T1 BIO S2
1/83
Earn XP
Description and Tags
askjgajehgalkwjhegjklew
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
84 Terms
DNA
deoxyribonucleic acid
structural unit of info in cells
stores and transmits genetic info, universala
Chromosomes
DNA in prokaryotes
found in the nucleoid region
circular dna, doube stranded
once chromosome
nucleoid region also contains RNA and proteins
plasmids - small amounts of circular DNA seperate from chromosomal
DNA in eukaryotes
found in nucleus
human somatic cells have 46 chromosmes (23 pairs) - 22 autosomal and one pair of sex
set of human chromosomes - karyotype
DNA in eukaryotes cont.
arranged in condensed linear strands - DNA molecules and histone protein
ends of chromosome have short DNA lengths called telomeres
telomere protects tips of chromosome from breakdown, stop them from binding to each other
chromosmes are only visible w light microscope when condensed (tightly coiled around histone)
chromosomes in eukaryotes - condensed and decondensed
decondensed - chromatin, important it can be accessed during drowth and syntheesis of cell so DNA, protein and RNA synthesiss can happen
must condense for cell division to prevent damage and ensure identical copies are transwered
membrane-bound organelles with DNA
mitochondria (mtDNA) and chloroplasts (cpDNA)
both double stranded, circular, not bound to histone
theorised that the reason they have DNA is because they were once free living unicellular organismsn
2 reasons for DNA strcuture
make identical copies so genetic info can be passed from cell to cell and inherited by next gen
provide code that can be used by cells to manufacture protein molecules
nucleotides include what? and are are bonded togehter by what type of bond?
bond - covalent bond
pentose (5-carbon) sugar - deoxyribose
phosphate group
one of 4 organic bases - adenine (A), thymine (T), cytosine (C), guanine (G)
DNA directionality, phosphate and sugar ends
phosphate of one nucleotide attatched to sugar of next - backbone alternates
one end of DNA has phosphat other sugar - giving directionality
end of strand with phosphate sticking is called 5’end (5 prime) - beginning of strand
sugar end with hydroxyl group is 3’ end (3 prime)
therefore sequence of dna written in 5’ to 3’ direction
phosphate is negatively charged, DNA has overall negative charge
bases
each form weak hydrogen bonds with complementary base - easy to break and reform
important for dna replication - break apart double helix to expose bases for base pairing
A to T, C to G
reading the strands and strands property
DNA assembly has 2 complementary strands linked by complementary base pairs
read in opposite direction, always start at phosphate then to sugar end
chromosome stuff
made of genes - unique sequence of bases on one strand of DNA (coding strand)
varies from one DNA molecule to other - enables it to be so versatile and diverse
diff species have diff characterists, therefore diff no of genes because they need hundreds of proteins to function
diff in chromosme no, base sequence, length of DNA molecules
DNA replication - general info (NOT THE PROCESS)
nucleoid region in prokaryotes, nucleus in eukaryotes
semi-conservative (see flashcard)
since the DNA strands of the parent are complementary, nucleotides are added in the opposite direction
5’ to 3’ direction by DNA polymerase to expose bases of old strands
product is 2 identical double-stranded DNA molecules
amount of DNA doubles during replication before chromosomes separate during cell division - no of chromosomes stays the same
semi conservative replication
each double strand consists of one old strand and one newly synthesised one
DNA replication summary
helicase enzyme breaks hydrogen bonds between complementary bases joining 2 strands, unwinds and exposes bases, creating a replication fork
