genetic engineering
1/11
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
12 Terms
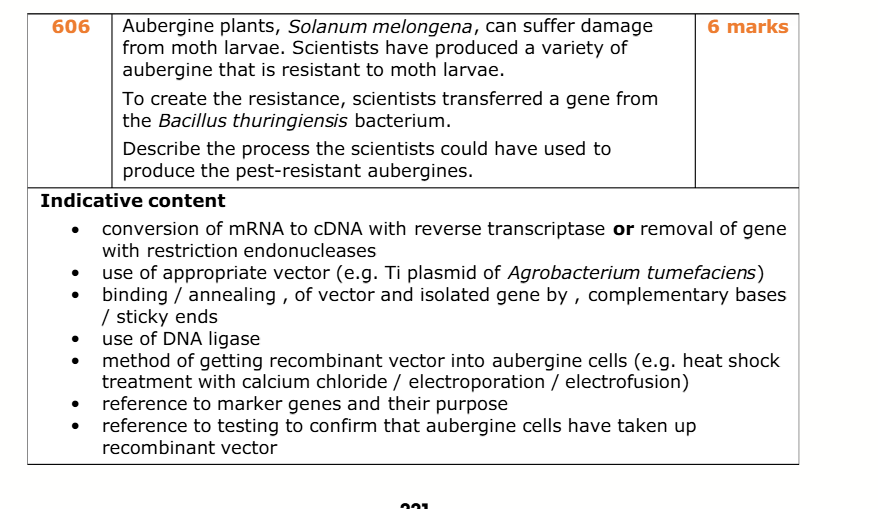
methods to isolate desired gene (2)
-use restriction endonucleases
-use reverse transcriptase
isolating gene using restriction endonucleases
-desired gene is isolated using restriction endonucleases
-the restriction endonucleases cuts DNA at specific recognition sequences, producing sticky ends
isolating gene using reverse transcriptase
-mRNA is extracted from cell that produces desired protein
-Reverse transcriptase converts mRNA into complementary DNA (cDNA)
recombinant DNA
DNA formed by combining DNA from 2 different sources
transgenic organism/ genetically modified organism
an organism that contains a gene from another species in its genome
e.g. Bacteria with human insulin gene → produce insulin
Formation of Recombinant DNA process
-desired gene is isolated using restriction endonucleases
-the restriction endonucleases cuts DNA at specific recognition sequences, producing sticky ends
-a plasmid vector is cut using the same restriction enzyme, creating complementary sticky ends.
-the complementary sticky ends anneal
-DNA ligase joins the gene and vector by forming phosphodiester bonds
-This produces a recombinant DNA molecule (plasmid + inserted gene)
how is recombinant DNA transferred into host cell [transforming cell]
the recombinant DNA is inserted into the host cell using electroporation
→it uses an electrical current that makes the membrane more permeable to allow the plasmid to pass through and enter cell
[Not all host cells take up the plasmid during transformation]
How are transformed cells identified?
using marker genes e.g. antibiotic-resistance markers, fluorescent markers, enzyme
How are transformed cells identified when the marker gene is on the plasmid?
→ antibiotic-resistance marker gene
→ fluorescent marker gene
antibiotic-resistance marker gene
-Plasmids contain an antibiotic-resistance marker gene (e.g. antibiotic resistance)
-After transformation, cells are grown on agar containing the antibiotic.
-Only cells that have taken up the plasmid survive because they are resistant→ these are the transformed cells
fluorescent markers
-Plasmids contain a fluorescent marker gene
-After transformation, cells are exposed to UV light.
-Only cells that have taken up the plasmid are fluorescent/glow
How are successfully transformed cells identified when the desired gene is inserted into a marker gene?
→plasmid has two genes for antibiotic resistance: to tetracycline and ampicillin→ desired gene is inserted into the gene for tetracycline resistance
-the desired gene disrupts the resistance to tetracycline
-cells are grown on agar containing ampicillin→ only cells that have taken up the plasmids survive and form colonies
-cells are grown on agar containing tetracycline→ cells with the desired gene cannot grow
-Therefore, cells that grow on ampicillin but not tetracycline contain the desired gene.
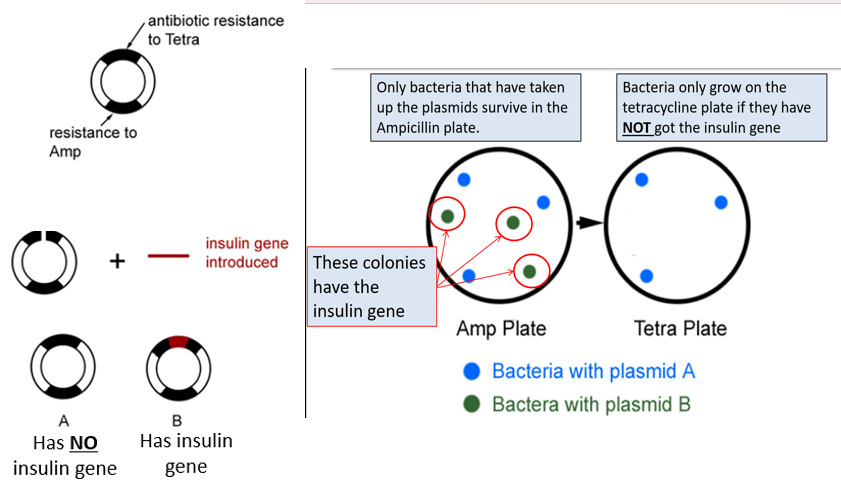
How are successfully transformed cells identified when the desired gene is inserted into a marker gene?
→desired gene is inserted into Green Fluorescent protein (GFP)
-the desired gene disrupts the GFP gene
-Cells are exposed to UV light
-the cells without the inserted gene are fluorescent (green) [are visible under UV light]
-the cells with the inserted gene are not fluorescent
![<p>-the desired gene disrupts the GFP gene</p><p>-Cells are exposed to UV light</p><p>-the cells without the inserted gene are fluorescent (green) [are visible under UV light]</p><p>-the cells with the inserted gene are not fluorescent</p>](https://assets.knowt.com/user-attachments/574c961b-9d0c-42e2-9091-7492ecdc2f5a.png)

exam q
