Cell Biology Exam 4: Chapter 13
1/74
Earn XP
Description and Tags
DNA Replication and Repair
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
75 Terms
DNA duplicates by a process called ____ _________
DNA replication
the DNA replication machinery is also used for ___ _______
DNA repair
sense strand (what is it also known as & what does it contain)
also known as coding, plus, or no-template strand
contains codons & is same as mRNA except the T in DNA is replaced by U in the RNA
antisense strand (what is it also known as, what does it serve as, what is it complementary to, & what does it contain)
also known as non-coding, minus, or template strand
serves as a template for mRNA synthesis
hence complementary to both the sense strands and mRNA (with U in RNA in place of T)
the antisense/non-coding strand contains anti-codons
what are the two strands of the double helix held together by
hydrogen bonds between the bases
A & T (2 H bonds)
C & G (3 H bonds)
Watson & Crick: replication
envisioned that replication occurred by gradual separation of the strands of the double helix
how does DNA replication take place & what is this called
DNA replication takes place by separation of the strands of the double helix, and synthesis of two daughter strands complementary to the two parental templates → semiconservative replication
what are the 3 proposed models of DNA replication
conservative model
semiconservative model
dispersive model
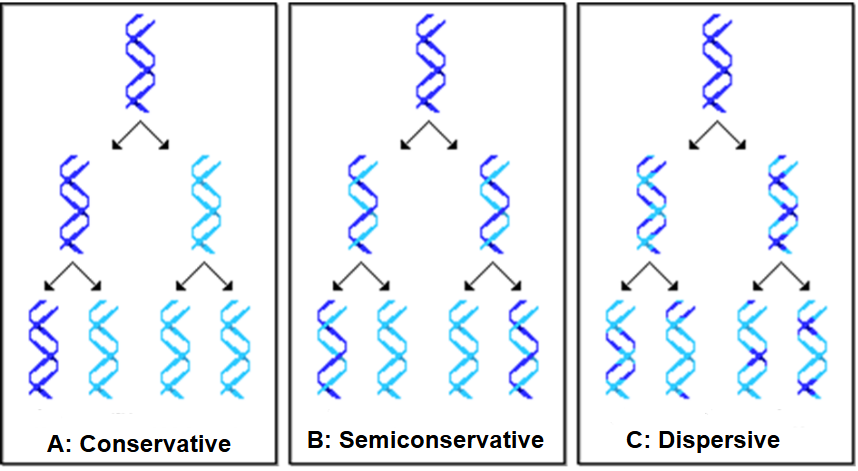
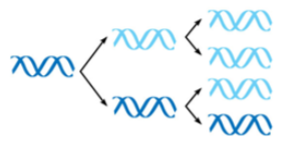
conservative model
parental strands re-associate after acting as templates for new strands
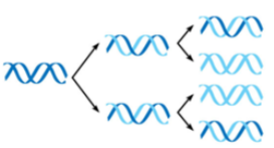
semiconservative model
the 2 parental strands separate; each acts as template for new, complementary strand
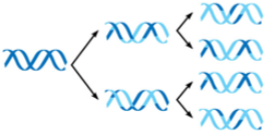
dispersive model
each daughter strand contains a mixture of parental and new DNA
what were the experiments performed to determine the currently accepted mode of DNA replication & prokaryote or eukaryote
Meselson and Stahl experiment (prokaryote)
Herbert Tylor’s experiment (eukaryote)
Meselson & Stahl experiment (what did they culture, what were the nitrogens, steps of experiment, what was concluded)
cultured E.Coli bacteria for several generations
15N was the heavy isotope of nitrogen and 14N was the lighter more common isotope of nitrogen
Steps
bacteria cultured in medium containing 15N
bacteria transferred to medium containing 14N
DNA sample centrifuged after 20 mins (1st replication → 1 band)
DNA sample centrifuged after 40 min (2nd replication → 2 bands)
conclusion: DNA replication follows the semiconservative model 1 band for 14N (higher) and another band with 14N &15N (lower)
why is DNA replication called semiconservative
half of the parent structure is retained in each of the daughter duplexes
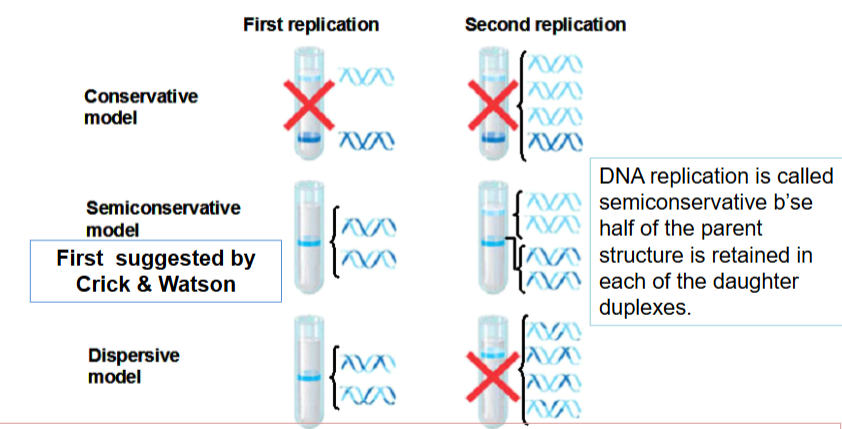
Herbert Tylor’s experiment (what was cultured, what compound was used, what were the findings)
cultured mammalian cells were allowed to undergo replication in bromodeoxyuridine (BrdU) a compound that is incorporated into DNA in place of thymidine
after 1 round of replication in BrdU → both chromatids of each chromosome conatined BrdU
after 2 rounds of replication in BrdU → one chromatid of each chromosome was composed of 2 BrdU-containing strands, & the other chromatif was a hybrid consisting of a BrdU-containing strand and a thymidine-containing strand
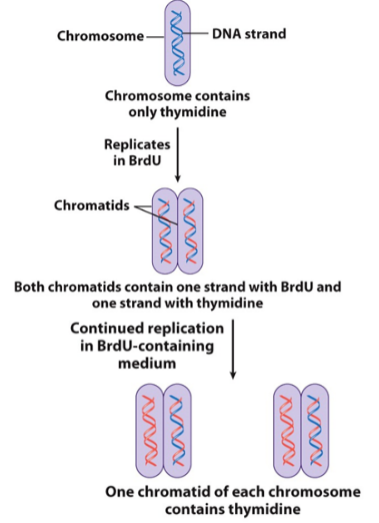
replication in bacteria: where does it start & what occurs
starts at the origin site (origin of replication) where a number of proteins bind to initiate replication
replication in bacteria: replication forks (what they are & what direction replication proceeds)
replication forks are points where a pair of replicating segments come together and join the nonreplicated segments
