final bsci222 exam 2 flashcards
1/100
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
101 Terms
Prokaryotic chromosome structure
Usually single, circular chromosome with one origin of replication per chromosome
Can also have plasmids, which are small, circular accessory DNA that contains extra genes like antibiotic resistance; these replicate independently, and multiple plasmids can be found in the bacterial cell
DNA located in nucleoid - region in the cytoplasm where DNA is condensed - (no nucleus) - prokaryotic DNA is organized by NAPs (nucleoid associated proteins) that anchor DNA into loops and compact it in the nucleoid
Eukaryotic chromosome structure
Multiple linear chromosomes that can have multiple origins of replication per chromosome
Located in nucleus
DNA is wrapped around histone proteins, forming chromatin. This allows compaction (2 meters of DNA in 10 um nucleus), organization (several layers, such as dna in double helix, histones, nucleosmes, etc), and regulation (loose chromatin = gene transcription ON, tight = off)
Explain the function of histones and nucleosomes in structuring chromatin
Histones are positively charged proteins that bind negatively charged DNA, allowing DNA to wrap around them to form nucleosomes, the basic unit of chromatin.
Nucleosomes are formed from the DNA double helix coiled around a structure with eight histones. Histone H1 locks the DNA onto the nucleosome.
Nucleosomes can interact and pack together, helping to compact DNA and regulate transcription.
Nucleosomes help compact, organize, and regulate access to genetic info/DNA.
Heterochromatin
Condensed, tightly packed DNA that is not transcribed (RNA machinery cannot access the DNA)
Types of heterochromatin
Constitutive (centromeres, telomeres) - permanently compact DNA
Facultative (Barr body) - developmentally regulated DNA that can switch between active and inactive; Barr bodies are X chromosomes that are inactivated in female cells, but then reactivated before meiosis; 1X chromosome is turned into heterochromatin
Euchromatin
Loosely packed DNA that decondenses during the cell cycle to allow transcription
Give examples of chemical modifications to histones and explain their consequences for chromatin structure and gene expression.
Histone methylation - histone methyltransferase (HMT) enzyme adds a methyl group (-CH3) to the histone tails, making them more nonpolar. This reduces the overall charge on the nucleosome, condensing nucleosomes together through hydrophobic forces.
Histone demethylation - histone demethylase (HDM) removes methyl groups
Histone acetylation - adds acetyl group (-COCH3) to histone tails, which neutralizes their charge aka reduces the positive charge, loosening their interaction with the negatively charged DNA and allowing transcription
Histone deacetylation - histone deacetylase (HDACs) removes acetyl groups, allowing tails to regain positive charge and interact stronger with DNA = no transcription
Define the function of centromeres on chromosomes
Centromeres are specialized, heterochromatic regions that join sister chromatids and serve as the site of kinetochore formation for spindle attachment during cell division.
They are constitutive (permanent) heterochromatin (tightly packed), and contain the CENP-A histone in place of the H3 histone that indicates the centromere’s location
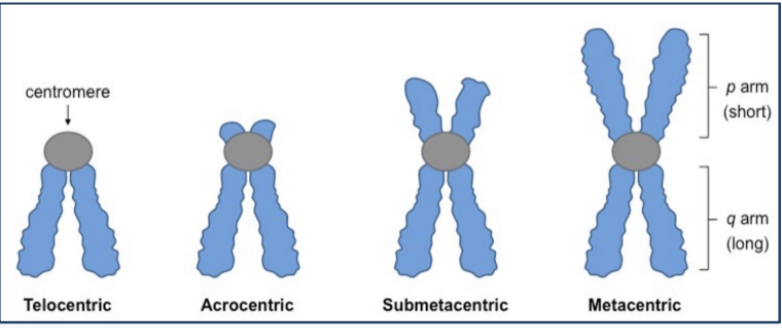
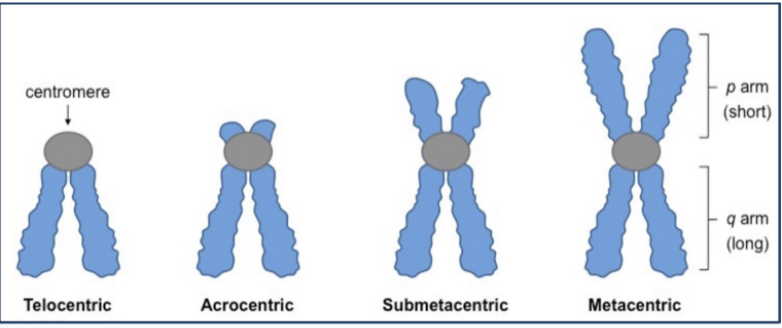
Different Chromosome types
based on centromere position
Telocentric - joins chromatids at tips (like bunny ears on bottom for q arm)
Acrocentric - joins chromatids near tips (short p arms; long q arms)
Submetacentric: joins chromatids near center
Metacentric: joins chromatids at center (symmetric arms)
Explain the structural features and function of telomeres and explain how they are maintained.
Telomeres are repetitive DNA sequences at the ends of chromosomes that protect chromosome ends and prevent degradation or fusion. They also act as a buffer during DNA replication
They are maintained by the enzyme telomerase, which adds to telomeres during gametogenesis - the creation of sperm and egg cells
Linkage
the tendency of genes or DNA sequences located close together on the same chromosome to be inherited together as a unit during meiosis
Linked genes violate Mendel's law of independent assortment because they do not separate independently, with closer proximity increasing the likelihood of co-inheritance.
Expectations for linked and unlinked genes in dihybrid crosses and test cross
Unlinked genes (independent assortment)
Dihybrid cross (AaBb x AaBb): Expect 9:3:3:1 ratio
Test cross (AaBb × aabb): 50% parental, 50% recombinant
Linked genes: more parental offspring than recombinant offspring
This is because the genes are inherited together most of the time
Expectations for linked and unlinked genes in trihybrid crosses
Unlinked genes (independent assortment)
Trihybrid cross (AaBbCc x AaBbCc): 27:9:9:9:3:3:3:1 ratio
Test cross (AaBbCc x aabbcc): 8 possible gametes, so 1 : 1 : 1 : 1 : 1 : 1 : 1 : 1
Linked genes: more parental offspring than recombinant offspring
This is because the genes are inherited together most of the time
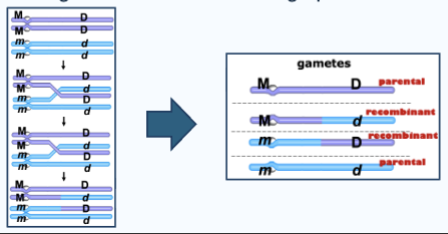
Crossover
aka recombination, is a fundamental process in meiosis that increases genetic diversity by exchanging genetic material between homologous chromosomes. This mechanism occurs during prophase I of meiosis I
Recombinants result from crossing-over during meiosis I
Homologous chromatids exchange pieces of DNA
Single crossover
One exchange happens between two chromosomes.
This produces: two recombinant chromatids and two parental chromatids
Double crossover
when two exchanges happen
Causes underestimates of recombination frequencies
Likelihood increases with distance between genes
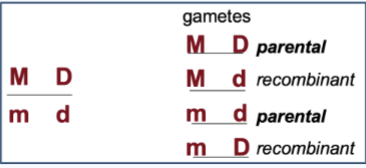
Coupling
how alleles are arranged on chromosomes BEFORE recombination
If the alleles are in cis (coupling), the dominant alleles are on the same chromosome
Example: AB / ab
This means the parental combinations are: AB and ab

Repulsion
how alleles are arranged on chromosomes BEFORE recombination
Each chromosome has one dominant and one recessive allele
Example: Ab / aB
This means the parental combinations are:
Ab and aB
Recombination frequency
how often crossing over occurs between two genes.
It is calculated as: (number of recombinant offspring ÷ total offspring) × 100
A low percentage means genes are very close together
A high percentage means genes are far apart
1% recombination = 1 map unit (centiMorgan)
Two genes are linked if
<50% recombination frequency
Two genes are not linked if
50% recombination frequency (independent assortment)
Coefficient of coincidence (COC)
Number of double crossovers is lower than expected because of crossing over at one region (chiasmata) inhibits crossing over at another region
Calculate by #observed DCOs/#expected DCOs
If COC is high, then that means there are more observed double crossover events
Ex. 10 expected DCO and 2 observed DCO
COC = 2 observed/10 expected = 0.2
Interference
1 - COC
If interference is high = low COC, so less observed DCOs than expected because crossing over alters the chromosome structure and interferes with crossing over at other chiasmata
Interference = 1 = 0 COC = almost no double crossovers
Interference = 0 = 1 COC = observed DCO = expected DCO = does NOT mean lots of crossovers; occurence of 1 crossover does not affect another
How to determine the linkage distance between two genes in a dihybrid cross
1) Identify which offspring are parental and which are recombinant.
