor DNA integrity, chromosome alignment, and cell size to ensure accurate division.
Ex: Rb protein inhibits cycle progress at G1 checkpoint
Retinoblastoma (Rb) Function
Tumor supression protein which inhibits uncontrolled cell division at G1 checkpoint.
Binds and inhibits E2F transcription factors until becomes phosphorylated and detaches allowing progress past G1.
Gene
A specific sequence of bases in a DNA molecule which codes for a specific polypeptide (protein).
Genome
Complete set of DNA in the cells of an organism.
Chromatin
Structure found in the nucleus made of DNA and histone proteins used to wrap DNA compactly.
Organized into more compact structures to create chromosomes.
Chromosome
Most compact form of DNA found in organisms.
Made of one very long DNA strand.
Homologous
Describing 2 or more chromosomes which contain different alelles of the same gene.
Have same size and same centromere placement.
Ex: Diploid cell containing one of each, ‘A’ and ‘a’ chromosomes.
Haploid
Describing a cell/nucleus having only one set of each chromosome.
Diploid
Describing a cell/nucleus having two complete sets of each chromosome, usually coming from each parent.
Sister Chromatid
Two identical copies of single replicated chromosome, formed during the S phase of cell cycle, and joined together by a centromere.
Centromere
Constricted point of chromosome where sister chromatids are connected with cohesin protein.
Binding site for spindle microtubules during cell division.
Allele
1 or 2 or more alternative forms of a gene that arise by mutation and are found at the same place on a chromosome.
Ex: ‘A’ and ‘a’
Mitosis
Process which makes 2 haploid daughter cells that are genetically identical to each other and the parent cell.
Made up of 5 different stages.
Phases of Mitosis
In order:
Prophase
Prometaphase
Metaphase
Anaphase
Telophase
Prophase
First step of mitosis characterized by chromatin condenses into chromosomes, nuclear envelope begins degredation, and spindle formation starts.
Before Prometaphase.
Prometaphase
Second step of mitosis characterized by full nuclear dissolution and completed spindle formation which then connects to centromere via kinetochores.
After Prophase, before Metaphase.
Metaphase
Third step of mitosis characterized by chromosomes lining up end to end at metaphase plate and spindles attaching to their centromere via kinetochores.
After Prometaphase, before Anaphase.
Anaphase
Fourth step of mitosis characterized by separase removing cohesin so sister chromatids can be pulled apart to poles by spindles via kinetochores and turn into daughter chromosomes.
After Metaphase, before Telophase.
Telophase
Fourth step of mitosis characterized by spindle dissolving, chromosomes decondensing into chromatin, and nuclear envelope reforming around both groups of daughter chromosomes seperately.
After Anaphase.
Separase
Enzyme responsible for breaking down cohesin during anaphase of mitosis to allow for proper chromosome segregation.
Activated by the Mitosis phase CDK.
Cohesin
Protein that holds sister chromatids in replicated chromosomes together from S phase until anaphase for mitosis and meiosis.
Metaphase Plate
Imaginary plane located at equator of a cell, halfway between both spindle poles, where chromosomes align during the metaphase.
Kinetochore
Multi-protein structure which attaches to centromere during mitosis/meiosis and connects to microtubules of spindle.
Depolymerizes microtubules plus end into tubulin which shortens them in order to create pull necessary to segregate chromosomes.
Spindle
Structure made of centrosomes and microtubules which is responsible for organizing, attaching to, and segregating chromosomes to poles of a cell.
Centrosome
Organelle responsible for the creation and orientation of spindle and microtubules.
Located at the 2 poles of cell and made of 2 centrioles each.
Centriole
Cylyndrical organelle found in centrosomes in cell.
Responsible for creation of spindle and organizing chromosomes.
Cytokinesis
Final process of cell division after telophase where the cell splits into 2 individual daughter cells.
Contractive ring of myosin and actin called “cleavage furrow” pull plasma membrane inwards until split occurs.
Somatic Cell
Any cell of an organism other than reproductive cells.
Replicates via mitosis.
Example: liver cell, skin cell, bone marrow cell
Meiosis
Specialized form of cell division used to create genetically diverse gametes (germ cells)
Results in 4 haploid daughter cells which are genetically unique from each other and parent cell.
Process takes place in 2 division steps known as meiosis I and meiosis II.
Chromosome Number (n)
Integer with unit ‘n’ representing total amount of unique chromosomes in cell.
haploid cells have ‘1n’ and diploid cells have ‘2n’.