replication proceeds bidirectionally
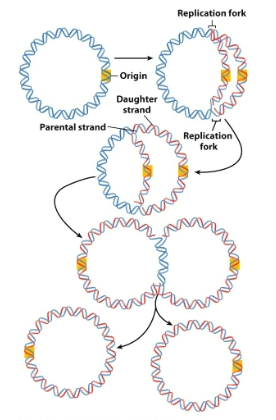
what occurs as DNA begins the unwinding process
tension is built up as DNA begins the unwinding process, and the DNA becomes positively supercoiled
DNA gyrase (what is it also called, what does it do, & what does it use)
topoisomerase II
relives tension by changing DNA into negatively supercoiled (underwound) DNA
uses ATP hydrolysis
torsional stress caused when unwinding DNA strands: what happens & what relieves tension
unseparated portion becomes more tightly wound
topoisomerase or gyrase breaks & rejoins coiled strands ahead of replication and thereby relives tension
DNA polymerase
responsible for synthesizing new DNA strands from a DNA template
what does DNA polymerase require
a primer
what do primers provide
provides the 3’-OH terminus on which to add new nucleotides
what direction does DNA replication/polymerization occur
5’ to 3’ direction
can any of the 3 DNA polymerases in bacteria initiate DNA chains?
no
how does DNA polymerization occur in terms of the OH group and the phosphate group
the -OH group at the 3’ end of the primer carries out a nucleophilic attack on the 5’-phosphate of the incoming nuceloside triphosphate
where can DNA replication only add new strands
the phosphate group always gets added to the 3’-OH end
what is at the 5’ carbon and 3’ carbon of DNA
5’ → phosphate group
3’ → hydroxyl group
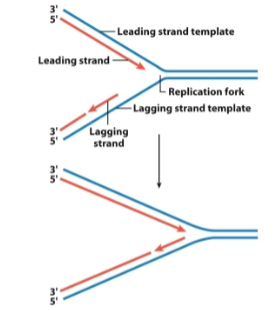
semidiscontinuous replication: how are both daughter strands synthesized
simultaneously
semidiscontinuous replication: the leading strand (direction & how it is synthesized)
in the direction of the replication fork movement
synthesized continuously
semidiscontinuous replication: the lagging strand (direction & how it is synthesized)
in the opposite direction of the replication fork
synthesized discontinuously
okazaki fragments (what they are & what are they joined by)
the small discontinuous fragments that the lagging strand is composed of which are joined by DNA ligase
DNA ligase
seals fragments/strands
seals Okazaki fragments into continuous strand
how is synthesis of each Okazaki fragment of the lagging strand intitated
short RNA fragments are used as removable primers in initiating synthesis of each Okazaki fragment of the lagging strand (primers introduce 3’-OH end)
why is synthesis of the lagging strand discontinous
during replication the phosphate group can only add at the 3’-OH end and the lagging strand has a 5’ end to start
that’s why the primers with 3’-OH ends need to be added
primase
an RNA polymerase that assembles short RNA primers
how are the RNA primers removed
by exonuclease activity of DNA polymerase I, which also fills in the gaps
what 2 things unwind the parental duplex and separate the two strands
Helicase
single-stranded DNA-binding (SSB) proteins
DNA polymerase III (what it is & what it does)
primary replication enzyme
synthesize successive fragments of the lagging strand
DNA polymerase II
DNA repair enzyme with a 3’ to 5’ exonuclease activity
what is DNA polymerase I involved in
DNA repair & also removes primers and replaces them with DNA
exonucleases
degrade nucleic acids by removing 5’ or 3’ terminal nucleotides
what are the exonuclease activities of DNA polymerase I
5’ → 3’: removes from 5’ end plays a role in removing the RNA primer
3’ → 5’: removes from the 3’ end removes mispaired nucleotides & maintains the accuracy of DNA synthesis
3’ → 5’ exonuclease of DNA polymerase I (what occurs during proofreading, what accounts for low error rates, how quick is replication)
during proofreading, mismatched bases are excised
careful selection of the nucleotide, proofreading, and mismatch repair account for low error rates in replication
replication is rapid
exonuclease activity DNA polymerase I vs II
DNA polymerase I
5’ → 3’ (removing RNA primers)
