Translation
1/80
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
81 Terms
Translation steps
Initiation
Elongation
Termination
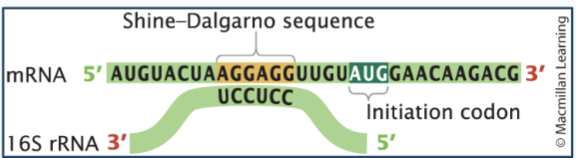
Translation initiation in prokaryotes
The ribosome binds to the Shine Delgarno sequence on mRNA, which positions the ribosome at the correct start codon.
An initiator tRNA, carrying the amino acid fMet (anti-codon), binds to the start codon (AUG) on MRNA with the aid of Initiation Factor proteins.
Initiator Factor (IFs) proteins also assemble the large subunit onto the small subunit with fMet in the P-site, forming the translation initiation complex.
Shine Delgarno sequence
5’-AGGAGG-3’
Translation initiation in eukaryotes
The small ribosomal subunit binds the mRNA at the 5'-cap and scans for the first 5' AUG codon to use as the start codon, which is part of a short consensus called the Kozak sequence.
As in prokaryotes, an fMet initiator tRNA (carrying amino acid fMet) binds at the start codon (AUG) and the ribosomal initiation complex assembles.
Kozak sequence
5’-ACCAUGG-3’
Protein function is determined by
protein structure including shape, ability to change shape, and interactions with other molecules at R-groups. A change in amino-acid sequence of a polypeptide can alter protein function
A mutation changes the amino acid sequence of a protein from: Valine → Aspartic acid
What is the most likely effect on the protein?
A. No effect because both are amino acids
B. Increased stability due to similar properties
C. Possible change in protein folding due to change in R-group properties
D. Protein will always be nonfunctional
C, possible change in protein folding due to change in R-group properties
Change in R-group → change in structure/function
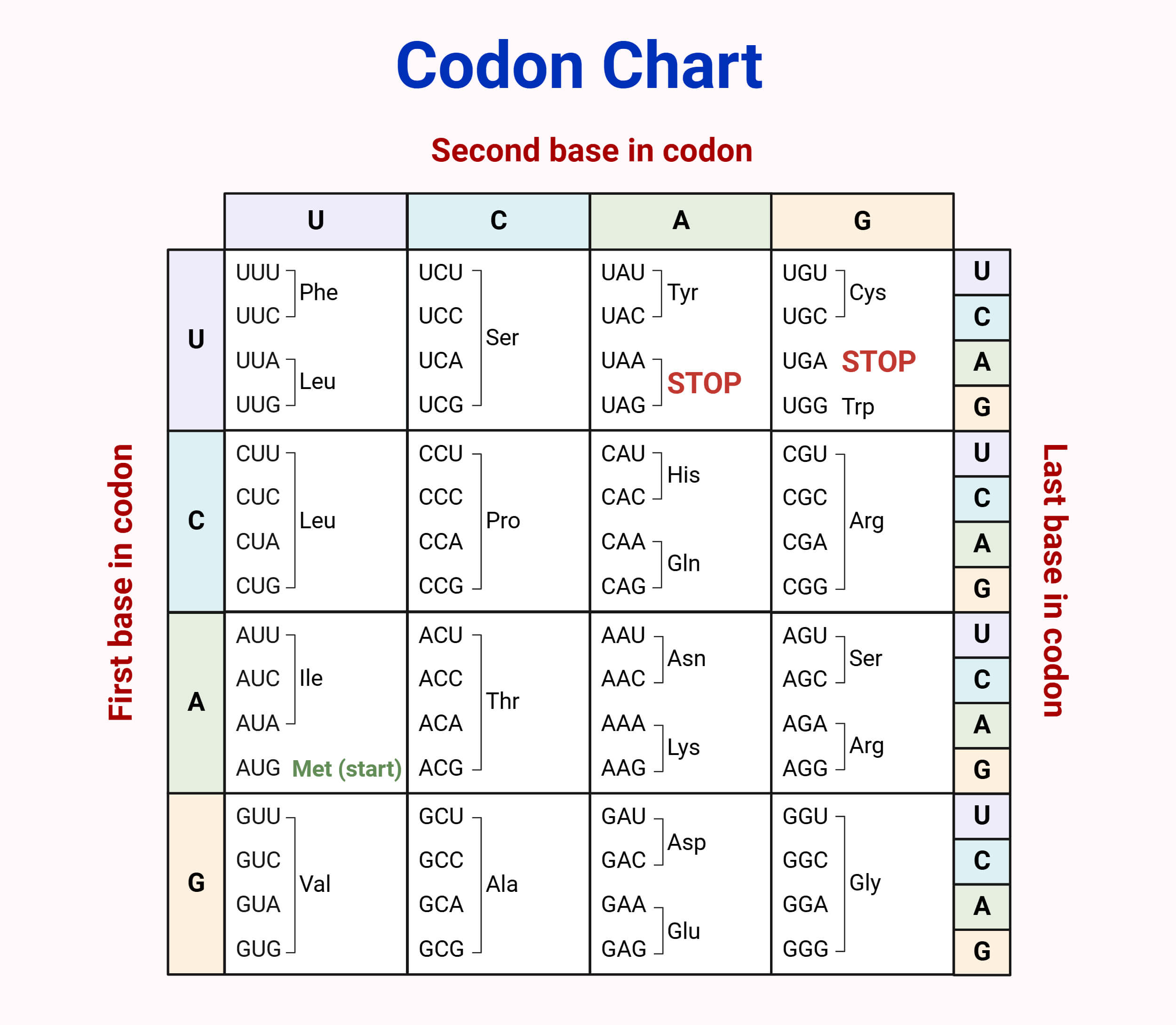
Given the mRNA sequence:
5’–CCGAUGGCUACCUAAUGG–3’
What is the correct polypeptide?
