Biology Unit 3 Test
1/50
Earn XP
Description and Tags
Biochemistry (metabolism, respiration, photosynthesis, lipid metabolism)
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
51 Terms
Laws of Thermodynamics
Conservation of energy, not created or destroyed but transferred
Energy transfers increase randomness (∆S)
∆G= …
∆H-T∆S
-∆G
exergonic
spontaneous
favorable
ex: ice melting
+∆G
endergonic
nonspontaneous
unfavorable
ex: climbing up
-∆H
exothermic
net loss in release
ex: burn CH4
+∆H
endothermic
net gain in consumption
ex: ice in water
-∆S
> organized
ex: synthesis
+∆S
> randomness
ex: break down
ATP and Energy Coupling
ATP hydrolysis is thermodynamically favored so it is paired with endergonic reactions to drive those reactions forward
ATP+ H2O —> ADP + Pi (∆G = -7.3 kcal/mol)
ex: Glucose + ATP → Glucose-P + ADP (-7.3 kcal/mol) →Glucose-P + Fructose →Sucrose + Pi. (+ 6.5 Kcal/mol)
Factors affecting enzymes
temperature (affects 3-D shape)
pH (affects 3-D shape)
salt concentration (affects 3-D shape)
cofactors (activates enzymes)
Substrate concentration (enzyme needs right substrate for rxn)
Vmax
highest velocity of the enzyme (maximum rate of reaction). Achieved when all the enzyme active sites are filled with the substrate
Km
is the substrate concentration at which the rate of reaction is half its maximum. indicates the [S] at which half the enzyme active sites are filled with substrates
Regulation of enzyme activity
Activators (+) and inhibitors (-)
Allosteric regulation
Feedback regulation
Chemical modification (chemically modified by themselves or by other enzymes to make them active or inactive)
Activators
bind to the enzyme, changing its conformation with a positive effect on its activity
Inhibitors
binds to the enzyme and changing their conformation, resulting in a reduced enzyme activity
Competitive inhibitors
compete with the substrate for the same active site on the enzyme. In the case of a competitive inhibitor, a higher concentration of the substrate can be added to overcome the competition
Km increases
Non-competitive inhibitor
binds to the protein at some place other than the active site, change the conformation of an enzyme and make it less active or inactive. Adding more of the substrate does not overcome the inhibition.
Vmax decreases
Uncompetitive inhibitor
Bind to the active site after the substrate binds to the enzyme
lowers Vmax and increases Km
Allosteric regulation
complex enzymes with separate catalytic (binding to substrate) and regulatory (binding to activator or inhibitor) subunits
An activator or inhibitor binds to the enzyme and changes its conformation to an active or an inactive form, respectively
They respond to the substrate concentration in a sigmoid (s) fashion.
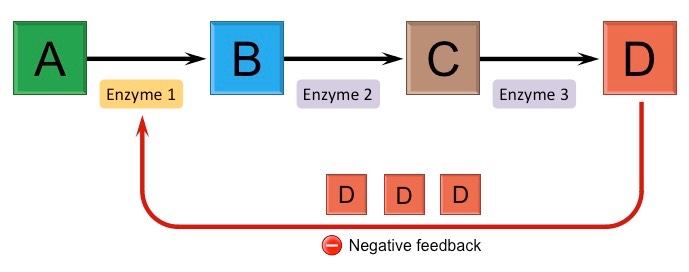
feedback regulation
The end product of a biosynthetic pathway or an intermediate of another but related pathway inhibits an earlier enzyme and stops the whole pathway
Keq= [products]/[reactants] when…
∆G = 0
what are examples of exergonic reactions?
respiration and passive transport
what are examples of endergonic reactions?
photosynthesis and active transport
glycolysis: place, inputs, outputs, key steps, key enzymes, # of ATP made (SLP or OP)
cytoplasm
Glucose, ADP pi, and NAD+
2 Pyruvate, 2 ATP, 2 NADH
Steps 1 (G+ATP→GP+ADP) and 3(F6P+ATP→FI6BP+ADP)
Hexokinase (step 1) and Phosphofructokinase (step 3)
2 ATP made by substrate level phosphorylation
Summary of glycolysis
Net inputs: Glucose, ADP pi, and NAD+
comes from: food, ATP hydrolysis, oxidative phosphorylation
net outputs: pyruvate, ATP, NADH
Go to: acetyl CoA formation, used in cytoplasm, goes to oxidative Phosphorylation via Electron shuttle
Acetyl CoA formation: place, inputs, outputs, key steps, key enzymes, # of ATP made (SLP or OP)
Mitochondrial membrane
CoA, 2 NAD+, 2 Pyruvate
2 NADH, 2 Acetyl CoA, 2 CO2
none
pyruvate dehydrogenase
none
what happens when there is too much Acetyl CoA?