Heterozygous
Describing a diploid cells containing 2 different alleles for a specific gene (s).
Ex: F1s of ‘AA’ + ‘aa’ cross are ‘Aa’
Homozygous
Describing a diploid cells containing 2 of same alleles for a specific gene (s).
Ex: Crossed Ps could be ‘AA’ + ‘aa’
Synapsis
Process of homologous chromosomes pairing together along their lengths and forming chiasmata to physically connect to each other.
Occurs during Prophase I (Meiosis I).
Non-Sister Chromatids
Describing relation of 2 chromatids from 2 seperate chromosomes formed from crossing over during synapsis in Prophase 1.
Chiasma
X-shaped, physical links between non-sister chromatids of homologous chromosomes that form during Prophase I
Meiosis I (1)
First half of Meiosis which creates 2 haploid daughter cells from 1 diploid parent by segregating homologous chromosomes.
Differs from Mitosis in Prophase, where Prophase I has synapsis.
Meiosis II (2)
Second half of Meiosis which creates 4 genetically distinct haploid daughter cells (gametes) from 2 haploid parent cells by segregating sister chromatids.
Differs from Mitosis in the amount of cells produced and the diversity of those cells.
Crossing Over
Exchange of genetic material between non-sister chromatids of homologous chromosomes during prophase I of meiosis.
Creates genetic diversity in the form of recombinant chromatids.
Recombinant Chromatids
Chromatids with new or different combinations of genetic material, formed from the process of crossing over when non-sister chromatids exchange DNA.
Phenotype
Set of traits/characteristics an organism possesses.
Ex: hair color, biological sex, inherited diseases, etc.
Genotype
Set of DNA in the form of genes and their alleles an organism possesses.
Ex: {AA, Bb, CcDD} as a set
Parental Generation (P)
First set of parents crossed in a genetic study, usually at the top of the pedigree.
Offspring of this generation are the F1 Generation.
Often chosen to be pure breeding for consistent results in F1.
Pure Breeding
Describing individual that consistently produces offspring with the same phenotype.
Ex: ‘AA’ crossed with ‘aa’ in P always make ‘Aa’ offspring.
F1 Generation (F1)
Immediate offspring from cross of P generation, found right under P generation.
Offspring of this generation are the F2 Generation.
If parents are true breeding for different alleles then this generation will always be heterozygous.
Dominance in Genes
Relationship between alleles of a gene where one allele takes precendence in gene expression over another allele.
Allele having precedence does NOT mean that it is more common.
Ex: heterozygous organism with ‘Aa’ genotype having only ‘A’ phenotype expressed.
Law of Segregation
States that during formation of gametes, 2 alleles of a gene separate from each other, so that each gamete carries only 1 of those 2 alleles.
Multiple gametes can be produced with the same allele, but they will only have 1 copy of it.
Ex: monohybrid heterozygous F1 generation in meiosis ‘Aa’ makes 4 gametes: ‘A’, ‘A’, ‘a’, ‘a’.
Law of Independent Assortment
States that alleles for different traits are sorted into gametes independently of one another.
OR
When homologous chromosomes pairs line up and stack in synapsis during Prophase I, there are 2 possible ways that each pair can stack in relation to another pair.
