bio 303 exam 3
1/101
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
102 Terms
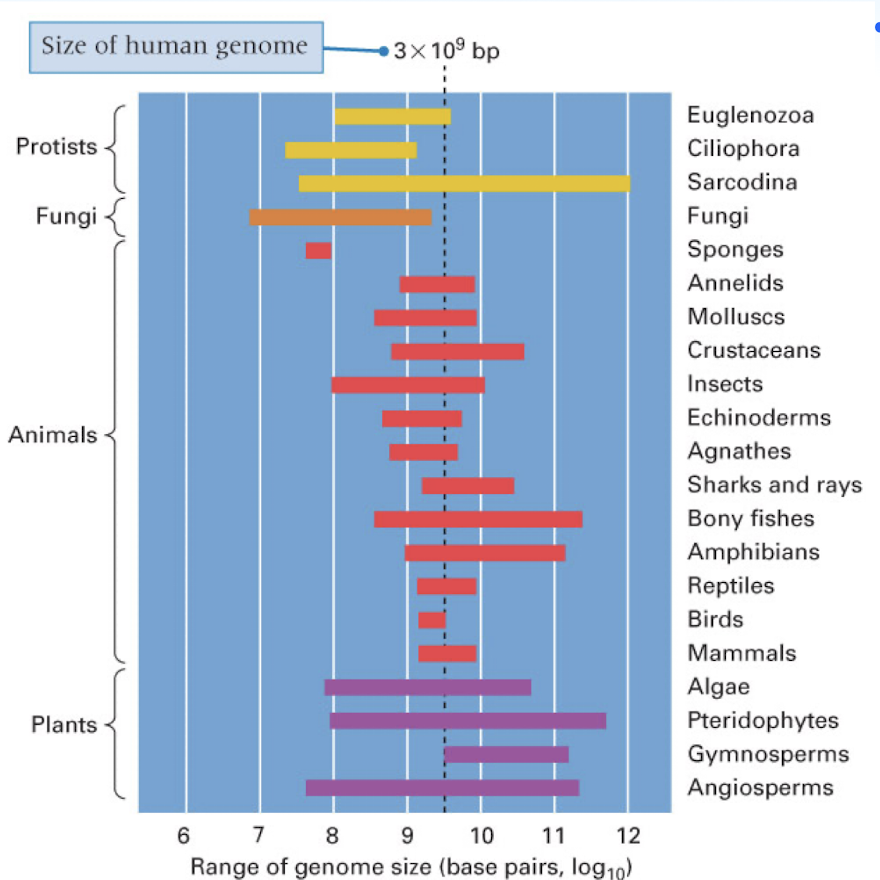
C Value paradox
the complexity of an organism is not related to its genome size (amount of DNA it contains or its “C Value”
genome size can range greatly in some groups of organism
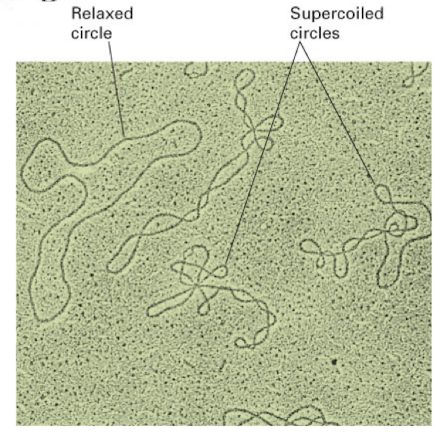
supercoiled DNA
DNA twisted beyond the typical double helix shape
does this to save space, relieve stress/pressure from the tight winding of DNA, and allows DNA to be opened up
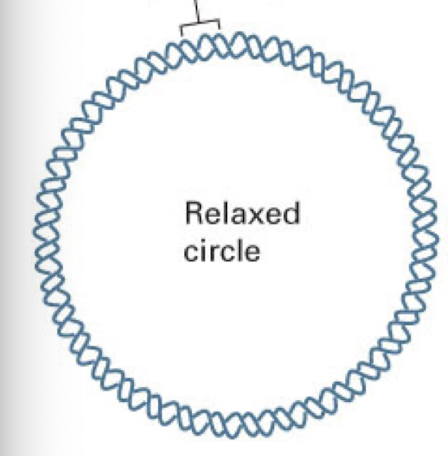
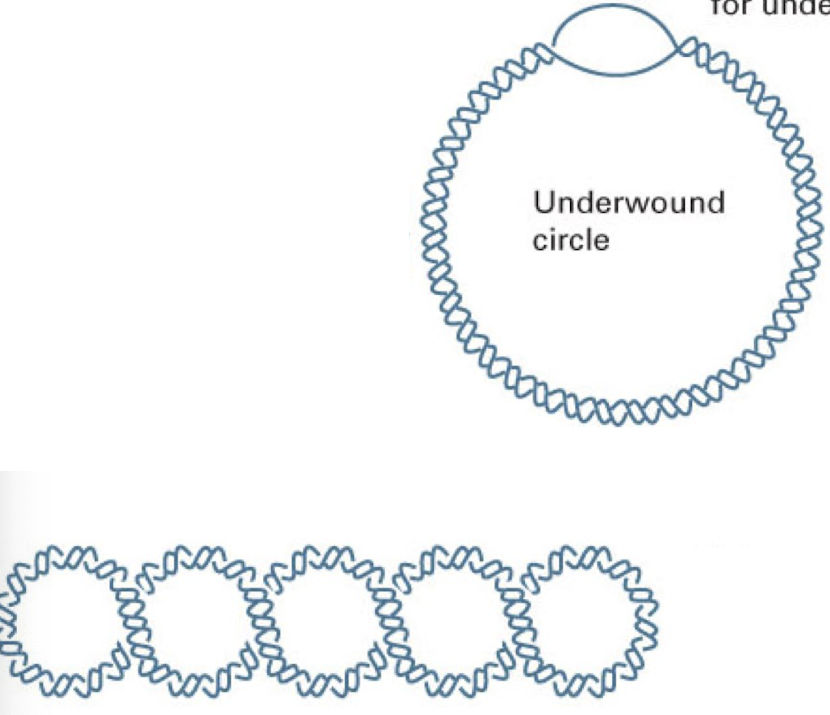
relaxed DNA circle
typical DNA structure with 10 base pairs per complete turn

underwound DNA circle
contain a bubble (or bubbles) on unpaired bases that compensate for underwinding
negatively supercoiled DNA- easier to open up DNA strand for replication, meiosis, mitosis, etc
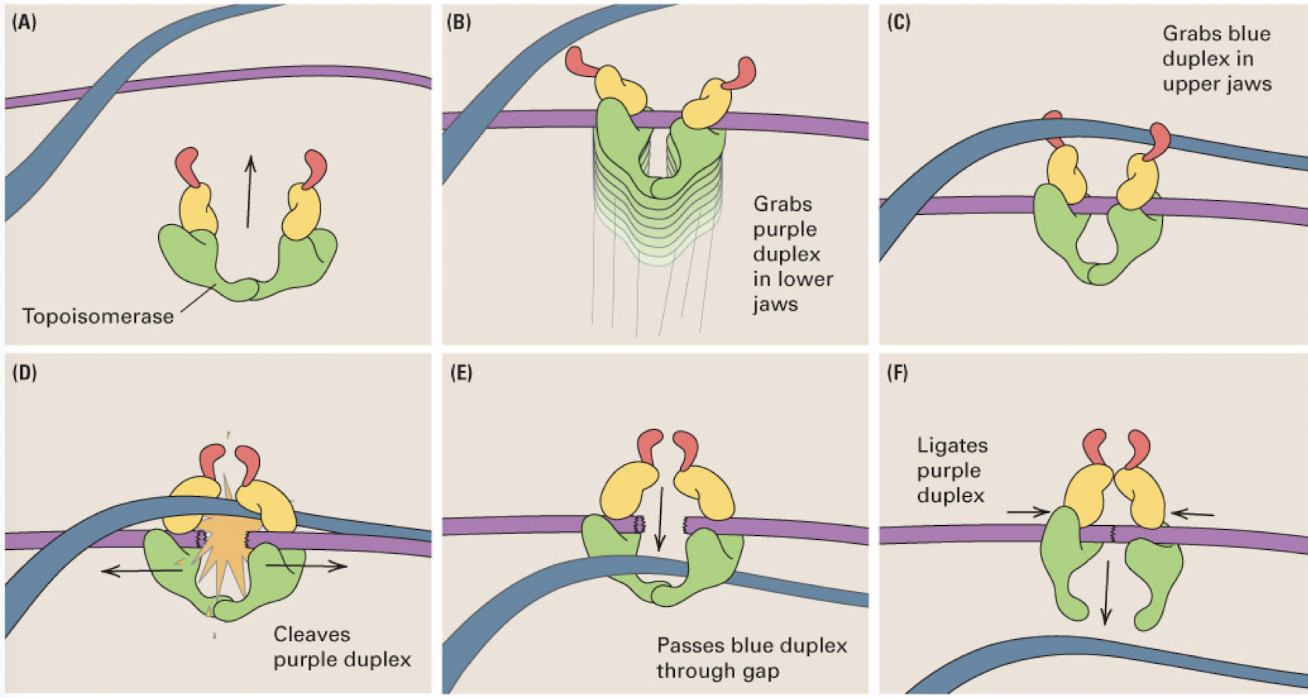
Topoisomerase II
enzyme that untangles a pair of DNA molecules by cleaving one DNA duplex and passing the other duplex through the gap
upper jaw grabs one duplex (not cleaved) and the lower jaw grabs the second duplex (which it cleaves) and passes the first through the split second DNA molecule
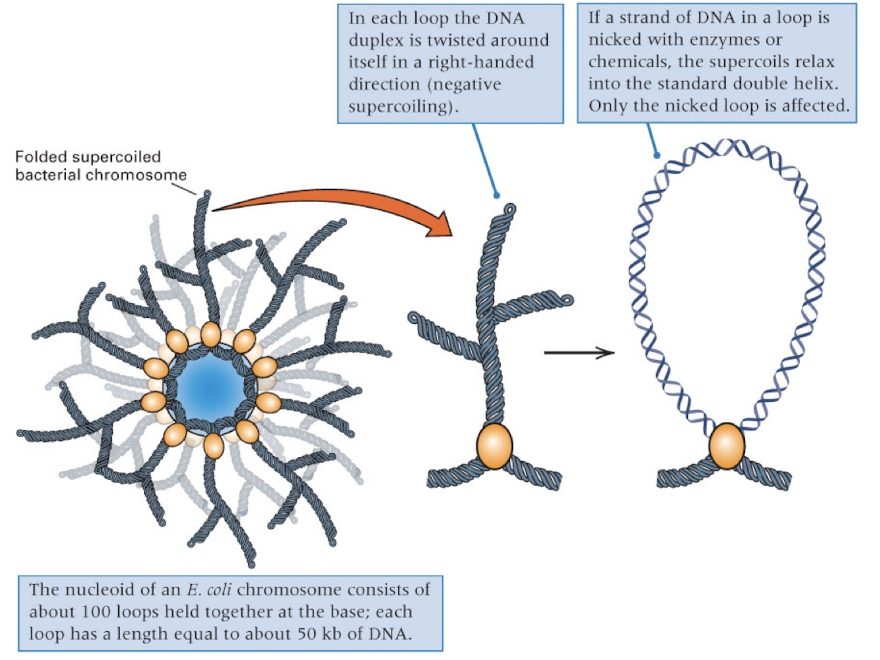
bacterial chromosomes are organized into…
supercoiled loops