2) Calculate the recombination frequency = is the number of recombinant offspring divided by the total number of offspring, multiplied by 100. That percentage is the linkage distance in centiMorgans (cM). One percent recombination equals one map unit.
For example, if a test cross gives 55 parental, 53 parental, 8 recombinant, and 7 recombinant offspring, then the recombination frequency is: (8 + 7)/(55 + 53 + 8 + 7) x 100% = 12.2%
So the two genes are 12.2 cM apart.
How to determine if the dominant alleles of a heterozygous parent are in cis or trans configuration
Ex. You perform a test cross:
AaBb × aabb
You get the following offspring:
40 AB
42 ab
9 Ab
9 aB
1) Identify the parental types (AB and ab because they are the highest frequency)
2) Since parental look like AB / ab, that means this is cis coupling, because the parental types have both dominant alleles together (AB) and both recessive alleles together (ab)
The alleles are in cis (coupling) configuration
Use the terms cis or trans and coupling or repulsion correctly when predicting offspring. Parent 1 is the genotype aabb is crossed with parent 2 of the genotype AaBb. Parent’s 2 genes are in coupling. What are the potential arrangements of alleles in the offspring? Which of these are the parental and which are the recombinant genotypes in the offspring?
Parent 1: aabb (homozygous recessive) —> can only produce ab gametes
Parent 2: AaBb, and the genes are in coupling (cis)
Since parent 2 is coupled, AB / ab
can produce AB, ab gametes (parental, most common) and Ab, aB gametes (recombinant, less common, from crossing over)
Combining gametes, we get
From parental gametes: AB × ab → AaBb and ab × ab → aabb (most common offspring)
From recombinant gametes: Ab x ab —> Aabb and aB x ab —> aaBb (less common)
Because the heterozygous parent is in coupling (cis configuration, AB / ab), it produces mainly AB and ab gametes, along with fewer recombinant gametes (Ab and aB). When crossed with aabb, the offspring will be AaBb and aabb as the most common parental types, and Aabb and aaBb as less frequent recombinant types.
Identify when DNA replication occurs in the cell cycle
DNA replication occurs during the S-phase of the cell cycle before cell division
Define semi-conservative synthesis.
Each daughter DNA molecule contains one parental strand and one newly synthesized strand
Origin of replication
Specific DNA sequence where replication begins, and DNA unwinds to form a replication bubble with 2 replication forks, one on each side
Replicon
A unit of DNA replicated from a single origin
Prokaryotes vs. Eukaryotes origin of repiclation/replicon
Prokaryotes: Single origin (oriC), One replication bubble, One replicon
Eukaryotes: Multiple origins, Many replication bubbles, Many replicons
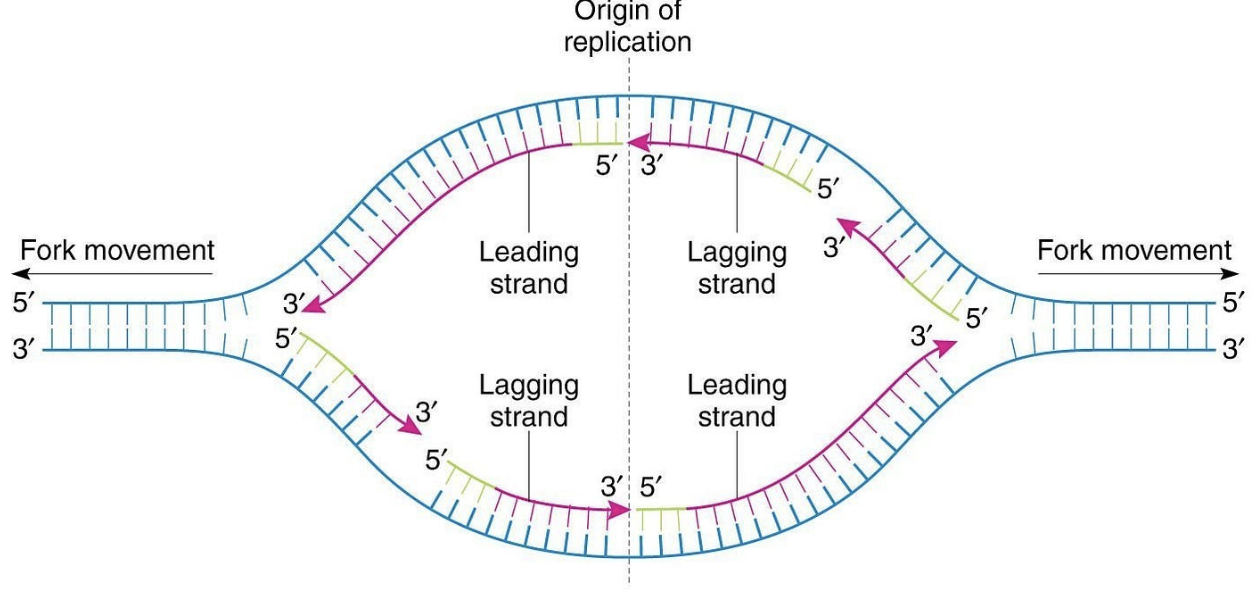
Draw a replication bubble, demonstrating why daughter strands are synthesized in opposite directions and where you would expect continuous and discontinuous synthesis.
Daughter strands are synthesized in opposite directions because DNA strands are antiparallel to each other (one is 5’ to 3’ direction, other is 3’ to 5’) and DNA synthesis by DNA polymerase can only occur in only 5′ → 3′ direction.
The leading strand is synthesized continuously while the lagging strand is synthesized discontinuously as Okazaki fragments.
Continuous synthesis on one strand (3’-end towards the fork)
Discontinuous synthesis on other strand (Okazaki fragments)
Replication fork
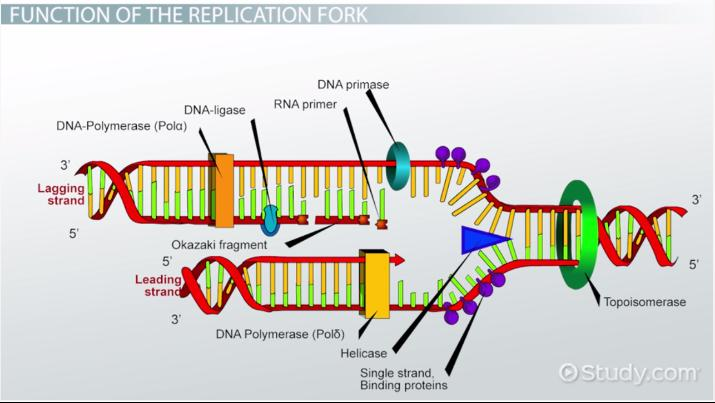
DNA replication prokaryotic enzymes
Helicase: unwinds the DNA by breaking hydrogen bonds between nitrogenous bases (not the backbone)
Gyrase (type II topoisomerase): relieves supercoiling ahead of the replication fork
Cuts both DNA strands, twists them, and then joins them together
RNA primase: binds to the single-stranded DNA and adds a primer to the 3’ end, providing a 3’ OH group DNA polymerase can add to
DNA polymerase III - adds complementary DNA nucleotides in the 5’ to 3’ direction, starting at the primer
DNA polymerase I - has a 5’ to 3’ exonuclease that removes the RNA primers and fills the gaps
Ligase - joins the distant lagging strands by phosphodiester bonds (joins 3’ OH and 5’ phosphate after DNA polymerase 1 removes primers); does not add nucleotide
DNA replication eukaryotic enzymes
Helicase: during G1, origin recruits MCM2-7 proteins that bind to DNA at the origin and form an inactive complex; during S phase, MCM2-7 proteins are activated to become helicase, and it begins unwinding
Topoisomerase (class I): relieves supercoiling by cutting ONE strand, allowing it to relax, and then rejoining strand ahead of fork
Dna polymerase has 3 different proteins
DNA polymerase alpha: acts like RNA primase, adding short RNA primers and adds DNA bases to 3’ end of template strands
DNA polymerase delta: lagging strand synthesizing (extend Okazaki fragments) - repeatedly restarts after each primer
DNA polymerase epsilon: leading strand synthesis
FEN1: exonuclease that proofreads
DNA ligase: joins fragments
Prokaryotic DNA replication steps
Initiation - origin of replication recruits initiator proteins (dnaA) that bind cooperatively to dnaA boxes in the oriC near DUEs (DNA unwinding elements) - also on the oriC begin to unwind (melt), allowing other enzymes to uwind the DNA - repliation fork created
Bubble expansion - helicase breaks the hydrogen bonds between nitrogenous bases, not the backbone, working in the 5’ to 3’ direction on the lagging strand
Single-stranded binding proteins (ssBPs) keep strands separate; bind to nucleotides and prevent re-annealing and secondary structure fomration
Gyrase (type II topoisomerase) relieves supercoiling ahead of the replication fork; cuts both DNA strands, twists them, and then joins them together
DNA strand synthesis - RNA primase binds to the single-stranded DNA and adds a primer to the 3’ end, providing a 3’-OH group DNA polymerase can add to
DNA polymerase III adds complementary nucleotides in the 5’ to 3’ direction, starting at the primer
Has both polymerase (binds DNA) and exonuclease activity (remove DNA), performs proofreading (if an incorrect base is placed, then the exonuclease in DNA poly III is triggered and cuts out the mistake, then replace it with the correct nucleotide)
Exonuclease works in the 3’ to 5’ direction of the newly synthesized DNA
DNA Polymerase I has a 5’ to 3’ exonuclease that removes the RNA primers and fills the gaps
Ligase joins the strands by phosphodiester bonds (joins 3’OH and 5’ phosphate ends after DNA polymerase I)
Termination
Prokaryotes have a terminator sequence (Ter) - Tus protein binds to the Ter sequence, acting as a roadblock. This blocks helciase from unwinding. When one replication fork hits the Ter-Tus complex, it stalls. When the other replication fork meets it, replication stops.
