Intro Molecular Genetics Exam Three
1/44
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
45 Terms
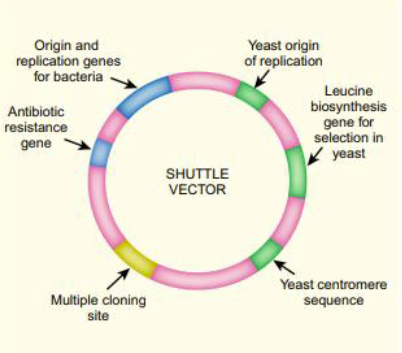
Describe the necessary features of shuttle vectors
→Origin for species 1
→Selectable marker for species 1
→Origin for species 2
→Selectable marker for species 1
→Multiple cloning site
Describe the induction/repression of the lacUV and tet promoter
Components of lacUV
lac promoter (lacUV varient)
LacI repressor
Isopropyl B-d-1-thiogalactopyranoside (IPTG) is an inducer
Default state if OFF. When you add IPTG it turns on. Very useful when you want inducible expression within bacteria
Tet system
Tet-Off system: tetracycline prevents the tTA transcription factor from binding at the promoter. Gene expression is inhibited in the presence of tetracycline
Tet-On system: Tetracycline binds the rtTA transcription factor and allows it to bind DNA at the promoter. Gene expression is induced in the presence of tetracycline.
Compare promoter systems
Components of lacUV
lac promoter (lacUV varient)
LacI repressor
Isopropyl B-d-1-thiogalactopyranoside (IPTG) is an inducer
Default state if OFF. When you add IPTG it turns on. Very useful when you want inducible expression within bacteria
Tet system
Tet-Off system: tetracycline prevents the tTA transcription factor from binding at the promoter. Gene expression is inhibited in the presence of tetracycline
Tet-On system: Tetracycline binds the rtTA transcription factor and allows it to bind DNA at the promoter. Gene expression is induced in the presence of tetracycline.
T7 expression system
T7 promoter, T7 RNA polymerase, Very strong transcription
Very useful when you want very high protein production in bacteria
Name 4 methods of getting vectors into mammalian cells
→Lipid-mediated transfection
→Calcium phosphate
→Electroporation
→Viral transduction
Compare/Constrast transient transfections and stable transfections
Transient Transfection
Transfected DNA not integrated into genome, but remains in the nucleus
Transfected genetic material is not passed onto the progeny; genetic alteration is not permanent
Does not require selection
Both DNA vectors and RNA can be used for transient transfection
high copy number of transfected genetic material results in high levels of protein expression
Cells are typically harvested within 24-96 hrs of transfection
Generally not suitable using vectors with inducible promoters
Stable transfection
Transfected DNA integrates into the genome
Transfected genetic material is carried stably from generation to generation; genetic alteration is permanent
Requires selective screening for the isolation of stable transfectants
Only DNA vectors can be used for stable transfection; RNA by itself cannot by stably introduced into cells
Single or low copy number of stably integrated DNA results in lower level of protein expression
Requires 2-3 weeks of selection for the isolation of stably transfected colonies
Suitable for studies using vectors with inducible promoters
Define transfection and transduction relative to mammalian expression vectors
Transfection: Introduction of vectors, without using viruses, into mammalian cells
Transduction: Introduction of vectors into mammalian cells using viral-mediated delivery
Design a basic mammalian expression strategy
pee pee poo poo
Define a DNA library and explain its purpose in molecular genetics research
Problem: A genome contains thousands of genes, but cloning usually isolates one fragment at a time
Solution: Create a DNA library- a collection of cloned DNA fragments that together represent an entire genome or transcriptome
Key Idea: A DNA library is a stored representation of genetic information
When is it used?: Gene discovery, genome sequencing, functional genomics, protein expression
