2.2 - genetics + gene function
1/22
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
reverse genetics
select gene to investigate
mutate that gene using chem/molecular bio
observe phenotype w/ mutated gene
compare mutant + wild-type; gene of interest is likely responsible for phenotype changes
forward genetics
gather/create mutant individuals that display the phenotype of interest
screen the genomes of the mutant individuals + look for a gene mutation that all of the individuals share
if all have a mutation in the same gene, this gene is likely responsible for the phenotype
RNAi process
dsRNA processed by DICER into siRNAs
RISC complex recruits siRNAs
siRNA hybridizes against target gene’s mRNA
mRNA degraded
RNAi characteristics
fast + easy way to study gene function through targeted mutagenesis
problem = off target effects, only temporary gene knockdown
conditional mutants
created by scientists that express a mutation under specific conditions
popular method = insert mutation downstream of a promoter which is controlled by transcription factors relevant to the condition
permit study of lethal mutations
polymorphisms
sequence variations at particular regions of the genome
most are SNPs, others include CNVs
tend to be inherited on haplotype blocks
polymorphisms and family history
distant relatives = diff haplotype blocks from meiotic recombination
close relatives = share numerous haplotype blocks
mendelian disorders
caused by mutations in a single gene + therefore the mutation segregates w/in families in a mendelian fashsion
dominant disorders
mutations in 1 allele is sufficient to cause the disease
recessive disorders
mutation in both alleles is required to cause disease
more prevelent in certain parts of the world bc of incest
multigenic disease
usually arise from cumulative effects of mutations in multiple genes that normally have a small effect size, but together create a large enough effect
tend to arise later in life
risk-enhancing alleles are inherited rather than eliminated; can become common
how are risk alleles identified
by looking for Single Nucleotide Polymorphisms that are statistically linked + more common to the development of the disease
genome-wide association studies (GWAS)
identify DNA variations that are significantly more frequent in ppl w/ age-related macular degeneration
alleles tend to be found in non-coding regions
and/or typically only mildly affect expression
once you identify a putative gene mutation, how can you model the mutation’s effects to determine if it is indeed the disease-causing variant?
identify the impact of the mutation on global gene expression in cells
RNA sequencing is one method
RNA-sequencing
RNA → cDNA, which are then sequenced
provides quantitative analysis of cell’s transcriptome
can also detect rare splice variants
in situ hybridization
can tell when and where a particular gene is expressed by hybridizing a fluorescently labeled, singlestranded probe against complementary RNA sequences within cells
reporter genes
used to determine the pattern of a gene’s expression
coding sequence of a gene is replaced with a reporter gene (GFP)
GFP is then controlled by the gene’s endogenous regulatory sequences
ensures that GFP expression will match the normal expression patterns of the gene of interest
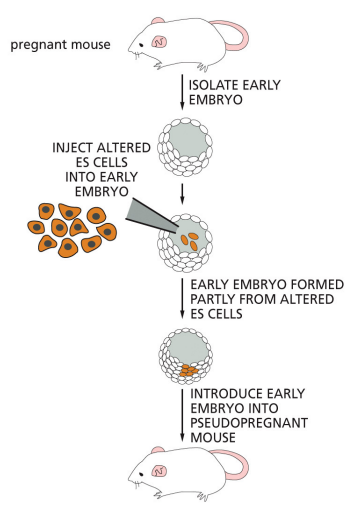
homologous recombination using ES cells
DNA creation
gene sequence altered in cultured ES cells
DNA plamids introduced into ES cells w/ mutated DNA @ gene of interest, flanked by homologous DNA seq corresponding to ES cell
plasmid also contains selection marker (antibiotic resistance gene)
ES cells cultured in presence of antibiotic — only cells that take up plasmid + correctly swap out the DNA fragment will survive
pick + inject each surviving clone into blastocyst
blastocyst (black) inplanted into pseudopegnant mother (brown)
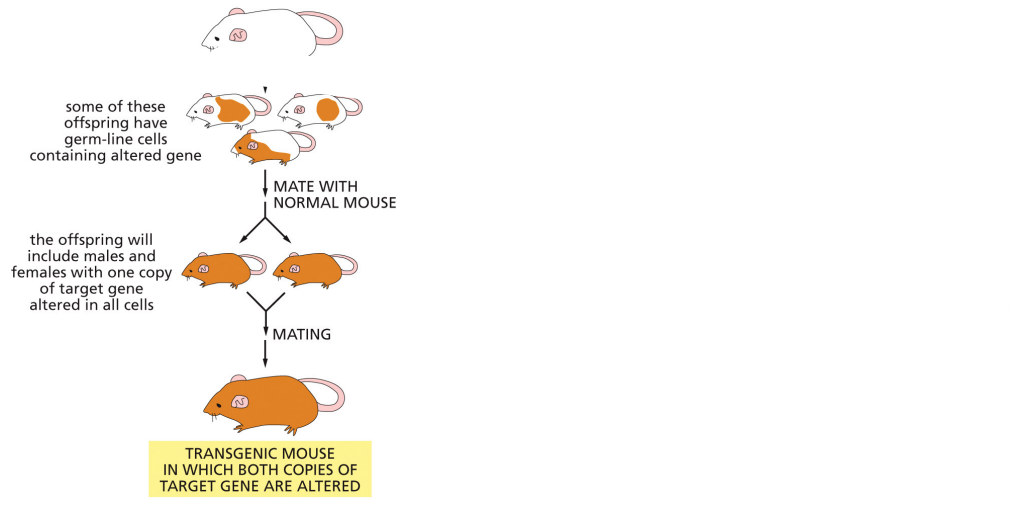
homologous recombination using ES cells
results
results are chimeras — fur is white + brown (most likely to have mutation in germ cells
chimeras mated w/ normal mouse to generate F1s — sequences for the gene mutation to check successful transfer
F1s can mate to product mutant mice that have 2 copies of the mutant allele
CRISPR
Cas9 can induce double-stranded breaks into DNA, but it needs a guide RNA to target specific sequences
Cas9 + guide RNA are expressed in a cell of interest
guide RNA targets Cas9 to the gene of interest so it can induce a double stranded break
donor DNA is also added, and this contains a altered piece of DNA
homologous recombination swaps the wild-type gene for the donor gene, and enzymes that repair double stranded breaks finish the process
this can be done in the blastocyst, eliminating the need for ES cell based homologous recombination!
how can Cas9 actively a dormant gene/ turn off an active gene
by using a mutant form that can no longer cleave DNA
conditional knockouts (CKO)
a gene can be selectively disabled in a particular target tissue
CKO in mice
mouse #1 = insert two pieces of DNA called LoxP sites into the flanking regions of the target gene
mouse #2 = insert a piece of DNA encoding for the enzyme Cre recombinase — inserted down-stream of a gene promoter that is known to direct gene expression to a particular type of cell or tissue
1 + 2 mate— offspring that inherit the gene sequence flanked by LoxP sites, in addition to Cre-recombinase, will have excision of that gene is all cells where the gene promoter directing the expression of Cre recombinase is active