topoisomerase prevents supercoiling at the fork, and SSB (single stranded binding proteins) proteins prevent seperated strands from coming back together
primase attatches to strands and creates a short RNA primer to show DNA polymerase where to start.
DNA polymerase adds free DNA nucleotides to exposed corresponding bases on seperate strands according to complementary base pairing rule
new DNA synthesised in 5’ to 3’ direction - DNA polymerases add DNA nucleotides to 3’ end of DNA strands
DNA polymerase deletes RNA primers and adds DNA nucleotides in their place
Ligase joins fragments on lagging strands together
DNA polymerase does a double check
new double-stranded DNA rewinds to double helix, joined at centromere
gene
unique sequence of DNA nucleotides that code for polypeptide chain, protein, or RNA molecule
heritable, control specific characteristics
located at specific locations (locus)
3 parts: promotor region, coding sequence, terminator sequence
genome
total no. of genes in an organism
genomics - study of genomes
1990 - internation effort to map entire human genome (location of all genes in all chromosomes) - first draft published 2001 and used in various fields
RNA
ribonucleic acid, essential for expression and regulation of genes
made of nucleotides
contains:
pentose (5carbon) sugar called ribose
phosphate group
adenine+URACIL (2 hydrogen bonds), cytosine+guanine (3 hydrogen bonds)
4 major types - messenger RNA (mRNA), transfer RNA (tRNA), ribosomal RNA (rRNA), micro RNA (miRNA)
all are single stranded
exons
coding sequences of DNA thats translated into polypeptides/proteins
2% of DNA is exons
highly conserved because changes to code can result in change to translated protein
introns
noncoding sections of DNA, do not code for polypeptides/proteins
can be transcribed into RNA molecules such as tRNA, rRNa, and miRNA
98% of human DNA is introns
less conserved as they are non-coding
most prokaryotes dont have introns but have exons bceause they need coding sections to code proteins
genes for exons and introns
contain both
trasncirbed into premature messenger RNA (pre-mRNA) strands in nucleus as part of protein synthesis
introns must be spliced out of pre-mRNA forming mature mRNA before transltated into amino acid sequence → process is called RNA splicing
messenger RNA (mRNA)
syntehsised using DNA as template in process known as transcription occurs in nucleus,
following some post-transcriptional modification to become mRNA it moves to cytosol
single stranded, consists of sequence of RNA nucleotides - varied length
codes for specifc sequence of amino acids
every 3 bases (RNA codon) codes for specific amino acid
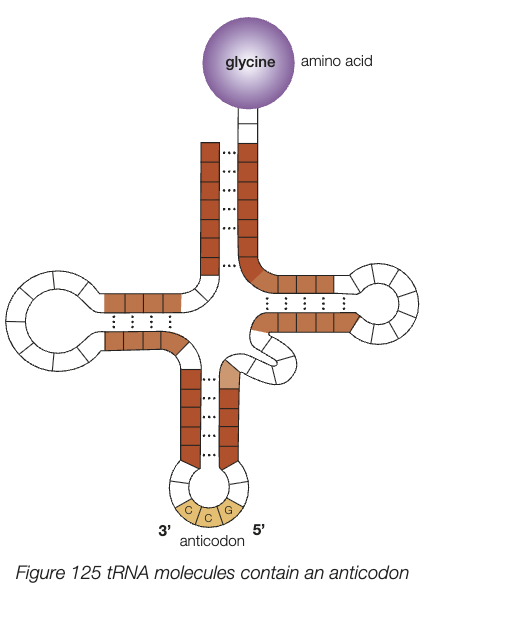
transfer RNA (tRNA)
roughly 80 nucleotides long, 3 dimensional clover leaf shape
contains anticodon - 3 bases complementary to specific mRNA codon
carry specific amino acid at opposite end of anticodon
function - place specific amino acid into correct sequence in polypeptide
after it transfers, it collects another corresponding amino acid in cytosol
amino acid tRNA attatchment involves enzymes with specific shapes so each amino acid is only ever attathced to their corresponding tRNA
ribosomal RNA (rRNA) and ribosome facts
essentially mRNA but when its at the ribosome
ribosomes are made of them and protein molecules
ribosomes - sites of translation in protein synthesis, found often attached to ER in cytoplasm
move along mRNA tranaslating mRNA into amino acid sequence (polypeptide molecule)
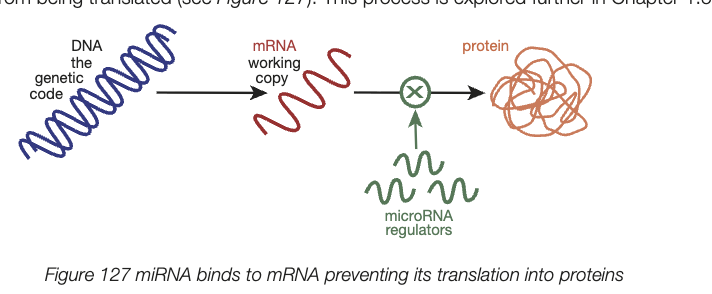
microRNA (miRNA)
non-coding RNA (ncRNA) molecules, approx. 22 nucleotides
regulate gene expression after transcription
exert regulatory action by binding to a small section of a specific mRNA molecule - prevents it from being translated
small section of mRNA becomes double stranded - prevents ribosome from accessing full mRNA code and expression of that gene
