10. Distinction Between Self and Non-Self: B-Cell Antigen Receptors
1/22
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
Broadly speaking, what is the B cell antigen receptor complex?
A molecular assembly of several polypeptides embedded in the membrane of B cells, consisting of:
IgM or IgD: Antigen-binding and recognition domain.
2 Ig-α/Ig-β (CD79a/CD79b) heterodimers containing ITAMs (Immunoreceptor Tyrosine-based Activation Motifs): Signal transmission.
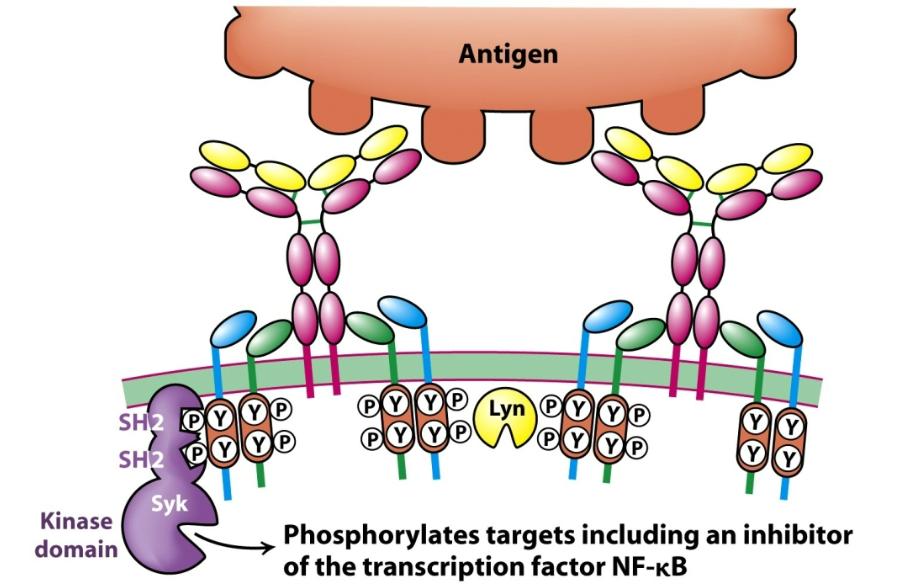
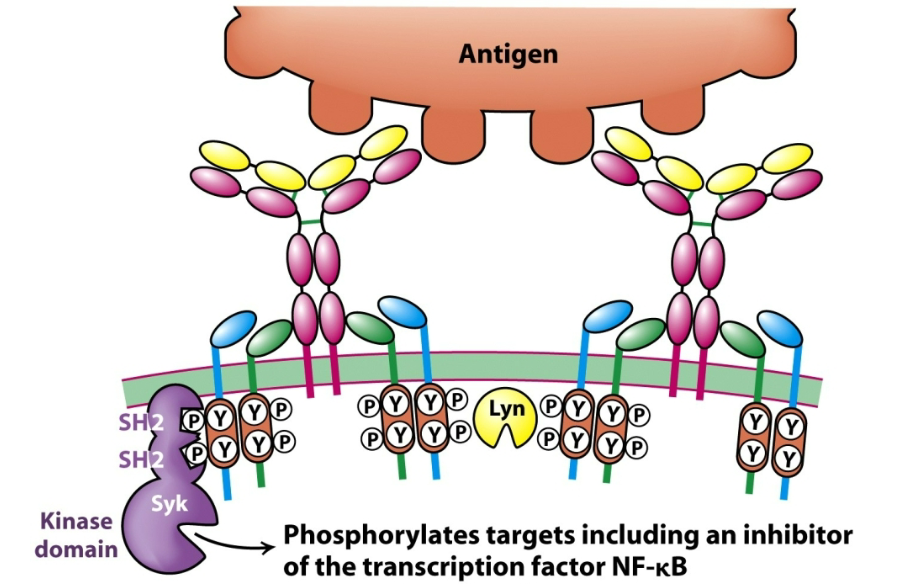
How are B cell receptors involved in the secretion of antibodies? Describe the general process.
The B cell receptor (BCR) binds its specific (cognate) antigen.
Antigen binding leads to cross-linking of BCRs, clustering them on the surface.
This clustering activates Src-family kinases (e.g., Lyn), which phosphorylate ITAMs on the cytoplasmic tails of Igα/Igβ.
Syk kinase docks to these phosphorylated ITAMs and becomes activated.
A cascade of intracellular signaling follows, leading to:
B cell proliferation,
Differentiation into plasma cells,
Secretion of antibodies, which are the soluble form of the BCR.