Distinguish between genomic DNA libraries and cDNA libraries based on the source of DNA and the types of sequences they contain
Genomic DNA library→ Contain fragments representing the entire genome of an organism. It includes both coding and noncoding DNA
cDNA library→ Collection of DNA fragments made from mRNA molecules. Represents genes that are being actively expression
Outline the major steps involved in constructing a genomic DNA library
Isolate genomic DNA
Fragment DNA (restriction enzymes or mechanical shearing)
Insert fragments into vectors
Transform host cells
Store cells
Describe how cDNA libraries are generated from mRNA and explain the role of reverse transcriptase in this process
→mRNA isolation
→Reverse transcription
→Creates cDNA: complementary DNA synthesized from RNA templates
Overall Steps
Isolate mRNA
Use reverse transcriptase to synthesize cDNA from mRNA
Covert to double-stranded DNA
insert into cloning vector
Transform host cells
RESULT: Library representing expressing genes in a specific cell type or condition
Compare advantages and limitations of genomic and cDNA libraries for studying gene structure and gene expression
Advantages:
Contain only expressed genes
Lack introns and regulatory regions
useful for protein expression in bacteria
Applications
Identifying expressed genes
Studying gene expression patterns
producing recombinant proteins
Limitations
Only genes expressed in sampled cells appear
Rare transcripts may be underrepresented
No information about
Promoters
Introns
regulatory regions
Identify common cloning vector used for DNA library construction and explain how insert size influences genome coverage
Coverage depends on:
Genome size
Insert size
Number of clones
Higher coverage increases the probability that any gene is represented
Large-insert vectors dramatically reduce clone numbers
Plasmids: ~5-10 kb
Lambda phage: ~20 kb
Cosmids: ~40 kb
BACs: ~100-300 kb
YACs: up to ~1 Mb
Explain at least two methods used to screen DNA libraries to identify clones containing a gene of interest
→Once library exists, we need to identify the clone of interest
Common screening methods
Hybridization screening
PCR screening
Antibody screening
Functional screening
Overall goal is to find one clone among thousands or millions
Examples One: Hybridization sequencing
→ Uses a DNA probe complementary to the gene of interest
Transfer colonies to membrane
Lyse cells
Hybridize labeled probe
detect signal
positive colonies contain target sequence
Example Two: PCR screening
→Used to scan complex libraries (genomic or cDNA) to isolate specific clones by producing visible DNA bands, which reduces false positives compared to hybridization
Pooling: Library to broken down into structured, hierarchical pool to minimize the number of reactions
Amplification: Primers designed to the target gene are added to the pools to identify positive locations, as described in this article from Intact genomics
Identification: Positive, smaller sub-pool are screened until the specific clone is isolated
Describe the purpose of expression libraries and how they differ from other types of DNA libraries
→Libraries designed so inserted DNA produced protein
Features:
Strong promoter
Ribosome binding site
Translation signals
Purpose: Screen clones based on protein activity or antibody binding
Define terms: dideoxy (chain termination) sequencing
Determines the order of bases by selectively terminating DNA synthesis using modified dideoxynucleotides (ddNTPS).ddNTPS stop strand elongation due to missing 3’-OH group, generating fragments of varying lengths to resolve the seuqnece
Describe the chain termination method for sequencing DNA and how it differs from normal cellular DNA replication
Chain termination: determines DNA sequence by intentionally halting DNA synthesis using modified nucleotides (ddNTPS).