each loop can be independently relaxed or condensed
four proteins (HU- wrapping, FIS and IHF- bending, HNS- compaction) involved in supercoiling into a higher order structure
EX: each loop of the DNA duplex is twisted around itself in a right handed direction- results in negative supercoiling
supercoiling can only be relaxed when the loop is nicked (and only the nicked loop will be affected)
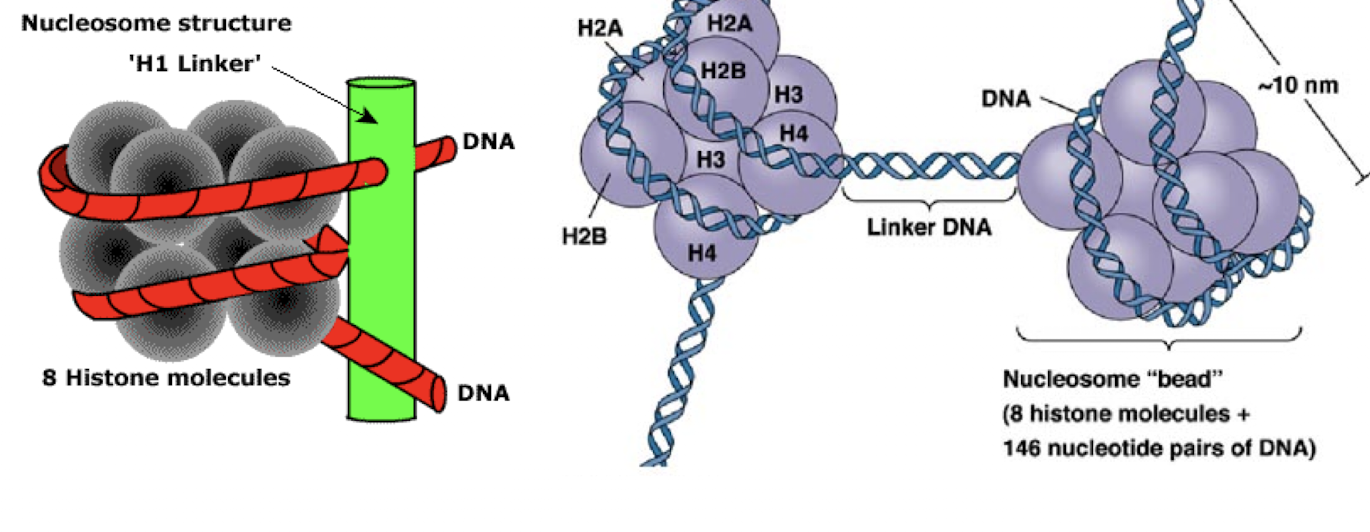
eukaryotic DNA is organized into nucleosomes
each nucleosome consists of two of each H2A, H2B, H3, and H4 histone proteins, a segment of DNA containing about 200 nt, and one molecule of H1
molecule of H1 is a linker- outside of 8 histone molecule complex and connects DNA wrapped around the histone proteins
histone proteins
small proteins (100-200 AA)
five major types: H1, H2A, H2B, H3, H4
20-30% of AA are lysine + arginine- positively charged (**DNA neg charged)
both DNA-histone and histone-histone binding are important for chromatin structure (bc they make up nucleosome structure)
sequence of histones are conserved during evolution
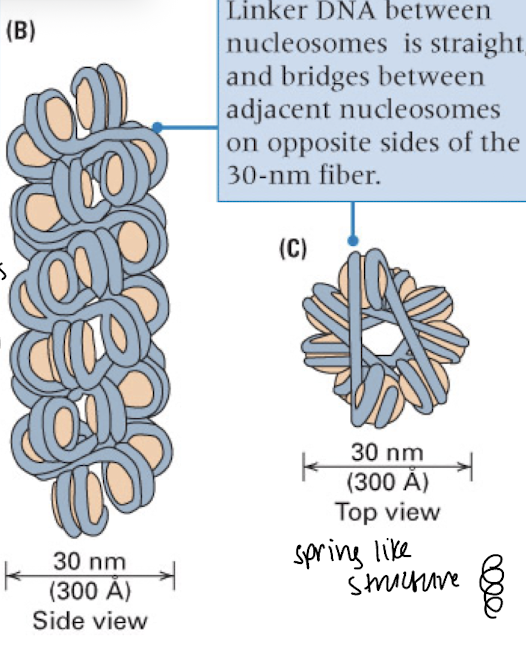
nucleosomes compact into…
chromatin fibers- spring-like/coiled structure
bead line structure of nucleosomes only seen in artificial conditions, typically it is compact into chromosome fibers in the nucleus
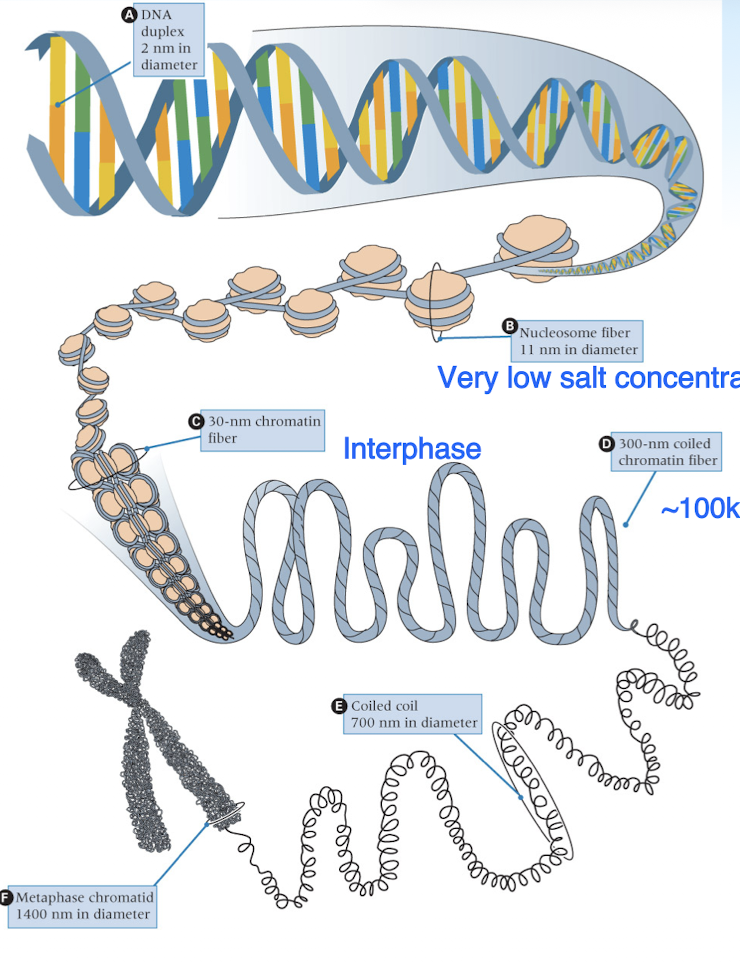