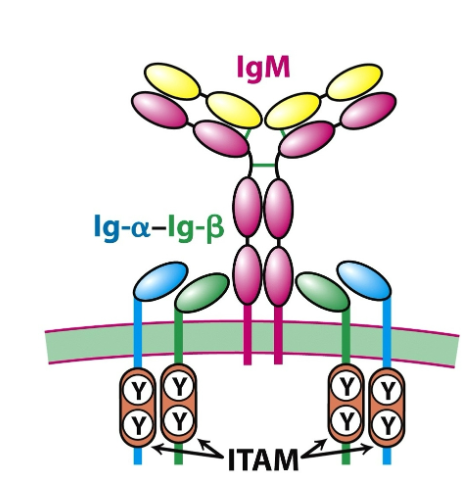
What is the structure of the BCR?
Effectively, a BCR is a membrane-bound antibody.
B-cell receptor (BCR) = membrane-bound antibody (usually IgM or IgD) + signalling components.
Core Components:
IgM or IgD: Antigen-binding and recognition domain of the BCR.
Ig-α and Ig-β (CD79a/CD79b): Signalling molecules non-covalently associated with the antibody recognition domain.
Key structural details:
Ig-α/Ig-β heterodimers contain ITAMs (Immunoreceptor Tyrosine-based Activation Motifs) in their cytoplasmic tails.
ITAMs transmit signals into the cell upon BCR engagement
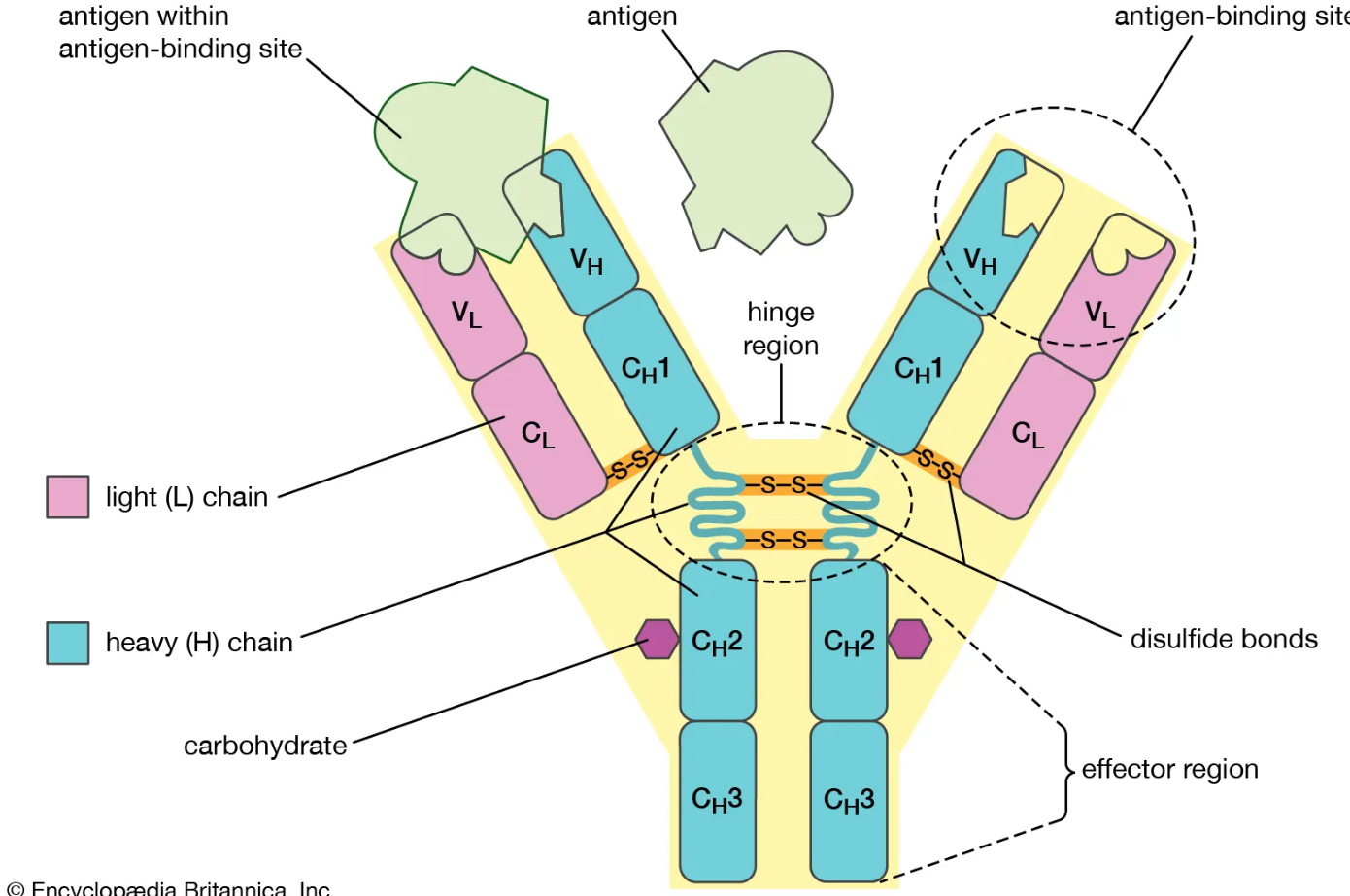
What are the basic components of an antibody?