Normal DNA replication contains only dNTPS and terminates when replication finihes, ends of linear chromosomes or colision. Also creates a full length copy of the template strand
Differentiate dideoxynucleotides versus regular deoxynucleotides
→dNTPs are normal bases while ddNTPS are fluorescently labeled
→These fluorescent ddNTPS can not be added onto the next, their 3’ hydroxyl is unavailable
→Each ddNTP has a unique fluorophore
Describe the advantage of sequenase over klenow polymerase
Sequenase: genetically modified T7 DNA polymerase. It has high processivity and efficient incorporation of dideoxy and deoxynucleotides. It also produces more uniform band intensities and has fewer false signals
Describe how DNA fragments separated and detected after chain termination sequencing
Each fragment is typically ran through a gel where it is separated based on size. Laser exites the dye-labeled terminal ddNTPS as they pass a detector, creating a chromatogram that shows the nucleotide sequence/
Describe how a dideoxynucleotide terminates a growing DNA chain
Dideoxynucleotide lacks a 3’-OH group, making it impossible for continue replication
Adapters
Short, synthetic DNA oligos ligated to the ends of DNA fragments to enable amplification on the flow cell, binding to the sequencer, and sequencing primer annealing
Next gen sequencing (NGS)
Set of modern DNA sequencing technologies that perform massive parallel sequencing, allowing for rapid and cost-effective sequencing of large genomes or multiple samples simultaneously
tagmentation
A streamlined library preparation method where a Tn5 transposase enzyme simultaneously cuts (fragments) and tags (ligates adapater sequences to) DNA in one step, replacing traditional multi-step fragmentation and ligation
Barcode sequence
A unique, short DNA sequence added to samples during library preparation that allows multiple samples to be pooled and sequenced together, with the barcode enabling the identification and sorting of reads back to their original sample
Consensus sequence
A representative sequence derived by comparing multiple overlapping sequencing reads (aligned sequences) of the same region, calculating the most frequent nucleotide at each position to resolve errors
Contig
A continuous, assembled seqeunce of DNA formed by aligning and joining together, overlapping sequencing reads, often representing a longer part of a chromosome
Read depth
The average number of times a specific nucleotide position is sequenced, often expressed as 30X or 100X. Higher depth increases accuracy
Emulsion PCR
A method used to amplify clonal, individual DNa templates in separate, water-in-oil micelle droplets. Used commonly in earlier NGS platforms to create polonies
Third gen sequencing
Newer, “longer-read” sequencing technologies, such as Pacific Bioscience and ONT, that sequence single molecules of DNA in real time without needing prior PCR amplification, allowing for much longer reads (thousands to millions of base pairs)
Describe 2 methods of fragmenting genomic DNA or NGS library preparation
Fragmentation by ultrasonic disruption: uses a focused high-frequency acoustic energy to shear DNA within a closed, airtight tube.
Fragmentation by tagmentation:Uses a cocktail of transposases to cleave DNA. Highly efficient and scalable, allowing fragmentation, end repair, and A-tailing to be done in a single tube. It is beneficial for low-input samples, although it may introduce some sequence bias
Describe two methods of creating a target sequencing library for next generation sequencing
PCR: PCR amplify the region of interest then add adaptors to sequence.
Biotinylated oligonucleotide probes: Fragmented DNA is added to biotinylated probes, creating a panel. The probes have sequences complimentary to specific genes or exons
Illumina sequencing
Breaks DNA into fragments, attaching them to a flow cell, and using fluorescently labeled nucleotides to identify bases, allowing millions of sequences to be read simultaneously
→ uses SBS, which tracks the addition of fluorescently labeled nucleotides as the DNA chain is copied, enabling accurate, large-scale, and rapid sequencing
Library preparation: DNA is fragmented and adaptors are added to end
Cluster generation: fragments are attached to a flow cell (glass slide) and amplified (bridge amplifcation) into distinct, clonal clusters to boost the signal
Sequencing: Fluorescently labeled nucleotides are incorporated; a camera records the emission, identifying the base
Data analysis: Billions of reads are aligned to a reference genome ,or assembled, to identify variants
it requires fluorescence, it is slightly slower than ion torrent, its very accurate and you need an optical imaging system
Ion Torrent sequencing
DNA isolation, fragmentation, adding adaptors
Partition each fragment into its own location using microbeads
Emulsion PCR: Separate the beads using oil and water to avoid contamination. Replicate DNA using PCR, so there are thousands of copies of each DNA fragment
Sequencing: Oil is removed and beads are separated into microwells. Primer and polymerase are added. dNTPS are added one at a time and if a dNTP is added, the addition is recorded
detects a pH change, it is faster, lower in repeats and requires a semiconductor chip
It is faster than illumina, but it can not differentiate multiple of the same dNTPS in a row
Nanopore sequencing
DNA/RNA passes through nanopore protein
Changes in ionic current are measured
Signal translated into nucleotide sequence
Developed my oxford nanopore
how does it work?
