Molec Bio Exam 3
1/82
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
83 Terms
Exon Shuffling
When exons are combined to create new genes
Vertical Gene Translation
From mother to daughter cell
Types of Horizontal Gene Translation
Transformation: Uptake from Environment
Transduction: Mediated by viruses
Conjugation: Bacterial “Sex”
Transposons
“Jumping Genes”, move around genome creating new genes at insertion
Requires “Transposase” enzyme to occur
Every organism has Transposons
Retrotransposons
Common in Eukaryotes
RNA intermediate phase
Converted to DNA via Reverse Transcriptase
Still requires Transposase
Virus Protein Covering
“Capsid”
Retroviruses/Proviruses
Must integrate into genome as dsDNA (antipararell)
Added to protein known as “Integrase”
LINEs/SINEs
Make up “Junk DNA”, often originating from pseudogenes, transposons, duplications
SNPs
Single Nucleotide Polymorphisms
Main way humans vary from one another
Used for DNA
Phospholipid Membrane Side Names
Outer leaflet
Inner leaflet
Most Common Plant/Animal Phospholipid
phosphatidylcholine
Choline = Phosphate = Glycerol = 2 Fatty Acid Tails
Double bonds saturate it
Phospholipid Movement Types
Flexion: Flexing of tails
Rotation: Spinning of Phospholipid
Flip-Flop: leaflet-to-leaflet movement, rare
Membrane Fluidity
Decreased by Tail Length, Sterol prescence, Saturated Fats
Flippases
Move phospholipids from outer to inner leaflet
Floppases
Move phospholipids from inner to outer leaflet
Scrambles Lipid Directions
Scramblases
Can be independent or dependent on Ca+
Smooth ER Phospholipid Depositing
Deposited into Cell Membrane’s Cytosolic (inside) Side before transport
Phospholipid Transport
Either independent or Done in vescicles
Membrane Protein Types
Transporters/Channels: Move materials through membrane
Anchors: Link cytoskeletal Proteins with extracellular matrix
Receptors: Involved in Cell Communications
Enzymes: Carry out reactions
Integral Membrane Protein Types
Transmembrane: Crosses bilayer
Monolayer (Leaflet) Associated: Associated with one side
Lipid Linked: Bonded to lipid in membrane
Pores
Made by Porins, Alpha Helices or Beta Sheets
Detergents
Amphipathic Molecules, best way to distrupt membranes
Cell Cortex
Part of Cytoskeleton
Has Spectrin Coils which meet at Actin Proteins
Attached via Attachment Proteins
Posttranslational Modifications
Proteins are often glycosylated with glycans, “high mannose sugars” to mature them