2 heavy chains + 2 light chains (identical pairs)
Form a Y-shaped structure
Held together by disulfide bonds
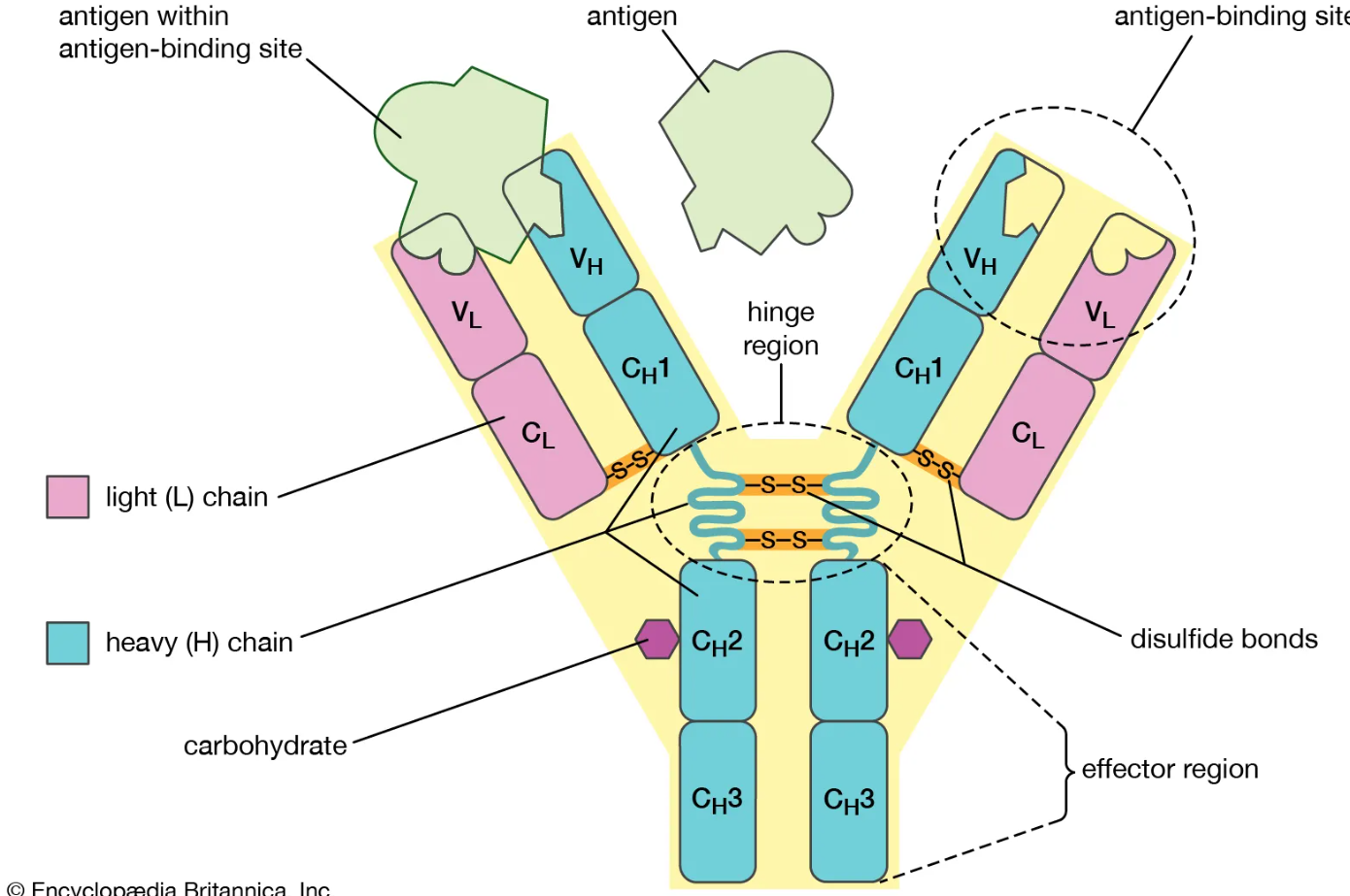
Where does an antibody bind antigen?
