Lecture 10: Phylogenetics I
1/38
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
39 Terms
Carl Linnaeus
1707-1778
developed lifeform classification system (used to understand how species are related to each other) which is NOT derived from evolutionary thinking
similar species can be grouped into hierarchical levels of similarity based on morphology
8 taxonomic ranks: DKPCOFGS
Binomial Nomenclature
genus followed by species
genus always capitalized, both always italicized
Phylogenetics
study of relationships among taxa based on their evolutionary history
goal: reconstruct evolutionary history of species and understand their pattern of descent
results are typically presented as a phylogeny/phylogenetic tree
phylogenies built using characters
Systematics
study of classifying organisms
Phylogenetic Systematics
modern approach to classification using evolutionary relationships between organisms
Character
any observable characteristics, or feature, of an organism
morphological, behavioural, genetic, etc.
ex. coat colour in oldfield mice, genetic sequence, etc.
Trait
the specific value of a character (i.e. the character state)
ex. brown or white coat, specific nucleotide (ACTG), etc.
used to build trees so that we may infer patterns of descent among species/populations
Traits in Phylogenetics
traits in phylogenetics are used to:
build trees
infer timing of evolutionary events by “mapping”
Phylogenetic/Evolutionary Trees
built by mapping traits of interest to infer a pattern of descent among species/populations
HYPOTHETICAL!