stages of DNA condensation
DNA duplex —> nucleosomes (histone proteins wrapped with DNA) —> chromatin fibers (DNA form during interphase) —> chromosomes (DNA form for meiosis/mitosis- higher order)
chromosome territories
less dense chromosome domains/contain fewer genes- outskirts of the nucleus or close to the nucleolus (very center of the nucleus)
more dense chromosome domains/contain more genes- further inside the nucleus (but between the nucleolus and the outskirts of the nucleus)
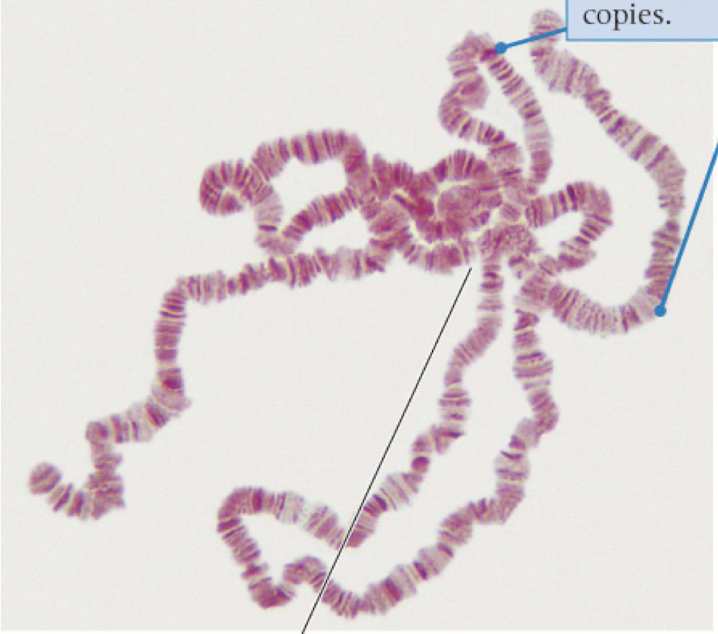
polytene chromosomes
highly repeated DNA sequences that never separate
produce large amounts of RNA and proteins quickly
nucleoid
bacteria cells (no nucleus + one circular chromosome as DNA)
DNA forms a multiply looped structure- supercoiled and compact
eukaryotic DNA consists of three major components
30-75%: unique sequences
one copy of each unique sequence
5-45%: highly repetitive sequences
as many as 10^5 copies per genome
1-30%: middle repetitive sequences
10-1000 copies per genome
some parts of DNA condense more/less than others because they are different sections of different complexity
repetitive nucleic sequences in eukaryotic genomes
some components have a base composition that is different from the average
satellite DNA: components with unusually low/high G+C contents, fairly short nucleotide sequences that may be tandemly repeated up to a million times