Fab regions (arms) contain variable regions (V regions)
Each arm has an antigen-binding site made from CDRs
Both arms are identical, so each antibody binds 2 identical epitopes
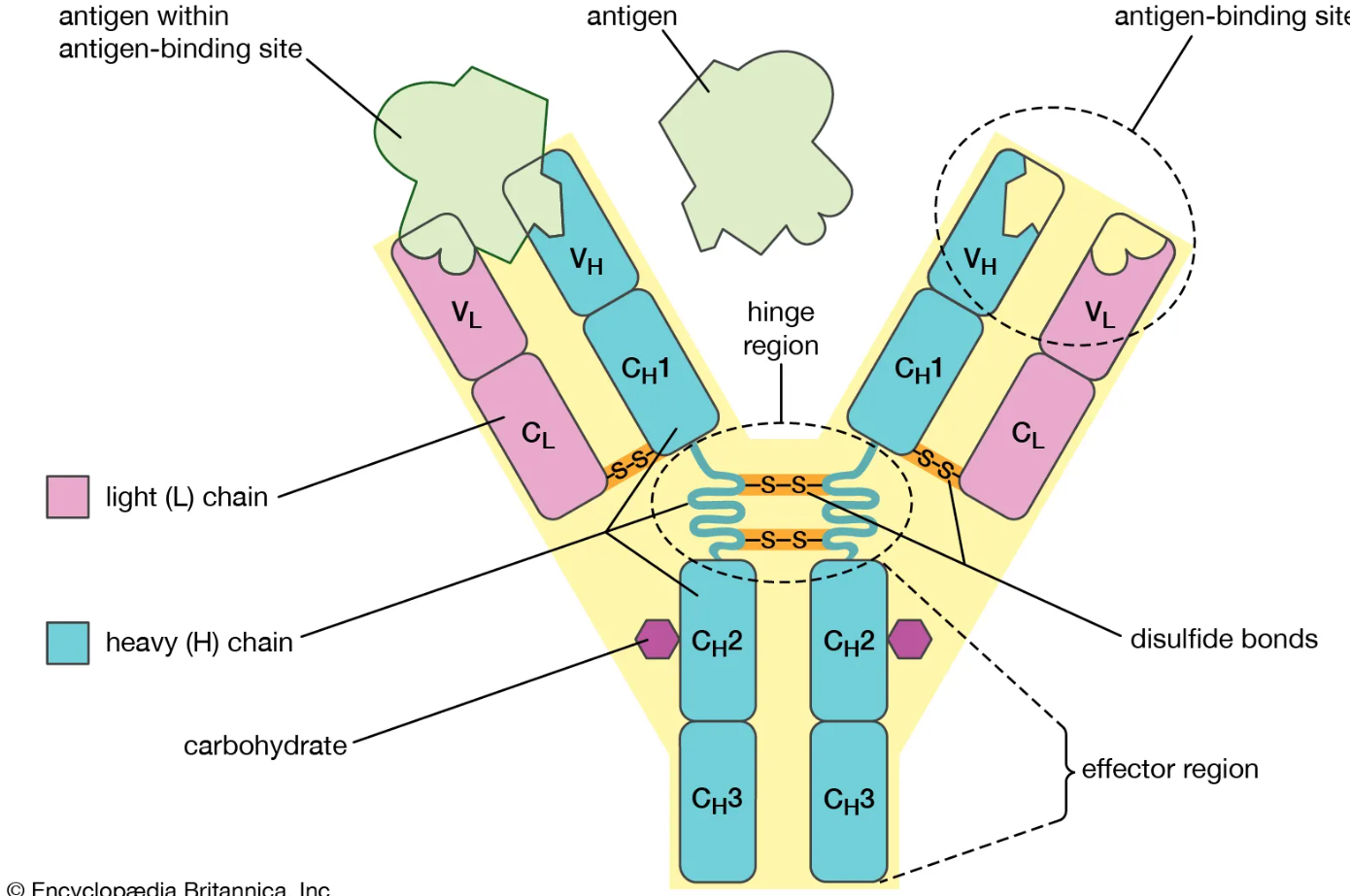
What is the function of the Fc region of an antibody?