amino acids (aa) +bond in polpeptide chain
20 different ones
building blocks of polypeptides and protein molecules
polypeptides have a specific sequence of amino acids linked together by aSTRONG PEPTIDE BONDS
proteins can be one chain folded into a precise 3D shape or more than one chain linked together
coding strand
aka non-template strand, sense strand
codes for gene of interest
determines correct mRNA sequence but doesnt take direct part in transcription
mRNA produced from transcription will have same code as coding strand (except T is replaced with U)
consists of triplets of DNA codons
template strand
aka anti-sense strand
complementary to coding strand
provides template genetic code for growing mRNA strand to form against
mRNA produced is complementary to template strand, except T is replaced with U
contains DNA anticodons
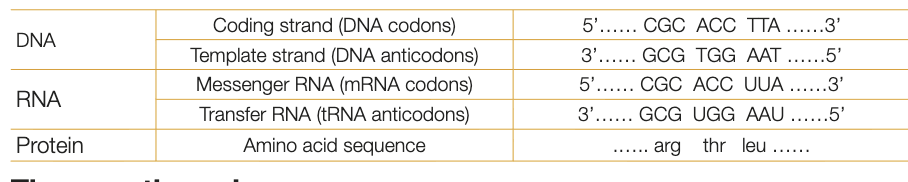
anticodons and codons in mRNA and tRNA and template and nontemplate strands
coding strand has DNA codons
template strand has DNA anticodins
mRNA molecules are synthesised against template strand in complementary pattern, therefore have mRNA codons (correspond to DNA codons in coding strand)
tRNA molecules with complementary anticodons to mRNA codons bring amino acids to polypeptide chain
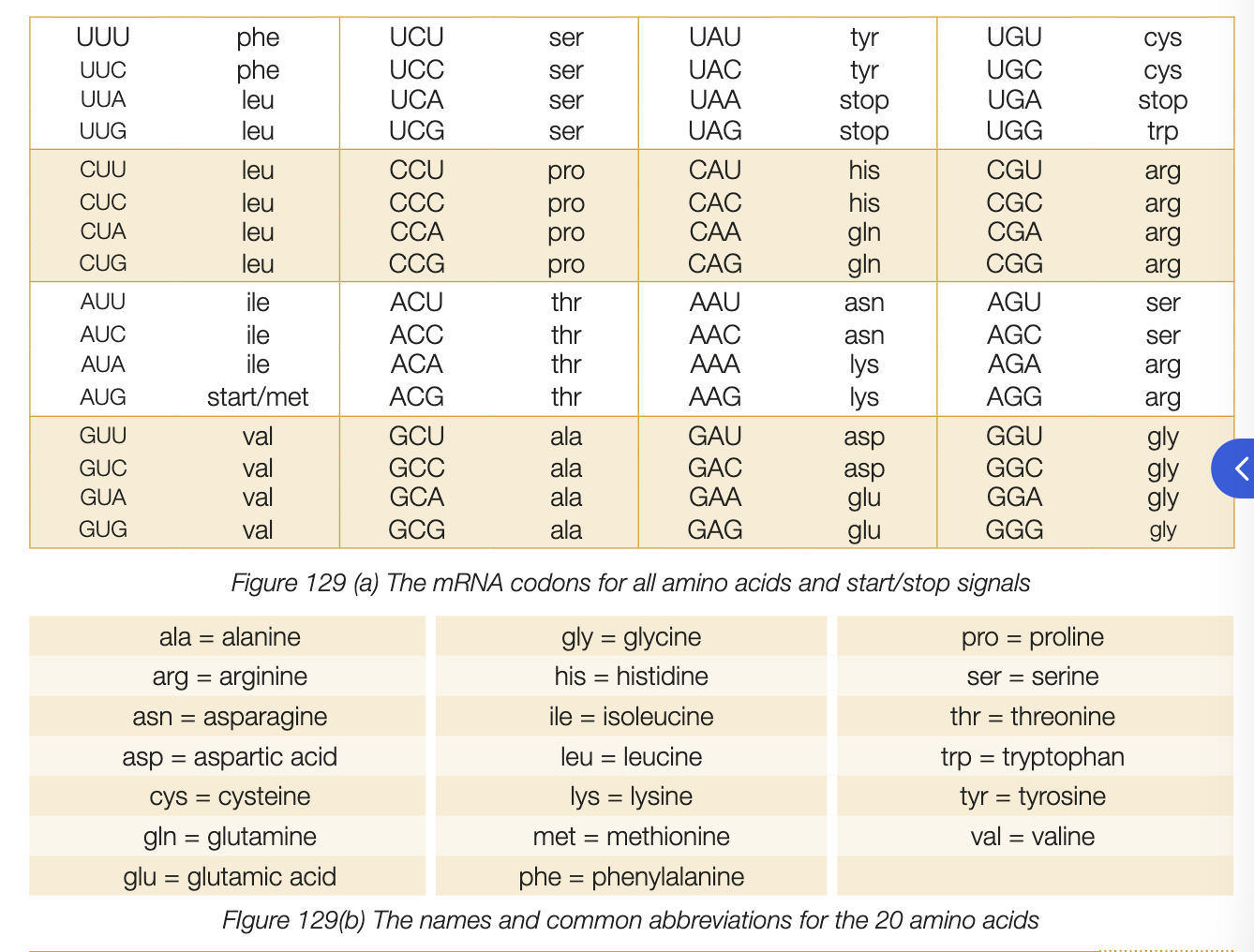
genetic code
codons are smallest unit necessary to code for all 20 amino acids
make up both coding DNA strand and mRNA strands
mRNA molecules transmit copy of gene from coding strand from nucleus to ribosome - sequence of codons is translated into amino acids
codons are base triplets thereofre, no of nucleotides making mRNA must be x3 no. of amino acids in protein
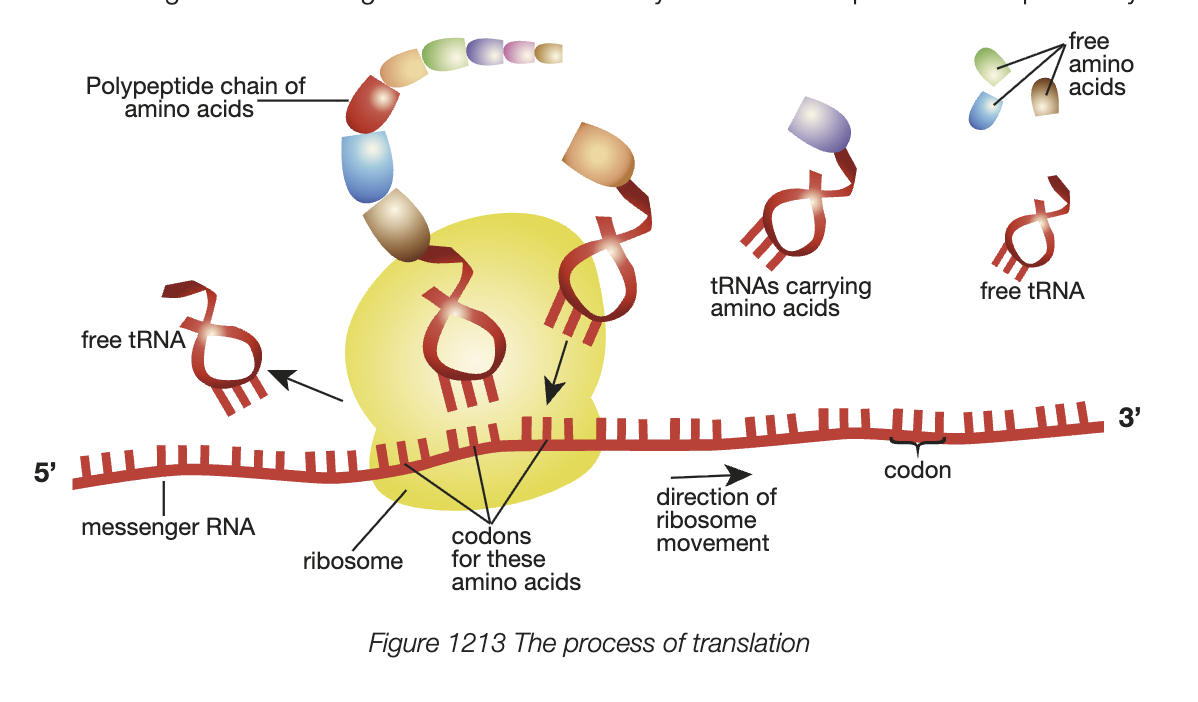
Transcription
occurs in nucleus of eukaryote
mRNA molecule is syntehsised from gene on DNA
1. initiation - RNA polymerase, once in contact with an activator protein, binds to the promoter region of DNA strand and begins to unwind it
2. elongation - RNA polymerase moved 5’ to 3’ direction (3’ to 5’ on the template strand), makes RNA transcript by adding nucleotides to growing RNA strand
3. termination - once RNA polymerase reaches termination site, transcript (mRNA) seperates from template, 2 DNA strands reform double helix
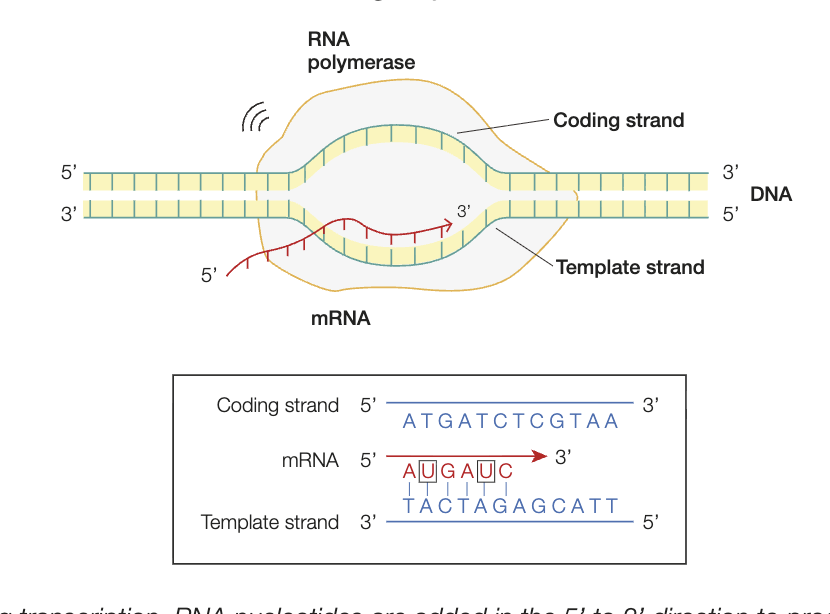
RNA splicing (part of transcription)
intial product of transcription is primary transcript - has both introns and exons
only exons are expressed as polypeptide, therefore introns in the pre-mRNA molecule (after transcription) must be removed
occurs in the nucleus, produces mature mRNA molecule
intron/exon junction recognised by snRNP proteins, cluster together to form splieceosome
splieceosome cleaves introns, joins 2 ends of exon togehter - mRNA now ready to leave for cytosol (mature mRNA)
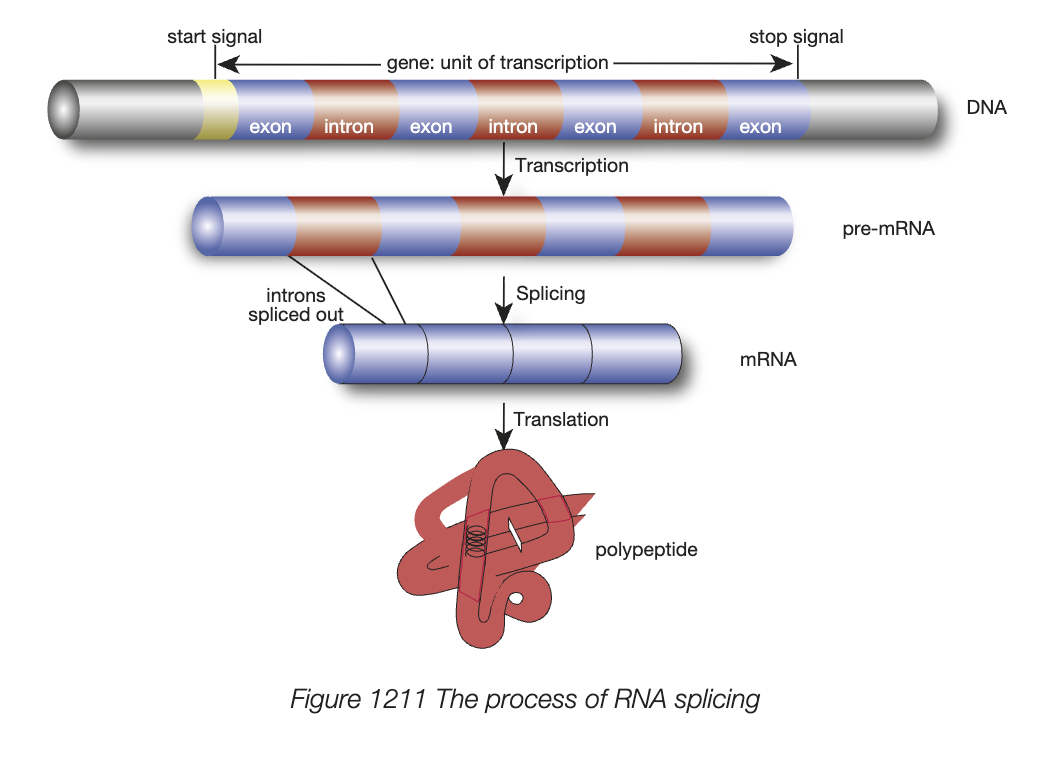
translation general info
polpeptide chain is built using sequence of codons in mature mRNa molecule
3 stages - initiation (use of start codon AUG), chain elongation (building amino acid sequence from codons), termination (completion of mRNA sequence with stop codon
happens at ribosome which consists of two tRNA binding sites
advantages of many ribosomes
can move along mRNA one after the other - leads to syntehsis fo many copes of polypeptide in short time period
large number of proteins are needed for multicellular organisms
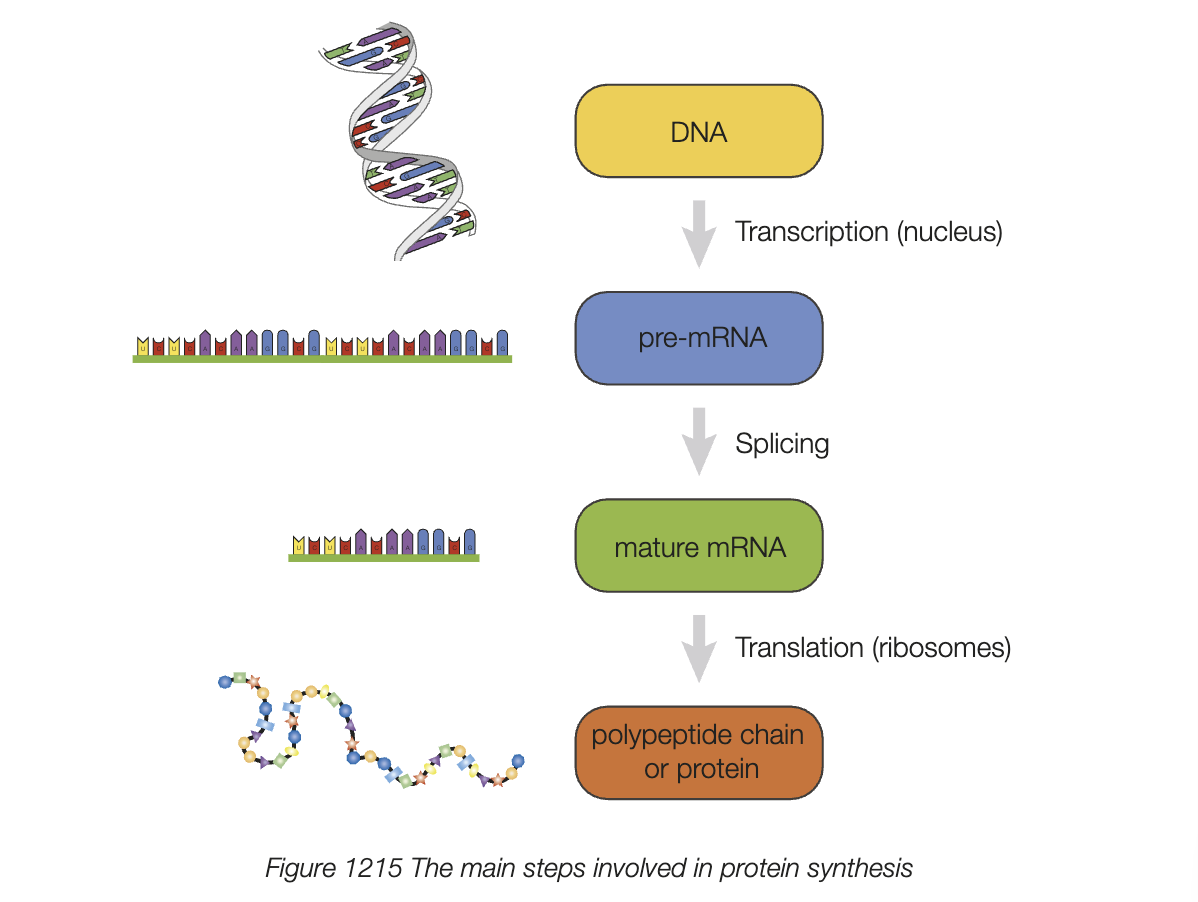
translation process
ribosomes read mature mRNA strand in 5’ to 3’ direction
two tRNA molecules move to ribosome sites, bond with mRNA (detmerined by complementary codon/anticodon)
ribsome faicllitates formation of peptide bond between 2 amino acids
ribsomes moves along mRNA - growing polypeptide chain
tRNA molecules keep bringing amino acids until stop codon is reached
mature mRNA continues to be translated, many copies of polypeptide chain are formed until miRNA message is sent