repeats are separated from other DNA because their density is different
renature more readily than nonrepetitive DNA because they consist of high similar or identical sequences
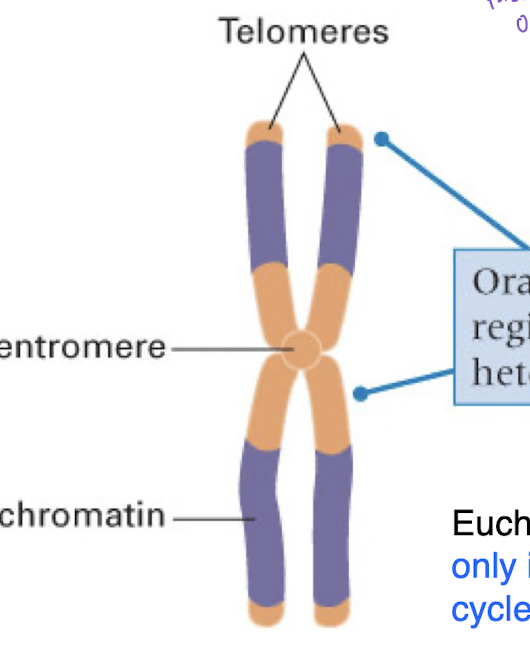
heterochromatin
highly repetitive and low complexity
condense faster because of the high amount of repeats
located on the middle of chromosomes near centromere and on the ends where telomeres are (orange section)
molecular structure of the centromere
specific region of the eukaaryotic chromosome that becomes visible as a narrow constriction along the condensed chromosome
holocentric chromosome
very large
have centromeric sequences spread throughout their length
also called a diffuse centromere
localized centromere
conventional type
microtubules attach to a single region on the chromosome- kinetochore
point centromeres: found in different yeasts, more repetitive
regional centromeres: found in other eukaryotes, may contain hundreds of kilobases or repetitive DNA
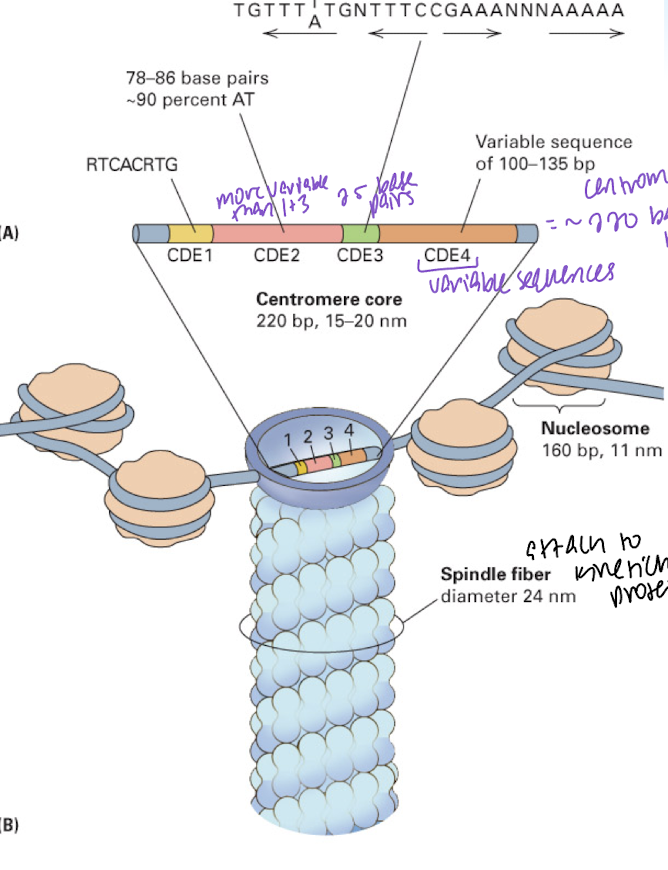
point centromere
yeast centromere
centromere DNA has four centromere determining elements (CDEs)
CDEs: CDE1, CDE2 (more variable than 1 and 3), CDE3 (~25 base pairs), CDE4 (variable sequences) = ~220 base pairs total (centromere)
spindle fibers connect t kinetichore proteins