1) Find first AUG sequence
5’–CCGAUGGCUACCUAAUGG–3’
2) Read in codons (set of three RNA nucleotides)
AUG - GCU - ACC - UAA - UGG
3) Translate using codon table
AUG → Met
GCU → Ala
ACC → Thr
UAA → STOP
Therefore, the final polypeptide is Met-Ala-Thr
Structural genes
Genes that encode proteins that do the actual work of the cell; metabolic and structural proteins
DO THE JOB.
Enzymes
Receptors
Structural proteins
Transport proteins
Hormones
Cytoskeletal proteins
Membrane proteins
Metabolic enzymes
Biosynthetic enzymes
Receptor proteins
etc.
What do structural genes make?
proteins (enzymes, receptors, etc.)
Regulatory genes
Genes that encode proteins that control expression of other genes
A gene encodes a protein that binds DNA and regulates transcription. What type is it?
A. Structural
B. Regulatory
C. Metabolic
D. Ribosomal
B - regulatory
Which one of the following would be considered a regulatory gene (not a structural gene)?
A. The TUBA gene product is the cytoskeletal protein tubulin alpha whic polymerizes with tubulin beta to make microtubules
B. The Max gene product is a DNA binding protein that promotes expression of metabolic proteins for cellular respiration
C. The GLUT2 gene product is a hexose sugar transporter that allows liver cells to increase glucose transport when blood glucose is high
D. The GLS gene product is an enzyme that removes the amino group from the amino acid glutamate so it can be metabolized in cellular respiration by mitochondria
B - The Max gene product is a DNA binding protein that promotes expression of metabolic proteins for cellular respiration
What do regulatory genes make?
transcription factors (TFs), which are proteins that control other genes
Types of transcription factors
Activators
Inhibitors/Repressors
Activators
promote transcription; Help RNA polymerase bind or work better = positive control
Inhibitors/Repressors
Block transcription; prevent RNA polymerase from working = negative control
Regulatory genes PRODUCE proteins, while transcription factors proteins carry out positive or negative control
Regulatory genes PRODUCE proteins, while TRANSCRIPTION FACTORS proteins carry out positive or negative control