miRNA binds to mature mRNA (complementary fasion) - deactivates expression of this gene
promotor region, coding sequence, and terminator sequence
the 3 parts of a gene
promotor region - non-coding sequence, initiates transcription, typically located immediately upstream coding sequence. binding of RNA polymerase to it is controlled by transcription factors (in eukaryotes)
coding sequence - region of DNA that is transcribed
terminator sequence - part that tells RNA polymerase to stop transcribing
functions of proteins
catalysing reactions (e.g. enzymes)
cellular communication (e.g. hormones)
inactivating or neutralising invading molecules, substances, or organisms (antigens) (e.g. antibodies)
providing structure
transport of substances (e.g. oxygen via haemoglobin)
contraction and movement (e.g. muscles)
regulation of genes (e.g. activator protein, repressor protein)
proteins (and functional protein def)
made of amino acids which make polypeptide chains - can be many chains (many genes to code for them)
each protein is unique as made of varying combos of aa and can vary in type number or sequence
unique shape determines specific function
consist of one or more polypeptide chain, folded to produce specific shape
functional proteins - recognise and bind to other molecules that are complementary in structure (4 different types - primary, secondary, tertiary, quaternary)
primary structure
unique, linear sequence of amino acids in polpeptide chain held togehter by covalent peptide bonds - result of interactions between amino acids in primary structure
aka polypeptide
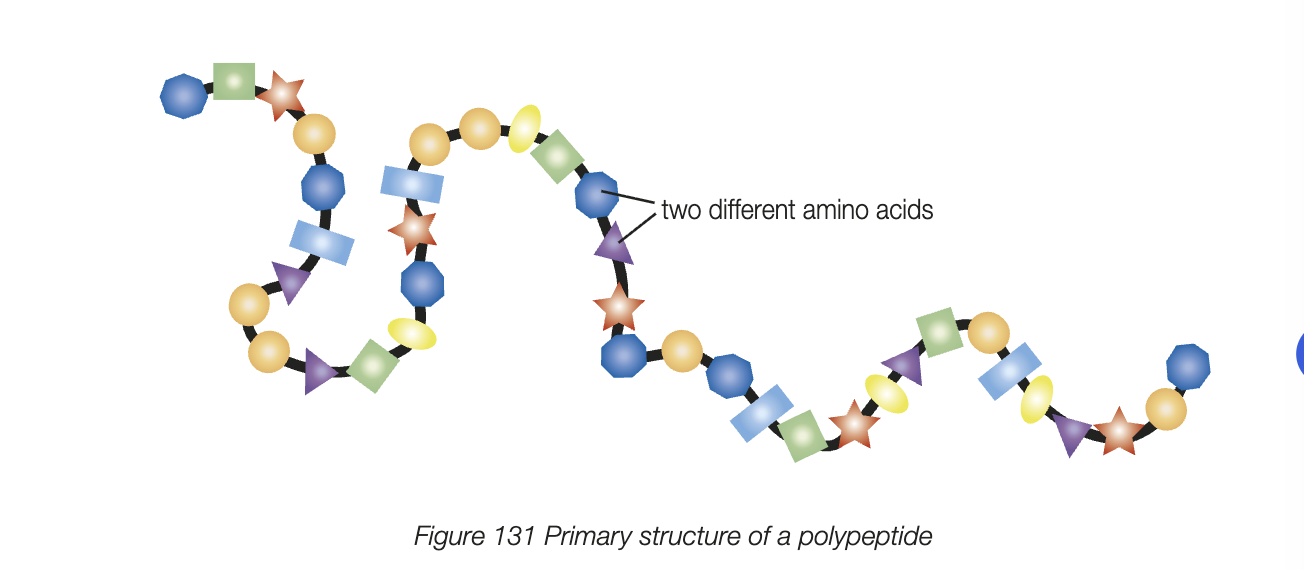
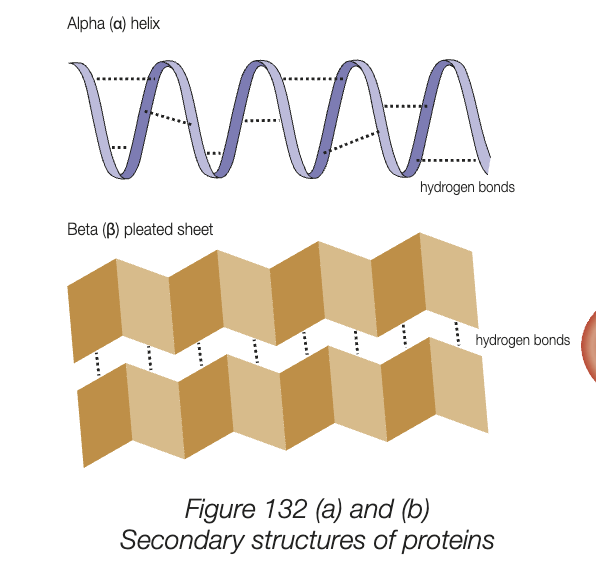
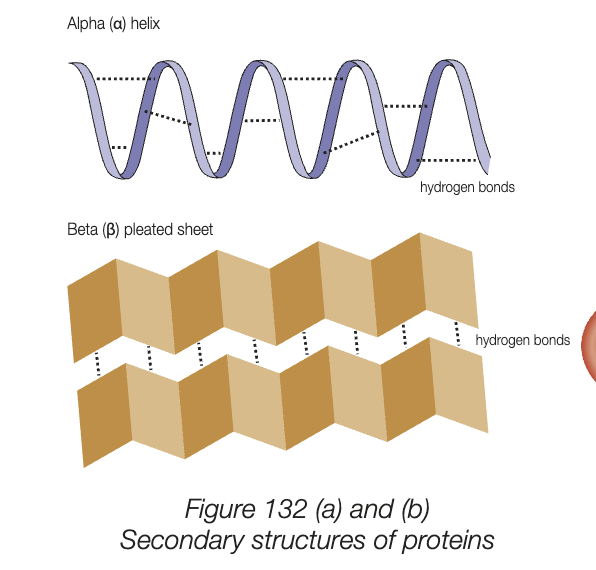
secondary structure
localised coiling or folding of polypeptide chain, caused by hydrogen bonding between amino acids
intermediated between primary and folding into 3D shape of tertiary
2 most common shapes - alpha helix (coiling) and beta pleated sheet (folding)
many antibodies have strucutre made of mostly beta pleated sheets, myoglobin is protein with mostly alpha helixes
tertiary strucutre
3D shape of a polypeptide caused by the way the polypeptide chain is folded
forms spontaneously because of forces of attraction between aa in the polypeptide chain
many types inc. hydrogen bonds, ionic bonds, strong covalent disulphide bonds
folded shape - globular proteins
3D shape with unique folds, grooves, and clefts determine function of protein, therefore 3D structure is critical to function
quaternary structure
protein consisting of 2 or more polypeptide chain, quaternary strucutre forms from polypeptide chains chemically bonding together
e.g. haemoglobin - 2 alpha and 2 beta chains bonded together
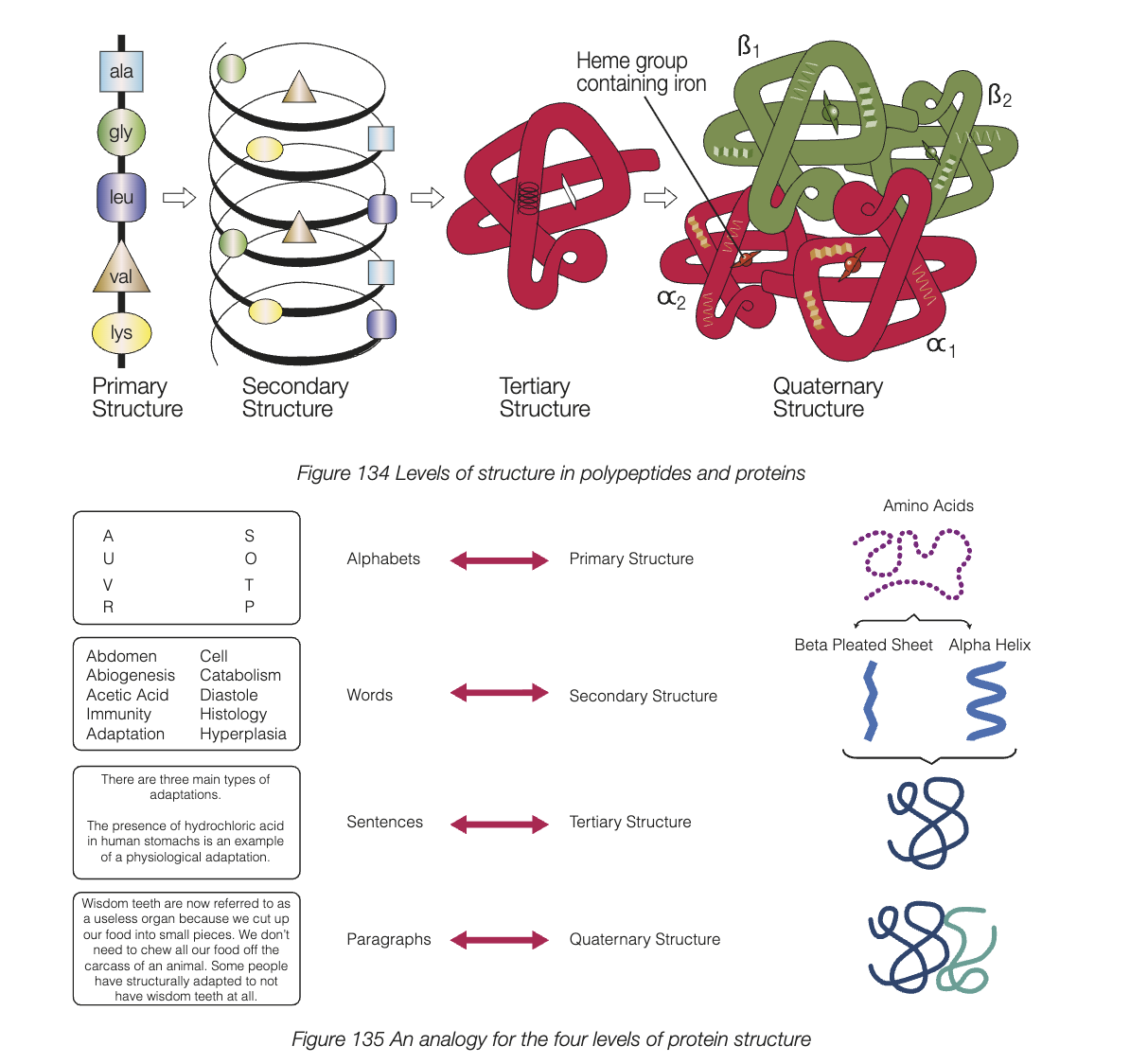
enzymes
most are globular proteins, increase the rate of a specific chemical reaction or catalyse it
have an active site - formed by folding and binding of protein chain - that substrate (s) or reactant (s) bind to
bc of the shape of the active site, enzymes have specificity - each enzyme only has one substrate
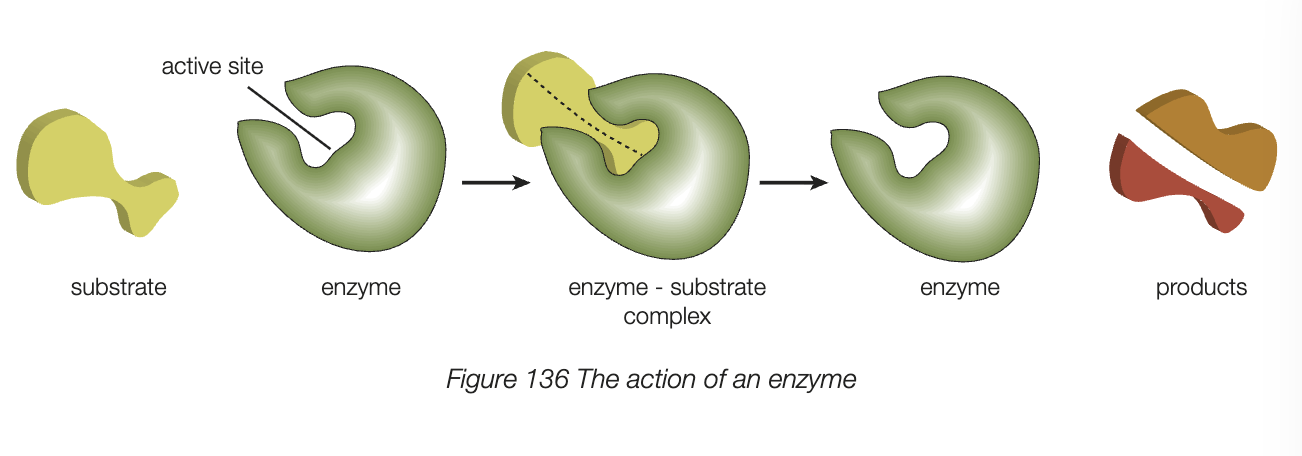
receptor proteins and peptide/protein hormones
proteins are found on inner and outer surface of cell membranes, play major role in membrane structure and function
receptor proteins have specific shape that bind toone specific messenger molecule with complementary shape (e.g. complementary hormone)
hormone - chemical messenger molecules
not all are proteins, some are lipid based
differences in primary, secondary, tertiary, and quaternary structures allow them to bind to specific cell membrane receptors
e.g. thyroxine - metabolic activity, insulin
hormone signalling includes processes of:
synthesis of hormone
storage of hormone in gland, and secretion into bloodstream
transport via blood to target cells
binding to complementary receptor protein in cell membrane of target cell