telomere
special DNA protein structure located at each end of a linear chromosome
consists of tandem repeats of simple sequences and associated rpoteins
essential for chromosome stability
this region cannot be synthesized in DNA replication
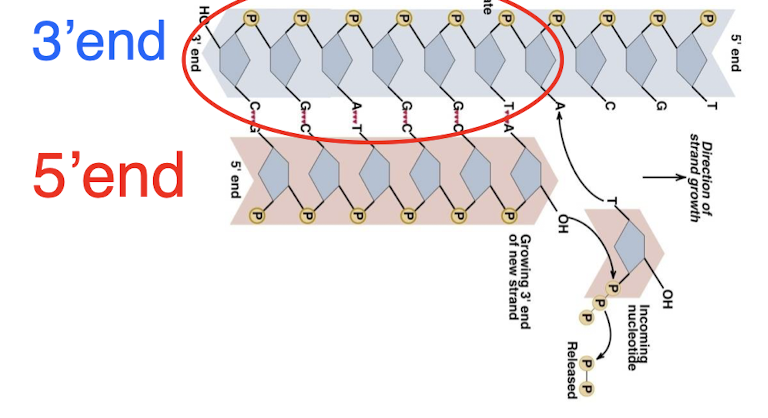
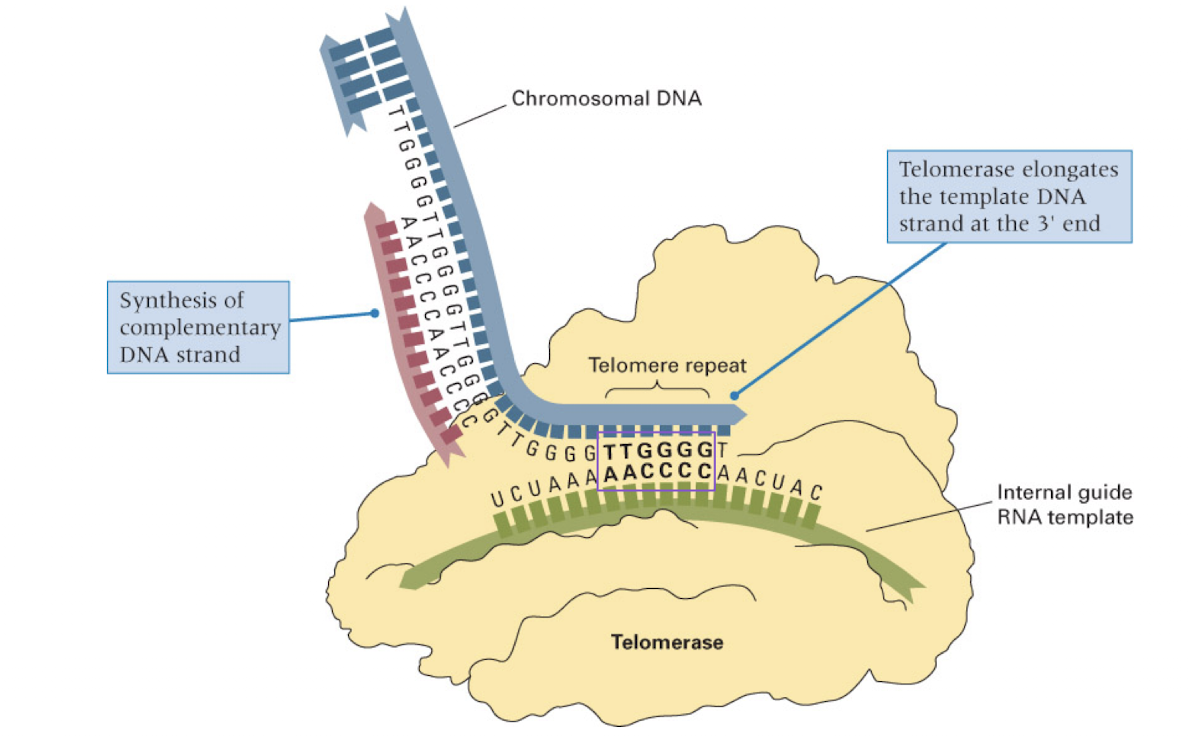
telomerase
RNA molecule which serves as the template for telomere DNA synthesis
adds telomeres to the end of the chromosome
mechanism to match age of cells (stops DNA from becoming too short over time as replication occurs and the telomere is lost)
complexity of a DNA sequence
the number of possible pairing partners for any DNA strand present in a given quantity of DNA
compare complexity of 2 strands using same amount of DNA
repetitive sequence = less complex than a random sequence
shorter sequence = less complex than a longer sequence
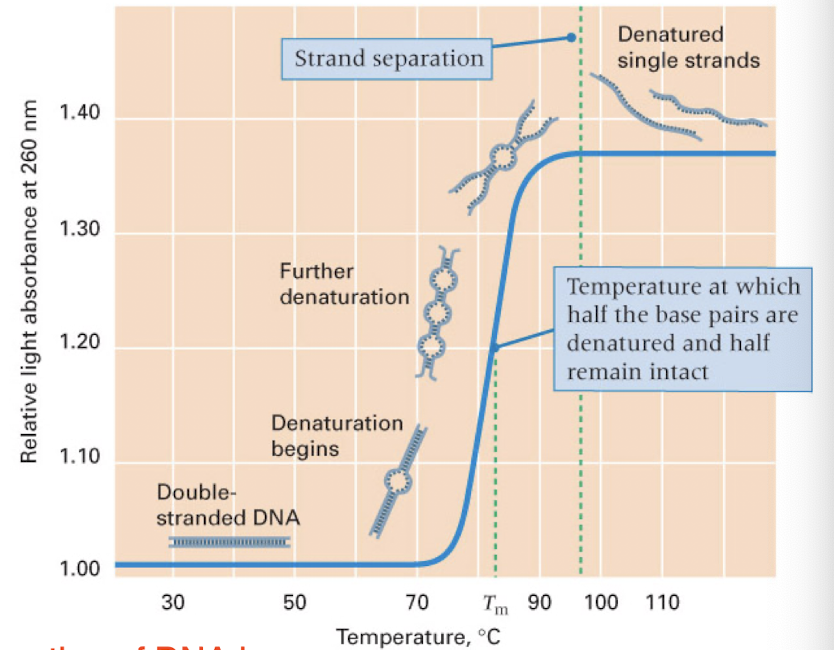
denaturation
heating of DNA leads to denaturation of the two strands
rate depends of the G+C content (3 Hydrogen bonds are harder to break than the 2 between A+T)
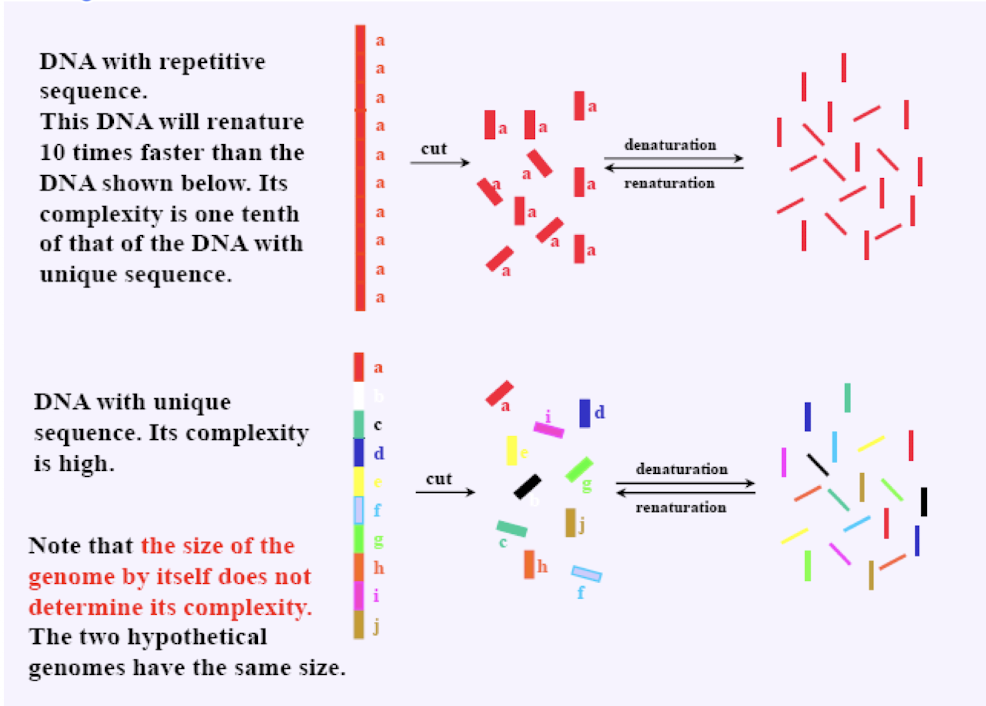
repetitive DNA sequences renature…
more rapidly than unique sequences (also denature faster)
easier to find pairing partners/arrange back into the proper sequence when repeated/less complex
rate of DNA renaturation depends on..