Eukaryotic DNA replication steps
Initiation - origins of replciation recruit the origin recognition complex (ORC) that anchors helicase to the origin; ORC binds during the G1 phase
Licensing factors determine which origins are active and undergo DNA replication
2. Bubble expansion - during G1, origin recruits MCM2-7 proteins that bind to DNA at the origin and form a compelx, but they are inactive
During S phase, MCM2-7 protein sare activated to become hecliase, and it begins unwinding
Class I topoisomerase relives supercoiling by cutting ONE strand, allowing it to relax, twisting, and rejoining strands
DNA strand synthesis
DNA polyemrase in eukaryotes has 3 different proteins
DNA alpha - actsl ike RNA polymerase, adds short RNA primer and DNA bases to 3’ end of template strands
DNA delta - lagging strand synthesis (extend Okazaki fragment); repeatedly restarts after each primer
DNA epsilon - leading strand synthesis
FEN1- exonuclease that proofreads
RNA Transcription
making RNA from DNA template
Why RNA transcription?
B/C DNA contains the genetic info for making protein, but isk ept in the nucleus (can’t leave)
mRNA = reverse-complement copy of DNA
mRNA can be delivered to the ribosomes that then build polypeptides based off the mRNA sequence
Example of RNA transcription
DNA template: 3’ — TAC —- 5’
What is the mRNA?
5’ —- AUG — 3’
Different types of RNA that need to be transcribed
Cells have several classes of RNA molecules that serve different functions, including:
mRNA - messenger RNA; provides information to ribosome to build a polypeptide.
rRNA - ribosomal RNA; allows ribosomes to bind to mRNA.
tRNA - transfer RNA; binds to mRNA in ribosomes to deliver amino acids.
snRNAs - small nuclear RNAs; allows snRNPs to interact with pre-mRNA for splicing.
miRNAs, siRNAs - involved in regulation of mRNA through degradation.
Promoter
Promoter - Upstream DNA sequence where RNA polymerase binds
Prokaryotes: -35 and -10 (Pribnow box)
Eukaryotes: TATA box -25
Template strand
The strand RNA polymerase reads; Always read 3′ → 5′
mRNA is complementary to this strand
Direction of transcription
RNA polymerase moves in the 5’ to 3’ direction along the template strand
Sequence of RNA product
Complementary to template strand
Same as coding strand except: U instead of T
EXAMPLE - DNA:
Coding strand (always 5’ to 3’ direction):
5′ — ATG CCG TTA — 3′
Template strand (always 3’ to 5’ direction):
3′ — TAC GGC AAT — 5′
mRNA product?
mRNA = 5′ — AUG CCG UUA — 3′
Prokaryotic RNA Polymerase
One RNA polymerase
Requires sigma factor for promoter recognition
Eukaryotic RNA polymerase
Multiple different RNA polymerases depending on what product is being made
RNA polymerase I - rRNA
RNA polymerase II - pre-mRNA, small RNAs
RNA polymerase III - tRNA, small RNAs
Require:
1) Basal transcription factors - Proteins required to initiate transcription by helping RNA polymerase bind to the promoter
2) Regulatory transcription factors - Proteins that control the rate of transcription by enhancing or inhibiting RNA polymerase activity by binding regulatory DNA elements like enhancers and proximal elements
Prokaryotic RNA transcription
requires RNA polymerase and sigma factors
Initiation -
sigma factor (bacterial transcription initiation protein) complexes with RNA polymerase at the promoter sequences 1) Pribnow (TATA box) -10 bp and at 2) second continuous sequence (-25 bp). The sigma factor positions the haloenzyme from -50bp to 20bp.
Formation of the transcription bubble: haloenzyme conformationally changes shape to bind tighter to the promoter sequence; at -10bp (Pribnow), unwinding occurs as as a result, and RNA polymerase only uses the template DNA (non-coding)
Initiation starts at +1 site.
Elongation - RNA polymerase travels downstream the template strand and synthesizes complementary RNA nucleotides in a 5’ to 3’ reaction in the transcription bubble
Termination - 2 kinds depending on the structure of RNA
Rho-independent - termination sequence is transcribed, the RNA folds back on itself due to reverse-complementary sequences on itself to form a hairpin loop structure - causes RNA polymerase to pause, and disrupts the RNA:DNA duplex, releasing RNA (normally, RNA is base-paired to DNA inside RNA polymerase; but the hairpin pulls the RNA away from the DNA and causes RNA polymerase to stop)
Rho-dependent - The RNA sequence does NOT form a hairpin; termination sequence is transcribed, with rut site where Rho protein binds. Rho moves in 5'->3' direction along RNA, and when it reaches the polymerase it disrupts RNA:DNA interaction, freeing the RNA (pulls RNA polymerase away from the DNA template)
Rho-independent termination
termination sequence is transcribed, the RNA folds back on itself due to reverse-complementary sequences on itself to form a hairpin loop structure - causes RNA polymerase to pause, and disrupts the RNA:DNA duplex, releasing RNA (normally, RNA is base-paired to DNA inside RNA polymerase; but the hairpin pulls the RNA away from the DNA and causes RNA polymerase to stop)
Rho-dependent termination
The RNA sequence does NOT form a hairpin; termination sequence is transcribed, with rut site where Rho protein binds. Rho moves in 5'->3' direction along RNA, and when it reaches the polymerase it disrupts RNA:DNA interaction, freeing the RNA (pulls RNA polymerase away from the DNA template)
basically Rho yanks RNA polymerase off
Differences in Eukaryotic RNA transcription
more complex
Key differences between prokaryotic and eukaryotic RNA transcription
Chromatin remodeling - modification of chromatin structure to affect the accessibility of DNA for transcription
Promoter diversity - diff rna polymerases (1 to 3) in eukaryotes mean they have diff protomers to attach to
Termination
Prokaryotes: rho-independent/rho-dependent
Eukaryotes: no formal termination sequence; relies on DNA processing and degradation
Explain how chromatin remodeling factors and histone modifications are involved in eukaryotic transcriptional regulation.
Eukaryotic RNA transcription
Initiation -
Chromatin remodeling - histones in nucleosomes are modified by de-methylation and acetylation to make DNA available for transcription
Basal transcription factors - required proteins that help RNA polymerase II bind to the promoter at the TATA box, -25 base pairs. In this case, the TATA binding protein
Regulatory transcription factors - modify RNA polymerase activity, binding upstream as activators or repressors
Activators make RNA polymerase work better
Repressors interfere with RNA polymerase function
2. Elongation - NA polymerase moves along the DNA template and synthesizes the RNA strand.
A → U
T → A
G ↔ C
3. Termination - no specific termination sequence at the end, eukaryotes rely on degradation
RNA polymerase II transcribes the Poly A signal, which is the stop sequence in the 3’ UTR
Rat1 enzyme recognizes the sequence and cleaves the mRNA transcript, degrading the mRNA after the PolyA in the 5’ to 3’ direction
This releases the pre-mRNA for further processing
RNA processing
eukaryotes modify transcript (pre-mRNA) into messenger RNA (mRNA).