evolutionary histories are rarely directly observed
classifications will change as hypotheses about relationships change
Downside of Phylogenetic Trees
do not account for strange things which happen in evolution:
horizontal gene transfer
hybridization
merges
sometimes, trees are not great representations of evolutionary hisotries
however, they are still very useful tools!
Phylogenetic Tree Components
all trees have:
branches
branch tips
internal nodes
an outgroup
a root
Taxon
group of related organisms, placed at branch tip
could be species, genera, family, clades, etc.
Nodes
hypothetical common ancestors
ancestors do not resemble present day species
ancestral population, not individual
Branch Tips
represent decendents (following a node) on a phylogenetic tree
Derived Trait
a trait that arises from changes to the ancestral state/trait
characters therefore have polarity:
direction, or orientation
Ancestral → Derived Trait
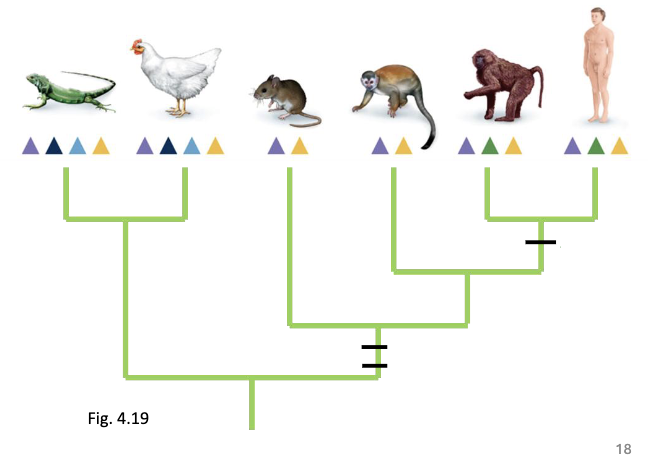
Mapping Example: Tetrapod Opsins
cone opsins: visual pigments
different animals have varied opsins, allowing them to see different wavelengths
Opsins are mapped at branch tips
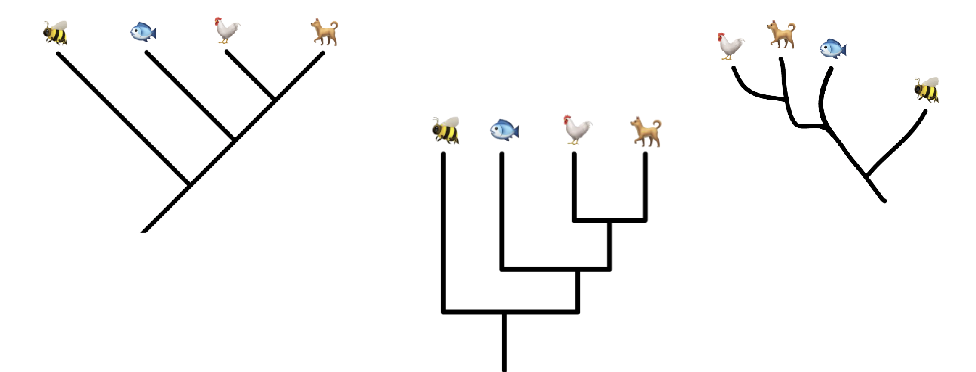
Orientation Phylogenetic Trees
there are several ways to draw trees, but the information conveyed about relationships among taxa is the same
relative positions from left to right do not indicate relatedness
distance to most recent common ancestor does (ie; the more recently that 2 groups share a common ancestor, the more closely related they are)