Made of constant regions of heavy chains
Does not bind antigen
Engages with Fc receptors and complement proteins for effector functions
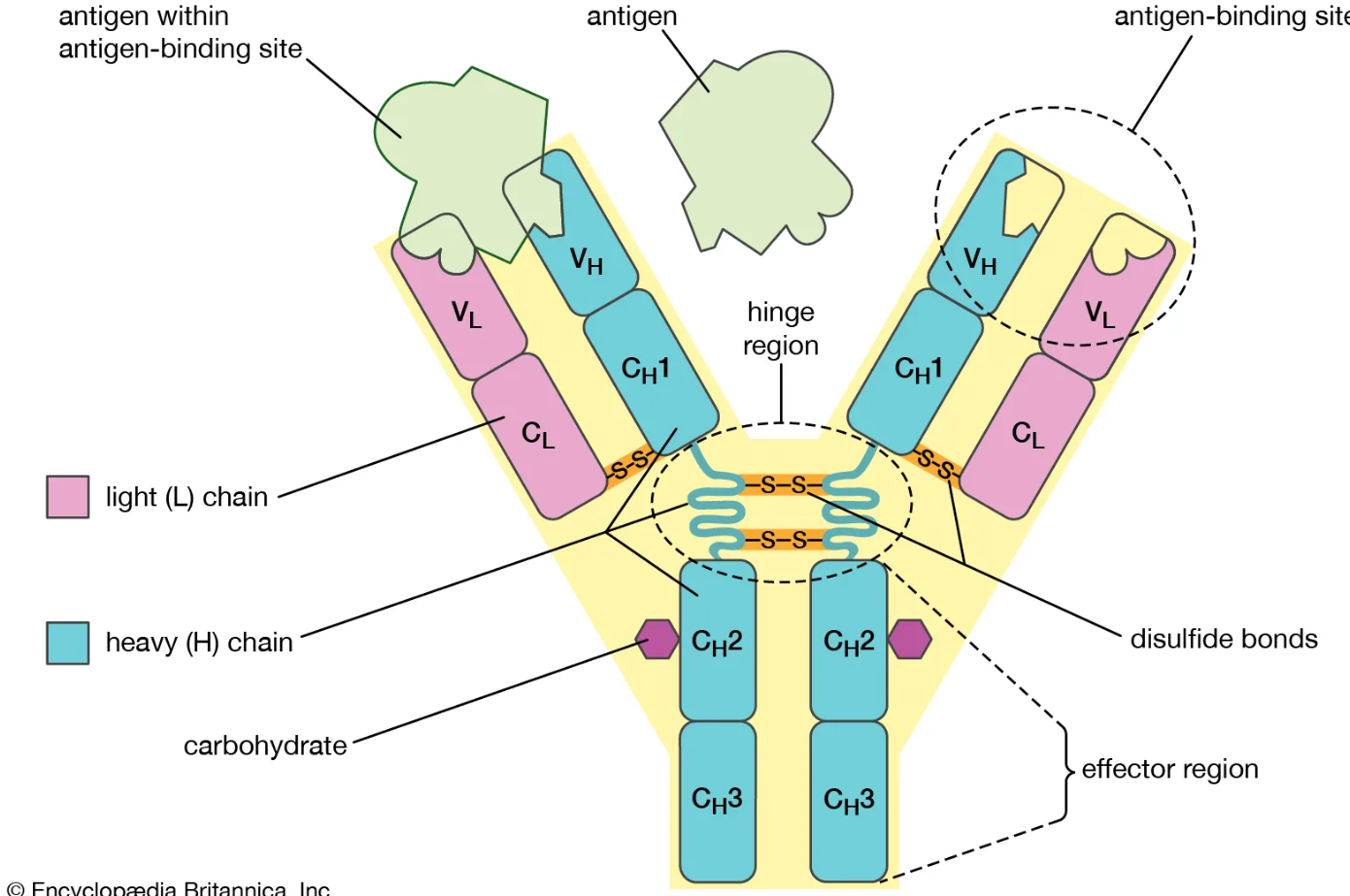
What gives an antibody its flexibility and stability?
Hinge region allows movement of Fab arms
Disulfide bonds stabilise structure but don’t affect specificity
Ig domains within the structure show structural conservation → align across antibody types
How are transmembrane regions of BCR stabilised?
Non-covalent interactions stabilise transmembrane (TM) helices in the BCR.
3 hydrogen bonds form between TM helices.
These are buried within the membrane, not exposed to water.
They help fix the relative positions of the helices, maintaining receptor structure.
Even in the membrane, specific hydrogen bonds (non-covalent) help organize and stabilise multi-pass proteins like the BCR.
How many Ig domains do light chains have?
2
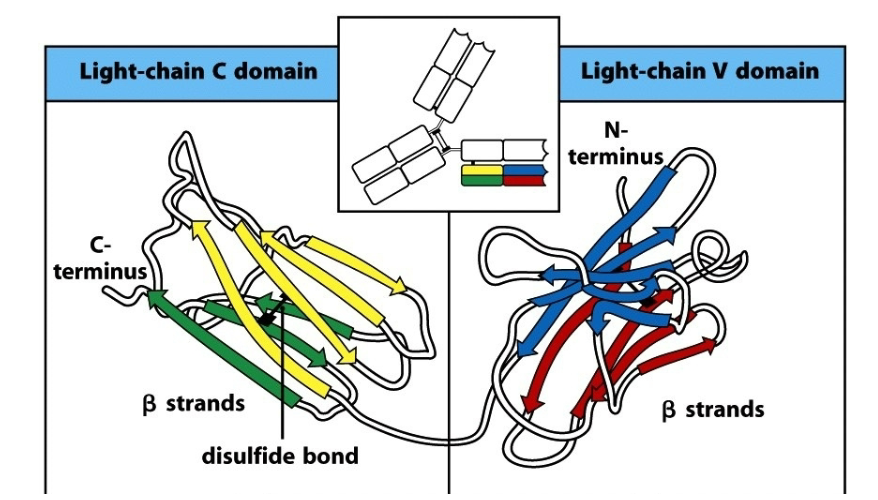
Describe the characteristics of the 2 Ig domains on the light chain in antibodies.