Describe how eukaryotic mRNA is modified by addition of a 5'-cap and 3'-polyA tail, and give functions for those structures.
Polyadenylation - adds a 3' PolyA Tail to mRNA, protecting the 3'-end of the protein coding sequence from early degradation, assisting ribosome binding, and contributing to nuclear export.
Capping - A 5'-cap is added that prevents degradation by 5'->3' exonucleases, assists in ribosome binding, and contributes to nuclear export.
Discuss why splicing is needed and how alternative splicing contributes to protein diversity in eukaryotes.
Splicing - removes Intron sequences from the mRNA, Required for export from nucleus to cytoplasm
Alternative splicing can shuffle exons making different mRNAs from the same RNA transcript (Different combinations of exons can be joined to produce multiple mRNAs from one pre-mRNA.); one gene → multiple proteins, increasing protein diversity without increasing gene number
Recognize that tRNA nucleotides are chemically modified to support structural demands.
tRNAs (transferRNAs that act a translators between ribosomes and mRNA sequence, bringing amino acids that match codons on mRNA to ribosomes)
uses more than 4 nucleotides/bases (Normally RNA = A, U, G, C, but in tRNA: some bases are chemically modified → “rare bases”) Ex. Uracil → Ribothymidine
tRNA-modifying enzymes chemically alter transcribed bases (after tRNA is made) to stabilize proper folding and aid secondary structure formation (how the RNA folds), ensuring the tRNA functions correctly during translation.
Base modifications are an important part of tRNA processing, aiding in formation of complex structures.
Translation
mRNA —> polypeptides
Recognize the importance of changes in amino acid sequences to functional changes in proteins as a consequence of changes to shape, ability to change shape, and R-group interactions with other molecules.
Changes in amino acid sequence → changes in protein structure → changes in function. Protein function is determined by protein structure including shape, ability to change shape, and interactions with other molecules at R-groups. A change in amino-acid sequence of a polypeptide can alter protein function
R-group (side chain) changes
Each amino acid has different properties: charge (positive/negative), polarity, size
hydrophobic vs hydrophilic
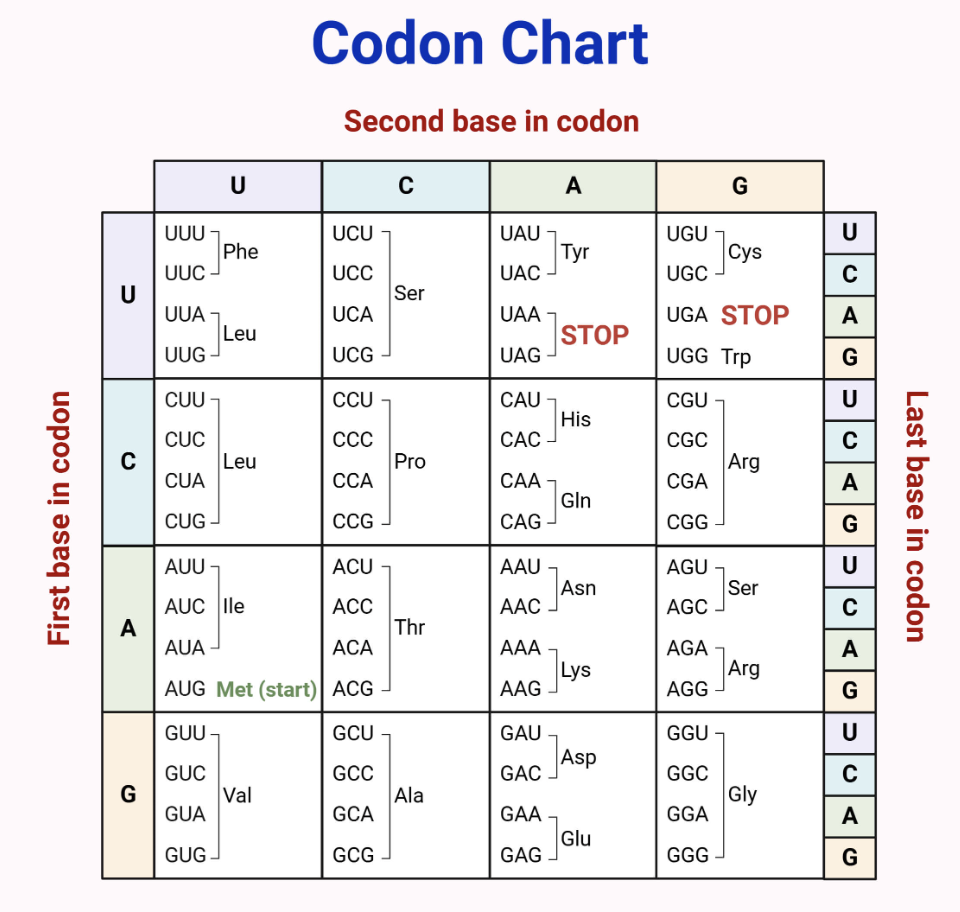
Given the mRNA sequence:
5’–CCGAUGGCUACCUAAUGG–3’
What is the correct polypeptide?
1) Find first AUG sequence
5’–CCGAUGGCUACCUAAUGG–3’
2) Read in codons (set of three RNA nucleotides)
AUG - GCU - ACC - UAA - UGG
3) Translate using codon table
AUG → Met
GCU → Ala
ACC → Thr
UAA → STOP
Therefore, the final polypeptide is Met-Ala-Thr
Translation initiation in prokaryotes
The ribosome binds to the Shine Delgarno sequence (5’-AGGAGG-3’) on mRNA, which positions the ribosome at the correct start codon.
An initiator tRNA, carrying the amino acid fMet (anti-codon), binds to the start codon (AUG) on MRNA with the aid of Initiation Factor proteins.
Initiator Factor (IFs) proteins also assemble the large subunit onto the small subunit with fMet in the P-site, forming the translation initiation complex.
Translation initiation in eukaryotes
The small ribosomal subunit binds the mRNA at the 5'-cap and scans for the first 5' AUG codon to use as the start codon, which is part of a short consensus called the Kozak sequence (5’-ACCAUGG-3’)
As in prokaryotes, an fMet initiator tRNA (carrying amino acid fMet) binds at the start codon (AUG) and the ribosomal initiation complex assembles.
Gene regulation - structural genes
Genes that encode proteins that do the actual work of the cell; metabolic and structural proteins; do the job;
Make proteins (enzymes, receptors, etc.)
Ex. Enzymes, Receptors, Structural proteins, Transport proteins, Hormones, Cytoskeletal proteins, Membrane proteins, Metabolic enzymes, Biosynthetic enzymes, etc.
regulatory genes
Genes that encode proteins that control expression of other genes
Ex. The Max gene product is a DNA binding protein that promotes expression of metabolic proteins for cellular respiration
Make transcription factors (TFs), which are proteins that control other genes. They do this because theyhey are DNA binding proteins that hydrogen bond to DNA. Transcription factors insert into the major groove and can hydrogen bond to particular base pairs.
2 types of TFs: Activators and Inhibitors/Repressors
Activators: promote transcription; Help RNA polymerase bind or work better = positive control (regulatory mechanism #1)
Inhibitors/repressors: Block transcription; prevent RNA polymerase from working = negative control (regulatory mechanism #2)
Regulatory genes PRODUCE proteins, while transcription factors proteins carry out positive or negative control
Positive (transcriptional) control
activators that boost transcription by helping RNA polymerase bind to DNA
Is something helping RNA polymerase bind?
→ Positive control
Negative (transcriptional) control
inhibitors/repressors that block transcription by interfering with RNA polymerase
Is something blocking RNA polymerase?