Pedigree vs. Phylogeny
nodes:
pedigree: individuals
phylogeny: populaitons
ancestors:
pedigree: 2 parents
phylogeny: 1 ancestral species
progression over time:
pedigree: expands backwards in time
phylogeny: expands forwards in time
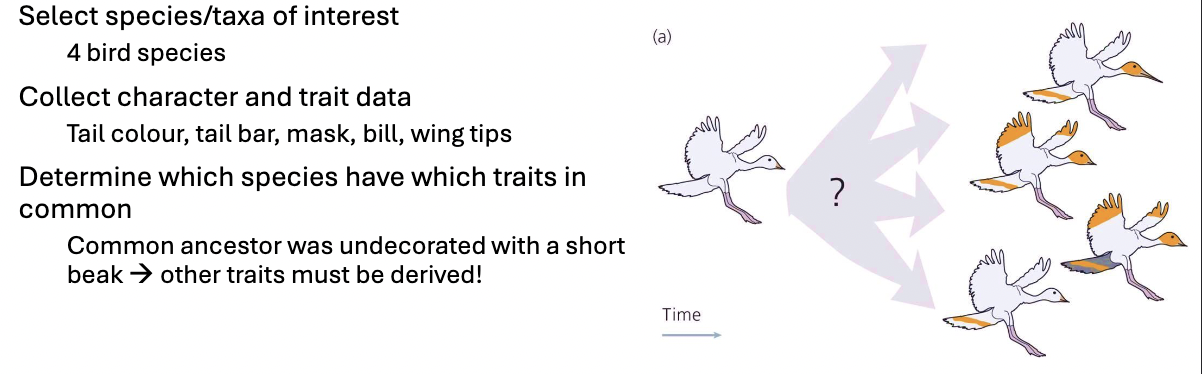
How to Build a Phylogeny
select species/taxa of interest
collect character/trait data
determine which species have which traits in common:
which traits are ancestral? which are derived?
logic: species with many traits in common are more likely to be related than species with few traits in common
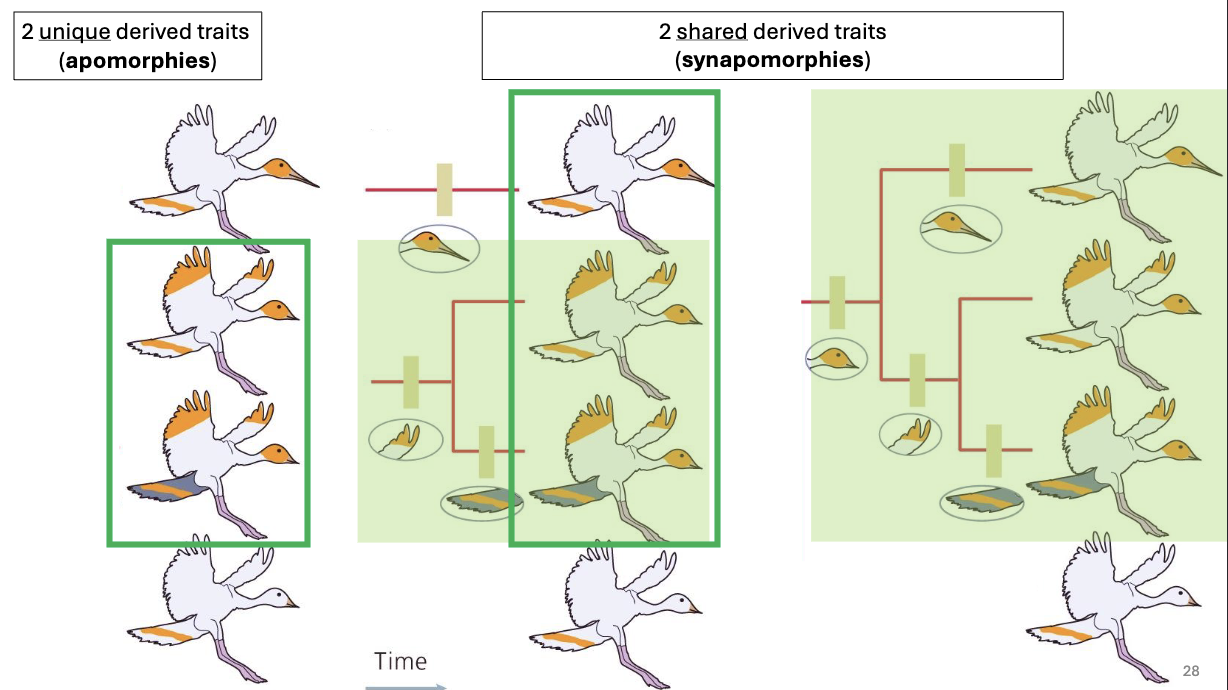
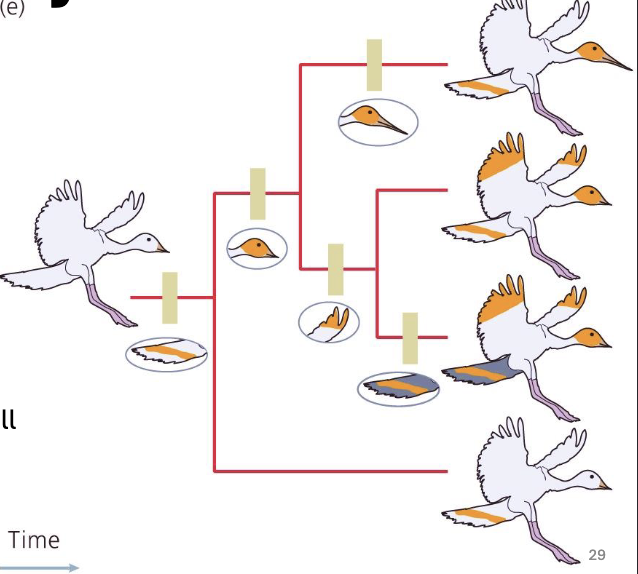

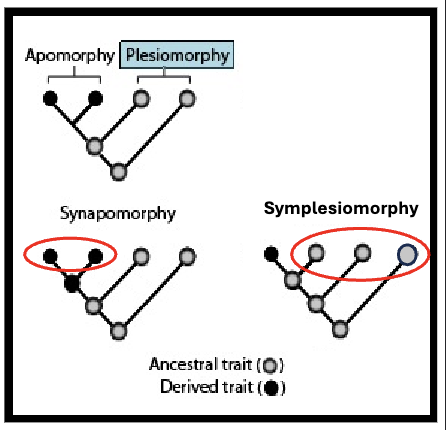
Apomorphy
evolutionary innovation
derived trait
what is called a plesiomorphy or apomorphy depends on context
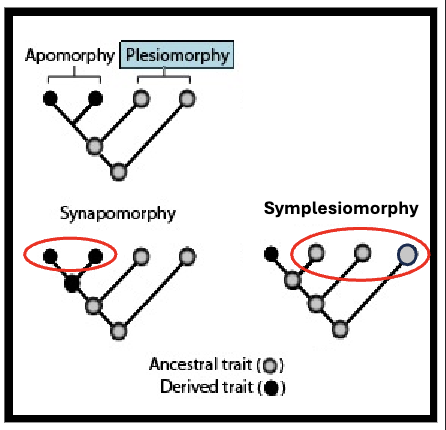
Plesiomorphy
pre-existing
ancestral trait
what is called a plesiomorphy or apomorphy depends on context
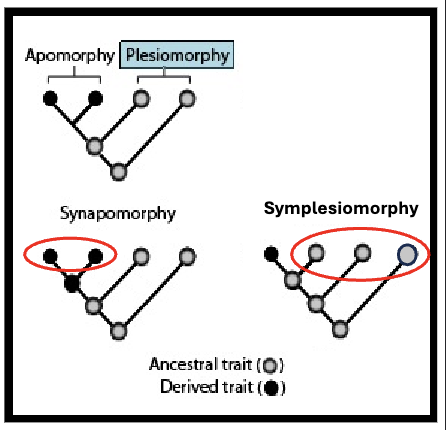
Synapomorphy
shared evolutionary innovation
derived trait
ideal to build trees with these, as the presence of synapomorphies (rather than the retention of ancestral characters) tells us about branch order
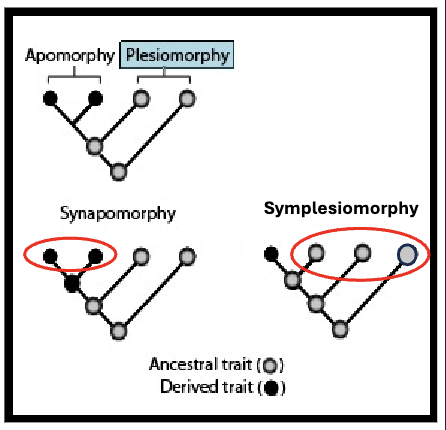
Symplesiomorphy
shared pre-existing
ancestral trait
Causes of Phylogenetic Similarity
homology
homoplasy
Homology
shared traits that are a result of common ancestry
homologous trait
ex. human eye and mouse eye
Homoplasy
similar trait but not because of inheritance from a common ancestor
analogy or reversals
Analogy
homoplasy