What are the types of regulatory mechanisms?
Positive control - activators that boost transcription by helping RNA polymerase bind to DNA
Negative control - inhibitors/repressors that block transcription by interfering with RNA polymerase
How do transcription factors carry out positive or negative control of transcription?
They are DNA binding proteins that hydrogen bond to DNA. Transcription factors insert into the major groove and can hydrogen bond to particular base pairs.
DNA-binding motifs
Specific structural features of transcription factors that allow them to bind specific DNA sequences, typically in the major groove, to regulate gene expression (Motif = shape that lets protein bind DNA)
Transcription factor DNA-binding motifs
Helix-turn-Helix proteins
Zinc Finger proteins
Leucine Zipper proteins
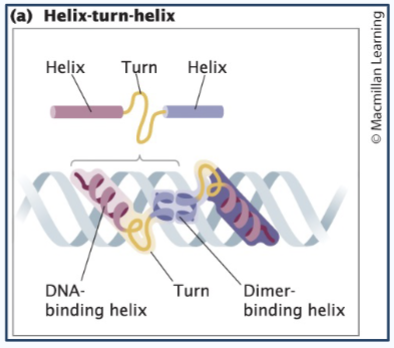
Helix-turn-helix
Two symmetrical helices = dimer that fits into the major groove
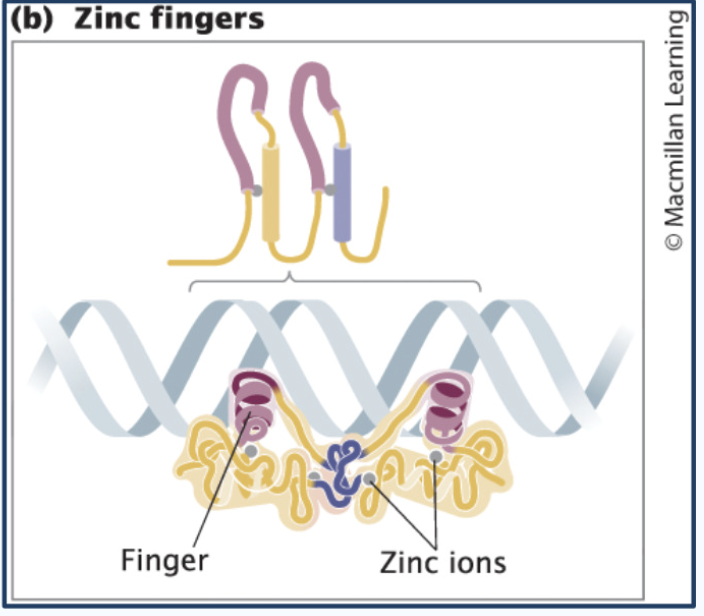
Zinc Finger proteins
Metal ion (zinc) can interact with four different amino acids and pull them together, folding the protein into a particular loop called a zinc finger that exactly fits into the major groove of the DNA
Uses zinc ion to stabilize shape
Forms a “finger” that inserts into DNA
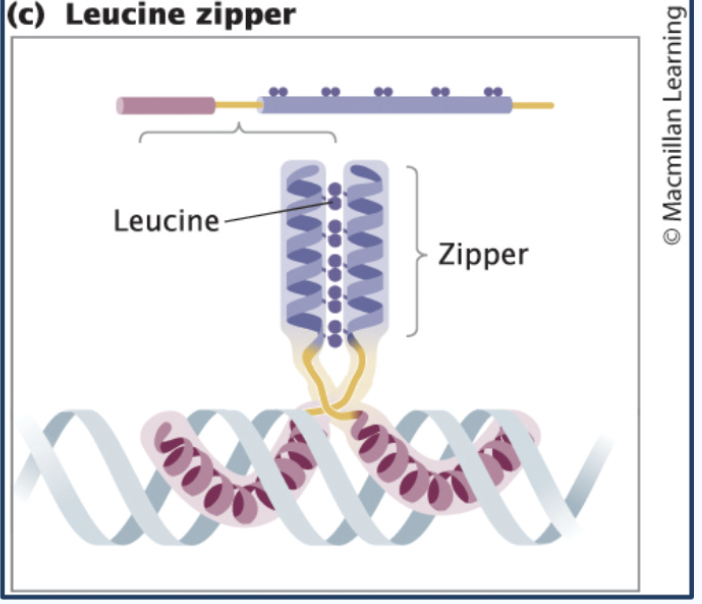
Leucine Zipper proteins
Two proteins stick together (dimer)
Dimer; purple has bunch of amino acid leucine which interacts with neighboring helix and stick together
called a zipper because the two helices are stuck together vertically
proteins zip together → then bind DNA
Prokaryotic gene regulation
Negative regulation used to stop RNA polymerase + sigma factor from attaching to DNA promoter and making RNA
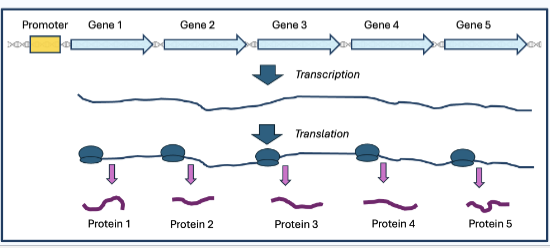
polycistronic RNA
prokaryotic single mRNA molecule that encodes for multiple different proteins at one time
One operon with a single promoter
Multiple protein-coding regions (genes)
Multiple Shine-Delgarno sequences = multiple ribosomes translate the RNA into protein
operon
prokaryotic feature; clusters of related genes under the control of a single promoter. It allows for the coordinated regulation of gene expression, producing a single mRNA molecule for multiple proteins to save energy