relaying message to cell nucleus, leads cellular response
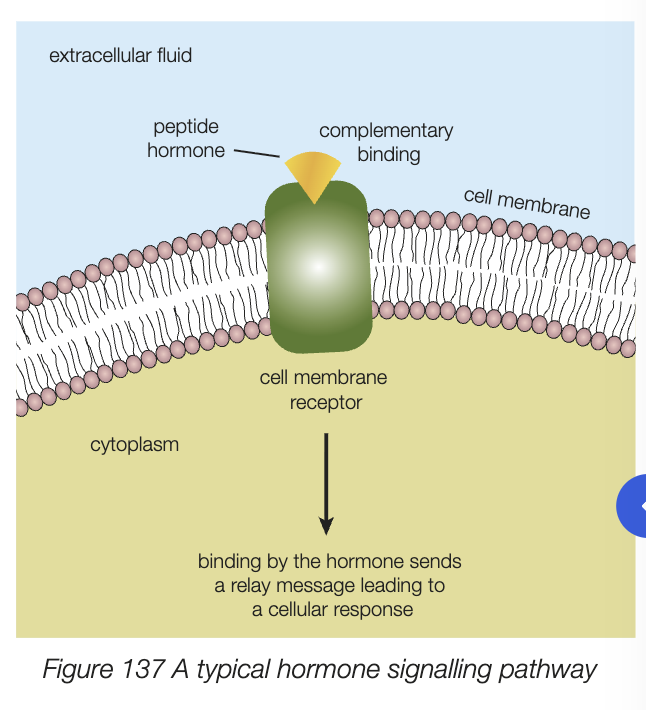
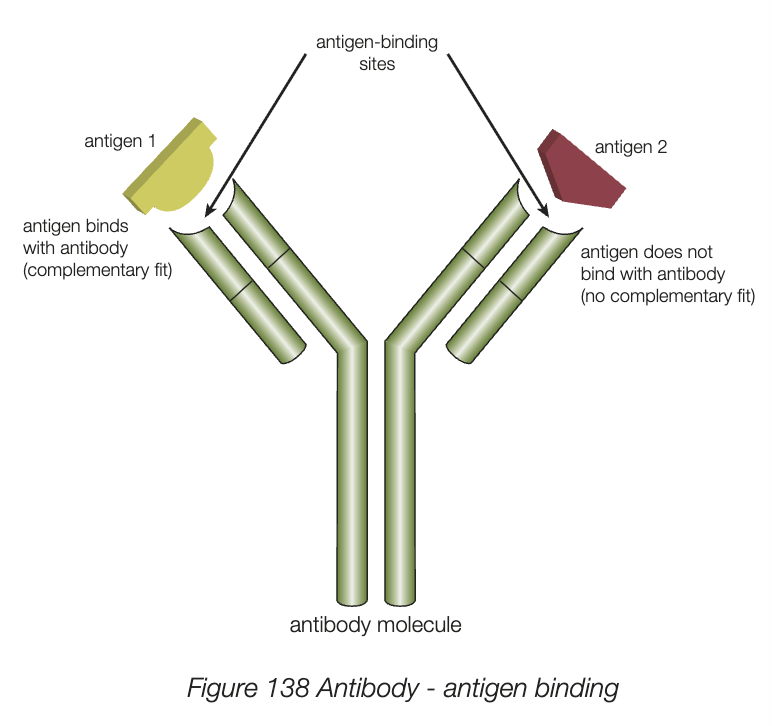
antibodies
antigens are foreign molecules (non-self) usually found embedded in surface of viruses or cells
lymphocytes produce protein molecule antibodies as part of specific defence mechanism
act by binding to specific antigen in complementary manner, deactivates or neutralises
particular 3D shape of quarternary structure allows antibody to bind with and inactivate antigen
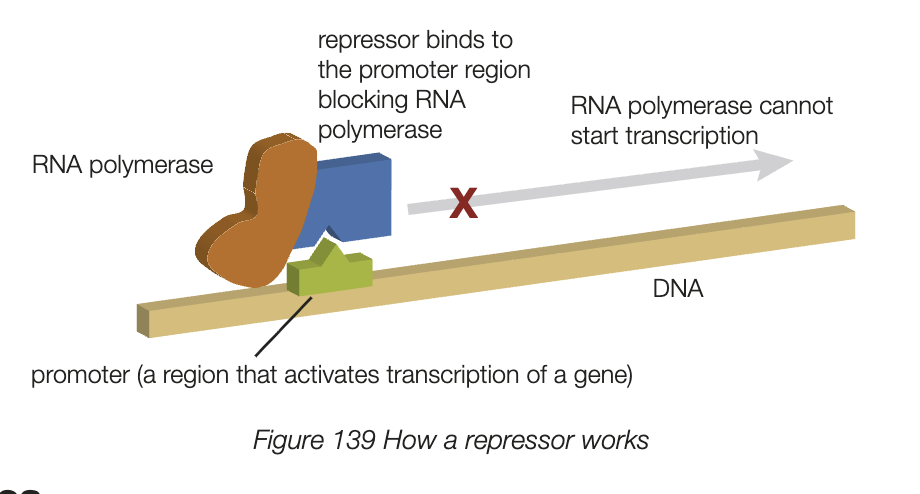
regulatory proteins
important area of contemporary research
cells have many ways toregulate/control swithcing genes on and off
genes are switched on when transcription can happen and opposite for off
repressor proteins bind to DNA section near gene (promoter region), preventing RNA polymerase from bindign and stopping transcription
activator proteins to DNA near gene assists RNA polymerase binding and promotes transcription
proteomics (and bit abt proteome, cells and diff proteins)
proteome - all proteins in cell or organism at specifc time
cell needs diff proteins at diff times of day and through life, no of proetins differ from cell to cell
each cell uses or expresses specific section of genome to make proteins it needs
proteomics - study of proteins (abundance, variations, modificatinos)
biomarkers - molecules (e.g. proteins found in blood) indicating abnormal processes of diseases
proteins are molecules of structure and funcion in cell
peptide/protein hormones
hormone - chemical messenger molecules
not all are proteins, some are lipid based
differences in primary, secondary, tertiary, and quaternary structures allow them to bind to specific cell membrane receptors
e.g. thyroxine - metabolic activity, insulin
enzyme
globular protein speeds up/catalyses specific chemical reaction within cells
reactants/substrates are chemically converted into new substances (products)
breaks chemical bonds in substrate and forms new bonds in product
are not used up in reactions
active site and enzyme specificity
region substrate binds to - groove or cleft on enzyme surface
binding is called enzyme-substrate binding; occurs due to active sites shape
shape is specific to one substrate
substate and active site shapes are complementary
enables specificity, ensuring that only the correct substrate can bind
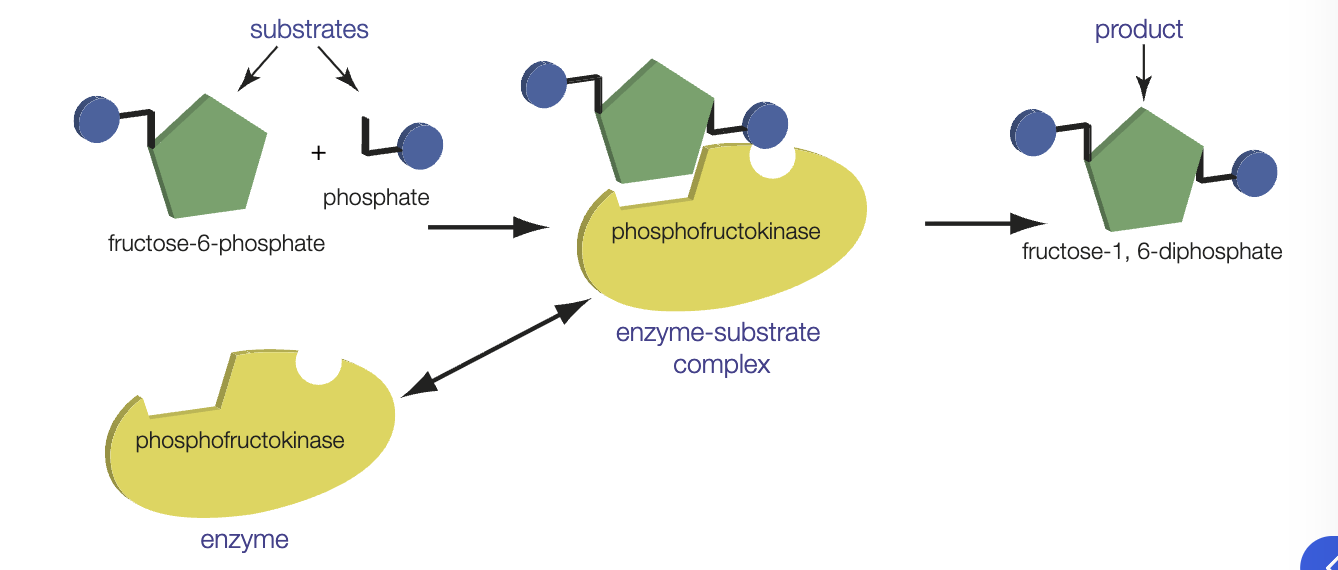
how do enzymes work
substrate molecules are bough into an orientation that facilitates breaking/forming of chemical bonds
bonds in substrate are stressed, lowering the amount of energy needed to break them and therefore the amount of energy needed for the reaction to proceed
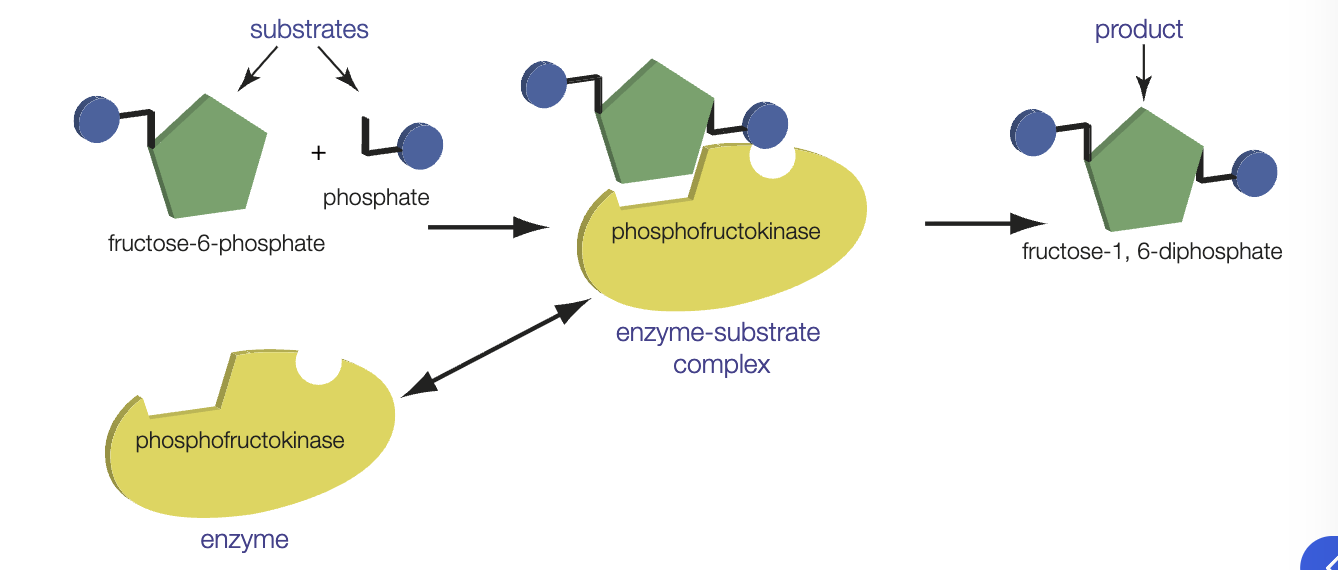
metabolism
metabolism - all biochemical reactions carried out by living organisms
rate of these reactions - metabolic rate
Separate biochemical reactions making up a cell’s metabolism are metabolic reaction
most reactions form part of metabolic pathways
each step is catalysed by a specific enzyme
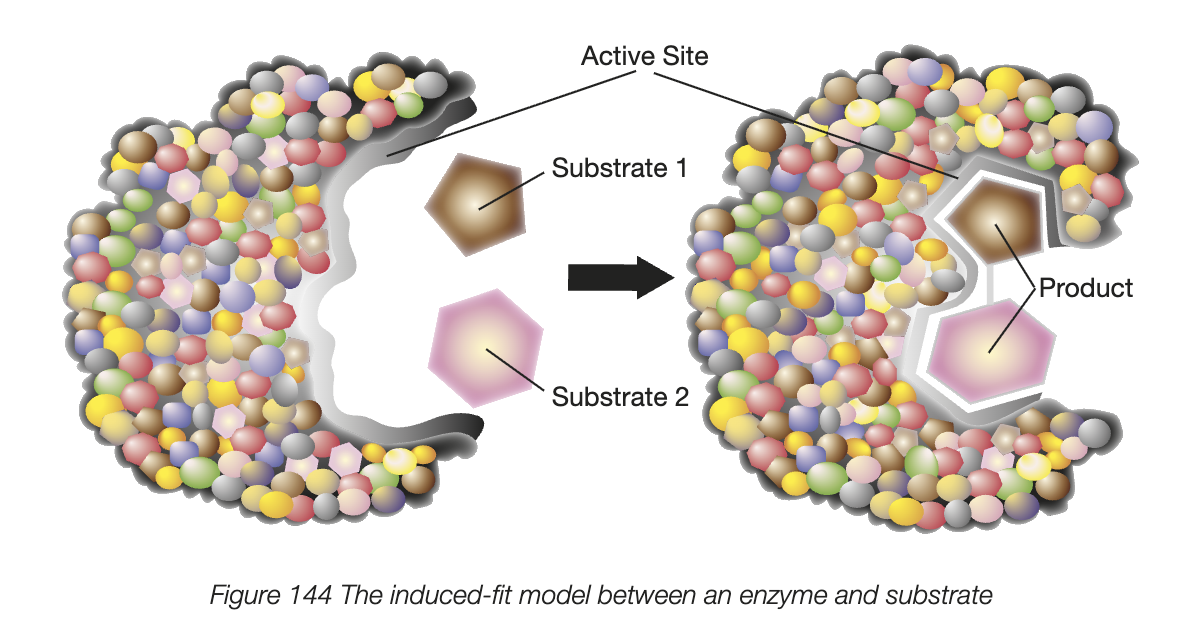
induced fit model
substrate binds to the enzyme by weak bonds, often causes a change of shape of both enzyme and the substrate
original model - lock and key - enzyme active site, which is like a keyhole to substrate (hence specificity)
induced-fit model - enzyme and substrate modify each other’s shape in the process of binding
active site continues to change until substrate is completely bound to it. this is when the final shape and charge are determined
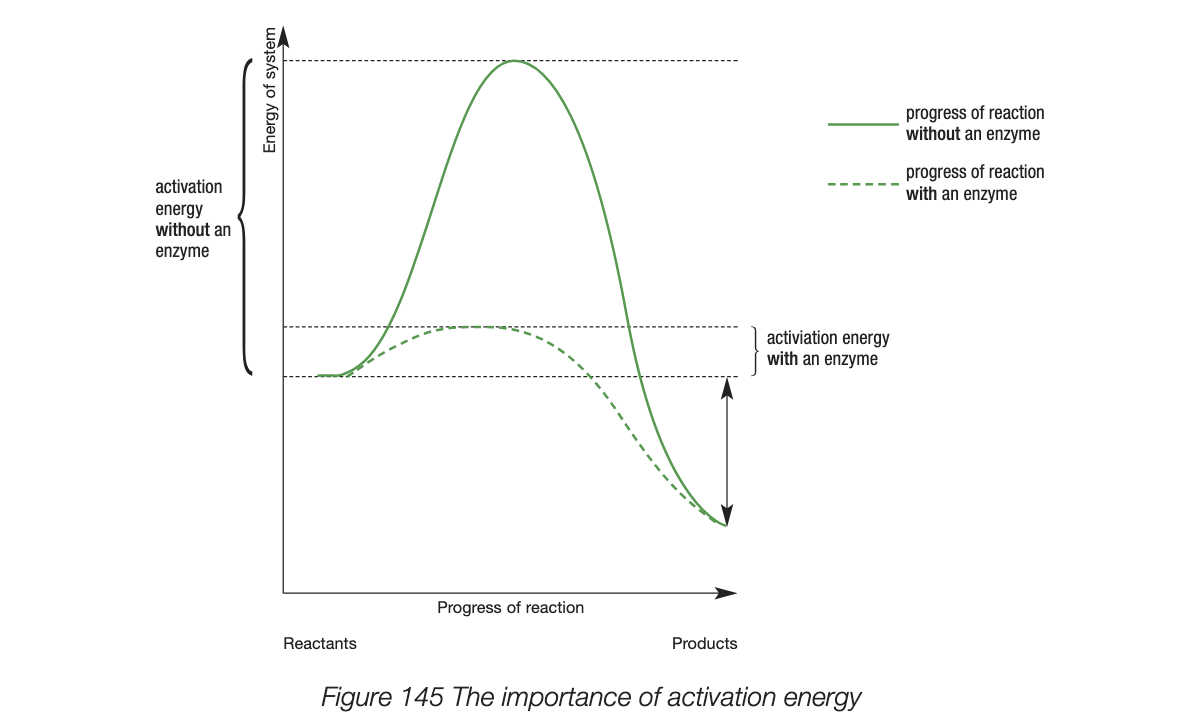
enzymes lowering activation energy
activation energy - energy required for the bonds of reactant molecules to break
the lower the activation energy the faster the reaction - less energy is needed to break bonds to initiate the reaction
factors influencing enzyme activity - temperature