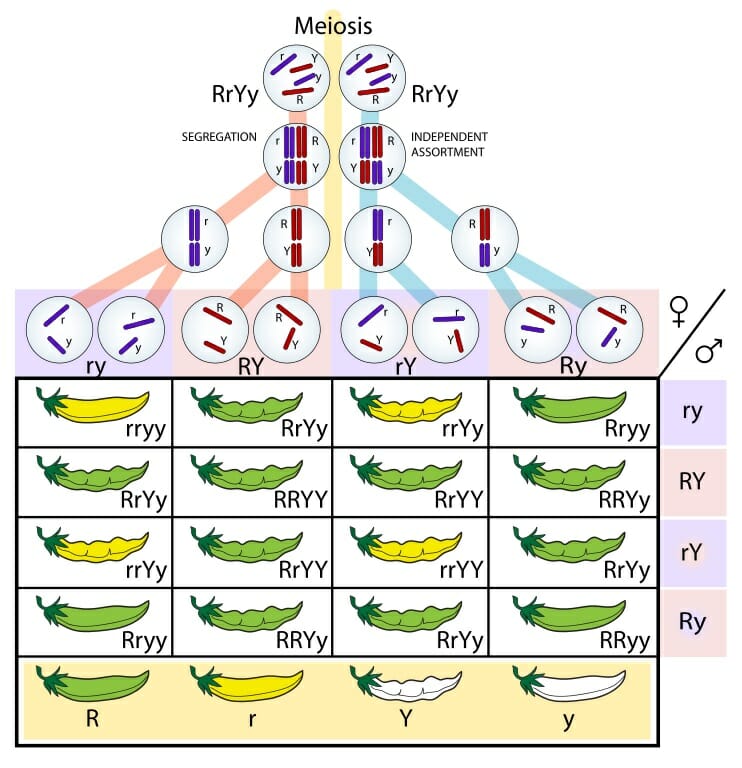
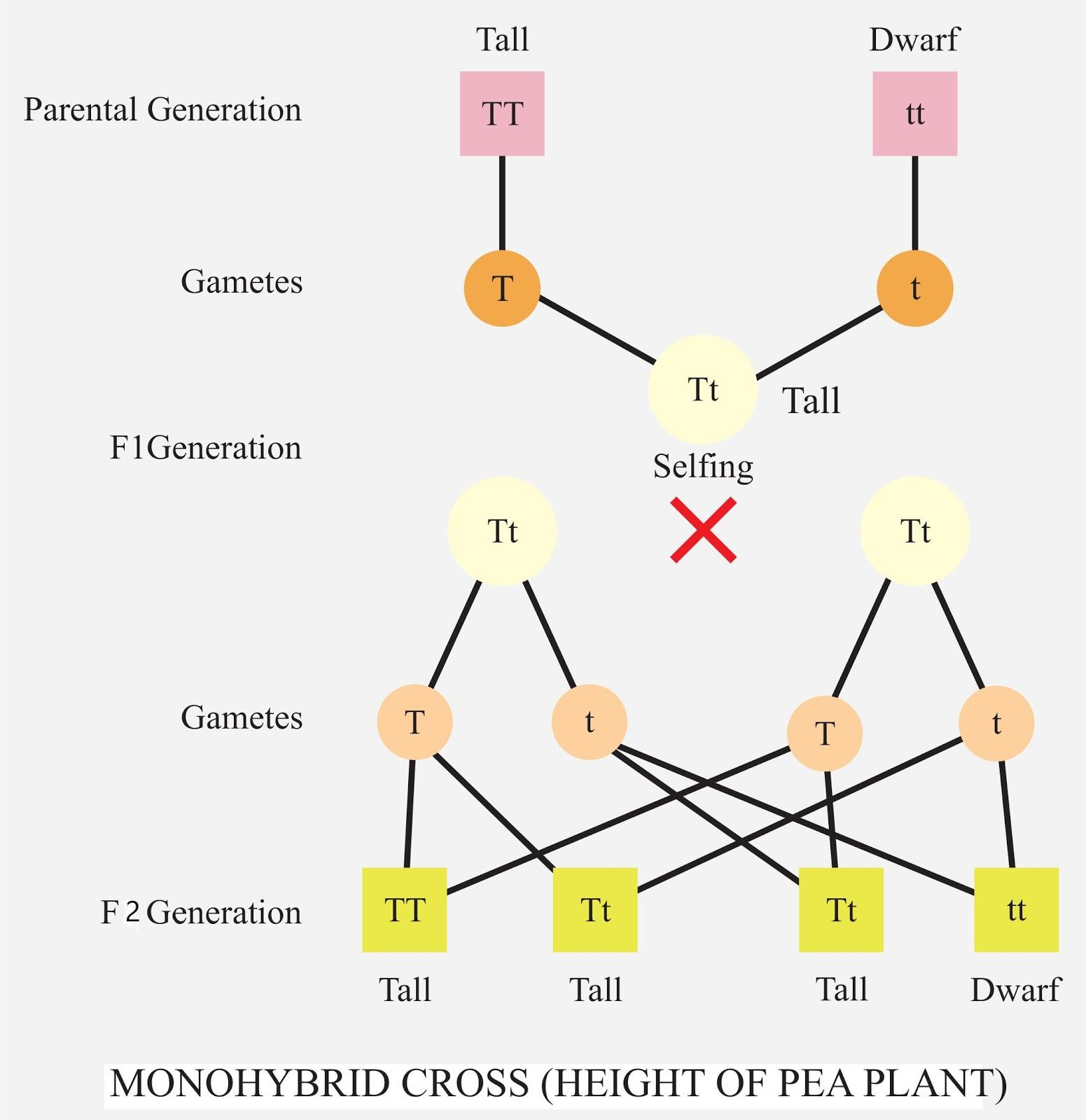
Monohybrid
Type of cross involving organisms heterozygous for ONLY 1 specific trait or gene.