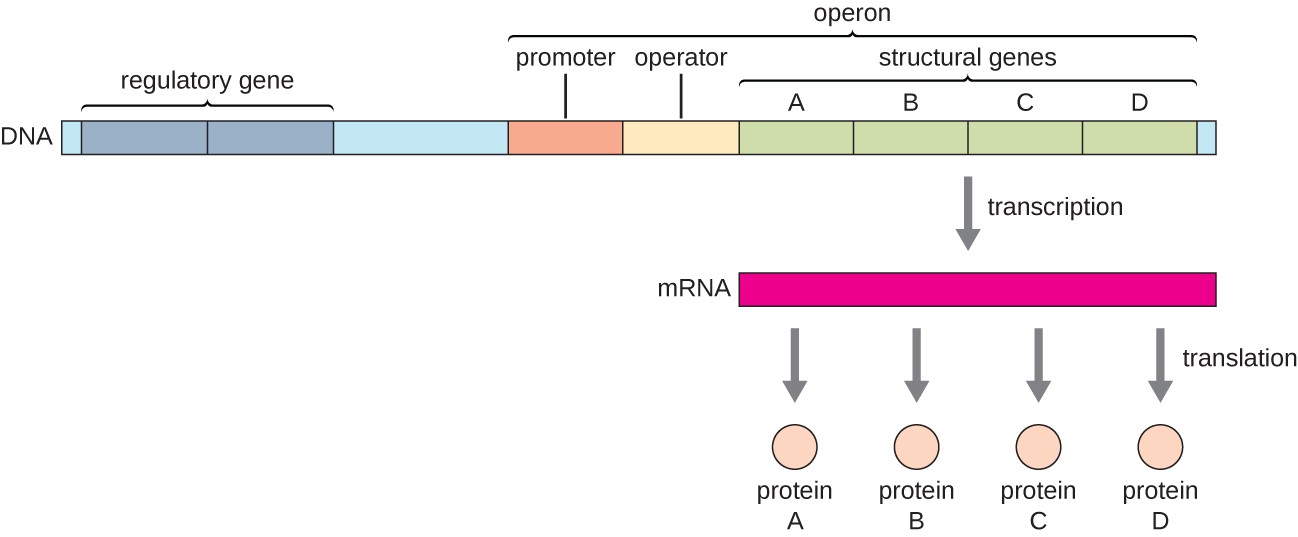
Structure of a bacterial operon
Promoter
Operator
Structural genes (the actual genes that code for proteins that will get transcribed into one mRNA)
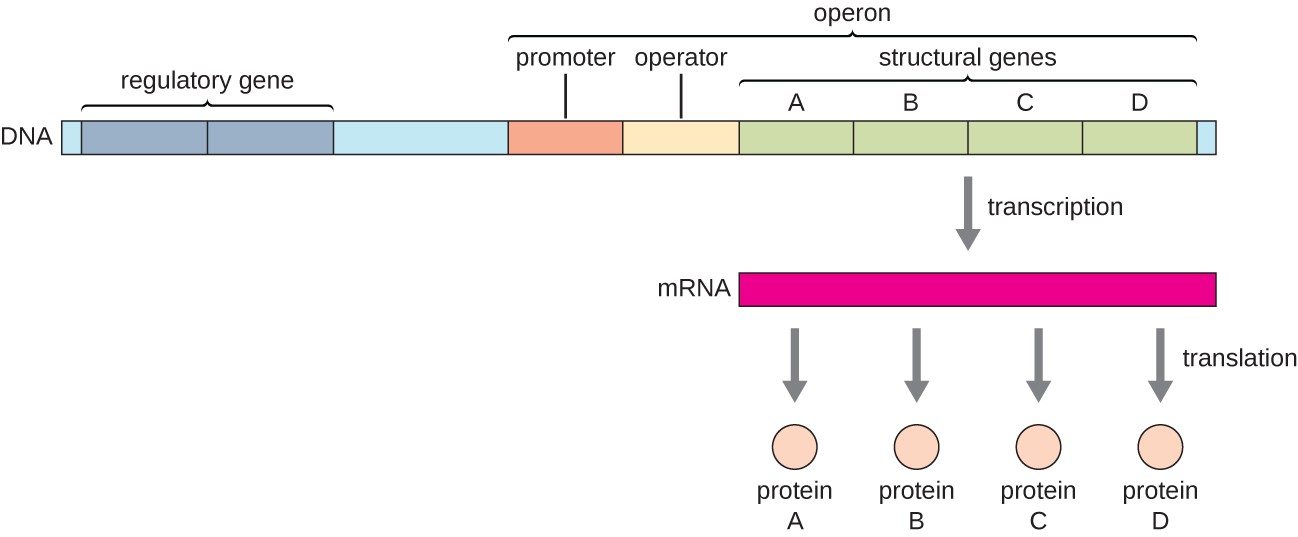
operon promoter
where RNA polymerase binds to transcribe DNA into RNA with sigma factors (holoenzyme); Same promoter controls all genes in the operon
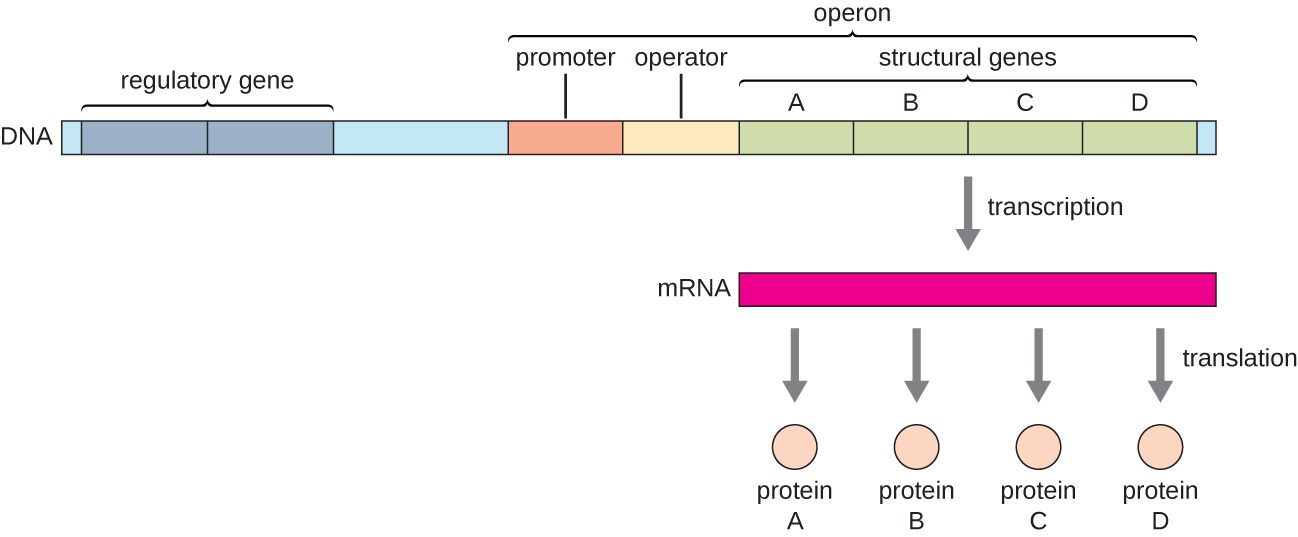
operon operator
Sequence in between the promoter and protein-coding sequences - regulatory region where a repressor may bind
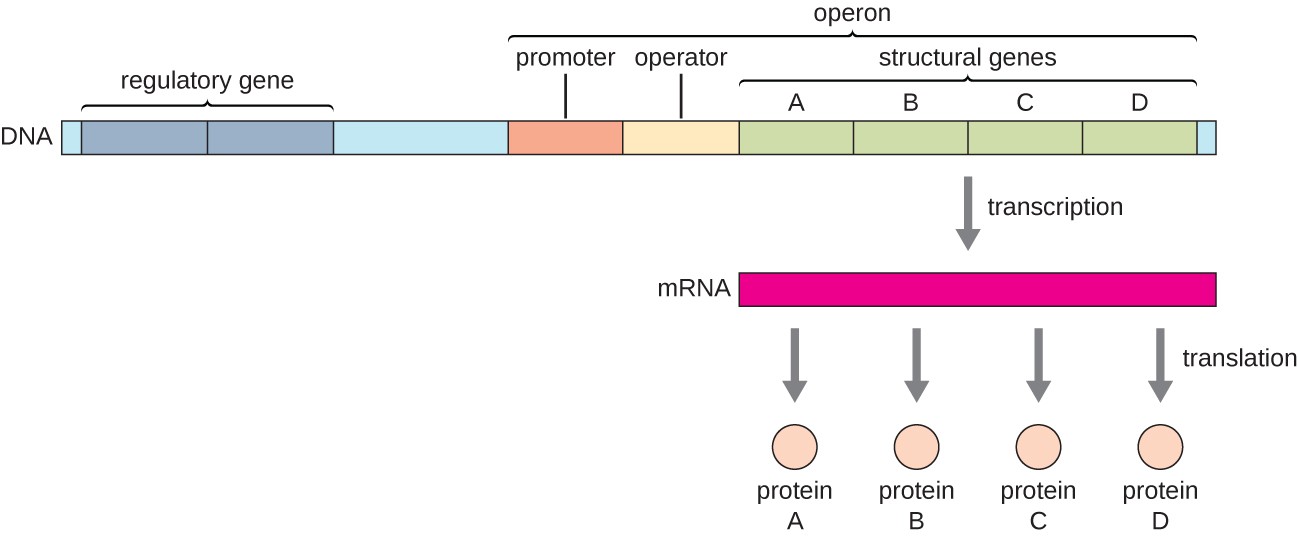
regulatory gene
NOT part of the operon itself, but ahead of the operon; encodes the repressor or activator protein that can regulate the operon by binding at the operator
Types of operons
Inducible
Repressible
Inducible operon
An operon that is normally repressed (default is “off”, no transcription/no RNA made) but can be turned on
Repressor is bound to the operator
However, if an inducer molecule is introduced, it interacts with the repressor and changes the repressor’s shape; the modified repressor falls off the DNA (not activation but DE-REPRESSION, aka removing a repressor), allowing transcription (DNA —> RNA)
Example of inducible operon
lac operon is a group of genes in bacteria, like E. coli, that are used to break down lactose ONLY when lactose is available
It contains structural genes (lacZ that cleaves lactose into its monomers glucose and galactose, lacY that transports lactose into the cell, and lacA that plays a minor role).
The lac operon itself has a promoter, which is where RNA polymerase binds to induce transcription, as well as an operator, which is where the repressor binds to stop transcription.
There is also a regulatory gene, known as lacI, which produces the repressor protein LacI, also sometimes called LacR (lac repressor). Because the lac operon is an INDUCIBLE operon, the LacR repressor is constitutively expressed (the LacI repressor protein is always being made; default is that the LacR repressor is bound to the operator and stopping transcription).