Motor protein unwinds DNA
Single strand passes through pore
Each base alters electrical current
Machine learning decodes signal
SMRT sequencing
High-molecular weight DNA is fragmented and hairpin adapters are ligated to both ends, forming a circular, single-stranded template called SMRTbell
The SMRTbell template is loaded into a SMRT cells, which contains millions of nanowells. DNA polymerase is anchored to the bottom of each well and binds to the template
As the polymerase incorporates fluorescently labeled nucleotides, each base emits a specific light pulse. This fluorescent label is cleaved off after incorporation, allowing the natural DNA synthesis to continue
The polymerase reads the circularized template multiple times in a rolling loop fashion. This multiple passes are analyzed to correct errors
Understand the principles of 3rd generation sequencing
Massively parallel method of sequencing
Genomic DNA (gDNA) isolation
NGS library construction
gDNA fragmentation
preparing fragments to interact with sequencing platform
Library partitioning
Separating fragments of DNA into clusters of identical copies
Cluster sequencing
Data analysis
Describe nanopore and pacbio technologies
Nanopore: measures changes in electrical current as single-stranded DNA/RNA molecules pass through a protein nanopore in a membrane. Each base causes a characeristic disruption in the current, translated into sequence via machine learning
SMRT: Single-molecule-real-time sequencing uses sequencing by synthesis. A DNA polymerase is anchored at the bottom of a nanopore plate. As it incorporated fluorescently labeled nucleotides, it emits light flashes detected in real time.
Compare Sanger/NGS/TGS methods
Sanger: low throughput, high accuracy, short reads
NGS(Illumina): High throughput, short reads
3rd Gen: long reads, real-time, single molecule
Application/advantages/limitations of Illumina
Advantages:
high accuracy
massive throughput
Cost effective
flexible
Limitations
Short read lengths
computational burden
Initial capital investment
Overclustering risks
No-real time data
Application/advantages/limitations of Ion Torrent sequencing
Advantages:
Rapid turnaround times
Lower captial equitment and operational costs compared to others NGS technologies
Simplicity and compactness
Targeted amplification
Scalability
Limitations
Homopolymer errors
Shorter read lengths
lower throughput
single-end reads
amplification bias
Application/advantages/limitations of Nanopore sequencing
Advantages
Ulta long reasons
Portable devices
real time sequencing
direct RNA sequencing
lost capital cost
Disadvantages
Higher error rates
Signal noise
Data analysis complexity
lower throughput than illumina
Application/advantages/limitations of SMRT sequencing
Advantages:
Long reads
High consensus accurancy (hiFi reads)
No amplifcation bias
Detects epigenetic modifications
Disadvantages
Higher cost than Nanopore
Lower throughput than illumina
Specialized equipment required
Differentiate Klenow polymerase and Taq polymerase
Klenow: lower optimal temp (37C), does proof read, high fidelity
Taq polymerase: high optimal temp (70-80C), low fidelity
Describe the following methods: Southern blotting and genomic library construction with both bacteria and phage
Southern blot→ to detect DNA samples
Bacterial libraries
Vector are plasmids
Genomic DNA and plasmids are cut with the same restriction enzymes. the DNA fragments are ligated into the plmids
Transformation: recombinant plasmids are introduced into E.coli cells via transformation
Baceria are grown on selective media to identify colonies containing the recombinant plasmid
Suitable for smaller fragments
Bacteriophage libraries
Lambda phage virus
Genomic DNA is partially digested and ligated into the phage genome, replacnig non-essential genes
Recombinant DNA is packaged into infectious phage particles in vitro