Glycoprotein/Glycolipid Complex
Glycocalyx, surface of cell
Cell Recognition, Protects Cell
Lubricates Cell
Recognize Carbohydrates on other cells
“Lectins”, transmembrane glycoproteins
Energy-Costing Cell Transport Means
Active Transport
Bulk Transport
Electrochemical Gradient
Cells are negatively charged
Have more salt/chloride in extracellular enviorn.
Have more potassium in intracellular enviorn.
Passive Transport Channels
Mechanically Gated
Ligand Gated (extra or intracellular ligands)
Voltage Gated
How water enters/leaves cell
Aquaporins
Active Transport Pumps
Coupled Reaction: Swap 2 substances, one thermodynamically favored
ATP Pump: ATP fights against gradient
Light Driven: Uses Light
Coupled Pumps:
Symport: 2 diff molecules in same direction
Antiport: 2 diff molecules in opposite direction
Globin
Allows for transport of oxygen molecule
Epithelial Cell Support Structure
Basal Lamina, an Extracellular Support Mat
Osmolarity
Amount of dissolved solutes in solvent
Where muscle cells store calcium
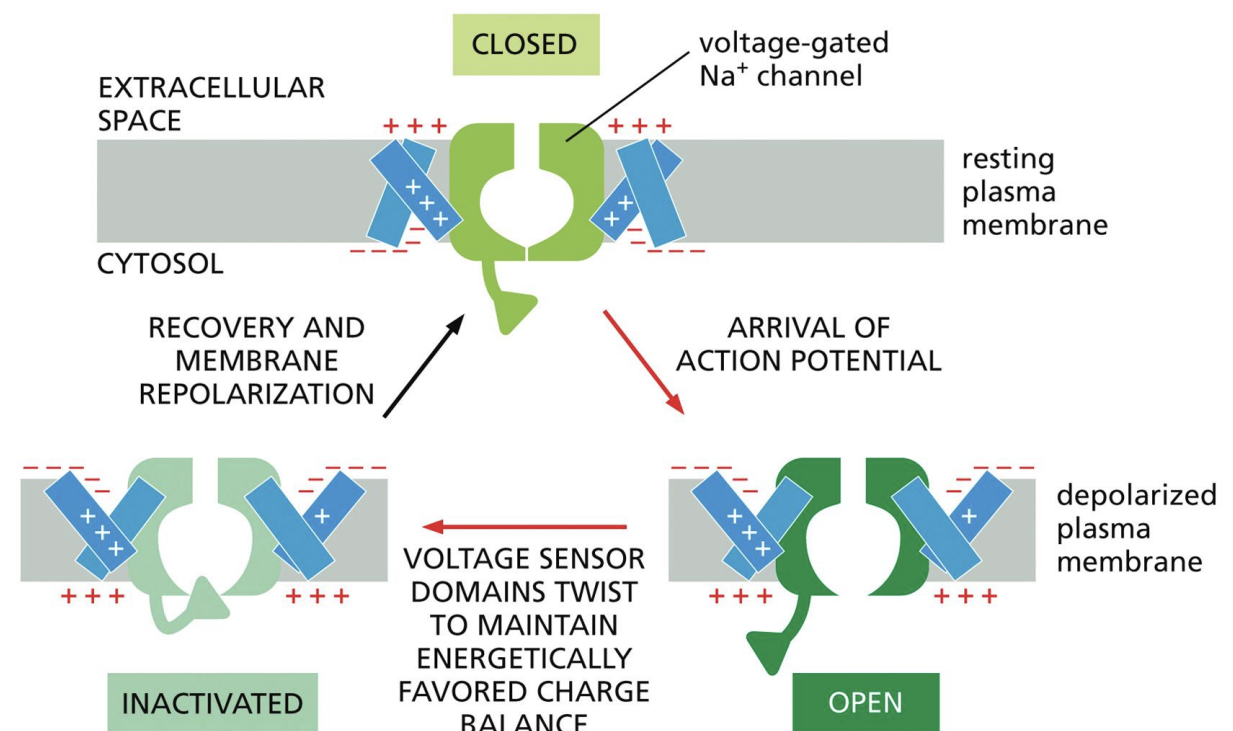
Action Potentials
Carried by voltage-gated channels
1: Cytosol starts off negatively charged