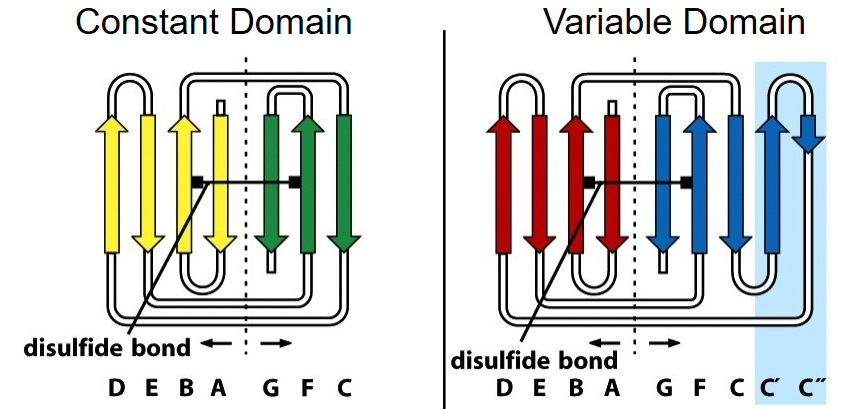
The basic Ig fold is defined as a 7-stranded β- sandwich, a fundamental building principle of Ig domains.
7 stranded β-structure for the C region.
V region includes 2 additional strands, C’ and C’’.
Intersheet disulfide bond enhances stability, but does not determine the fold.
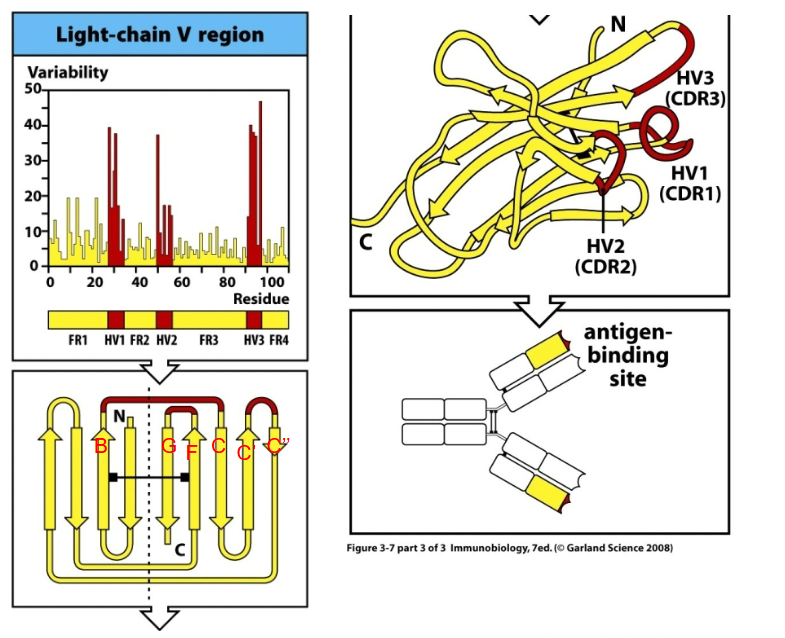
How are antibodies structurally diverse?
Attributed to 3 surface loops (in both heavy and light chains) that are close to each other in space.
CDR1 (B-C)
CDR2 (C’-C’’)
CDR4 (F-G)
CDR = complementarity determining region.
High sequence variability, determining specificity. Antigen binding is determined by 2 sets of CDR loops.