LacR possesses a helix-turn-helix transcription factor (motif), which means it has the right shape to bind to the major groove of DNA at the operator and prevent transcription as a repressor. The LacR repressor acts as a sensor for lactose; when we add the lactose sugar, aka the activator, it binds to the LacR repressor protein and bends it, causing a negative conformational change. The LacR “fingers” no longer fit into the major groove. This causes the LacR protein to pop off from the operator region, allowing transcription.
The lactose is transported into the cell and is processed by the cell - transforms into allolactose (the true inducer).
Inducible operon is…
De-repression (removal of the LacR) repressor protein so that transcription can be turned “on”
Inducer disables repressor via allosteric inhibition
Repressible operon
An operon that is normally transcribed (default is “on”). Usually genes that are commonly needed but in some situations may be turned off.
The regulatory gene makes a nonfunctional repressor protein (wrong shape). A second molecule (a co-repressor) is needed to bind to it to change its shape and make it work.
Once the co-repressor binds to the repressor protein, the repressor protein becomes functional and attaches to the operator and therefore turns off transcription.
Example of repressible operon
trp operon (tryptophan = essential amino acid)
Cells will make trp when needed for protein synthesis (requiring energy). They usually do this by converting one amino acid to another. If too much, or dietary trp available, the pathway is inhibited (more efficient/less costly to use trp when it’s consumed from diet vs. having to make it from scratch, which requires energy)
The trp operon is normally active (transcribed)
The co-repressor is tryptophan (activates repressor) - bacteria no longer needs to waste its own energy to make tryptophan by itself by the normal tryptophan pathway
The trytophan binds to the nonfunctional repressor and changes its shape, causing it to become active (works) and bind to the operator, where it inhibits transcription of the trp operon that contains a cluster of genes that normally produce tryptophan.