at lower temperatures, activity is low - fewer substrate colliding with enzymes ac’s active site, don’t have enough energy to overcome the activation energy barrier
as temperature increases, so does enzyme activity - substrate and enzyme molecules will move faster, resulting in more collisions at the active site and hence a greater rate of reaction
enzyme activity is maxed out at optimum temp
temp inc above optimum temperature causes the active site structure to be altered (denaturation). substrate can’t bind to the denatured active site, thus activity is 0
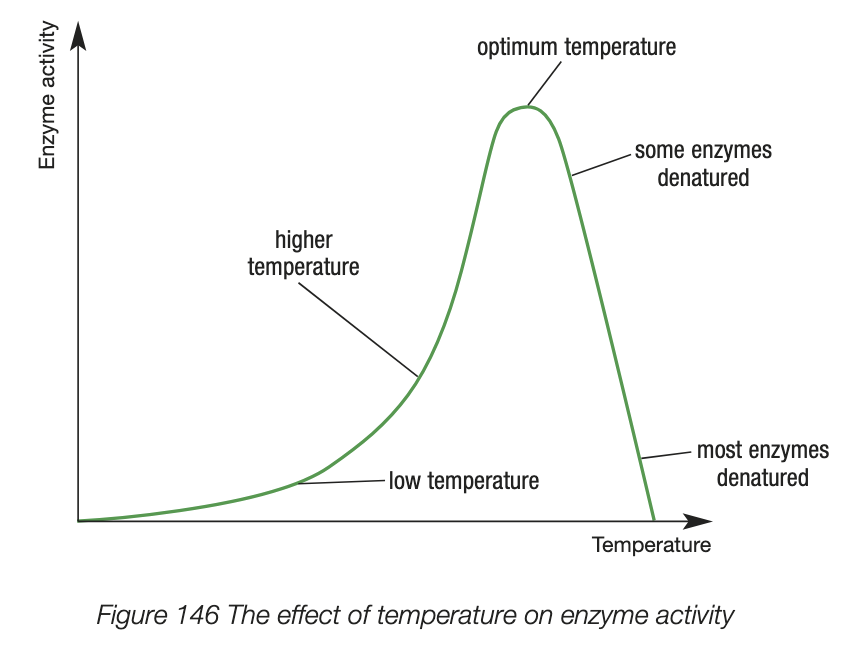
factors influencing enzyme activity - pH
optimum pH - activity is at maximum
pH change above or below optimum temp denatures the enzyme - substrates can’t bind to the denatured active site, enzyme activity decreases to 0 quickly
enzyme-substrate binding is reduced when a change in pH alters the shape of the enzyme or substrate - no longer complementary
optimum pH is different for enzymes
Chemical buffers in the body help stabilise pH and allow enzymes to catalyse reactions
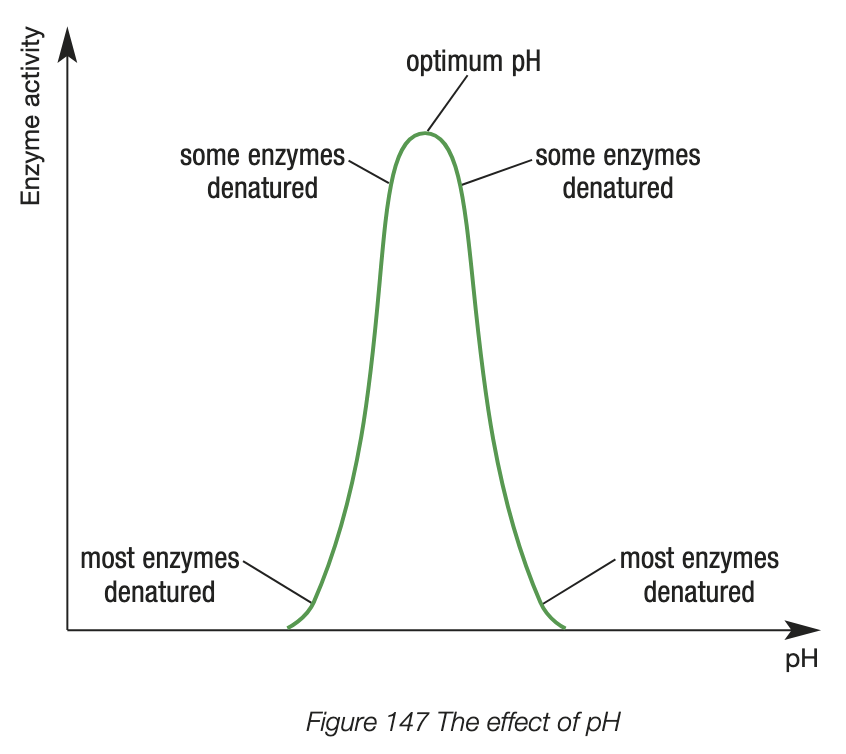
factors influencing enzyme activity - concentration of reactants
low substrate concentration - fewer substrates to collide with the enzyme active site, enzyme activity is low
as concentration of substrate increases, enzyme activity increases - more frequent collisions between enzyme and substrate at active site
higher substrate concentrations - most active sites on the enzyme are occupied by substrate, increases in substrate concentration begin to cause smaller and smaller increases in enzyme activity
increase in substrate concentration above optimum substrate concentration don’t lead to greater enzyme activity - all active sites are occupied by substrate (saturated)
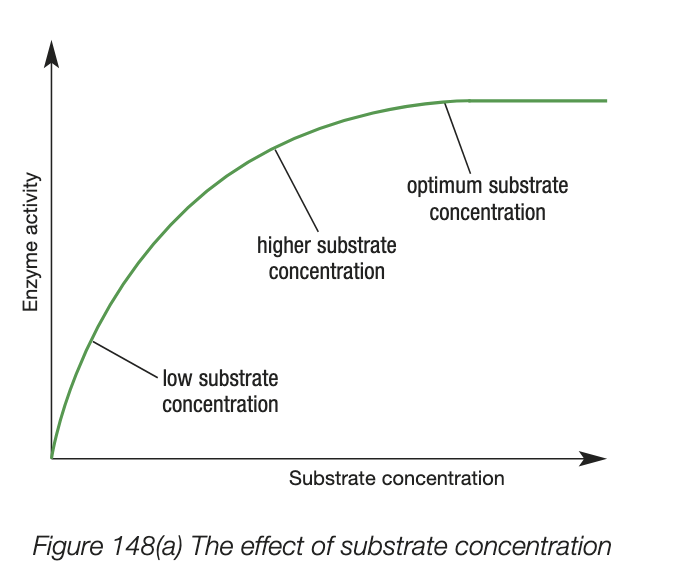
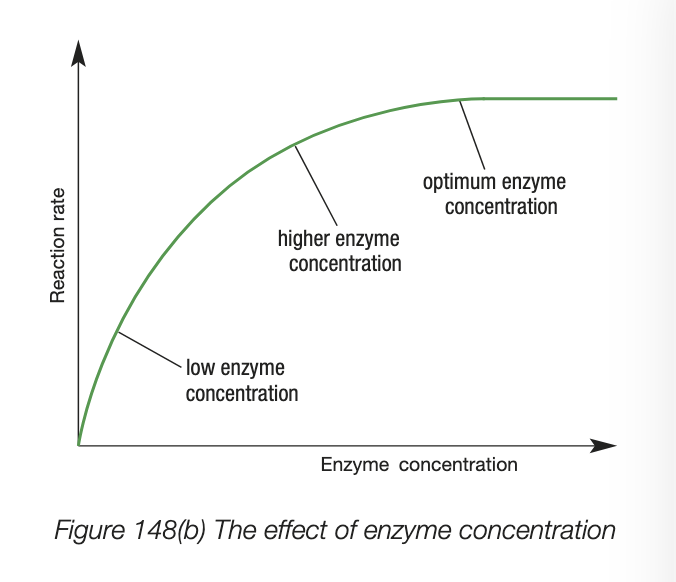
factors influencing enzyme activity - concentration of enzyme
same logic as substrate
difference is that substrate is the limiting factor
factors influencing enzyme activity - concentration of products
The activity of some enzymes is regulated by substances that bind to the enzyme NOT AT THE ACTIVE SITE but in the ALLOSTERIC SITE
causes a reversible change to the active site and alter binding
end products of metabolic pathways can act as inhibitors of pathway action associated with the control of metabolic pathways in cells
known as end product inhibition - occurs when the presence of a high concentration of the end-product of a metabolic pathway inhibits the production of products that stop the production of the end-product
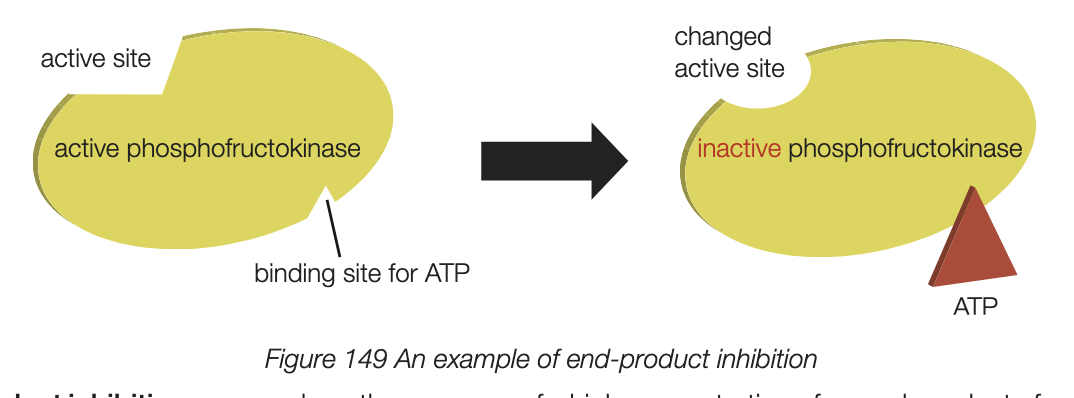
factors influencing enzyme activity - inhibitors (competitve and noncompetitive)
molecules that reduce rate of enzyme-catalysed reactions that can lead to the accumulation of substrates
COMPETITIVE INHIBITORS: mimic the structure of the substrate - compete with the substrate by binding to the active site of the enzyme. enzyme-substrate binding is reduced and enzyme activity is decreased
NON-COMPETITIVE INHIBITORS - do not compete with substrate, bind elsewhere on enzyme at allosteric site. structure/shape of active site changes, preventing binding of enzyme and substrate at active site and reducing enzyme activity
phenotypic expression of genes
gene pairs of homologous chromosomes control phenotypic characteristics (physical factors expressed from genes)
alleles - genes of corresponding pairs, alternative forms of same gene
different alleles exist because genes are subject to mutations
2 organisms can look similar but have different genotypes
2 organisms can have same genotype but appear different because they’ve been exposed to different environmental conditions
epigentics
study of how the environment influences gene expression
