Cell Bio chp. 15 - Intracellular compartments and Protein Transport
1/59
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
60 Terms
What occurs in different membrane enclosed compartments within the cell
Different metabolic processes
Organelles surrounded by at least one membrane and function
nucleus
Rough ER
Golgi
Mitochondria
lysosome
endosome
peroxisome
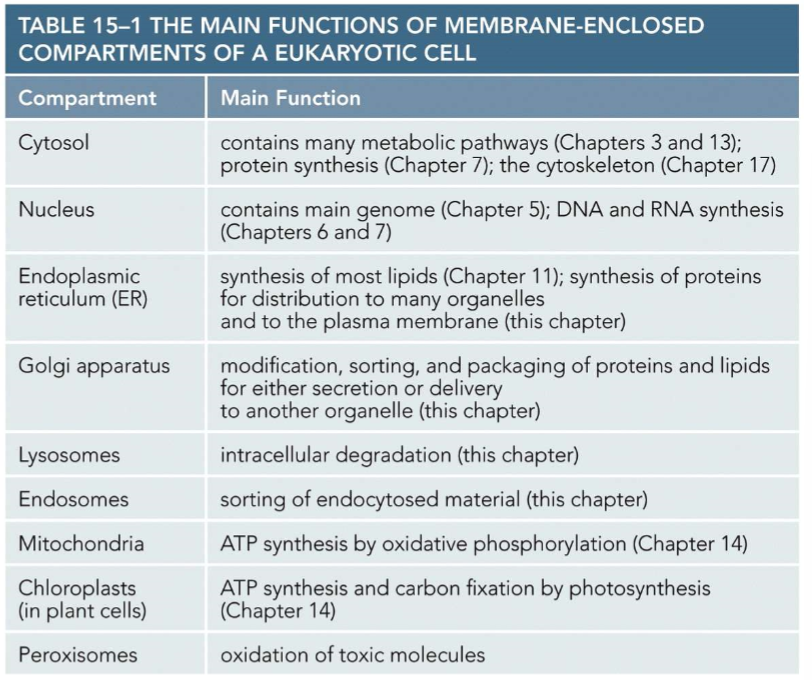

Where you may find some cell components (In class)
Enzymes for Krebs - Mitochondria
Enzymes for photosynthesis - Chloroplast
Enzymes for unraveling DNA, DNA replication - Nucleus
Enzymes that are receptors - cell membrane
Enzymes for detoxification - lysosomes and peroxisomes
Cytoskeleton - Cytoplasm
How do proteins move (In class)
proteins traffic naked
pack proteins into vesicles
Where are all proteins made (In class)
They are all translated in free cytoplasmic ribosomes
(Later flashcards mention either continuing translation in the cytoplasm or going to the ER)
Path of proteins traveling in ribosomes (In class)
Translation starts in cytoplasm then moves to ER where it can be transported as a vesicle
Path of naked protein travel (In class)
starts in cytoplasm and gets to translocators except for peroxisomes and nucleus
Liver cell name
hepatocyte
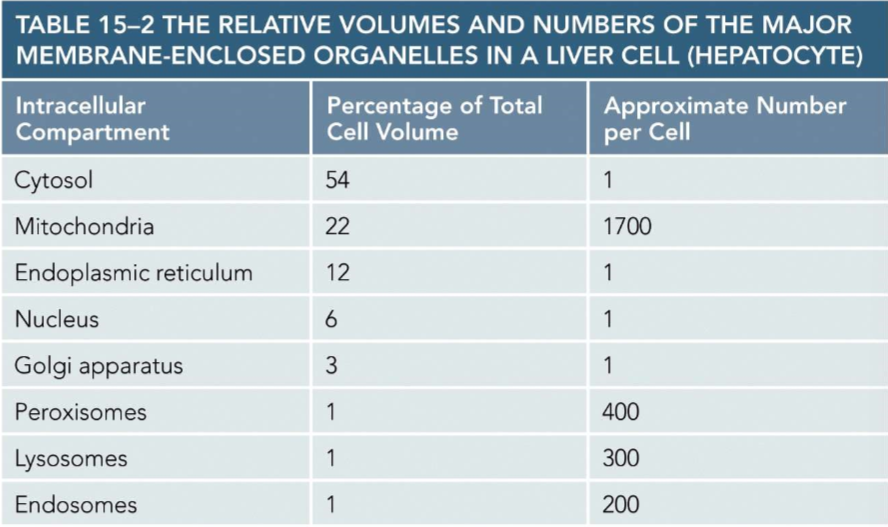
Volume and number per cell of membrane enclosed organelles within liver cells
Do not need to remember specific numbers. Just know generally which make up majority of cell
Two ways membrane bound organelles evolved
invagination: Cell membrane folded in on itself to form a membrane within the cell (Formation of nucleus, ER, and Golgi from ER folding itself)
Endosymbiosis theory: pre-eukaryotic cells engulfed other prokaryotic cells to create membrane bound organelles (formation of mitochondria and chloroplast)
3 mechanisms in which proteins are transported into organelles
1) Transport through nuclear pores (entering nucleus)
2) Transport across membranes via protein translocators guiding unfolded proteins across membranes(entering mitochondria/chloroplast)
3) Transport by vesicles (ER → Glogi)
(Peroxisomes can do vesicles and translocators)
Signal sequence
Amino acid sequence that directs protein to a specific location/compartment within the cell.