Both repressible and inducible operons are examples of what?
negative control, because a repressor protein is used to inhibit transcription
positive control
A regulatory mechanism where activator, binds to DNA to turn on or increase gene expression
Example of positive control
activator protein increases transcription - CAP system (lac operon)
cells prefer glucose because it costs less energy to use; lactose needs to be modified
CAP is the activator
Don’t have a lot of glucose = lots of cAMP because importing glucose is what consumes cAMP= CAP works and can bind to cAMP = cAMP + CAP binding to lac operon upstream of promoter bends DNA, making promoter easier to access = boost transcription
Lots of glucose = less cAMP (b/c costs energy to transport it inside) = CAP doesn’t work, no cAMP to bind = lac operon transcription decreases
In positive control, an activator protein (e.g., CAP in the lac operon) enhances RNA polymerase binding, increasing transcription and allowing efficient expression of operon genes.
If only glucose is high, cells will metabolize glucose
• If only lactose is high, cells will metabolize lactose
• If glucose and lactose are both high, cells will metabolize glucose preferentially
- High glucose makes low cAMP, so CAP is disabled
Attenuation
a backup system that stops transcription EARLY after it starts to STOP leakage
leakage
small amounts of RNA transcript are still made in presence of repressor/when repressor briefly falls off DNA (repressor is dynamic) - producing unwanted mRNA when gene is not needed (ex. when trytophan levels are high)
Example of attenuation (trp operon)
trp operon - trp mRNA transcript contains a leader sequence with 4 different regions (1-4)
Recruitment of ribosome initiates leader peptide translation
When there are HIGH levels of tryptophan, bacteria don’t need to make their own tryptophan. Therefore, the operon should be repressed. The ribosome covers region 2, so that Regions 3 and 4 hydrogen bond together, forming a strong terminator hairpin that halts transcription. RNA polymerase falls off RNA. TRANSCRIPTION WAS STARTED BUT STOPS EARLY BEFORE FULL GENES ARE MADE = ATTENUATION
When there are LOW levels of tryptophan, bacteria need to make their own tryptophan. Therefore, the operon should be induced. The ribosome “covers” region 1, allowing regions 2 and 3 to hydrogen bond together to form a weak hairpin called an antiterminator that halts translation, but transcription of the trp operon continues. (not attenuation)
STRONG HAIRPIN FORMED WHEN TRYPTOPHAN CONCENTRATION IS HIGH TO STOP LEAKAGE AND TERMINATE TRANSCRIPTION.
Why is attenuation important for prokaryotic gene regulation?
Prevents linkage, where unwanted mRNA is not needed when conditions change (ex. high levels of trytophan = stop transcription early because the protein is no longer needed to be made by the bacteria).
Prevents unnecessary gene expression
Eukaryotic gene regulation similarities with prokaryotes
Transcription factors modify RNA polymerase activity
Both positive and negative control
Combinatorial control through regulatory elements (combination of repressors and activators that determine how much RNA is being made)
Eukaryote gene regulation differences with prokaryotes
1) Eukaryotes generally do not use operons
2) Individual eukaryotic genes have dedicated regulatory elements in addition to basic promoter
3) Combinatorial control is more complex
4) mRNA processing (capping, splicing)
5) Chromatin structure (bacteria lack chromatin. eukaryotes have heterochromatin + euchromatin)
6) Compartmentalization of cell also allows new kinds of control (eukaryotic DNA is located in the nucleus. ribosomes are in the cytoplasm = two different compartments, which allows us to control the relationship between transcription and translation. in bacteria, transcription and translation are coupled in the cytoplasm)
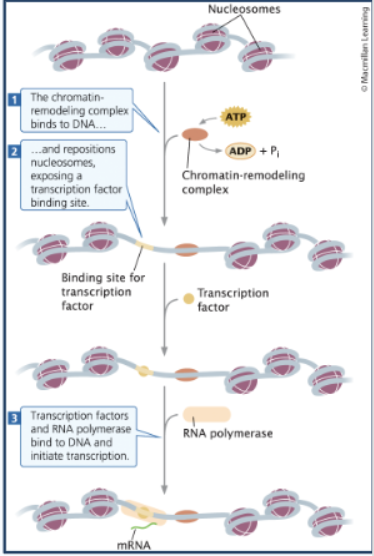
Chromatin remodeling
Chromatin remodeling complexes bind to sites of DNA near promoters, reposition nucleosomes to make chromatin available for transcription
Histone modification
Positively charged histone tails bind to negatively charged DNA phosphate backbone
Modifications to histone tails alter interactions with DNA
Methylation
Acetylation
Ubiquitination
Phosphorylation
Sumoylation
Methylation
Methylation - histone methyltransferase (HMT) adds methyl groups (-CH3) to histone tails, making them more nonpolar; nucleosomes condense into heterochromatin, becoming inaccessible for the cell - PREVENT TRANSLATION
Demethylation - histone demethylases (HDM) remove methyl groups from histone tails, returning histones to their polarity and allowing transcription
methylation is associated with
long term inhibition (barr bodies, heterochromatin)
Acetylation