Any excess acetyl CoA formed from the glycolysis of carbohydrates are converted to fats for storage
Krebs Cycle: place, inputs, outputs, key steps, key enzymes, # of ATP made (SLP or OP)
Mitochondrial Matrix
6 NAD+, 2 FAD, 2ADP 2pi, 2Acetyl CoA
6 NADH, 2 FADH, 2 ATP, 4CO2, 2 CoA
1 (citrate synthesis) and 3 (Oxidative decarboxylation of isocitrate)
Citrate synthase (step 1) and Isocitrate dehydrogenase (step 3)
2 ATP substrate level phosphorylation
Oxidative phosphorylation: place, inputs, outputs, key steps, respiratory poisons , # of ATP made (SLP or OP)
Mitochondrial matrix
NADH, FADH2, ADP pi, O2
NAD+, FAD, ATP, H2O
none
CO cyanide, oligomycin, DNP
32-34 ATP by oxidative phosphorylation
Alcohol Fermentation: place, inputs, outputs, # of ATP made (SLP or OP)
cytoplasm
Glucose and ADP Pi
2 ATP, 2 CO2, 2Ethanol
1 and 3
Hexokinase and phosphofructokinase
2 ATP by substrate level phosphorylation
Acetate fermentation: place, inputs, outputs, # of ATP made (SLP or OP)
cytoplasm
glucose, 2ADP pi
2 lactate, 2 ATP
1 and 3
Hexokinase and phosphofructokinase
2 by substrate level phosphorylation
phloem
transport sugars to other parts of the plant for use or storage
Xylem
transport water and minerals
Photophosphorylation
Electron transfer through a series of thylakoid membrane proteins results in the generation of a proton gradient inside the thylakoid, which becomes acidic.
When protons return to the stroma through ATP synthase, they help in the synthesis of ATP, as in the chemiosmosis of respiration.
cyclic light reaction
Involves only PS I
Electrons return to PS I reaction center
ATP synthesized in PS I
No NADPH made
No O2 is evolved
Electrons are recycled
Occurs when NADPH level is high
noncyclic light reaction
Use both PS I and PS II
Electrons do not return to reaction center
ATP synthesized in PS II
Electrons are used to reduce NADP+ to generate NADPH
O2 is evolved from the splitting of water
The ultimate electron source is H2O
Most common in plants
key enzyme in Calvin cycle:
ribulose bisphosphate carboxylase/oxygenase (rubisco)
RuBisCo is regulated by…
CO2 and O2 concentration
NADPH concentration
Mg++ cofactor
pH (high in stoma and low in thylakoid)
Net inputs and outputs of Calvin Cycle
inputs: CO2, ATP, NADPH
outputs: CH2O (C6H12O6), ADP pi, NADP+
C3 plants: where does Calvin cycle occur, C4 pathway, examples
mesophyll
none
rice and wheat
C4 plants: where does Calvin cycle occur, C4 pathway, examples
bundle sheath
mesophyll
corn and sugarcane
CAM plants: where does Calvin cycle occur, C4 pathway, examples
day time
night time
cacti and pineapple
photosynthesis: place, inputs, outputs
chloroplasts
6CO2 + 6H2O
C6H12O6 + 6O2
cyclic light reactions: place, inputs, outputs
thylakoid
light, ADP pi
ATP
noncyclic light reactions: place, inputs, outputs
Thylakoid
light, H2O, NADP+, ADP pi
O2, NADPH, ATP
Calvin cycle: place, inputs, outputs
mesophyll (c3 plants) or bundle sheaths (c4 plants)
CO2, ATP, NADPH
CH2O, ADP pi, NADP+
C4 pathway: place, inputs, outputs
mesophyll
CO2, ATP, NADPH
CH2O, ADP pi, NADP+
Fats: composition, precursor, key enzyme, break down
glycerol and fatty acids
acetyl CoA
acetyl CoA carboxylase
beta oxidation
Steroids: composition, precursor, key enzyme, break down
4 fused rings
acetyl CoA
HMG CoA and HMG CoA reductase
Receptor mediated endocytosis
phospholipids: composition, precursor, key enzyme, break down
glycerol, fatty acids, and phosphates
G3-P, Acetyl CoA
Acetyl CoA carboxylase and phosphatidic acid
beta oxidation
carotenoids: composition, precursor, key enzyme, break down
hydrocarbons (40-c) and plastids
isoprene unites
NA
other pigments