Which side of polypeptide is signal sequence towards
N-terminus
How many amino acids typically make up a signal sequence
15-16 amino acids
Features of signal sequences that determine which organelle they will go to
length of sequence
amino acid properties
what to remember for sequence that directs to nucleus
Has a bunch of lysine’s (Lys)
signal sequence for protein going to cytoplasm
Does not have a signal sequence
Signal sequence experiment
ER protein with an ER signal sequence resides in the ER while a cytosolic protein without a signal protein resides in the cytosol. Genetic engineers can remove the signal sequence and attach it to the cytosolic protein so that the ER protein is in the cytosol and the cytosol protein is in the ER.
Nuclear envelope
double membrane that encloses nuclear DNA and is made up of the inner nuclear membrane and outer nuclear membrane
Inner nuclear membrane
contains proteins that are binding sites for chromosomes and proteins that anchor nuclear lamina
nuclear lamina
mesh of protein filaments lining the inside of the inner membrane to provide structural support.
(the cell cortex of the nucleus)
outer nuclear membrane
resembles the ER and is continuous with the ER membrane
nuclear pores
penetrate the nuclear envelope and can allow proteins to enter the nucleus. Is specific to what is allowed in.
nuclear pore structure
cytosolic fibrils: fibrils on outside of nuclear envelope
nuclear basket: fibrils on nuclear side form a basket structure
pore complex proteins: Proteins within the membrane that create a gel like structure with fibrils that allow for specified molecules to enter
Nuclear transport steps
cargo proteins contain a nuclear localization signal that is recognized by a nuclear import receptor
nuclear import receptors guide the protein to a pore by interacting with cytosolic fibrils
nuclear import receptors along with the protein go through the unstructured, specific pore protein region until entry where the protein is released
nuclear import receptors go back into the cytosol through pores to be reused
Do proteins change structure when passing through nuclear pores into the nucleus
Proteins structures remain unchanged
(Only organelle where proteins do not need to unravel to pass through)
Is energy needed for proteins to go through nuclear pores
Yes, GTP is used
Nuclear import receptor process with GTP
After entering the nucleus with a protein, Ran-GTP binds to the receptor and facilitates the release of the protein.
The receptor is transported back into cytosol through a pore, still bound to Ran-GTP
In the cytosol, Ran hydrolyzes GTP and Ran-GDP and Pi detach, allowing the nuclear import receptor to bind to another protein
What must proteins do before entering the mitochondria and chloroplast
They need to unfold
Mitochondrial precursor protein entry process
Precursor proteins signal sequence binds to a receptor protein on the outer membrane
The receptor is associated with a translocator protein (TOM) that transports the signal sequence from the outer membrane to the intermembrane space
The complex of signal sequence, receptor, and TOM diffuses laterally until the signal sequence is recognized by a second translocator protein (TIM)
The two translocator proteins diffuse the protein across the outer and inner membrane while unfolding the protein
When inside the matrix a signal peptidase will cleave off the signal sequence
Chaperone proteins will refold proteins in the matrix using the hydrolysis of ATP
Why must proteins unfold when passing through translocators
Translocators have a narrow opening
Two places proteins can enter peroxisomes from
Cytosol and ER
(Only organelle that can allow proteins to enter through translocators and vesicles)
Process of Protein transport from cytosol to Peroxisome
Peroxisome proteins contain three amino acid signal sequences that are recognized by receptor proteins in the cytosol
The protein is delivered and enters the peroxisome by the receptor protein that then returns to the cytosol after
There are also translocator proteins on the peroxisome protein but these do not unfold the protein
Process of protein transport from ER to Peroxisomes
Vesicular transport
Bud off ER and fuse with Peroxisome membrane
Or become Peroxisomes themselves
Zellweger syndrome
abnormalities in the brain, liver, and kidney due to not being able to detoxify
will not make it past 6 months
Zellweger syndrome causation
Mutations that block peroxisome protein import due to lack of translocators
What happens to polypeptides (proteins) as they are transferred to the ER