Pure breeding P generation crossed this way lead to F2:
Genotype ratio of 1 : 2 : 1 (hmzg dom : htzg : hmzg rec)
Phenotype ratio of 3 : 1 (dom : rec).
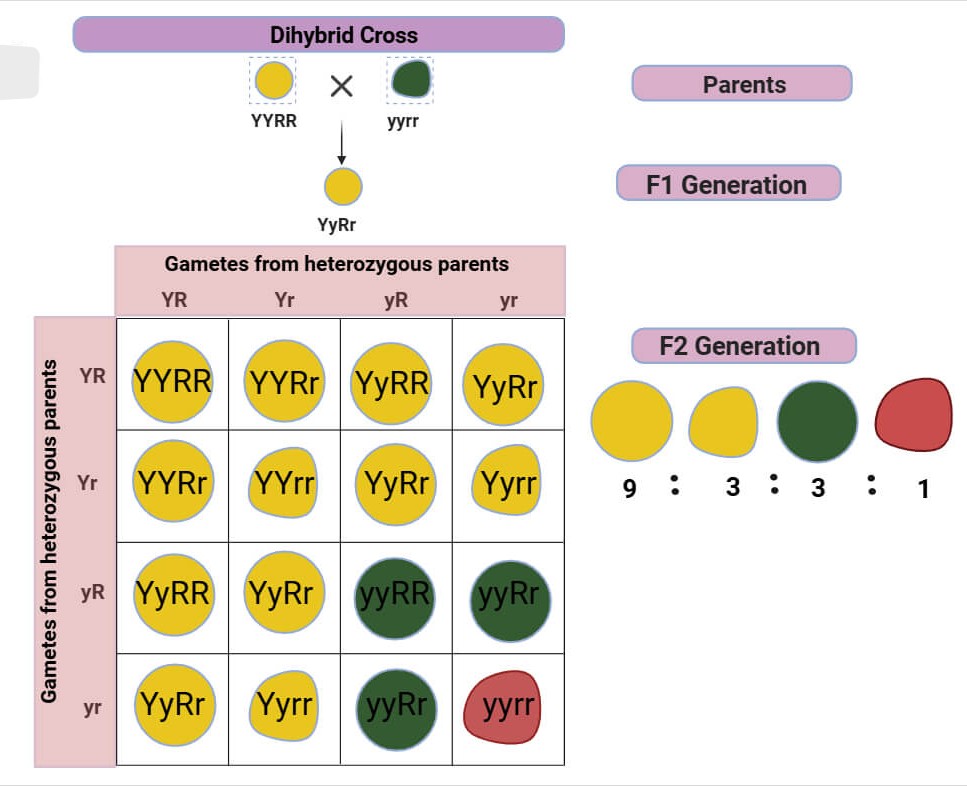
Dihybrid
Type of cross involving organisms heterozygous for 2 specific traits or genes.
Pure breeding P generation crossed this way lead to F2:
Genotype ratio calculated via punnet square
Phenotype ratio of 9 : 3 : 3 : 1 (both dom : 1 dom : other 1 dom : both rec).
Multiplication Rule
Rule Stating that to find probability of 2 independent events happening together, multiply individual probabilites together.
Key word is ‘and’.
Ex: roll one d6 (1/6) and flip one quarter (1/2), getting 3 and heads is a 1/12 chance (1/6 × 1/2).
Addition Rule
Rule Stating that to find probability of 2 mutually exclusive events happening together, add together individual probabilites together.
Key word is ‘or’.
Ex: When finding P of htzg offspring (‘Aa’) from htzg parents (‘Aa’ and ‘Aa’) progeny can get ‘A’ from 1 parent OR the other and the same with ‘a’ (1/4 + 1/4 = 1/2 = 50%).
Pedigree
Diagrammatic family tree symbols to map inheritance of specific phenotypes, traits, or genetic diseases across multiple generations.