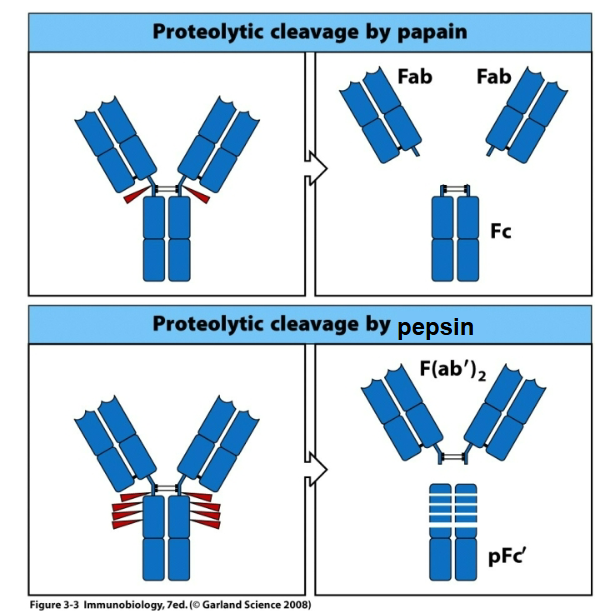
How are antibodies used experimentally?
Using artificial entities produced from proteolytic antibody digest. They do not occur in vivo.
Crystallographic studies of antibody-antigen binding complexes only include the Fab fragment.
What is significant about shape complementarity in antibody binding?
Shape complementarity is a key to molecular recognition and specificity of binding between antibodies and their cognate antigens.
What is specificity?
Small structural changes at the binding interface that result in ‘large’ changes in binding affinity.
Determines discrimination between alternative ligands, thus the immune system is able to distinguish self from non-self.
Describe antibody binding in the context of ligand binding.
The antibody-antigen interaction essentially depicts the interaction of a protein with its ligand.
A large Ka/ small Kd represents a strong interaction. The inverse represents a weak one.
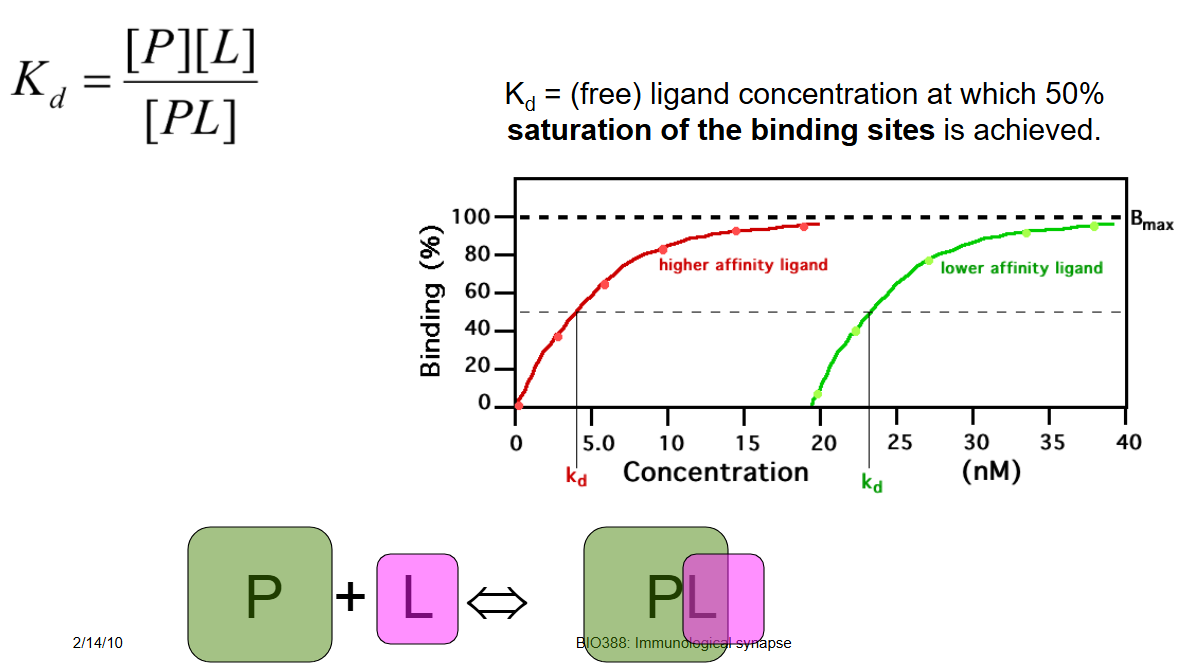
Define Kd
Free ligand concentration at which 50% saturation of the binding sites is achieved.
A lower Kd implies that not much ligand is required to reach 50% occupancy, thus reflecting tight binding of the ligand to its protein.
