Uworld Biochem incorrects
1/24
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
25 Terms
which of the following peptides would exhibit the greatest steric constraints
amino acids with large side chains will feature greater steric constraints
steric constraints are when electrons clouds that are near each other repel, they usually occur in molecules that contain many atoms or atoms with large electron clouds
delta G describes what
relate this to amino acids
wether a reaction favors reactants or products
negative delta G means products are favored
positive delta G means reactants are favored
under physiological settings like PBS solution, proteins will spontaneously fold
folding is the product so equilibrium lies towards the products making delta G negative
unfolded is the reactant so equilibrium lies toward infilding making delta G positive
What are stacking (pi–pi) interactions, which amino acids can form them, and are they part of secondary or tertiary structure?
which aminos can form them
Stacking interactions (also called pi–pi interactions) are non-covalent attractions between aromatic rings.
They occur when the pi electron clouds (from double bonds in the rings) of two aromatic side chains interact and stabilize each other — like cards stacked in a pile.
Amino acids that can form pi–pi stacking interactions:
Phenylalanine (F) – benzene ring
Tyrosine (Y) – benzene ring with –OH
Tryptophan (W) – large indole ring
Histidine (H) – imidazole ring (aromatic, but weaker stacking)
Histidine is sometimes included because it has an aromatic ring, but:
Its ring is smaller, less hydrophobic, and partially charged at physiological pH.
So, its pi–pi stacking is weaker and less common.
Structural level:
These are tertiary structure interactions because they involve side chain–side chain attractions, not backbone hydrogen bonds.
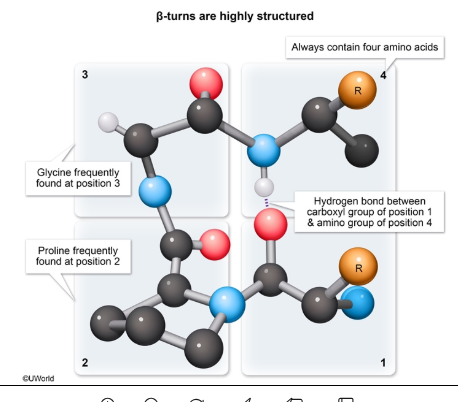
What are beta turns, what stabilizes them, and which amino acids are common in them?
Beta turns are short, 4-residue loops that reverse the direction of the polypeptide chain.
Part of secondary structure.
Stabilized by a hydrogen bond between the carbonyl of residue 1 and the amide of residue 4.
Proline (rigid) and glycine (flexible) are commonly found.
Often connect antiparallel beta sheets and help compact the protein
Why don’t parallel beta sheets use beta turns to connect strands?
Parallel beta sheets have strands running in the same direction, so no sharp reversal is needed.
Beta turns are used in antiparallel beta sheets to reverse chain direction.
Parallel sheets are connected by longer loops or crossover regions, not short 4-residue turns.
protein folding primarily driven by
hydrophobic effect
this causes proteins to
adopt a conformation that hides as many hydrophobic residues on the interior of the protein as possible
changes that cause a protein to misfold often lead to
exposure of hydrophobic residues to the aqueous envirnoment
these misfolded proteins often
aggregate or group together to hide the exposed residues
aggregation leads to an
decrease in protein solubility
Does protein solubility depend more on the peptide backbone or the amino acid side chains (residues)? Why?
Protein solubility depends primarily on the hydrophilicity/hydrophobicity of amino acid side chains, not the peptide backbone. The backbone is generally polar and buried in folded proteins, while side chains vary in polarity and determine how the protein interacts with water—hydrophilic residues increase solubility, hydrophobic residues decrease it.
beta turn
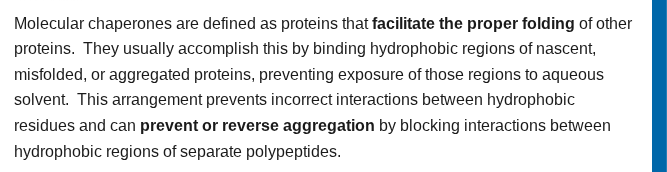
what are chaperone proteins

what is a degenerate genetic code
The mRNA code is degenerate because multiple codons (triplets of nucleotides) can code for the same amino acid, allowing some redundancy in the genetic code.