concentration and sequence complexity
low complexity = faster
high concentration = faster
short DNA strand = faster
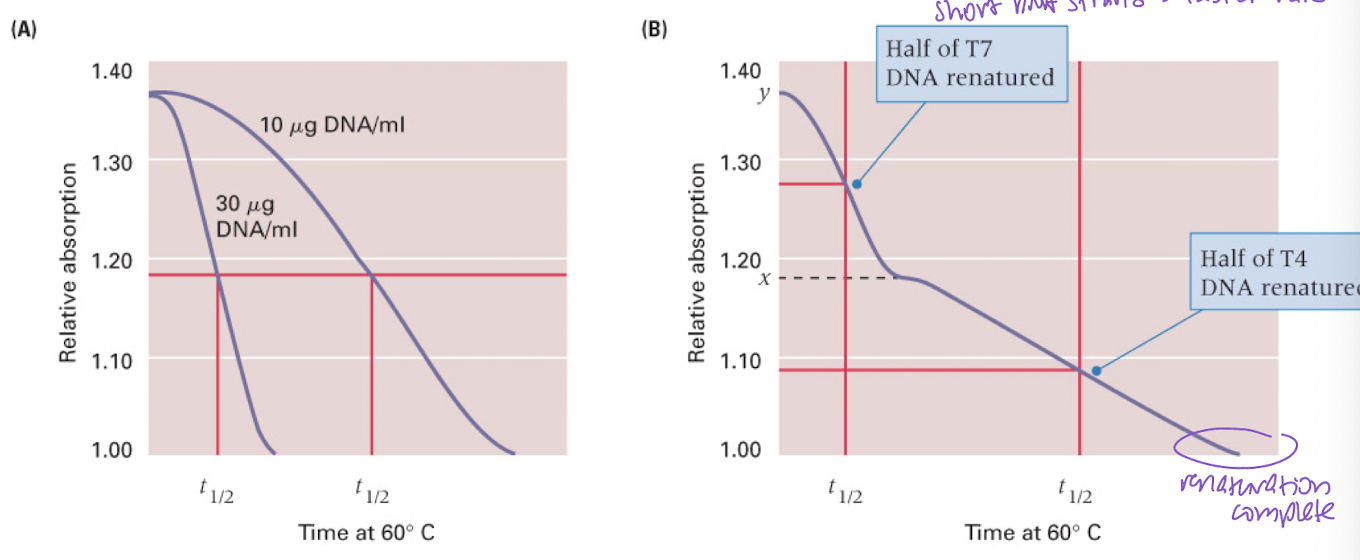
rates of renaturation of DNAs with different kinetic complexities
longer sequences = more DNA = longer to renature
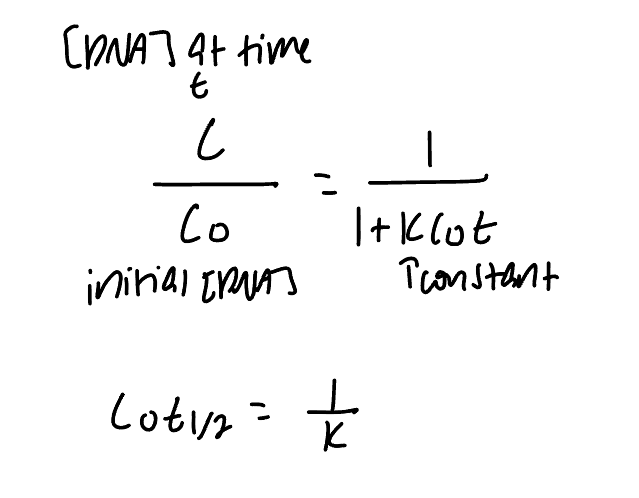
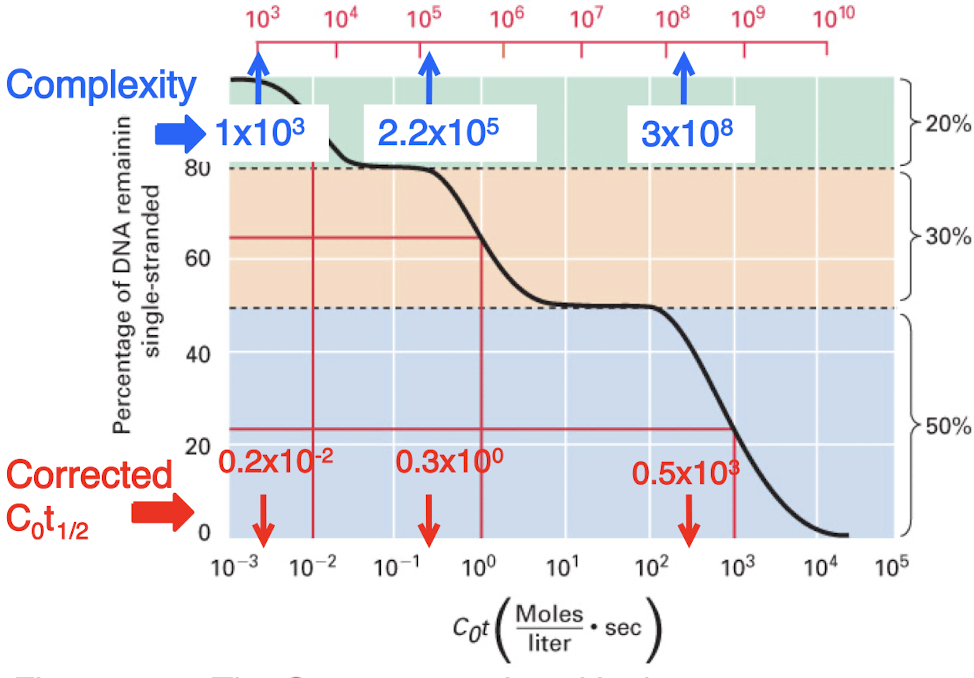
C0t curves for eukarytoic genomes typically have three distinct steps
highly repetitive, middle repetitive, and unique sequences
number of copies of a sequence is inversely related to the obersved (uncorrected) C0t1/2 values for each class
fundamental concepts for semiconservative DNA replication
replication of DNA by strand separation
each parental strand remains intact
each parental strand serves as a template for the synthesis of a complementary strand
copying of the 2 template strands using A-T and G-C base pairing
end result of DNA replication is the formation of two daughter duplexes that are identical to the parental duplex
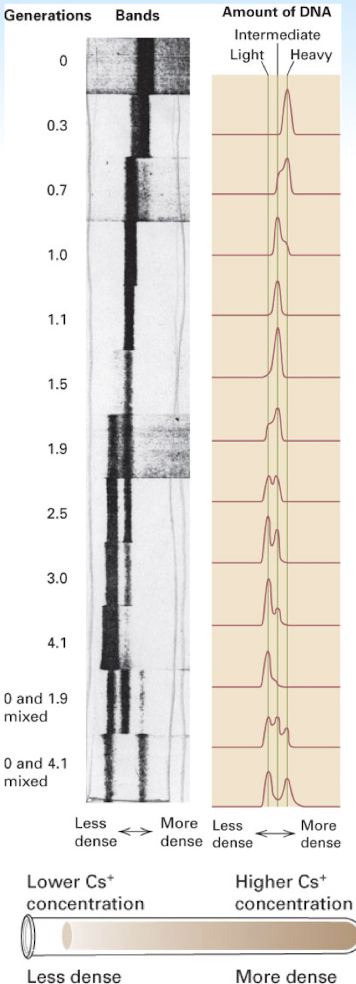
experimental proof of semi conservative replication of DNA