→ Negative control
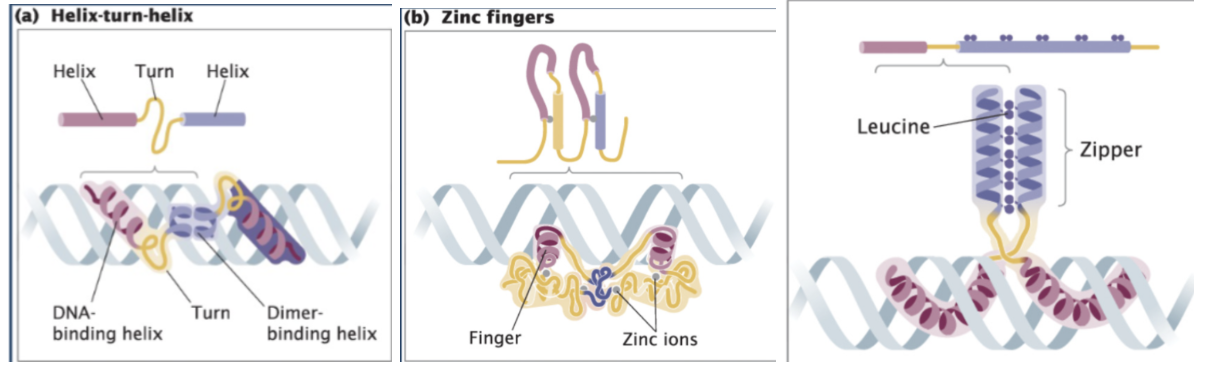
Recognize examples of different types of DNA-binding proteins (Helix-turn-Helix, Zinc- finger, and Leucine Zipper proteins).
DNA-binding motifs - Specific structural features of transcription factors that allow them to bind specific DNA sequences, typically in the major groove, to regulate gene expression (Motif = shape that lets protein bind DNA)
Helix-turn-helix - Two symmetrical helices = dimer that fits into the major groove
Zinc Finger proteins - Metal ion (zinc) can interact with four different amino acids and pull them together, folding the protein into a particular loop called a zinc finger that exactly fits into the major groove of the DNA; Uses zinc ion to stabilize shape; Forms a “finger” that inserts into DNA
Leucine zipper - Two proteins stick together (dimer); purple has bunch of amino acid leucine which interacts with neighboring helix and stick together; called a zipper because the two helices are stuck together vertically; proteins zip together → then bind DNA
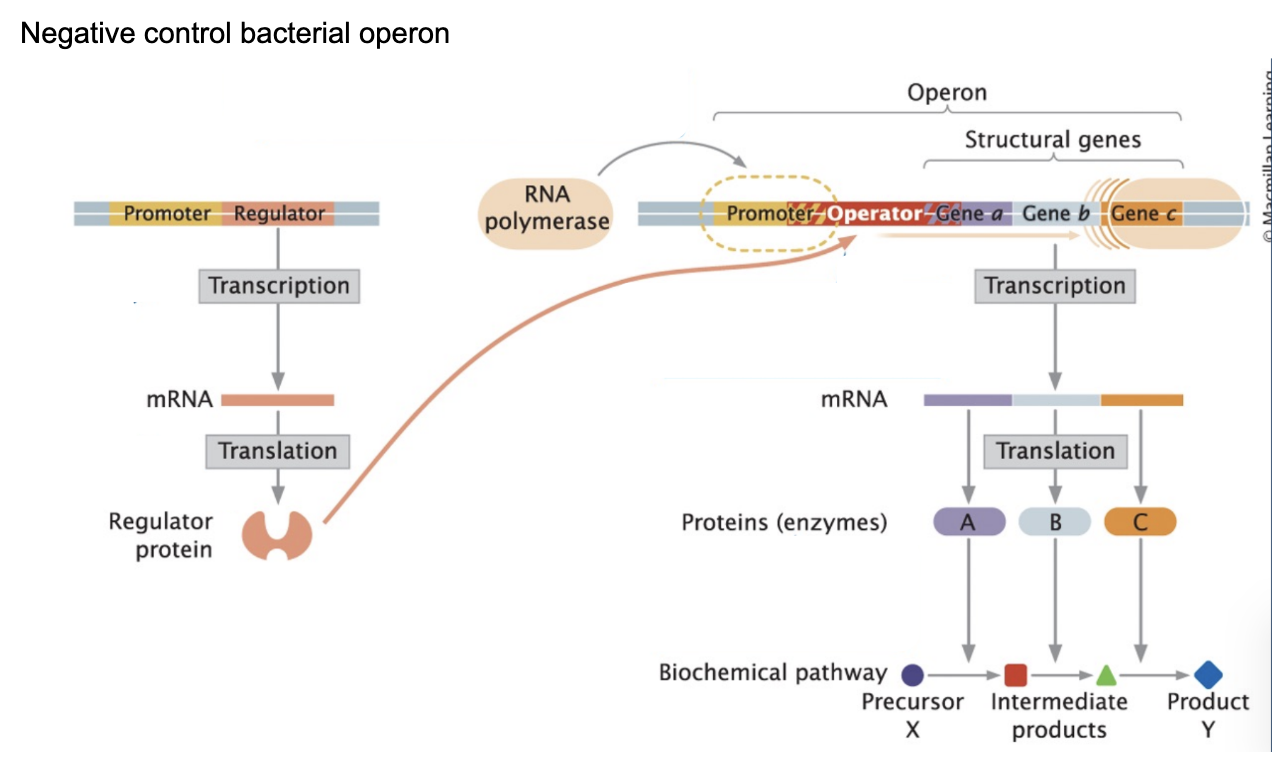
Operon
A group of genes controlled by one promoter that are transcribed together into a single mRNA
Structure of an operon
Regulatory gene → Promoter → Operator → Structural genes (A, B, C)
Regulatory gene → makes repressor protein
NOT part of the operon/operator
Located outside the operon
Codes for the repressor protein
Has its own promoter
It just makes the regulator protein
Transcription occurs when repressor not on DNA
Promoter →where RNA polymerase binds
Operator → repressor binding site (operator adjacent to the promoter)
Inhibits RNA polymerase by acting as a roadblock
Structural genes → encode proteins
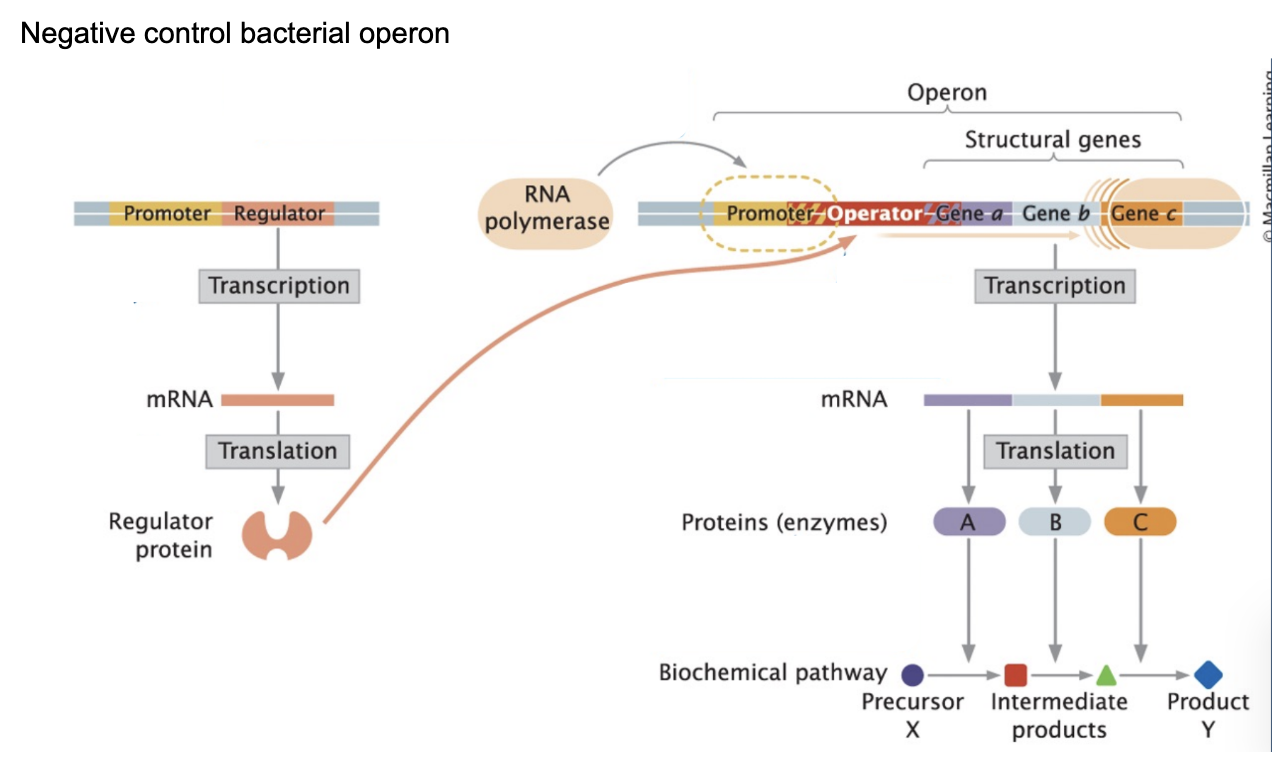
Polycistronic RNA product is made by…
One mRNA contains multiple genes
“one promoter controls several protein coding regions (genes)”
Each gene has its own: Shine-Dalgarno sequence → multiple proteins made from one mRNA
A polycistronic RNA is produced when a single promoter drives transcription of multiple adjacent genes in an operon. This results in one mRNA molecule that contains multiple coding regions. Each gene within the mRNA has its own Shine-Dalgarno sequence, allowing ribosomes to independently initiate translation at each site and produce multiple different proteins from the same mRNA.