2: Change in charge balance opens gate, allowing potassium out
3: Extremity of voltage sensor is bent to block gate, closing it
4: Gate eventually reverts back to original state
This process repeats within a tube structure
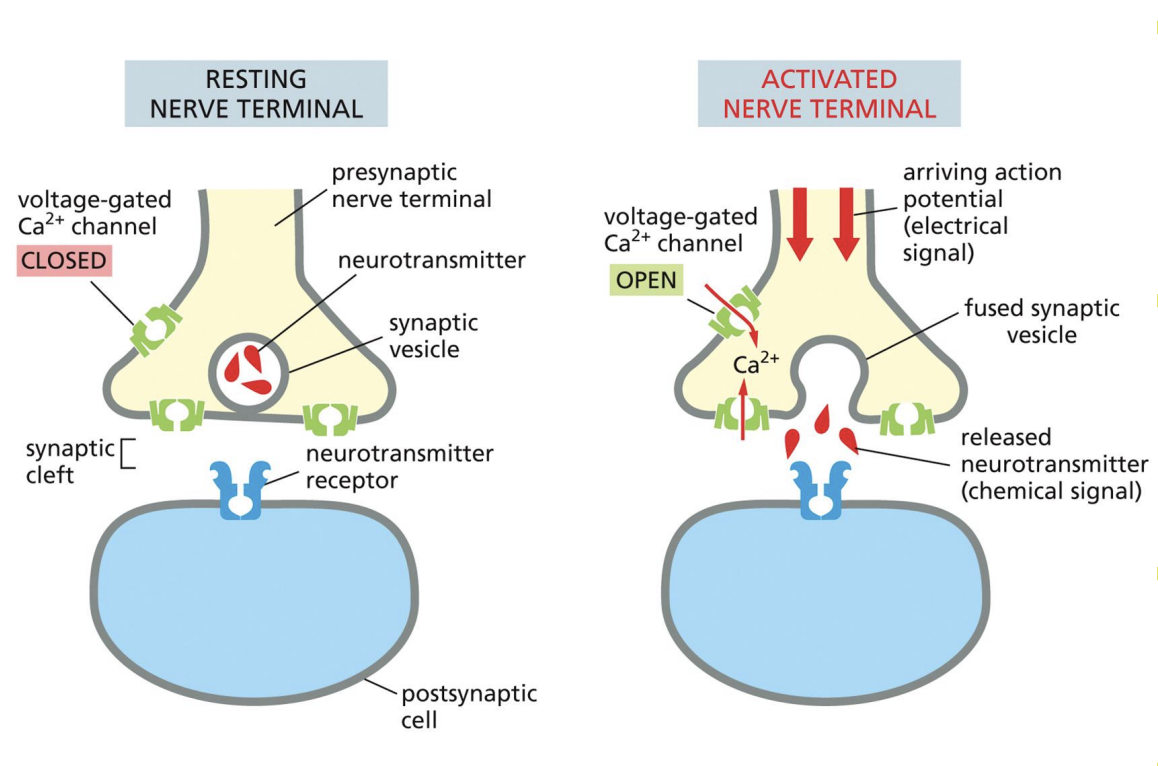
Synaptic Vesicle
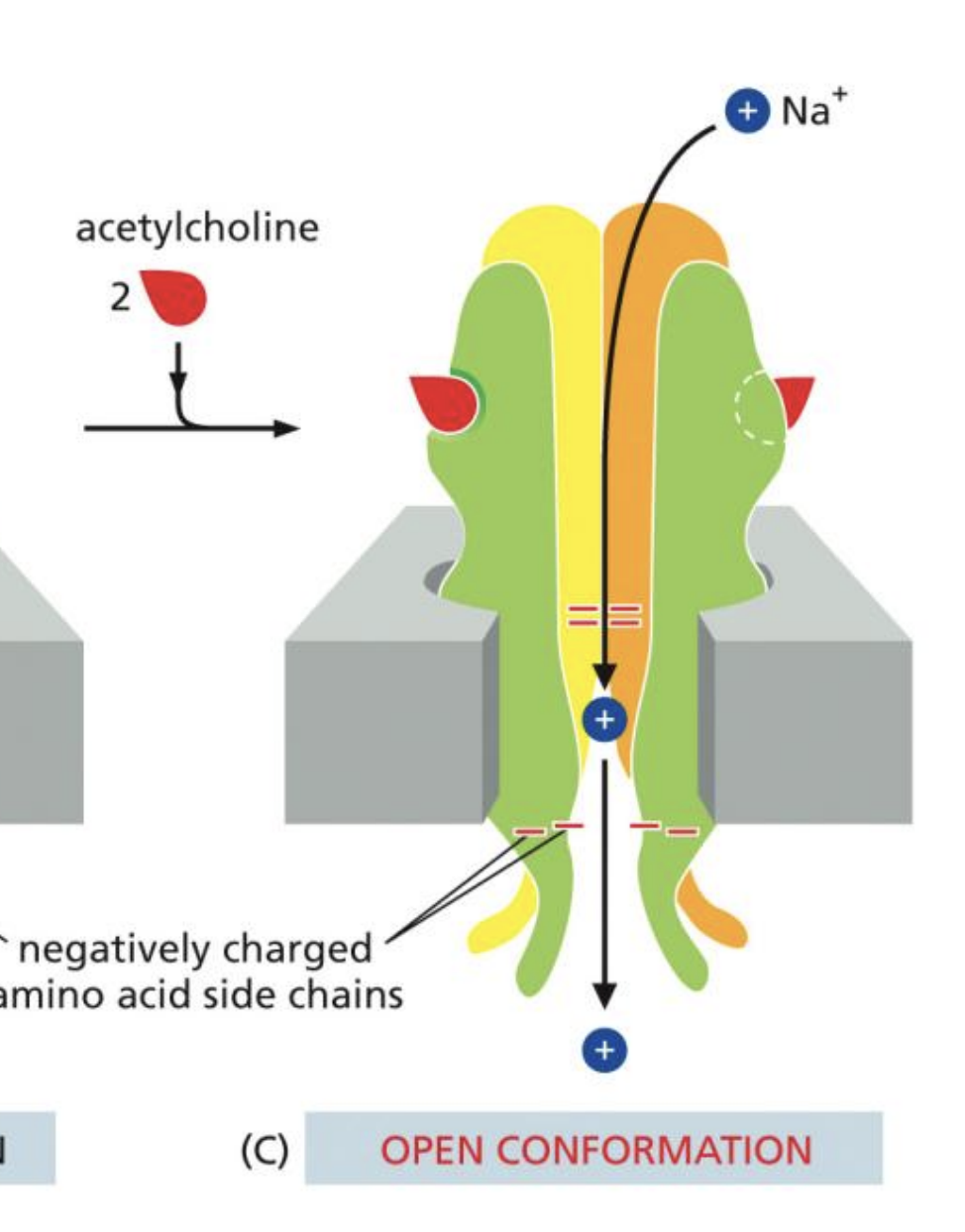
Acetyl-Choline Receptors
5 Transmembrane subunits, 2 bind to Acetylcholine
When both are bound, sodium-potassium transfer occurs
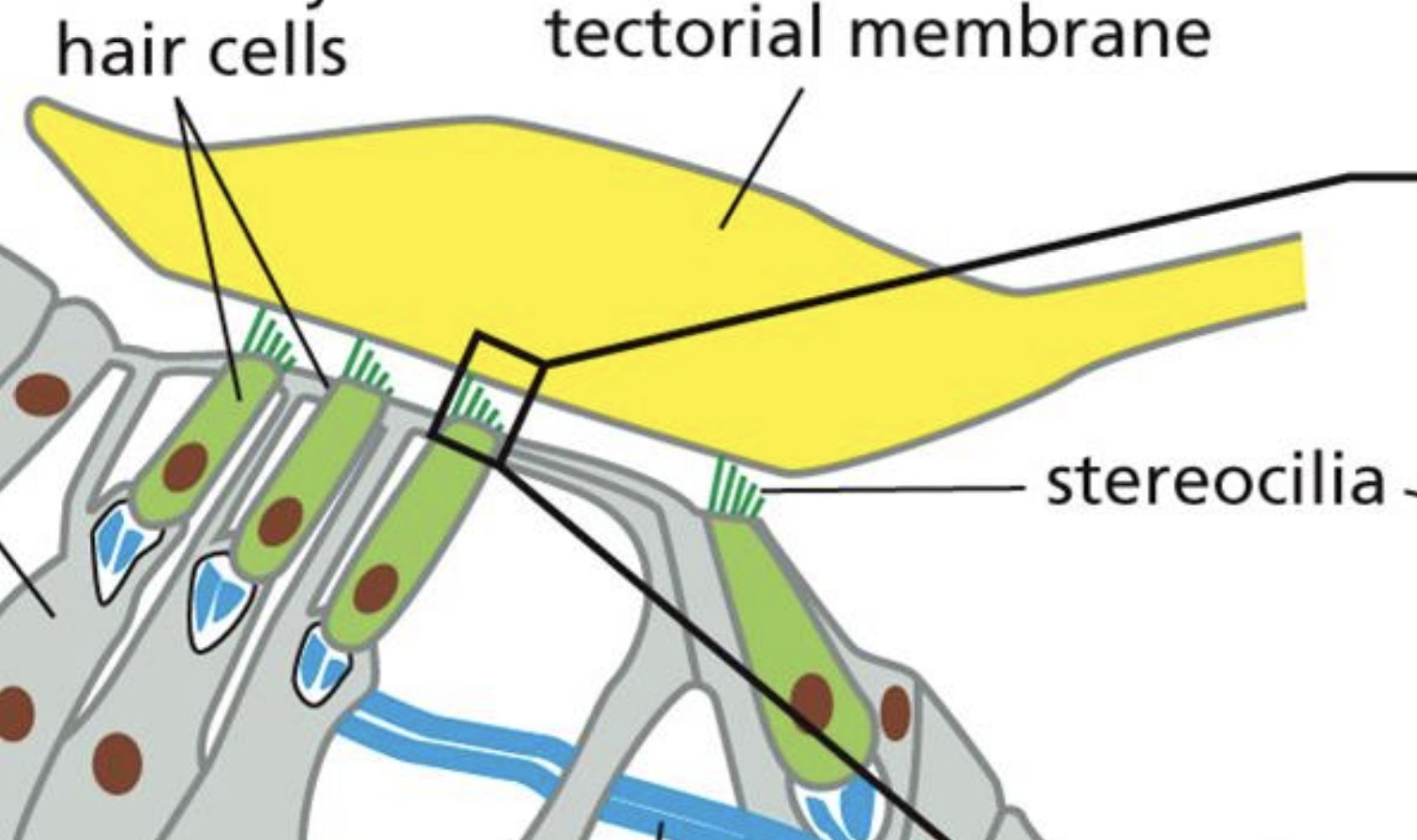
Hearing Gated Channels
“Fingertips” of stereocilia are linked by spring proteins, opened by stretching
This allows potassium inside vesicle
Depolarization causes vescicle output releasing neurotransmitters
Signal Sequence
A “Signal” sequence of RNA at the ends or center of proteins that signals the location of transport
Cytoskeleton of Nucleus
Nuclear Lamina, formed by Intermediate Filaments
Broken down by Kinases during Mitosis, reformed by Phosphotases
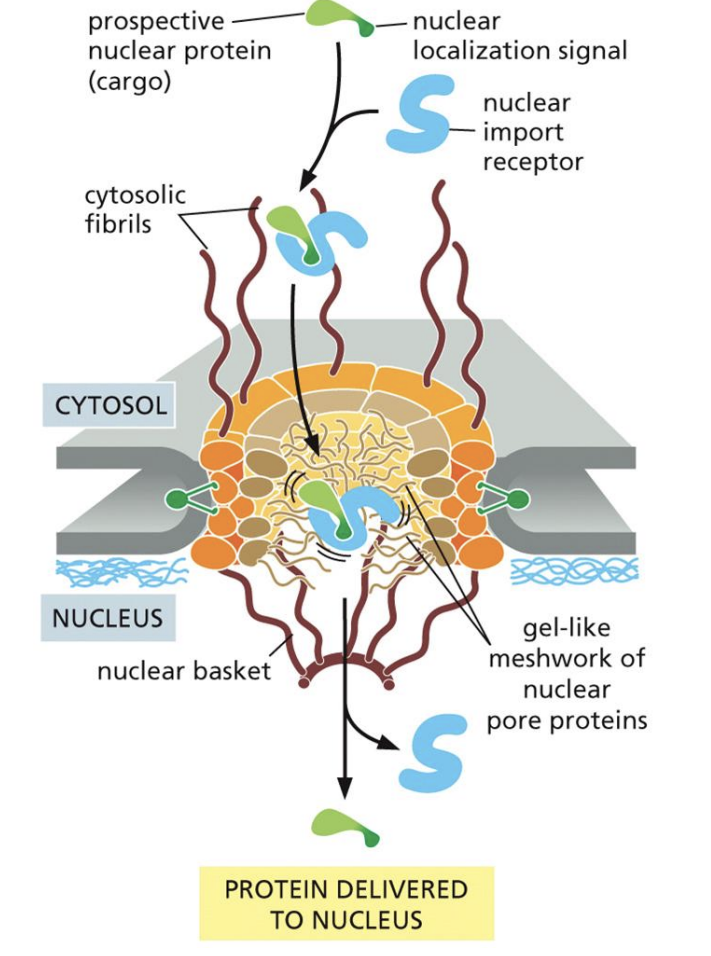
Nuclear Import Receptors
1: Bind to Nuclear Localization Signals, guide them to pores
2: Once inside cell seperate, then bond to Ran-GTP
3: Carries Ran-GTP out of cell, then dissociates
Nuclear Transport Energy
1: Ran-GDP in cytosol enters cell