Acetylation - histone acetyltransferase (HAT) adds acetyl groups (-COCH3) to histone tails, neutralizing the positive charges, and reducing histone tail affinity with DNA = nucleosomes are looser, BOOSTING TRANSCRIPTION
Deacetylation - histone deacetylases (HDA) removes acetyl groups, reintroducing positive charges and increasing histone tail affinity with DNA = nucleosomes are compact, preventing transcription
CpG islands
areas near promoters (transcription start sites) that get heavily methylated
Core promoter
Eukaryotic binding site for basal transcription factors near the transcription start site (+1) that recruit RNA polymerase II for transcription (part of the promoter)
Basal transcription factors
proteins that bind to promoter regions on DNA to activate transcription
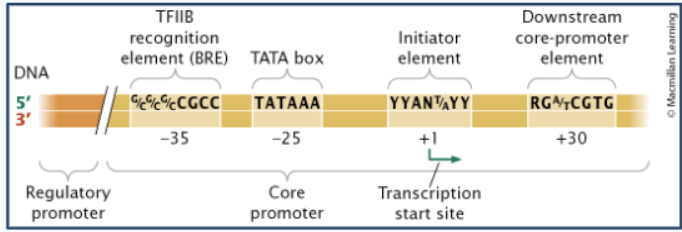
Core promoter components
Basal transcription factors
Initiator (+1 bp)
TATA box (-25 bp)
TFIIB recognition element/BRE (-35 bp)
Downstream promoter element/DPE (+30 bp)
Regulatory promoter
EUKARYOTIC upstream region of the promoter that contain proximal control elements that bind transcription factors (repressors and activators) - so NOT operators
Proximal control elements
Short DNA sequences located CLOSE TO THE PROMOTER that bind transcription factors (activators = increase transcription, repressors = decrease transcription) to regulate transcription
Ex. CAAT, OCT
Enhancers
DNA sequences that ARE FAR AWAY FROM THE PROMOTER but increase transcription by binding activator proteins (helping RNA polymerase)
Regulatory neighborhoods
However, we want to avoid jacking up transcription of the wrong thing with enhancers. Therefore, we can take millions of bases of DNA and organize them into what are called regulatory neighborhoods
regions where within a certain span, all of the proteins/stuff inside can interact with each other; but does not affect adjacent neighborhoods
Insulators
Elements (sequences of DNA) that limit enhancer-promoter interactions and create the regulatory neighborhoods by organizing DNA
Prevent enhancers of one neighborhood from bending over and interacting with the enhancers of another neighborhood
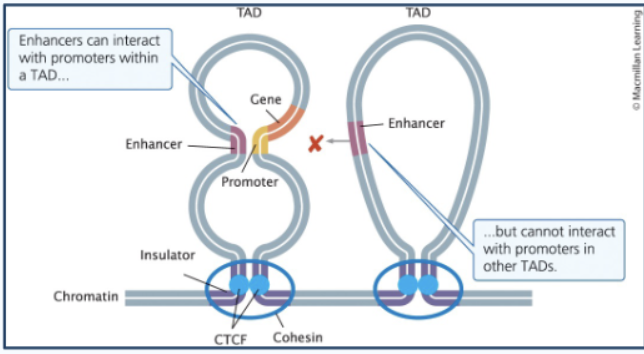
TADs
topologically associated domains - big loops of DNA sticking out of the chromosome - regions of DNA that form loops so that genes and regulatory elements within the same loop (TAD) aka “neighborhood” can interact
Transcriptional stalling
When RNA polymerase II starts transcription but then pauses (20–50 nucleotides)
Why? because the gene is already “primed” - so if the cell needs the gene, polymerase can quickly continue
RNA regulation
control of gene expression by RNA molecules that influence transcription, translation, or mRNA stability
What are some of the ways RNA is regulated?
Riboswitch
Anti-sense RNAs (sRNA)
CRISPR
Micro RNAs
Riboswitch
Prokaryotic stem-loop (hairpin) that may form on the 5’ UTR of mRNA that regulates translation
Discuss the mechanism by which riboswitches regulate translation of prokaryotic mRNA.
They modify translation efficiency by blocking the Shine-Dalgarno sequence (location where prokaryotic ribosomes normally bind) in a negative feedback regulatory mechanism
The hairpin isn’t super strong, so an effector molecule binds to the riboswitch to stabilize its structure, making it a negative feedback regulatory mechanism
The stabilization of the riboswitch’s structure allows it to mask the ribosome-binding site = no translation takes place
Without the enhancer (regulatory protein), the riboswitch assumes an alternative secondary structure, that makes the ribosome-binding site available = translation takes place
Riboswitches regulate translation in prokaryotes by binding effector molecules that induce conformational changes in the mRNA/stabilize its structure which blocks the ribosome binding site and blocking translation. However, without the effector model, the riboswitch assumes a different secondary structure, allowing translation.
CRISPR
Defense system in bacteria against foreign DNA; RNA molecule that guides Cas enzymes (like Cas9) to cut specific, complementary viral (foreign) DNA
Describe the function of the CRISPR system in the degradation specific DNAs in prokaryotic cells
CRISPR - RNA molecule that guides Cas enzymes (like Cas9) to cut specific, complementary viral (foreign) DNA
Provide defense against foreign DNA molecules in prokaryotes
Foreign DNA is cleaved by Cas proteins (can bind to CRISP RNA that is made to specifically hydrogen bond to a sequence)
And a small fragment added to CRISPR array on chromosomes (so the bacteria remembers that same foreign DNA)
crRNAs combine with Cas proteins to cleave foreign DNA
antisense RNAs (sRNAs)
Prokaryotic cells can have non-coding (no translation) regulatory genes that make antisense RNA that is the reverse-complement of an mRNA sequence. The sRNA will bind to an mRNA from a structural gene, blocking translation.