Inducible operon
operons that are normally repressed (default = OFF), as the repressor protein is bound to the operator site
The repressor is functional in absence of the INDUCER - a small molecule that triggers gene expression by disabling a repressor protein, allowing RNA polymerase to transcribe specific genes’
Transcription is turned on by de-repression - removing the repressor
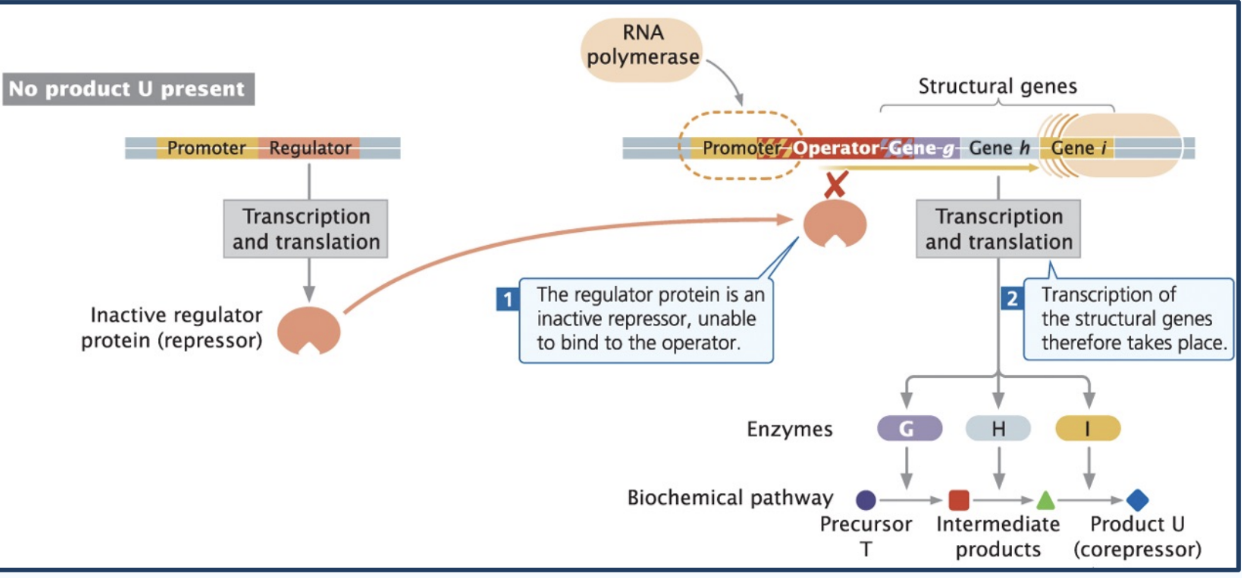
Ex. of inducible operon
Ex. Lac operon = inducible operon
LacR repressor constitutively expressed (always made in the cell)
Cells use glucose when available, preferring it over lactose (lac operon repressed
If no glucose, but lactose available -> derepress operon, so that lac operon can be used and lactose can be cleaved
Lactose (allolactose) = inducer; when lactose is present, some of it is converted into allolactose, which then binds to the regulatory protein (LacR repressor), and induces a conformational change in the regulatory protein
As a result of this conformational change, the LacR repressor can no longer bind to the operator region, which allows RNA polymerase to transcribe the structural genes
Repressible operon
operons that are normally transcribed (default = ON) because the repressor is inactive by default
Co-repressors activate a repressor so that it is functional, and can bind at the operator site to block transcription
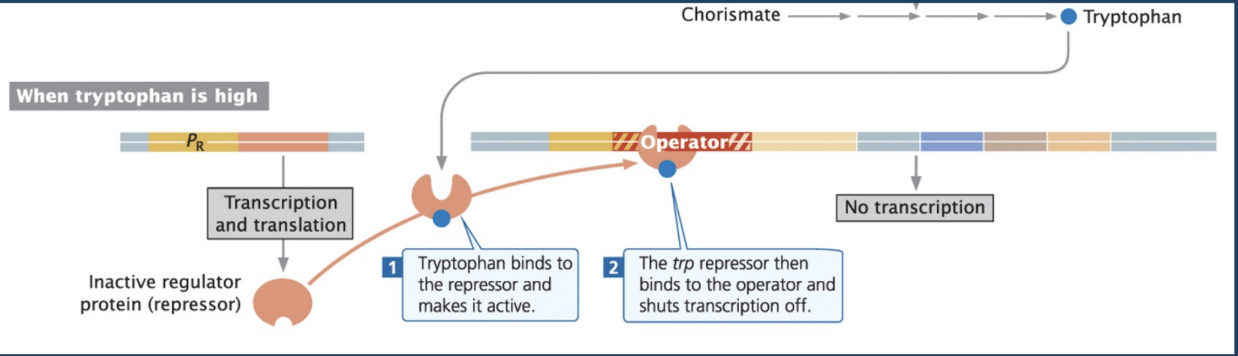
Example of repressible operon
Ex. trp operon = repressible operon
Trp operon is normally active (transcribed)
Tryptophan = Co-repressor (activates repressor)
When there is excess tryptophan in the environment, or the cell produces too much tryptophan, the tryptophan acts as a co-repressor by activating the inactive repressor protein
The trp repressor then binds to the operator and shuts transcription off
INDUCIBLE AND REPRESSIBLE OPERONS ARE BOTH EXAMPLES OF NEGATIVE CONTROL, BECAUSE A REPRESSOR PROTEIN IS BEING USED TO REGULATE TRANSCRIPTION.
INDUCIBLE AND REPRESSIBLE OPERONS ARE BOTH EXAMPLES OF NEGATIVE CONTROL, BECAUSE A REPRESSOR PROTEIN IS BEING USED TO REGULATE TRANSCRIPTION.
Negative control
inhibitors/repressors that block transcription by interfering with RNA polymerase
Example of positive control
Ex. CAP + cAMP system = positive control
The CAP–cAMP system is a mechanism that enhances transcription of the lac operon when glucose levels are low.
Why? Because cells prefer glucose as it is less costly to use, whereas lactose needs to be modified
Low glucose = high cAMP because importing glucose is what consumes cAMP
High cAMP means that CAP, the activator protein, binds to cAMP. The CAP-cAMP complex binds near the promoter and helps RNA polymerase bind more effectively by bending the DNA - making promoter easier to access = boost transcription of the lac operon = lac operon is transcribed when glucose levels are low
Lots of glucose = less cAMP (b/c costs energy to transport it inside) = CAP doesn’t work, no cAMP to bind = lac operon transcription decreases
If only glucose is high, cells will metabolize glucose
If only lactose is high, cells will metabolize lactose
If glucose and lactose are both high, cells will metabolize glucose preferentially
High glucose makes low cAMP, so CAP is disabled
Attenuation
A mechanism that prematurely stops transcription
a backup system that stops transcription EARLY after it starts to STOP leakage
Leakage
small amounts of RNA transcript are still made in presence of repressor/when repressor briefly falls off DNA (repressor is dynamic) - producing unwanted mRNA when gene is not needed (ex. when trytophan levels are high)
Example of attenuation
trp operon - trp mRNA transcript contains a leader sequence with 4 different regions (1-4); Recruitment of ribosome initiates leader peptide translation
Before the actual trp genes, the mRNA has a short “leader region”
It does NOT code for the main proteins
It contains 4 segments (regions 1–4)
These regions can fold into different hairpin structures
These regions can pair in different ways:
2 + 3 → anti-terminator (transcription continues)
3 + 4 → terminator (transcription stops)
➡ This is how the cell decides ON vs OFF
“Recruitment of ribosome”
👉 In bacteria:
translation starts while transcription is still happening
So:
Ribosome binds the leader sequence
Starts translating a short leader peptide
The ribosome’s position determines which regions can pair
🔴 High tryptophan = bacteria want to repress trp operon
Ribosome moves quickly
Covers region 2
Region 3 + 4 pair (hydrogen bond together), forming a strong terminator hairpin that STOPS transcription
RNA polymerase falls off RNA
TRANSCRIPTION WAS STARTED BUT STOPS EARLY BEFORE FULL GENES ARE MADE = ATTENUATION
🟢 Low tryptophan = bacteria want to transcribe trp operon
Ribosome stalls at Trp codons
Region 2 + 3 pair (hydrogen bond) =
➡ weak hairpin called anti-terminator forms → transcription continues, but TRANSLATION is halted
Key idea
👉 The ribosome acts like a sensor for tryptophan levels
Eukaryotic Transcriptional Regulation - Recognize examples of gene regulation by histone modification or DNA methylation and determine if changes will promote or inhibit gene transcription.