They are simultaneously being synthesized
Where do proteins with no ER signal end up
In the cytosol
How do ribosomes end up on the ER to make up the rough ER
they are dragged onto the ER surface by the signal sequence of a polypeptide it is translating
Polyribosome
Multiple ribosomes binding and translating a single mRNA
Process of ER protein entering ER
Ribosome in a pool of subunits within the cytosol begins translating mRNA
If the protein has an ER signal sequence it will guide the ribosome to the ER
The proteins signal sequence will associate to a translocator on the ER membrane where it can enter the ER lumen
multiple ribosomes may begin synthesizing the same mRNA sequence (polyribosome)
Do proteins need an energy source to enter the ER
No, they are translocated as they are being made so no additional energy is needed
SRP
Signal recognition particle
Soluble protein transport into the ER process (More detailed process)
SRP binds to both the ER signal sequence and the ribosome, slowing protein synthesis
The SRP-ribosome complex (SRP, ribosome, and polypeptide) then binds to an SRP receptor in the ER membrane
SRP is displaced and can be reused
ribosome and ER signal sequence are passed to a protein translocator in the ER membrane
Growing then proceeds normally and the translocator transfers the newly synthesized protein into the ER lumen as a loop
A signal peptidase will eventually cleave the signal sequence from the polypeptide
The signal sequence stays in the bilayer and is degraded
The finished soluble protein enters the ER lumen and the translocator closes
What determines the arrangement of proteins in the lipid bilayer
Start and stop signals
hydrophobic Stop-transfer sequence
hydrophobic start-transfer sequence
(Not soluble)
Transport to the ER of transmembrane proteins
Start-transfer sequence binds to a protein translocator initiates transfer
when a stop transfer sequence enters the translocator protein, both sequences are transferred to the lipid bilayer
a signal peptidase will cleave off the start-transfer sequence
A transmembrane protein anchored by the stop-transfer sequence is left
Side that N-terminus and C-terminus face for transmembrane proteins
N-terminus: Faces lumen and outside of cell
C-terminus: Faces cytoplasm always
Inward endocytic pathway
Endocytosis ingests extracellular molecules in vesicles derived from plasma membrane
Vesicles are delivered to early endosomes then late endosomes and finally lysosomes
Outward secretory pathway outcome 1
Proteins from the ER pass through the Golgi to early endosomes then to late endosomes and finally a lysosome
Outward secretory pathway 2
Proteins from ER go through Golgi and are then passed to the plasma membrane where they go through exocytosis
Can proteins enter the Golgi from the cytosol directly
No, must come from ER or plasma membrane as a vesicle
Protein coated vesicle
membrane enclosed vesicles are surrounded by proteins on its cytosolic surface formed by budding off protein coated regions
clathrin
Protein that makes up coat of transport vesicles budding off the Golgi (outward secretory pathway) or the plasma membrane (inward secretory pathway)
Dynamin
GTP binding protein that closes the coated vesicle when budding off membranes
Adaptins
assemble clathrin on surface of vesicles and help capture cargo molecules and bind them to cargo receptors
transport signals on cargo molecules
Protein coated vesicle formation process
Adaptin binds to a cargo receptor with a cargo molecule which binds adaptin to cytosolic exterior of membrane
vesicle formation begins and dynamin wraps around budding vesicle neck
Dynamin will hydrolyze its GTP and other proteins will help to pinch of vesicle
when the vesicle is formed the protein coat of clathrin and adaptin leaves
the naked transport vesicle can fuse with its target membrane
Types of coated vesicles
Clathrin coated (From Glogi): made up of clathrin and adaptin 1 and end up at the lysosome via endosomes
Clathrin coated (From Plasma Membrane): made up of clathrin and adaptin 2 and end up at endosomes
COP-coated: Come from ER, Golgi cisterna, or Golgi apparatus, made of COP proteins, and arrive at the Golgi apparatus, Golgi cisterna, or ER (respectively)
What does vesicle docking depend on
Tethers and SNAREs
Rab proteins
GTP-binding proteins present on surface of transport vesicles and organelles to serve as a marker that ensures transport vesicles fuse with the correct membrane
tethering proteins
Filamentous transmembrane protein involved with docking of transport vesicle with target membranes