Q: What is the difference between antiport and symport transporters?
Antiport: Transports two different molecules or ions in opposite directions across a membrane.
Symport: Transports two different molecules or ions in the same direction across a membrane.
What is a uniport transporter?
A uniport transporter moves one type of molecule or ion across a membrane in one direction, usually down its concentration gradient. It can be passive or facilitated diffusion.
primary and secondary active transport
Primary active transport uses energy directly from ATP to move molecules against their gradient.
Secondary active transport (which includes most symporters and antiporters) uses the energy stored in the gradient of one molecule (usually ions like Na⁺ or H⁺) to drive the movement of another molecule against its gradient.
Q: Voltage-gated channels open in response to what change in electric potential?
They open when the membrane depolarizes—meaning the inside becomes less negative (voltage value increases from around –70 mV toward zero). This is described as a decrease in the magnitude of the electric potential difference across the membrane, even though the membrane potential value numerically increases.
Q: What are ligand-gated, mechanically gated, voltage-gated, and ion-gated channels?
Ligand-gated channels: Open or close in response to a specific chemical (ligand) binding.
Mechanically gated channels: Open or close in response to physical deformation or stretch of the membrane.
Voltage-gated channels: Open or close in response to changes in membrane potential (depolarization or hyperpolarization).
Ion-gated channels: (Less commonly used term) Typically refers to channels that open or close in response to specific ion binding.
how do amino acids exist as pH 7
At pH ~7.4, amino acids exist as zwitterions—molecules with both a positive and negative charge but overall neutral:
Amino group (–NH₂) is protonated → –NH₃⁺ (positive)
Carboxyl group (–COOH) is deprotonated → –COO⁻ (negative)
what are nucleophilic amino acids
Nucleophilic amino acids have side chains with lone pairs or pi electrons that can act as electron donors
read and interpret the movement of proteins through reducing sds gel
Multimers are proteins made of multiple polypeptide chains called subunits (dimers = 2, trimers = 3, tetramers = 4, etc.).
Reducing SDS-PAGE:
SDS denatures proteins and imparts a uniform negative charge.
The reducing agent breaks disulfide bonds linking subunits.
Each subunit runs independently through the gel.
Subunits of different sizes separate into distinct bands.
For homomultimers (identical subunits), all subunits migrate the same distance, producing one single band.
For heteromultimers (different subunits), subunits migrate differently, producing multiple separate bands.
According to the gel:
Each protein is a dimer.
Lanes A and C show single bands → these are homodimers.
Lanes B and D show two distinct bands → these are heterodimers.
Molecular weight of native dimer = sum of weights of individual subunits.
For homodimers, the weight of the single band is doubled to get the total protein weight.
Q: What is the difference between reducing and non-reducing SDS-PAGE?
Flashcard:
Q: What is the difference between reducing and non-reducing SDS-PAGE?
A:
Reducing SDS-PAGE: Uses a reducing agent to break disulfide bonds, separating protein subunits.
Non-reducing SDS-PAGE: Does not break disulfide bonds, so linked subunits stay together.
Used to determine whether proteins are single chains or linked by disulfide bonds.
Q: What makes up tertiary and quaternary protein structure, and how do they differ?
Tertiary structure: The 3D folding of a single polypeptide chain, stabilized by interactions like hydrogen bonds, ionic bonds, hydrophobic interactions, and disulfide bridges within that chain.
Quaternary structure: The assembly of multiple polypeptide chains (subunits) into a functional protein complex, held together by similar interactions (including disulfide bonds) between different chains.