similarity due to convergent evolution
ex. human eye and octopus eye, bird and bat wings
Reversals
homoplasy
loss of a derived trait through a return to the ancestral state
Divergent vs. Convergent Evolution
divergent: homologous traits can take on different appearances or functions
convergent: development of analogous triat
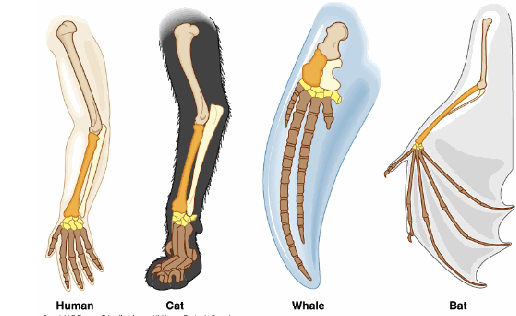
Divergent Evolution
closely related organisms diverge from each other due to differing selective pressures
homologous traits can take on different appearances or functions
ex. tetrapod limbs
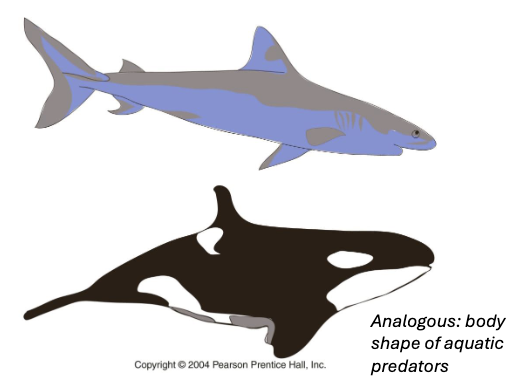
Convergent Evolution
two or more populations/species develop similarities due to similar selective pressures
develop analogous traits
ex. body shape of aquatic predators
Issue of Homoplasy
ideally build trees using synapomorphies, however convergence (or reversals) can be hard to identify
how to avoid this issue:
including multiple outgroups
provides more info about trait polarity and the ancestral state
including many characters
reduces chance that all traits are reversals or analogous
How do we distinguish homology from homoplasy?
comparative embryology (look at traits before extensive modification during development)
fossil record (transitional fossils linking past and present)
agreement with other phylogenetic hypothesises (assume homoplasy vs. logy if ‘stronger’ evidence indicates groups are not sister taxa)
number of features (inconsistent character most likely homoplasious)