epigenome - chemical tags, an additional level of coding on top of the DNA base sequence. do not affect the base sequence, but change how cells read and express DNA
2 main types - methylation and acetylation
epigenetic tags can be inherited and altered by environmental factors
affected by positive and negative experiences, can be temp or permanent.
different experiences of individuals as they develop alter the location of epigenetic tags
promoter sequence
region on DNA that is not transcribed but plays a role in controlling transcription
RNA polymerase binds to initiate transcription
transcription factors can also bind to specific base sequences in promoter regions and assist in the binding of RNA polymerase
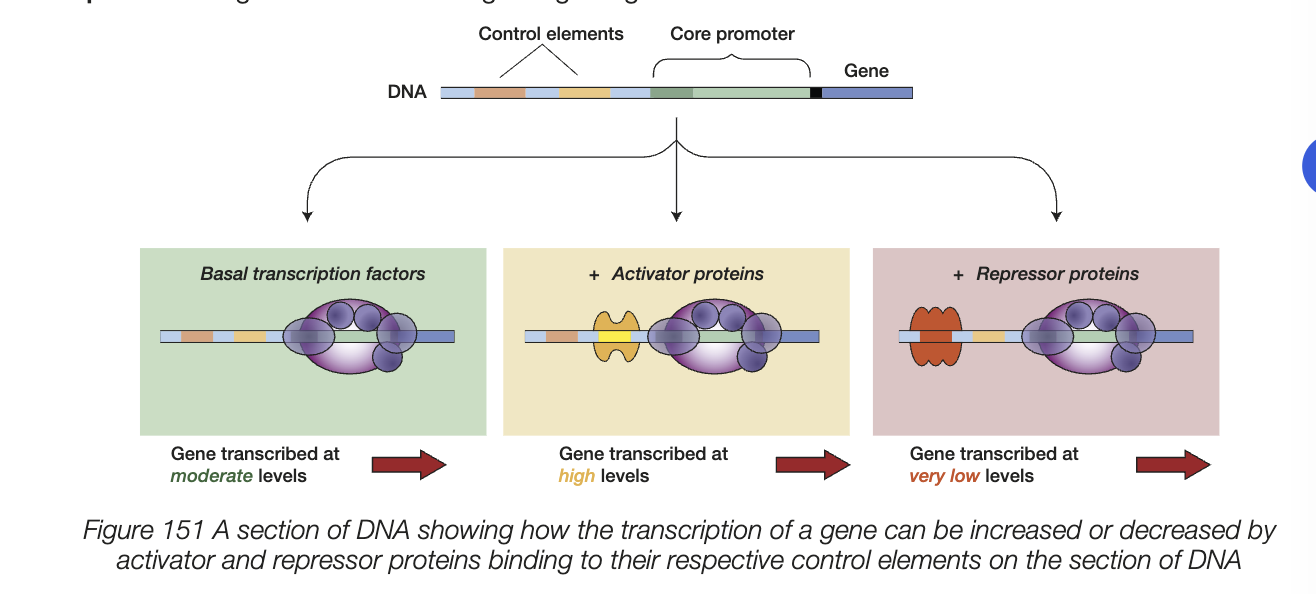
activators
bind to DNA and activate/increase rate of transcription
can bind to other base sequences upstream of the gene and promoter sequence
activator proteins binding to base sequence can stimulate gene expression
e.g. of binding it can initiate - triggering RNA polymerase to release from promoter region and start moving along gene
repressors
binding to DNA and slow/stop rate of transcription
can bind to other base sequences upstream of the gene and promoter sequence
repressor binding to other region silences transcription by inhibiting RNA polymerase from binding to promoter region
methylation
process where a methyl group is added to the DNA strand
usually occurs at a cytosine base
does not alter DNA sequence but can influence gene expression
inhibits RNA polymerase from binding to promoter regions, stopping transcription
after methylation, genes usually retain methyl groups when cell division occurs and the pattern of methylation is passed onto daughter cells
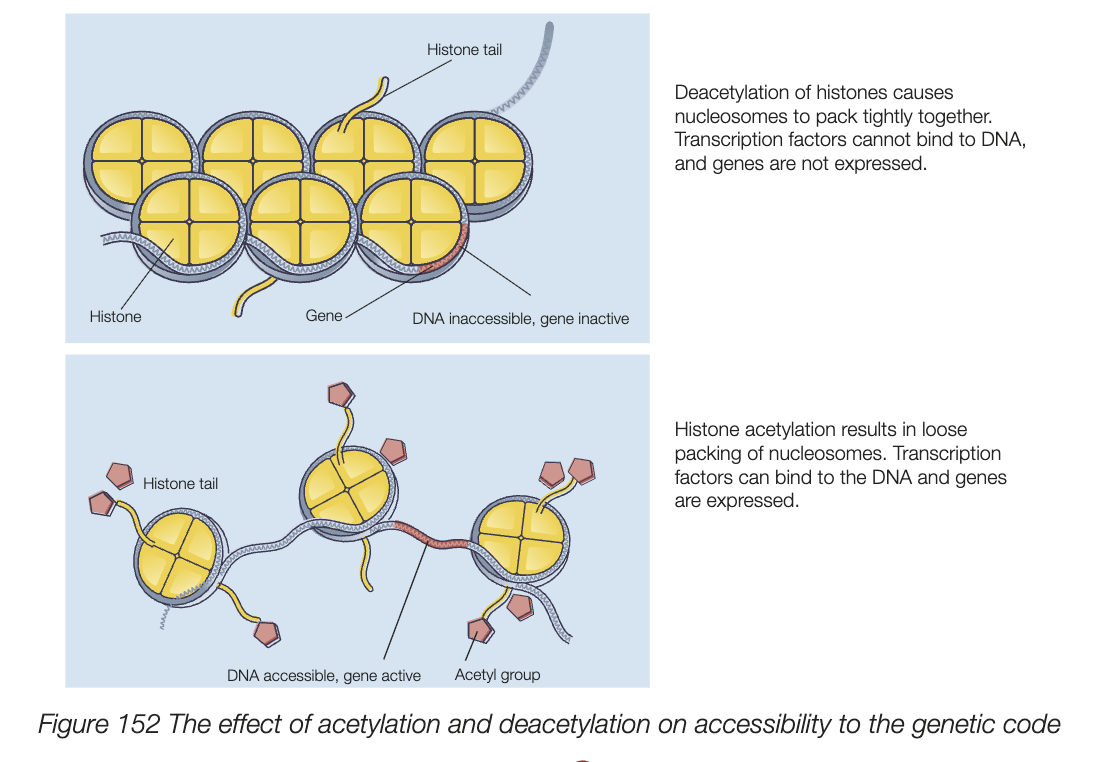
histone modification/acetylation
DNA is packed with histone proteins to form chromatin
acetylation of histone proteins changes RNA polymerase accessibility to genes
acetylation: chromatin to decondense - loose packing of histone proteins - RNA polymerase can bind to DNA and express genes
deacetylation: chromatin condenses, histone proteins are more tightly packed, RNA polymerase can’t bind, and transcription won’t happen
histone proteins can also be methylated - consequences depend on the amino acids within the histone protein being methylated and the number of methyl groups added to that amino acid
translation factors
translation is controlled as valuable resources like energy and amino acids are wasted, and increases in certain proteins can be detrimental for cells
translation factors help coordinate the synthesis of polypeptides and proteins
generally act on mRNA or miRNA
can increase or decrease translation on mature mRNA
miRNA, siRNA, lncRNA are focus in s2
all factors focused in s2 bio are synthesised from non-coding DNA and are caleld ncRNA (non-coding RNA)
small interfering RNA (siRNA)
aka silencing RNA
class of translation factors, act on mRNA, preventing translation
similar in size to miRNA, generally consist of 21 nucleotides
binds to a specific mRNA molecule in a complementary fashion, promoting the destruction of the mRNA strand
A specific mRNA strand cannot be translated by ribosomes, and gene expression is stopped
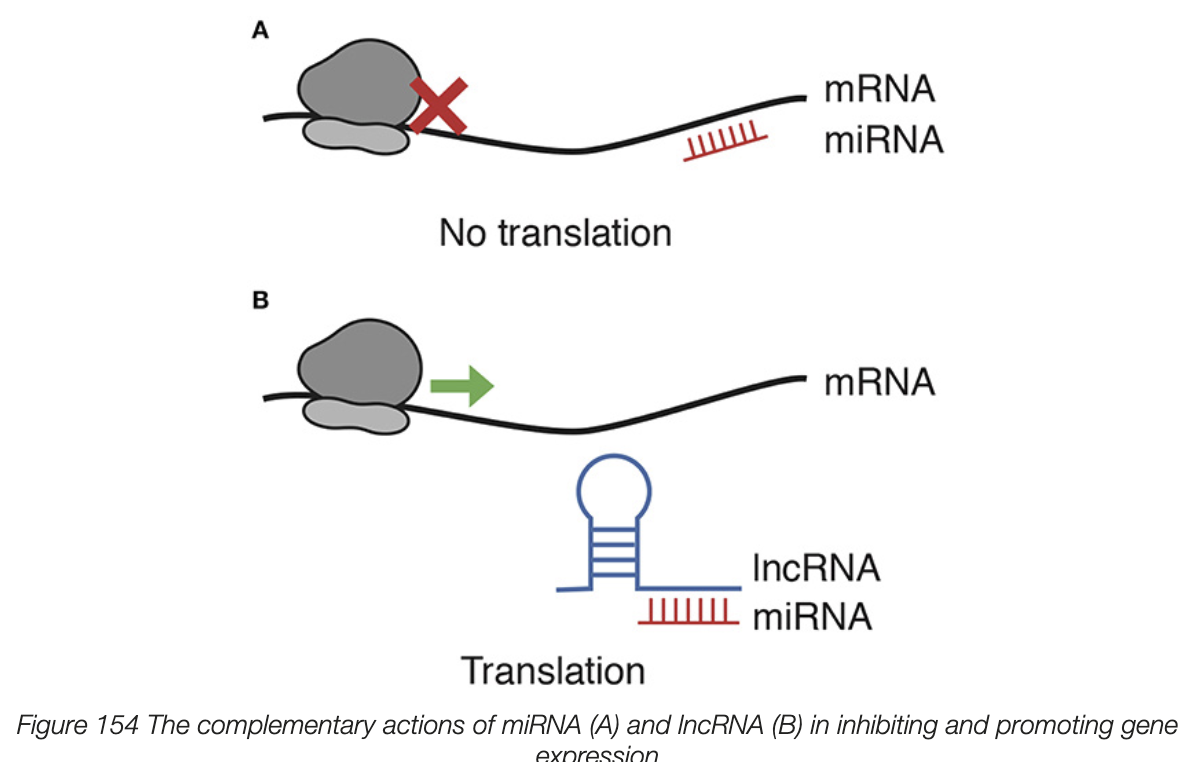
long non-coding RNA (lncRNA)
have many functions that control gene expression
longer than miRNA and siRNA, approx. 200 nucleotides
bind to specific miRNA molecule in a complementary fashion
as lncRNA binds to miRNA and not mRNA, miRNA cant bind to mRNA and prevent translation
lncRNA promotes gene expression by inhibiting specific miRNA
factors influencing gene expression ADDD
gender - some genes are expressed differently in males and females
chemicals
temperature
diet and lifestyle
trauma
CANCER - pro-oncogenes,
normally code for proteins involved in cell division
decreased methylation to DNA can therefore promote cell division and if uncontrolled leads to cancer