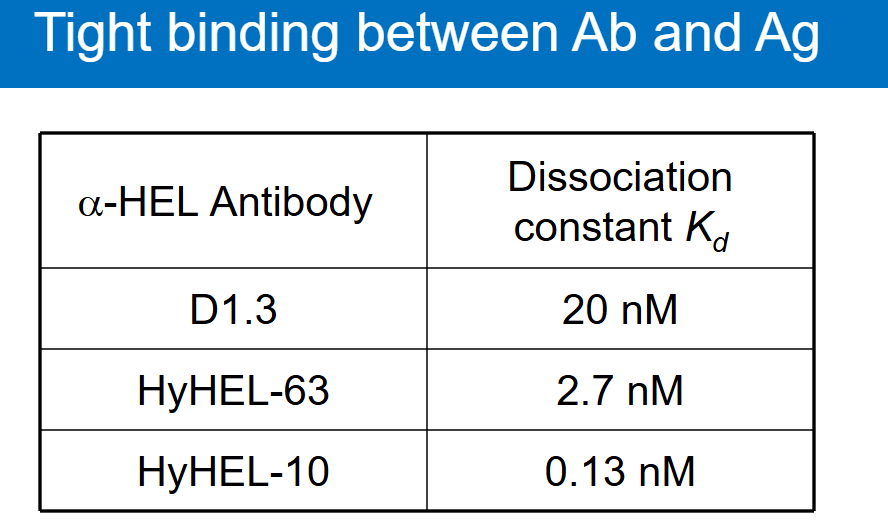
What is a hallmark of antibody binding?
Binding affinities in the low nM range.
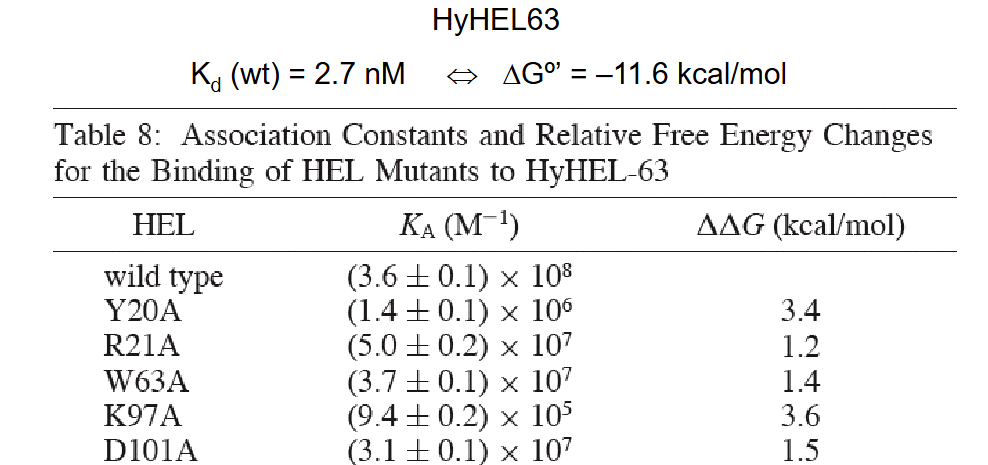

Describe an experiment exemplifying how residue mutations in CDR loops dramatically influence affinity.
Amino acid mutations were introduced in lysozymes though Ala-screening.
Some mutations had more profound changes in Ka than others in response to the mutations.
What are broadly neutralizing antibodies (bNAbs)?
Antibodies that recognise and block a wide range of pathogen strains (e.g., HIV, influenza) by targeting conserved regions (like viral spike proteins) to prevent cell entry.
Key Features:
Target highly conserved (non-mutating) parts of pathogens.
Effective against multiple variants (e.g., HIV clades, flu strains).
Important for vaccine design (e.g., HIV research).
Describe the domains on hemagglutinin.
Receptor domain (variable)
Very immunogenic. Changes only slightly to maintain infectiousness.
HA mediates host cell invasion
HA abundant in viral (membrane envelope)
Stem domain (conserved)
Required for membrane fusion and cell entry
Why are broadly neutralising antibodies (bNAbs) and hemagglutinin (HA) studied together?
HA (especially its conserved stem domain) is a key target for bNAbs in viruses like influenza.
Most antibodies bind HA’s variable head (strain-specific), but bNAbs bind conserved regions (e.g., stem), blocking multiple strains.
Research goal: Design vaccines/therapeutics that induce bNAbs against HA’s conserved parts for broader protection.
How could bNAbs against HA lead to a universal flu vaccine?
By targeting HA’s conserved stem (not the variable head), bNAbs could:
Neutralise multiple flu strains (not just one seasonal variant).
Reduce the need for annual flu shots.
How are researchers designing vaccines to preferentially induce antibodies against conserved pathogen regions (e.g., HA stem)?
Wild-type HA vaccines mainly elicit antibodies against the variable head domain (strain-specific, less protective).
Solution: Use engineered HA stem-only immunogens:
Isolate the conserved stem domain (bNAb target).
Fuse it to a scaffold protein (e.g., ferritin nanoparticles) to maintain its natural trimeric structure.
Scaffolds mimic the stem’s native conformation → immune system produces bNAbs against conserved regions. They avoid distracting immune responses to the variable head.
Goal: Universal vaccines (e.g., for influenza/HIV) that induce broad protection.