complexity of features (if 2 similar complex structures match in many details, it is unlikely they evolved independently)
Outgroup Analysis
outgroup are a species (or group) that is less closely related to any member of the ingroup than the ingroup members are to each other
branched off earlier in history
used to make inferences about common ancestor of ingroup
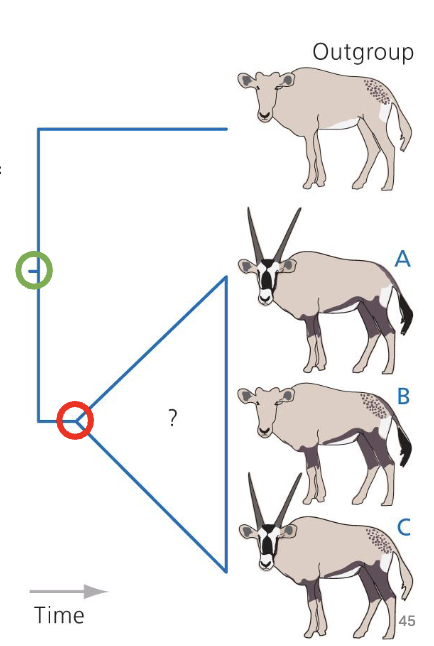
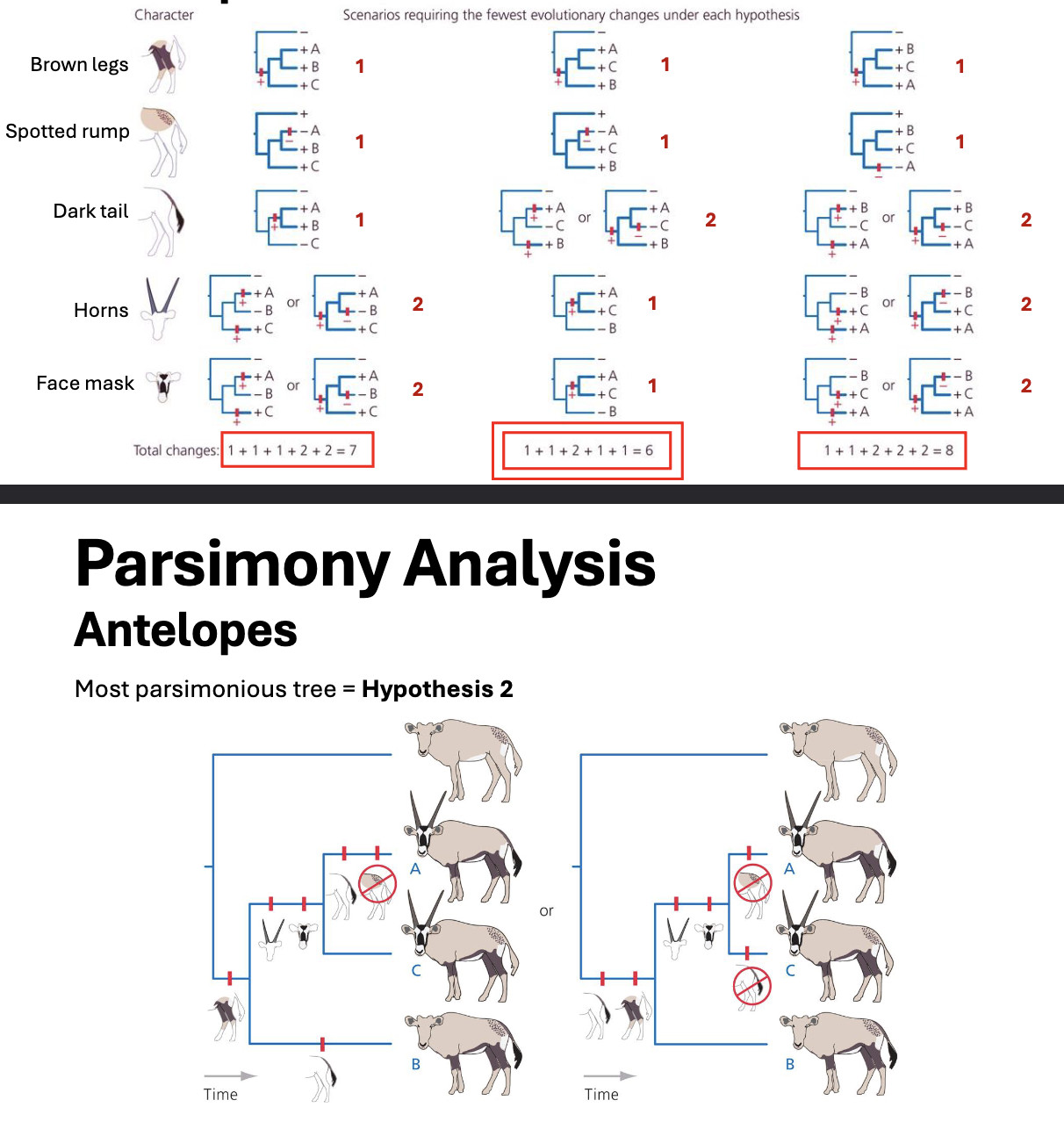
Parsimony Analysis
under parsimony, our most likely tree is that which minimizes the # of evolutionary changes
maximum parsimony= fewest changes
this approach gives no other cause for preferring one tree over another
Phylogeny
a phylogeny is a hypothesis about inferred evolutionary relationships
allow us to ask questions about trait origins
based on characters that are homologous and emphasize shared derived characters
synapomorphies
challenges include recognizing homology and direction of evolution
which is often unknown in ancestral states
Shoebills Evolutionary History
phylogeny from morphological characters (80s)
shoebill and heron: sister taxa
phylogeny from molecular genetics (2000s)
shoebill and pelican: sister taxa
phylogeny from current morphological studies (20s)
shoebill and heron: sister taxa (still!)
key point: phylogenies are hypotheses about evolutionary relationships, which can be continuously tested
Vestigial Traits
traits that serve no known current function
problem with Darwin’s theory
why do they remain?
not costly to remain
on its way out, just slowly
useful in phylogenetic reconstruction, as they can infer common ancestry