2: Ran-GEF seperates GDP from Ran
3: Ran binds to GTP, leaves cell
4: Ran-GAP hydrolyzes Ran-GTP
Mitochondria/Chloroplast Protein Transfer
1: Proteins made in Translocators, TIM (inner) and TOM (outer)
2: Proteins with signal sequence enter translocator into membrane
Protein Production in ER
1: Ribosome with “Signal Sequence” bonds to other ribosomes in cytosol
2: Polysome line translocates RNA into membrane
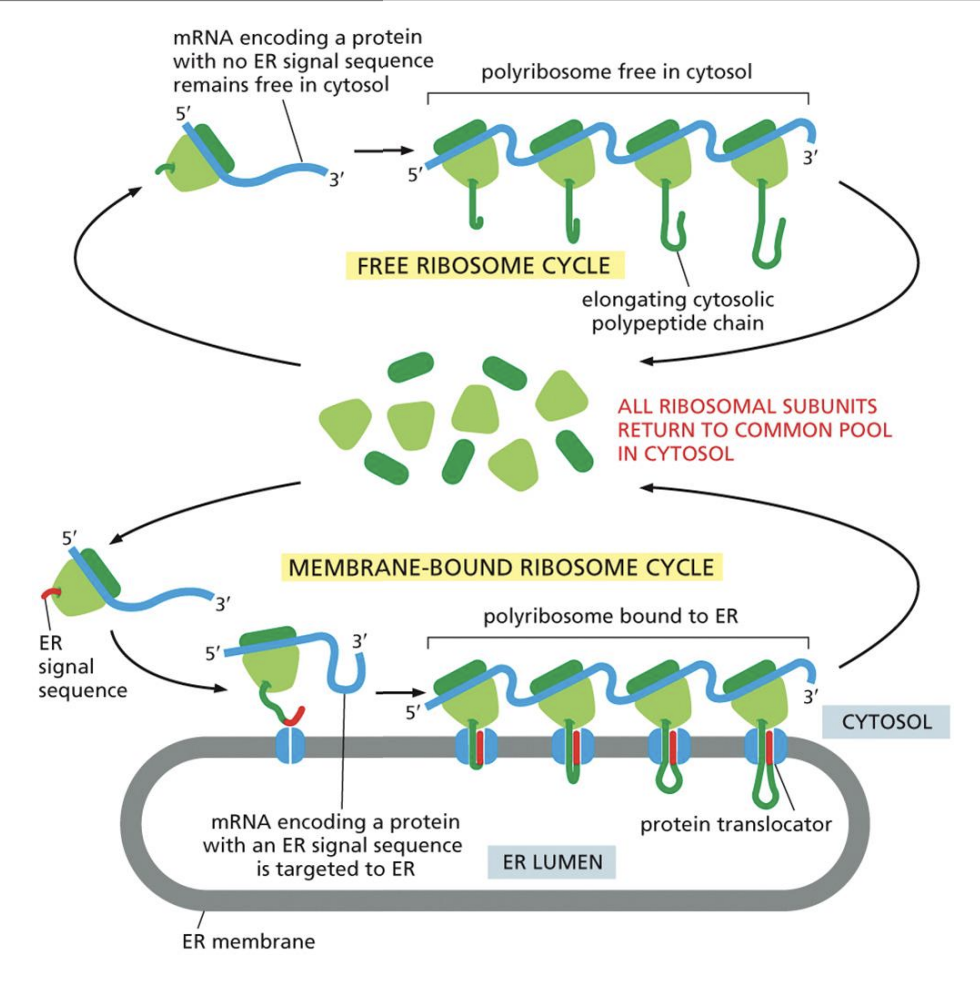
ER Translocation
1: Signal Recognition Particle binds Ribosome Signal Sequence to Translocator location
2: After protein is formed, “Signal Peptidase” cuts it off
Double+ Pass Membrane Proteins
1: Hydrophobic Start-Transfer Sequence enters first, stopping at translocator
2: Stop-Transfer Sequence enters, stops at same place
Endosomes
Large vesicles containing nutrients, broken down by lysosomes
Exocytosis
Done by vesicles budding from Golgi
Cell budding process
1: Adaptin is binded to by Chlarathin coating, creating round shape
2: Bud is cut off by dynamin to create vesicle
Vesicle Docking
1: Rab Protein molecular markers bind to vesicles
2: Tether Protein binds onto vesicle
3: V-snare (on vesicle) binds to C-Snare (on membrane), allowing fusion