Illustrate the process of RNAi, showing the function of Dicer and the RISC complex.
RNA interference (RNAi) is a gene silencing process that uses small regulatory RNAs in eukaryotes that have an antisense sequence, means they are complementary to mRNA, that bind to mRNA.
RNAi (RNA interference) = the overall system/process
👉 miRNA and siRNA = two types of small RNAs used in that system
Dicer: An enzyme that cuts long dsRNA into siRNA or mature microRNA (miRNA).
RISC Complex (containing Argonaute): A complex that uses a single-stranded guide RNA to identify, bind, and cleave complementary mRNA, preventing protein production.
siRNAs
Eukaryotic cells use siRNA as a defense system. It destroys harmful RNA from things like virsues. Eukaryotes also have a defensive small RNA system for RNAi that uses double-stranded siRNA molecules, made by Dicer cleaving double-stranded RNA in the cytoplasm from retroviruses or transposons.
siRNA combines with Ago proteins to form a RISC complex that cleaves complementary mRNA made by viral or transposon DNA, leading to degradation
siRNA molecules can also initiate transcription silencing of specific genes by recruiting histone methyl transferase enzymes to nucleosomes.
Micro RNAs (miRNA)
small, double-stranded RNA molecules made by cutting a hairpin RNA with the enzyme Dicer.
Eukaryotes transcribe miRNAs that are complementary to sequences on other genes from regulatory genes.
miRNA combines with Argonaut proteins to form a RISC complex, that binds to a target mRNA and inhibits translation.
miRNAs have an imperfect sequence match, and when binding to near the 5'-end do not trigger mRNA cleavage.
miRNAs binding near the 3'-end of an mRNA may trigger "Slicer independent" mRNA by recruiting de-capping and de-tailing enzymes.
The mRNA now loses protection:
No tail → unstable
No cap → exposed
Enzymes quickly degrade the mRNA
Describe the structure and processing of miRNAs and siRNAs, and their role in RNA interference.
miRNAs are derived from hairpin RNAs and bind imperfectly to target mRNAs to inhibit translation or promote degradation, whereas siRNAs originate from double-stranded RNA and bind perfectly to induce direct cleavage of mRNA.
Epigenetics
Phenotypic changes in response to environment; heritable changes to gene expression WITHOUT changing DNA sequence
Mediated by changes in chromatin structure
Can be passed on to new generations but also reversable
DNA methylation/histone modification in epigenetics
DNA methylation is a form of epigenetics
Why?
DNA methylation - add methyl groups to histone tails by histone methyltransferase, usually cystine (C) in a CpG pair, which reduces transcription (not affecting DNA) - Methylation passed down through cell division
DNA methylation is maintained by DNA methyltransferase enzymes that methylate daughter DNA at methylated parental CpG locations (When DNA is copied, the cell copies not just the DNA sequence, but also the methylation pattern)
Example: DNA methylation determines fate of female bees. Larvae fed a diet of ryal jelly, which inhibits methylase enzymes, become Queens, while other female larvae that undergo normal DNA methylation become workers
How epigenetic changes in cells can influence differentiation, metabolism, and behavior
Epigenetic changes alter gene expression without changing DNA sequence → different cell functions and phenotypes
1) Differentation - All cells have the same DNA, but epigenetic regulation like DNA methylation turns some genes ON, and some OFF; (we methylate the genes that are no longer needed, which creates specific, differentiated cells)
2) Metabolism - Epigenetics can control for enzymes and metabolic pathways; ex. DNA methylation can reduce enzyme production, slowing metabolic pathway
3) Behavior - Environmental experience (mother’s behavior) changes gene expression via epigenetics
Mother mouse licks pup a lot —> triggers DNA demethylation of the glucicorticoid receptor (GR) gene; gene becomes more accessible, so there is greater GR expression and pups are able to respond to stress better
Why can methylation be maintained in germ cells (passed on)? (Transgenerational Epigenetics)
Methyl group is added on top of cytosine
It does NOT:
change base pairing
change nucleotide identity
C is still C
Paramutation through epigenetic changes to DNA
epigenetic interaction between two alleles where one allele heritably changes the expression of the other allele without altering the DNA sequence.
You have two alleles:
One is normal (active)
One is silenced (epigenetically modified)
The silenced allele converts the normal allele into a silenced state
The newly silenced allele can now silence other alleles in future generations
through mechanisms such as DNA methylation and chromatin modification, without altering the DNA sequence.
Use the X-inactivation process as an example to illustrate RNA-mediated epigenetic regulation
A long non-coding RNA (Xist) silences one X chromosome in females by recruiting epigenetic modifications (methylate the X chromosome, so the X chromsome becomes heterochromatin). The X chromosome is now silence as a Barr Body and does not code for protein
Genetic imprinting
Genomic imprinting occurs when alleles are differentially silenced depending on which parent they come from.
ex. Angelman syndrome
Problem with maternal allele
Paternal allele is imprinted (silenced)
👉 So:
No functional gene expression
LEFT SIDE: Mutation from the FATHER
You inherit:
Mutant allele (A¹) from father
Normal allele (A⁺) from mother
👉 BUT: The maternal allele is silenced
👉 So: Only A¹ (mutant) is expressed
➡ Result:
👉 Mutant phenotype