DNA is wrapped around histones → like thread around spools
If DNA is tight (heterochromatin) → inaccessible → OFF
If DNA is loose (euchromatin) → accessible → ON
TWO MAJOR TYPES - HISTONE MODIFICATION (methylation/acetylation), and then DNA MODIFICATION
Histone modification
Methylation - inhibits transcription
Histone methyl transferase (HMT) adds methyl groups
Histone demethylase (HDM) removes methyl groups
Acetylation - promotes transcription
Histone acetyl transferase (HAT) adds acetyl groups
Histone deacetylase (HDAC) removes acetyl groups
2. Direct DNA modification- Directly on DNA. Methylation of cytosine bases = transcription inhibition.
CpG islands - areas that get heavily methylated near promoters
Direct DNA modification is associated with LONG TERM INHIBITION (Barr Bodies, heterochromaitn)
Why? Because Methyl groups are added to cytosines (CpG sites). During DNA replication: enzymes copy the methylation pattern. So the “OFF signal” is passed to daughter cells
Ex. Barr Bodies
In females: one X chromosome is silenced. How?
heavy DNA methylation
histone modifications
Becomes a barr body (condensed, inactive X chromosome)
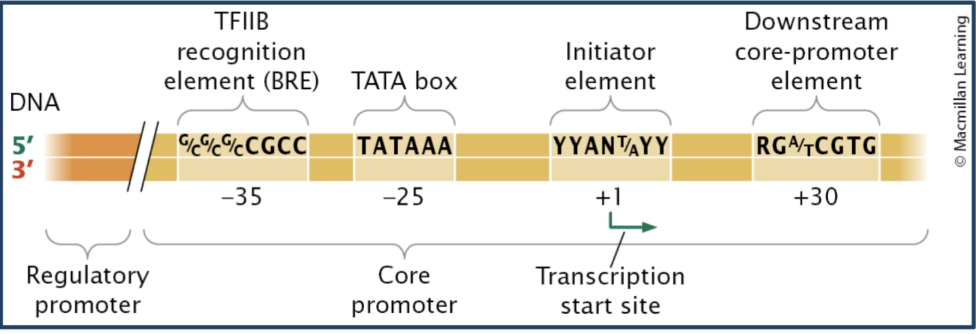
Identify important sequence components of the eukaryotic core promoter.
Eukaryotic core promoter- landing pad for transcription machinery, has “motifs” for basal transcription factor binding (Basal transcription factors ON THE EUKARYOTIC CORE PROMOTER recruit RNA pol)
Initiator / Inr (+1 site) - start site
TATA box (-25bp) - helps position RNA polymerase
TFIIB recognition element / BRE (-35bp) - binds TFIIB
Downstream promoter element (+30bp) - downstream binding site
This region recruits basal transcription factors and determines where transcription begins
Exam examples
“Where does RNA pol bind?” → core promoter
“Mutation in TATA box → ?” → ↓ transcription
Proximal control elements
Short DNA sequences close to promoter that bind specific transcription factors (activators and repressors); they fine tune transcription (Multiple TFs = combinatorial control)
Proximal control elements are part of the regulatory promoter - extended upstream region of the core promoter that regulates transcription and includes proximal control elements
In other words:
Regulatory promoter = region
Proximal control elements = specific sequences within that region
Enhancers
positive elements that act from a distance (upstream or downstream) to promote transcription (increase rate of transcription), works regardless of orientation
Enhancers bind to activator transcription factors, and then the activator binds to the mediator complex. The mediator complex binds to the transcription initiation complex.
Activator transcription factors bind enhancers and interact with the mediator complex, which bridges them to RNA polymerase II at the promoter (transcription initiation complex) through DNA looping
Review of transcription factors
A protein that binds DNA and regulates transcription
Activators → increase transcription
Repressors → decrease transcription
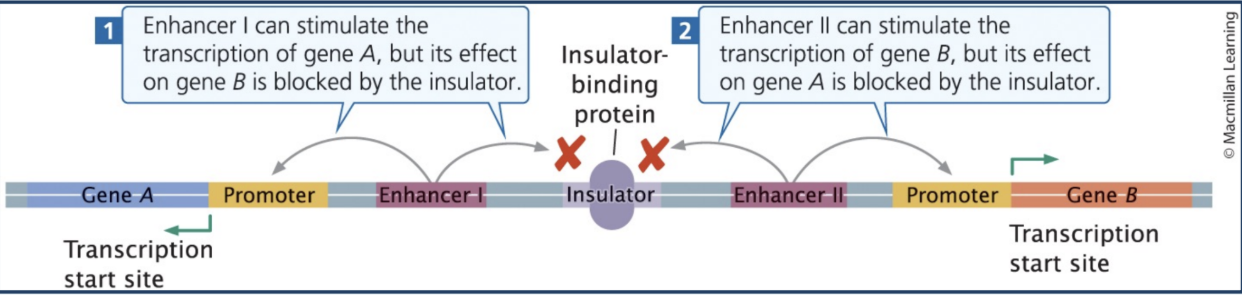
Insulators
Elements that limit enhancer-promoter interactions to a "regulatory neighborhood"
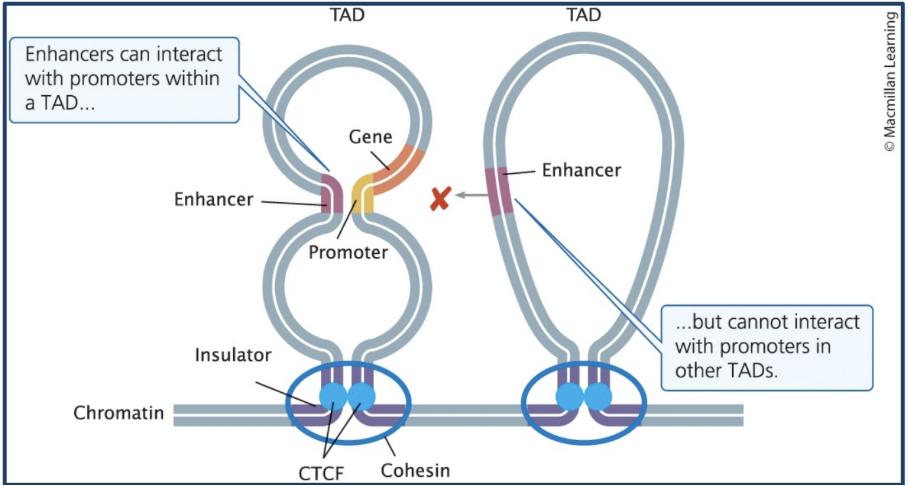
Topologically associated domains (TADs)
Transcriptionally active loops of DNA
Enhancers can interact with genes in the same TAD (elements within a loop (TAD) can interact)
Transcriptional stalling
Transcription starts but RNA pol stalls 20-50 nucleotides downstream
Regulated process
NELF (negative elongation factor) causes stalling
Phosphorylation of NELF by P-TEFb relieves stalling
Possibly allows “pre-loading” of RNA pol for rapid transcription
Why? because the gene is already “primed” - so if the cell needs the gene, polymerase can quickly continue
RNA Regulation
control of gene expression by RNA molecules
Riboswitch
Anti-sense RNAs (sRNA)
CRISPR
Micro RNAs
Discuss the mechanism by which riboswitches regulate translation of prokaryotic mRNA.