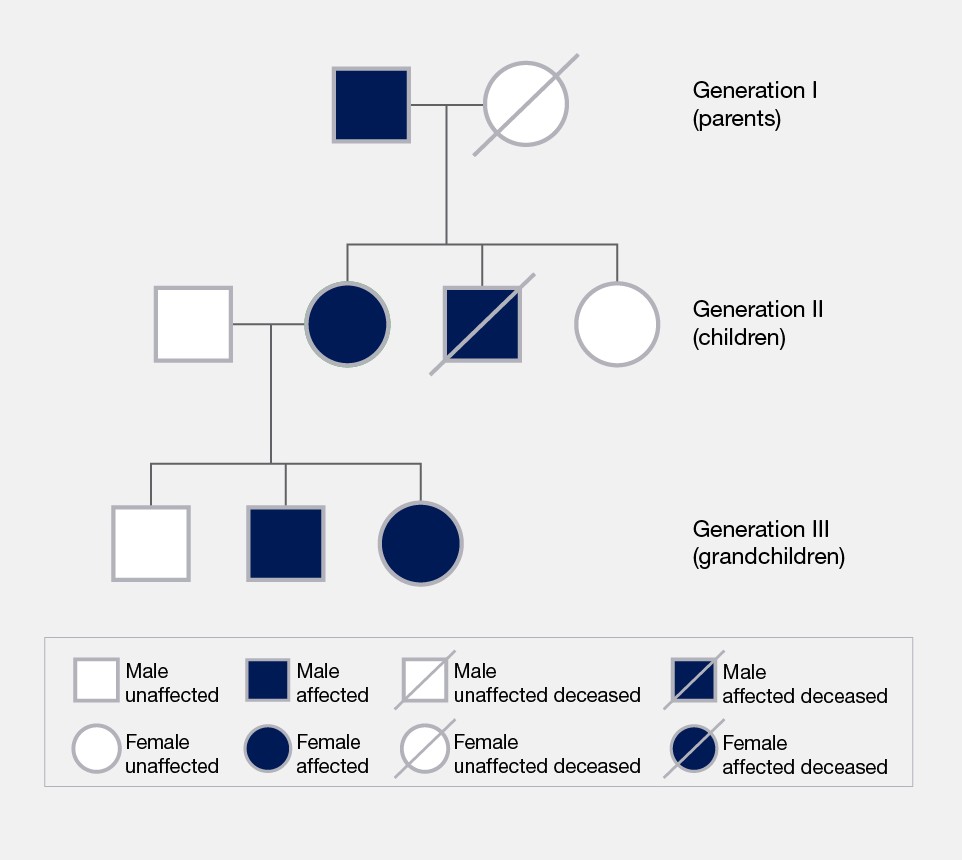
Affected
Describing an organism in a pedigree displaying a specific phenotype or trait.
Usually represented by a shaded shape.
Unaffected
Describing an organism in a pedigree NOT displaying a specific phenotype or trait.
Usually represented by an unshaded shape.
Autosomal Recessive
Pattern of inheritance where a trait or disorder requires two copies of an allele, one from each parent, to manifest.
If 1 parent is affected (‘aa’) and the other is a carrier (‘Aa’) and the offspring is also affected.
If both parents are unaffected (‘Aa’) but the offspring is affected (‘aa’)
Autosomal Dominant
Pattern of genetic inheritance where a single copy of an allele, located on a non-sex chromosome, is sufficient to cause a trait or disorder.
If 1 parent is affected (‘AA’ / ‘Aa’) and the offspring is also affected.
If both parents are affected carriers (‘Aa’) but the offspring is unaffected (‘aa’)
X-Linked Recessive
Pattern of genetic inheritance where a mutation on X chromosome causes a trait to be expressed fully in males (XY) but usually only in females (XX) who are homozygous for the mutation.
Wild Type Allele
Most common, naturally occurring version of a gene in a population.
NOT NECESSARILY DOMINANT!!!
Usually codes for a functional phenotype.
Compared to usually rare and nonfunctioning “mutant” genes in populations.
Mutant Allele
Variant form of a gene that has undergone a permanent change in its DNA sequence, differing from standard more common "wild-type" allele.
Sometimes codes for nonfunctioning phenotype.
Allelic Series
Set of 3+ different gene variants for a single locus, often causing a graded spectrum of phenotypes.
Can be ordered according to relative variant dominance.
Ex: ‘IA’ & ‘IB’ & i code blood type where ‘IA’ & ‘IB’ are codominant to each other but completely dominant to ‘i’
‘IA’ = ‘IB’ > ‘i’
Karyotype
Complete, organized set of chromosomes in 1 cell of an organism, ordered by size, number, and shape on a sheet.