CANCER - tumour supression genes
under normal circumstances, they code for proteins that suppress or inhibit cell division - keeps no. of new cells in a tissue in check and suppress tumour formation
increased DNA methylation of tumour suppression gene switches them off and proteins suppressing tumour formation can no longer be produced/produced in smaller amounts
uncontrolled cell division and tumour could develop due to lack of normal cell divison suppression
CANCER - DNA repair genes
can be activated to produce proteins involved in the repair of damaged DNA
increased methylation to DNA repair genes results in them being turned off. less damaged DNA is repaired and it accumulates - increases risk of uncontrolled cell division and cancer
cell differentiation
all organisms that reproduce sexually start as a fertilised egg or zygote, a single cell divides and grows to form blastocyst, then gastrula, then different cells type develop
every somatic cell contains all genetic info needed to carry out every function but only small portion of this is activated in each cell
differentiation - process where cells with identical chromosomes and genes become different and specialised in both structure and function
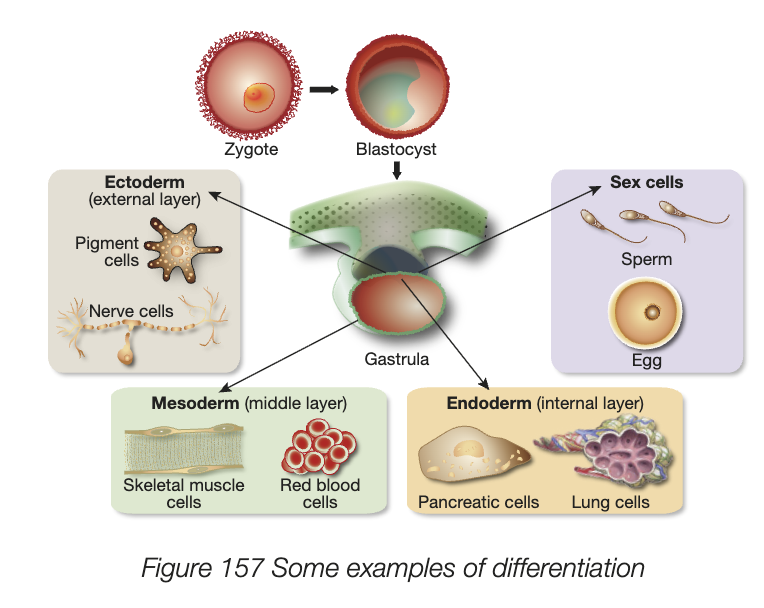
specialised cells
have specific epigenetic tags, help activate and silence specific genes
once specialised, cannot differentiate into another type of specialised cell
epigenetic tags can be passed from parent cells to daughter cells through mitosis - inheritance of these tags ensures daughter cells also become the same type of specialised cell and will be the correct type of cell for that specific organ/tissue
stem cells
unspecialised and can differentiate into specialised ones
2 types - pluripotent and multipotent
pluripotent - stem cells that have the potential to develop into any human cell
multipotent- stem cells that can develop into some cell types only
cell differentiation results in regulation of gene expression; some cells have particular genes switched on and off
housekeeping genes - certain genes that are always turned on
reasons why cells in different regions or positions develop differently
signal molecules released from one cell bind to surface cell receptors on target cells nearby and the signal molecules act to regualte genes in nucleus
interaction of membrane proteins on cells initiates signal molecules. signal molecules include transcription factors
cell signalling mechanisms in differentiation
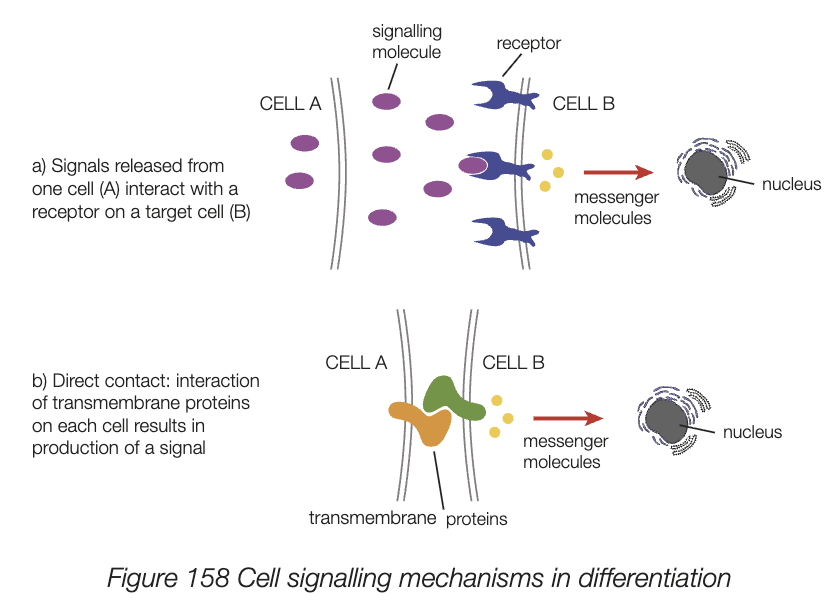
cloning
clone - group of cells or tissue derived from single cell or group of genetically identical organisms
two clones are genetically identical but are epigenetically different as they are exposed to different environmental conditions throughout life
therapeutic cloning
use of cloned stem cells in medicine
stem cells are unspecialised, so scientists can use different techniques to control gene expression and control the type of cell they differentiate into
research has great potential and is possible to produce a wide range of tissue and enable diseased cells to be replaced with healthy ones
highly regulated due to ethical considerations - creating new life just to extract stem cells, ending potential life
possible methods of acquiring stem cells for research:
extraction from leftover fertilised ovum from an IVF program (embryonic stem cells)
removal of tissue like bone marrow and umbilical cord (adult stem cells)
reprogramming ordinary somatic cells in a way that induces them to become stem cells (IPS cells)
once obtained, used to produce specific tissues in lab, transplanted to patients to replace or repair damaged ones in body or injected directly in specific region of body
limits - producing tissue that is structurally correct, rejection, development of wrong type of tissue, tumour formation
ction