Riboswitch - Prokaryotic RNA stem-loop (hairpin) in the 5′ UTR of mRNA that regulate translation by changing shape when a small molecule binds
They modify translation efficiency by blocking the Shine-Dalgarno sequence (location where prokaryotic ribosomes normally bind) in a negative feedback regulatory mechanism
The hairpin isn’t super strong, so an effector molecule binds to the riboswitch to stabilize its structure, making it a negative feedback regulatory mechanism
The stabilization of the riboswitch’s structure allows it to mask the ribosome-binding site = no translation takes place
Without the enhancer (regulatory protein), the riboswitch assumes an alternative secondary structure, that makes the ribosome-binding site available = translation takes place
Describe the function of the CRISPR system in the degradation specific DNAs in prokaryotic cells.
CRISPR system - Defense system in bacteria (prokaryotes) against foreign DNA; allows bacteria to recognize and destroy specific foreign DNA (like viruses).
CRISPR (RNA) - RNA molecule that guides Cas enzymes (like Cas9) to cut specific, complementary viral (foreign) DNA
Mechanism of CRISPR system:
Foreign DNA enters cell
Small fragment of foreign DNA is inserted into CRISPR array (so the bacteria remembers that same foreign DNA)
CRISPR RNA (crRNA) is made
crRNAs combine with Cas proteins (enzymes) to cleave foreign DNA
Illustrate the process of RNAi, showing the function of Dicer and the RISC complex.
RNA interference (RNAi) = eukaryotic defense system that uses small RNAs to silence genes; small RNAs find/bind to a matching FOREIGN messenger RNA (mRNA) and stop it from making protein.
There are two main types of small RNAs:
siRNA → defense (viruses, transposons)
miRNA → regulation (your own genes)
Step 1: FOREIGN Double-stranded RNA (dsRNA) appears in the cell. This can come from: viruses, transposons (jumping DNA), or normal cellular processes (like hairpin RNA). The cell recognizes this as something that needs to be controlled.
Step 2: Dicer cuts the RNA
An enzyme called Dicer cuts this long double-stranded RNA into short pieces.
These short pieces are called: siRNA (if from foreign RNA) and miRNA (if from the cell’s own RNA)
So, Dicer’s job is to produce the small RNA molecules used in RNAi.
Step 3: The RISC complex forms
siRNA combines with Argonaute (Ago) proteins to form a RISC complex
These small RNA pieces are loaded into a protein complex called RISC (RNA-induced silencing complex).
RISC contains proteins called Argonaute (Ago).
Inside RISC:
one strand of the RNA is removed
the remaining strand is kept as a guide
Step 4: The guide RNA finds its target
The guide RNA inside RISC searches for an mRNA with a complementary sequence.
It binds to that mRNA based on base pairing.
The mRNA is silenced
What happens next depends on the type of RNA:
If it is siRNA (perfect match):
The RISC complex cuts the mRNA, and the mRNA is destroyed ➡ No protein can be made. It can also:
recruit enzymes that modify chromatin (like histone methyltransferases)
cause DNA to become tightly packed
If it is miRNA (imperfect match):
The RISC complex: blocks translation or causes the mRNA to be degraded slowly ➡ Protein production is reduced or stopped.
Define miRNA and siRNA
miRNA and siRNA = two types of small RNAs used in the eukaryotic CRISPR system to silence gene expression by targeting specific mRNA molecules
Dicer: An enzyme that cuts long double stranded RNA into siRNA or microRNA (miRNA).
RISC Complex (containing Argonaute): A complex that uses a single-stranded guide RNA to identify, bind, and cleave complementary mRNA, preventing protein production.
RNAi is the whole system
Dicer makes the small RNAs
RISC uses those RNAs to find targets
The target mRNA is destroyed or blocked
miRNAs and their role in RNA interference.
MicroRNAs (miRNAs) are small RNA molecules produced from the cell’s own genes, which form hairpin structures and are made by the enzyme Dicer that cuts the hairpin RNA into a short, single-stranded RNA (they are complementary to sequences on other genes). miRNA joins Argonaute protein, and together they form RISC (RNA-induced silencing complex). Once RISC finds matching mRNA: Ribosome can’t translate properly → less protein is made
However, miRNAs bind imperfectly to target mRNA, SO THEY DO NOT CUT THE MRNA. INSTEAD, they usually block translation or promote gradual degradation rather than cutting the mRNA. Their main role is regulation of a cell’s own genes (gene expression).
miRNAs have an imperfect sequence match, and when binding to near the 5'-end do not trigger mRNA cleavage.
miRNAs binding near the 3'-end of an mRNA may trigger "Slicer independent" mRNA by recruiting de-capping and de-tailing enzymes.
If miRNA binds near the 3′ end (poly-A tail region)
It recruits enzymes that:
remove the 5′ cap
remove the poly-A tail
Why this matters
mRNA needs: 5′ cap and poly-A tail to stay stable
Removing them makes mRNA: unstable and degraded
miRNAs are made by the cell to regulate its own genes → They reduce protein production (they usually don’t completely destroy mRNA)
siRNAs and their role in RNA interference.
siRNAs - Eukaryotic Small interfering RNAs (siRNAs) are short double-stranded RNA molecules that usually come from foreign double-stranded RNA, such as viruses or transposons. They are made by Dicer cleaving double-stranded RNA in the cytoplasm from retroviruses or transposons and are incorporated into the RISC complex by combining with Ago proteins. Because siRNAs bind perfectly to their target mRNA, they cause direct cleavage and degradation of the mRNA, leading to complete gene silencing. Their main role is defense. The RISC complex cleaves the complementary mRNA made by viral or transposon DNA. siRNA molecules can also initiate transcription silencing of specific genes by recruiting histone methyl transferase enzymes to nucleosomes
Epigenetic regulation
HERITABLE changes in gene expression without altering the DNA sequence, often through DNA methylation and histone modifications.
roles of DNA methylation and histone modifications in the process
DNA methylation - Add methyl groups to cytosine (in CpG pairs, aka CpG islands near promoters)
Block transcription
CpG pair = one methylation site/DNA sequence: 5’ - C - G - 3’ (cytosine followed by guanine) p = phosphate backbone connection
CpG island = many CpG sites clustered together near gene promoters
Histone modifications - Change chromatin structure
Acetylation → open → ON
Methylation → closed → OFF
control DNA accessibility
Describe how epigenetic changes in cells can influence differentiation, metabolism, and behavior.
How epigenetic changes influence:
Differentiation
All cells have same DNA BUT different genes ON/OFF
epigenetics locks in cell identity by turning off genes
Example: neuron vs muscle cell
Metabolism
Genes for enzymes can be turned ON/OFF
Ex. DNA methylation can reduce enzyme production, slowing metabolic pathway
Behavior
Mother mouse licks its pups a lot —> triggers DNA demethylation of the glucicorticoid receptor (GR) gene; gene becomes more accessible, so there is greater GR expression and pups are able to respond to stress better
Discuss the phenomenon of paramutation through epigenetic changes to DNA.
Paramutation - epigenetic interaction between two alleles where one allele can permanently silence another allele epigenetically without changing the allele’s DNA sequence
The silenced state can be inherited and passed to future generations
You have two alleles:
One is normal (active)
One is silenced (epigenetically modified)
The silenced allele converts the normal allele into a silenced state. The newly silenced allele can now silence other alleles in future generations through mechanisms such as DNA methylation and chromatin modification, without altering the DNA sequence.
Example of paramutation
Example of paramutation
Maize (corn) kernel color is determined by the pigment gene
R allele → purple kernels (active gene)
r* allele → silenced version (lighter/colorless)
If you cross: Active allele (R) x Silenced allele (r*), you unexpectedly get kernels that are lighter / colorless, because r* silences the active allele R
The R allele becomes silenced
Now BOTH alleles behave like r*
The newly silenced allele can now silence OTHER active alleles in future generations (this effect is heritable)
How?
“Silenced” allele produces regulatory RNA”
The paramutagenic allele (silenced one) makes small RNAs (siRNA-like)
These RNAs target the active allele - They bind to the active allele’s DNA or RNA and guide silencing machinery to it
Epigenetic changes are added
DNA methylation
histone methylation
chromatin becomes closed (heterochromatin) and gene is silenced
New allele becomes “paramutated”
The previously active allele is now: silenced and able to silence others
➡ effect is heritable
Use the X-inactivation process as an example to illustrate RNA-mediated epigenetic regulation.
Dosage compensation - epigenetic X chromosome inactivation
In females, one X chromosome is turned OFF
The Xist RNA (a long non-coding RNA (lncRNA)) binds X chromosome and recruits DNA methylation and histone modification (methylate the X chromosome, so the X chromosome becomes heterochromatin)
The X chromosome is now silenced as a Barr Body and does not code for protein