density of daughter DNA duplexes less than that of the parental strand- becomes “lighter” (15N of parental DNA —> 14N of daughter DNA)
after one round of DNA duplication: the molecules contain one template/parental strand and one new/daughter strand, so they have an intermediate density due to the combination of the red 15N and blue 14N strands (red and blue DNA molecules)
after two rounds of DNA duplication: the molecules containing only the 14N appear lighter (all blue DNA molecules)
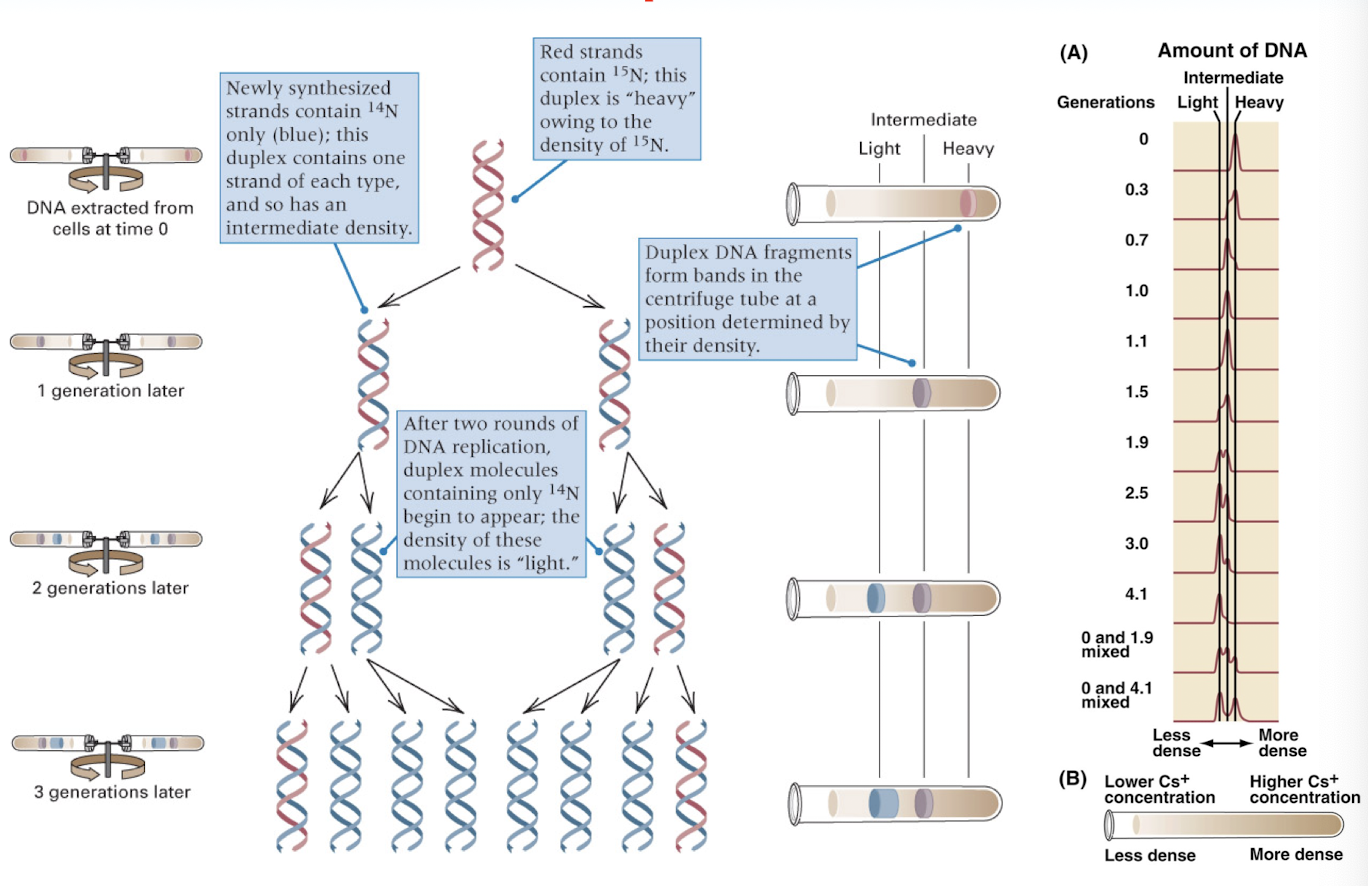
proved DNA replication is semi conservative
Meselson and Stahl
DNA becomes less dense over time due to semiconservative replication
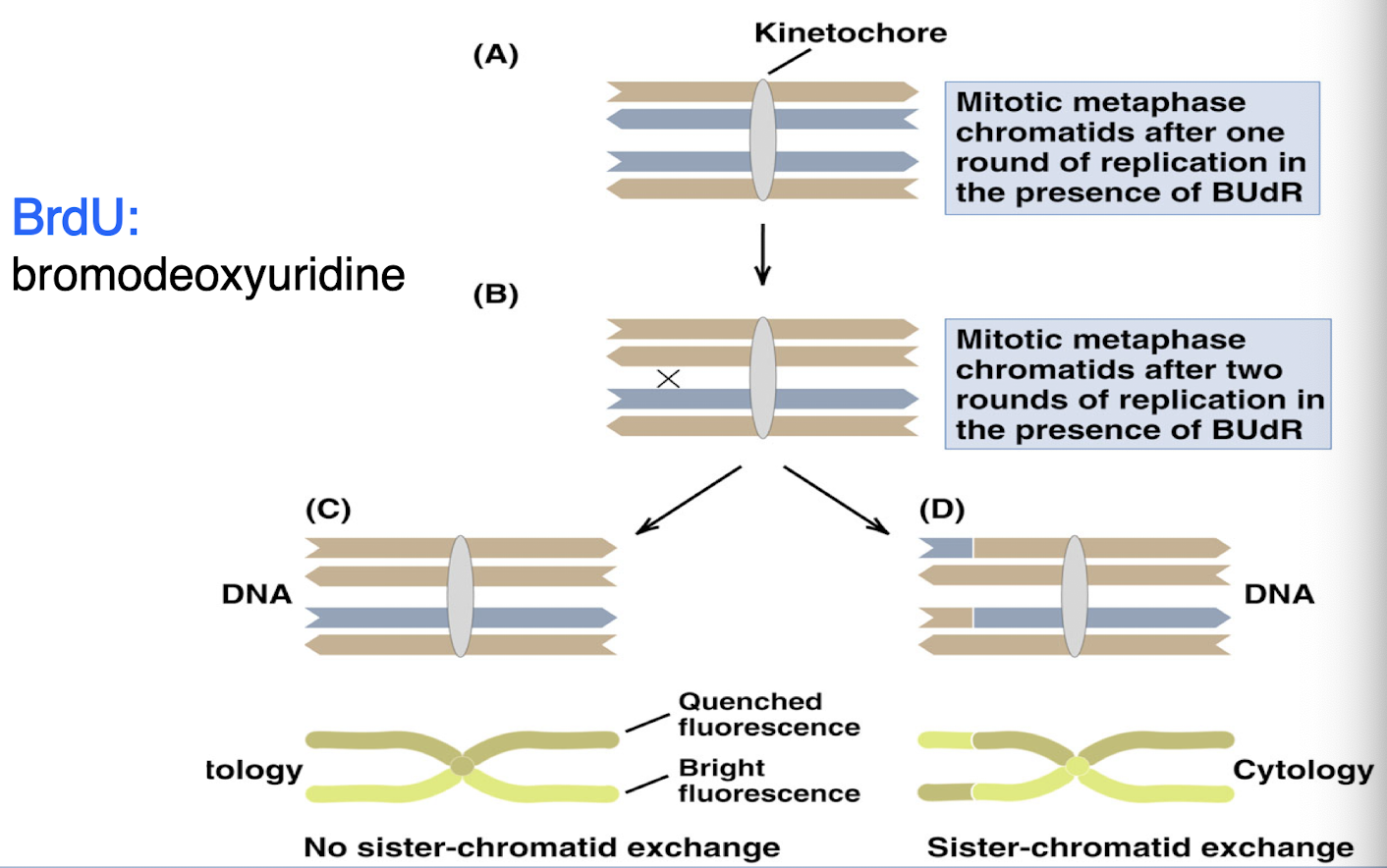
use of a thymidine analog (BRdU) provides cytological proof that DNA replicates in a semi conservative manner
BRdU detects sister chromatid exchange during DNA replication
one round of replication: each chromatid has one old strand (normal DNA) and one new strand (contains BRdU)
two rounds of replication: one chromatid is fully contains BRdU and other has one strand of each