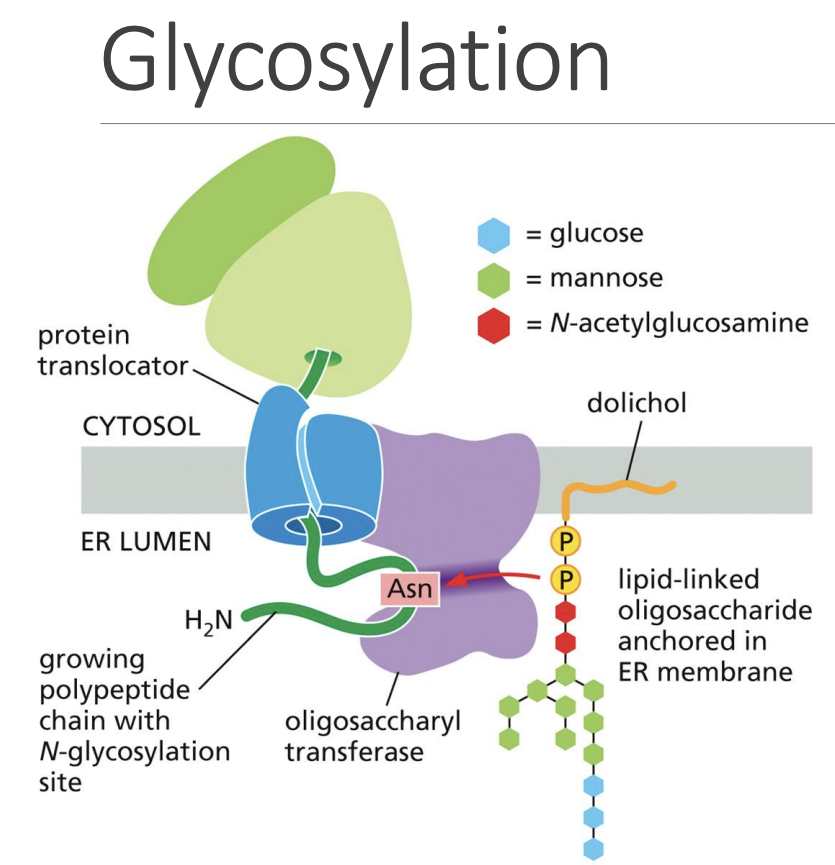
Glycosylation
1: “Dolichol” membrane lipid holding Oligosaccharide binds to Asparagine in “Oligosaccharyl Transferase” protein
Cis Vs. Trans Golgi
Cis Golgi is entry point for proteins, ER materials
Trans Golgi is exit point for proteins
Golgi Hole name
“Cisterna”
Golgi function
Folds proteins, adds high-mannose sugars, glycans
Constitutive Secretion
vs. Regulated, where Phospholipid Secretion is regulated via hormones/transmitters
Pinocytosis
“Cell Drinking” of nutrients in vesicles
Phagocytosis Steps:
Prokaryotes only
1: Particle/Body is enveloped by new membrane to create “Phagosome”
2: Phagosome fuses with the Lysosome, which degrades it
Autophagy
How cells get rid of damaged organelles
1: Organelle covered in membrane, “Autophagosome”
2: Merges with lysosome, which breaks it down
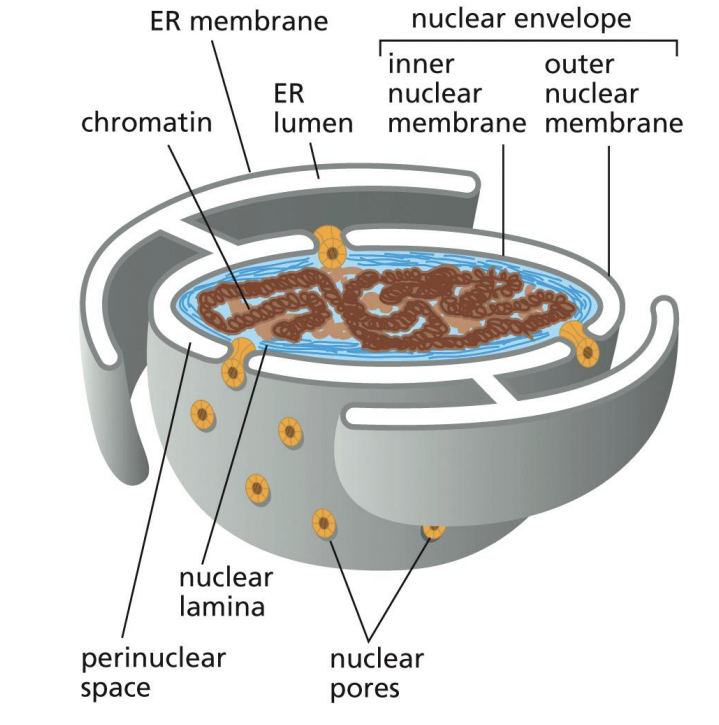
Nuclear Envelope
Bilayer Structure around Nucleus
Inner Layer is “Nuclear Lamina”, made from Intermediate Filaments
Intermediate Filaments
Nonpolar Antipararell Alpha Helices
N terminus: Unstructured, NH2
C terminus: COOH
Can form massive Alpha Helices of 8 tetramers or 32 Monomers
Intermed. Filament Types