Complete Dominance
Form of inheritance where dominant allele completely masks the presence of recessive allele in a heterozygote's phenotype.
Creates a 3:1 dominant : recessive mendelian ratio.
Ex: ‘AA’ & ‘Aa’ have same phenotype.
Incomplete Dominance
Form of inheritance where alleles are neither dominant nor recessive to each other and heterozygous phenotype is an intermediate “blend” of both homozygous phenotypes.
Heterozygotes not actually “blended”, can make homozygous offspring when crossed.
Ex: ‘FF’ = white flower, ‘ff’ = purple flower, ‘Ff’ = violet flower
Codominance
Form of inheritance where alleles are neither dominant nor recessive to each other and both alleles fully express simultaneously in heterzygous phenotype.
Ex: blood type ‘A-phenotype’ x ‘B-phenotype = ‘AB-phenotype’
Pleiotropy
Describing when 1 allele influences multiple seemingly unrelated phenotypic traits.
Counterpoint: Often, many genes contribute to just one trait like dog fur color.
Ex: Marfan Syndrome caused by 1 defect of FBN1 & causes long limbs AND heart defects
Epistasis
Form of non-mendelian inheritance where phenotype is determined by interaction of 2+ genes where 1 gene masks/inhibits expression of another gene.
Ex:
Dog fur color controlled by 3 proteins: {A} = switch on (dark), {K} = “switch” off (default yellow), {E} = “switch” that {A} & {K} bind to.
If {E} = gone/dysfunctional b/c ‘ee’ (homz. rec.), neither ‘A’ or ‘K’ can bind so defaults to yellow coat
So when = ‘ee’, ‘E’ gene overrides ‘A’ gene even when ‘A’ is dominant.
If put through true breeding dihybrid cross then final “9:3:3:1” ratio is actually “9:3:4” (4 = recessive yellow phenotype)
Sex Chromosome
Specific pair of chromosomes (X and Y in humans) that determine an organism's biological sex and gamete production.
Also carry genes other than for sex determination (e.g color blindness on X).
Linked Genes
Describing when 2+ genes are located near each other on same chromosome and how they interact in inheritance.
Overrepresentation of parental genes (less recombinant).
Can be determined by having recombinant frequency RF <50.
Gene Linkage
Tendency of linked genes to be inherited as 1 unit (parental genotypes), violating the law of independent assortment by being less affected by crossing over.
Closeness of genes to each other on chromosome determine how likely they are to inherit together.
Parental Genotype
Specific allele combinations (genotypes) that tend to be overrepresented because linked genes often stay together.
Gametes with these allele combinations usually match the parent cell’s genotype.
Ex: The more common ‘AB’ & ‘ab’ genotypes in the gametes of an organism with ‘AB/ab’ genotype where ‘A’ & ‘B’ are linked.
Recombinant Genotype
Specific allele combinations caused by crossing over (genotypes) that tend to be underrepresented because of linked genes.
Gametes with these allele combinations usually don’t match the parent cell’s genotype, and follow independent assortment.
Ex: The less common ‘Ab’ & ‘aB’ genotypes in the gametes of an organism with ‘AB/ab’ genotype where ‘A’ & ‘B’ are linked.
Unlinked Genes
Genes located on different chromosomes or very far apart on same chromosome causing them to assort independently during meiosis and inherit independently.
Parental and recombinant genes are represented equally.
Can be determined by having recombinant frequency RF = 50.
Recombinant Frequency (RF)
Measure of the likelihood that loci of 2 genes will be seperated during meiosis, calculated by proportion of recombinant offspring to total offspring. (#recomb. / # total) x 100 = RF (%)
Measured in centimorgans & map units. 1% RF = 1 cM = 1 mu
Must use a test cross in order to calculate.
Maximum value is 50%.
Test Cross
Cross used to determine if an organism displaying a dominant trait is homozygous or heterozygous by breeding with homozygous recessive individual.
Can determine presence of gene linkage if offspring phenotypic ratio deviates from 1:1:1:1 ratio of independent assortment.
Genotype Writing Styles
Alleles seperated by comma (,) = on same chromosome.
Alleles seperated by slash (/) = on different homologous chromosome.
Alleles seperated by semicolon (;) = on completely different chromosomes.
Ex: ‘A,b / a,B’ = ‘A’ & ‘b’ on same chromosome, ‘a’ & ‘B’ on different but homologous chromosome.