3’ → 5’ (proofreading)
DNA polymerase II
3’ → 5’ (proofreading & backup repair)
beginning of replication prokaryotes vs eukaryotes
prokaryotes
singe origin of replication
eukaryotes
multiple origins of replication (the genome is very big so one origin of replication would be too slow)
how do eukaryotes replicate their genome
in small portions (replicons)
there are _____ DNA polymerases in eukaryotes
several
how do eukaryotic DNA polymerases elongate (what direction, what is required, & what activity do some DNA polymerases have)
elongate in the 5’ to 3’ direction
require a primer
some have 3’ to 5’ exonuclease activity
functions: gyrase, primase, DNA ligase
gyrase (topoisomerase I/II): relieves positive supercoils ahead of replication fork
primase: synthesizes RNA primers
DNA ligase: seals Okazaki fragments into continuous strand
where is the replication machinery stationary
in the nuclear matrix
replication foci (what they are & what do they demonstrate)
replication foci are the sites in which replication forks are located
demonstrate that replication activities do not occur randomly throughout the nucleus but are confined to distinct sites
Helicase
unwinds DNA
single-stranded DNA binding proteins
keep strands apart
what is DNA repair essential for
cell survival
what can create spontaneous alteration (lesions) in DNA
ionizing radiation
common chemicals
UV radiation
thermal energy from normal metabolism
fidelity
how accurate DNA copying is
3 distinct categories that the fidelity of DNA can be traced to
accurate selection of nucleotides (base pairing rules)
immediate proofreading
post-replicative mismatch repair
what is formed within a DNA duplex following UV radiation
a pyrimidine dimer → tanning beds & prolonged sun exposure
what are the 4 DNA repair mechanisms
nucleotide excision repair
base excision repair
mismatch repair
double stranded break repair
base excision repair: what it does & what enzyme
removes altered nucleotides that produce distortions of the double helix
DNA glycosylases recognize the alterations and cleave the base from the sugar; they are specific for a particular type of altered base
once removed, an endonuclease cleaves the DNA backbone & a polymerase fills the gap by inserting a nucleotide complementary to the undamaged strand
DNA ligase seals the strand
DNA glycosylases: what they do & example
DNA glycosylases recognize the alterations and cleave the base from the sugar; they are specific for a particular type of altered base
ex.) hOGG1: DNA glycosylase; detects the oxidized form of guanine and it is able to fit into the active site of the enzyme
mismatch repair
the correction of mistakes that escape
what causes double-strand breaks
ionizing radiation (X-rays, gamma rays) along with some chemicals
by what pathways can double stranded breaks be repaired
non-homologous end joining
homologous recombination
double stranded breaks repair: nonhomologous end joining (NHEJ)
ku proteins bind to the free, broken ends and catalyze a reaction to rejoin the broken ends
mediated through DNA-dependent protein kinases
double stranded breaks repair: homologous recombination
requires a homologous chromosome to serve as a template for repair of the broken strand
more accurate than NHEJ
ATM protein kinases (what is it activated by, what does it do, & what does it activate)
activated by double-strand breaks
stops cell cycle
activates DNA repair proteins
ATR protein kinases (what is it activated by, what does it do, & what does it prevent)
activated by replication stress/single-stranded DNA
stabilizes replication fork
prevents collapse
what are skin cells with optimal levels of repair enzymes subject to
lesions that fail to be excised and repaired
skin cancer
disease promoted by deficiency or overworked DNA repair systems
colon cancer
due to mutations in mismatch repair genes