Nuclear: Nuclear Lamina in animals
Cytoplasmic:
Keratin Filaments in epithilial cell
Vimenten-related in muscle, glial (support) cells
Neurofilaments in Nerve Cells
Nail Beds are made of
Secreted Keratin Filaments
Provide Stability to Axon Nodes
Neurofilaments
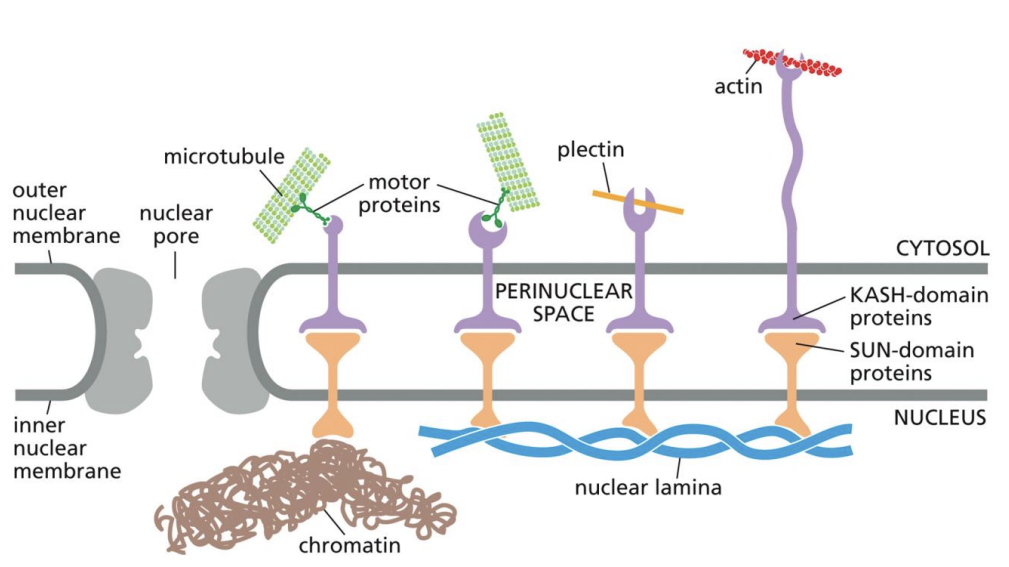
Plectin
Lets Intermediate Filaments bind to Cytoskeletal Proteins
KASH and SUN proteins
Bind to each other within membrane
KASH domain proteins bind to external particles
SUN domain proteins bind to Chromatin or Nuclear Lamina
Microtubules
Built by alternating alpha-beta “Tubulin” protein chains in a “Dimer”
Polar, Plus End is Positive, Minus End is Negative
13 Protofilaments make one Tubule
Tubilin Dimer Formation
Called “Nucleation”
Made from 13 Microtubules, form around “y-Tubulin” around “Centrosome”
Centrosome
Microtubule Organization Center
Contains 2 “Centrioles”, next to Matrix
Positive Microtubule End is inside Cytoplasm
Microtubule Instability
Composed of GDP-Tubulin with GTP-Tubulin “Cap”
If cap is ever degraded, whole structure falls apart
Protein-Microtubule Interactions
Nucleating Proteins
Sever Proteins
Stabilizing Proteins
Branching Proteins (augmin
Axon Terminals
Ridden by:
Kenisins (→ positive end)
Dyneins (→ negative end)
Become more tightly bound with ATP
Become more loosely bound on hydrolysis
Connect to Cargo by Adaptor Proteins
Cilia Vs. Flagella
Cilia move lipids, particles
Flagella move whole cells
Flagella/Cilia Structure
Ask what’s important about it
Allows Microtubule sliding
Dyenin
Microfilaments
“Actin Filaments”
Help with cell structure, division, mobility
“+” end at front, “-” end at back
Highly flexible
Actin “Treadmilling”
ATP bound Actin filaments break are hydrolyzed (+ end) after addition, eventually breaking off (- end), but stay same length
Forms Actin Filaments
ARP Complex
Myosin I
“Boots” move cargo across actin fibrils towards “+” end
Eukaryote Cell Movement
Cell sends out frontmost protrusions, crawling along substrate like slug
Driven by “Lamellipodium” actin filament network
“Filopodium” used to sense area in front of cell
Myosin II
2 “golf club” heads, and coiled tails which associate with one another in large structures
Branches out in both directions on actin filaments when uncapped, allowing for muscle stretching
Actin Filament Formation
“ARP Complex” Encourages the growing of actin filaments
Need to memorize other info