BRdU can visually detect recombination events:
uniform staining = no exchange
patchy staining = sister chromatid exchange occurred
homologs can undergo crossing over
recombination between sister chromatids of homologous pairs
three different replication models
theta replication: e. coli
rolling circle replication: DNA phages
multiple origins and bidirectional replication: eukaryotic chromosomes
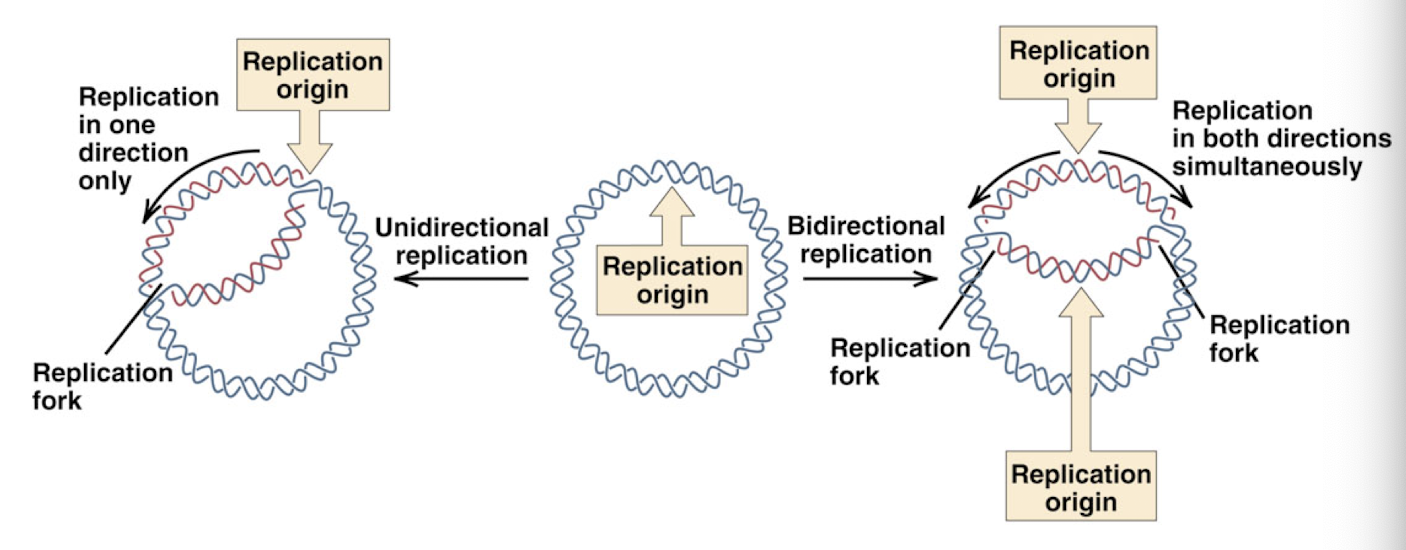
theta DNA replication
both parental DNA strands remain intact
DNA replication begins at specific sequences: “origins of replication”
replication fork: where the DNA is unwinding to allow new strand to form complimentary to template strand
can be
unidirectional: moves in one direction
bidirectional: moves both directions simultaneously
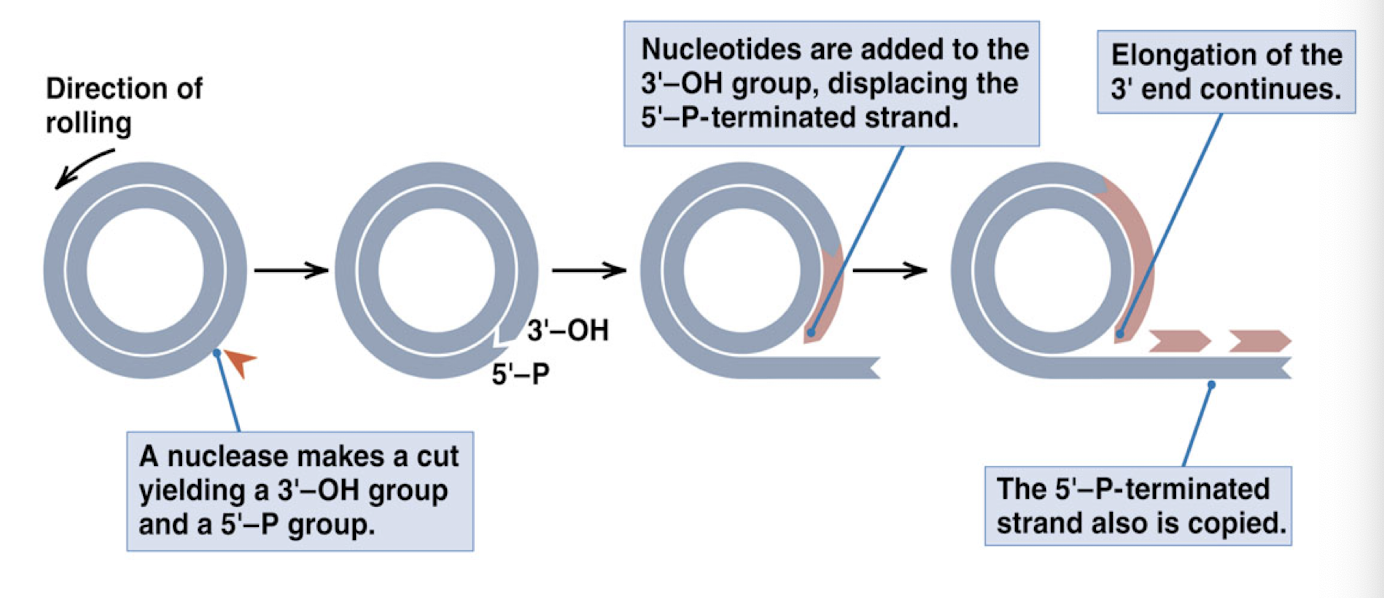
rolling circle DNA replication
one of the template DNA strands is cut by enzyme nuclease to create a primer 3- OH end
nucleotides are added to the 3’ end of the cut strand (bases added in 5’ —> 3’ direction)
for every round of replication, the tail of the rolling circle becomes one unit longer
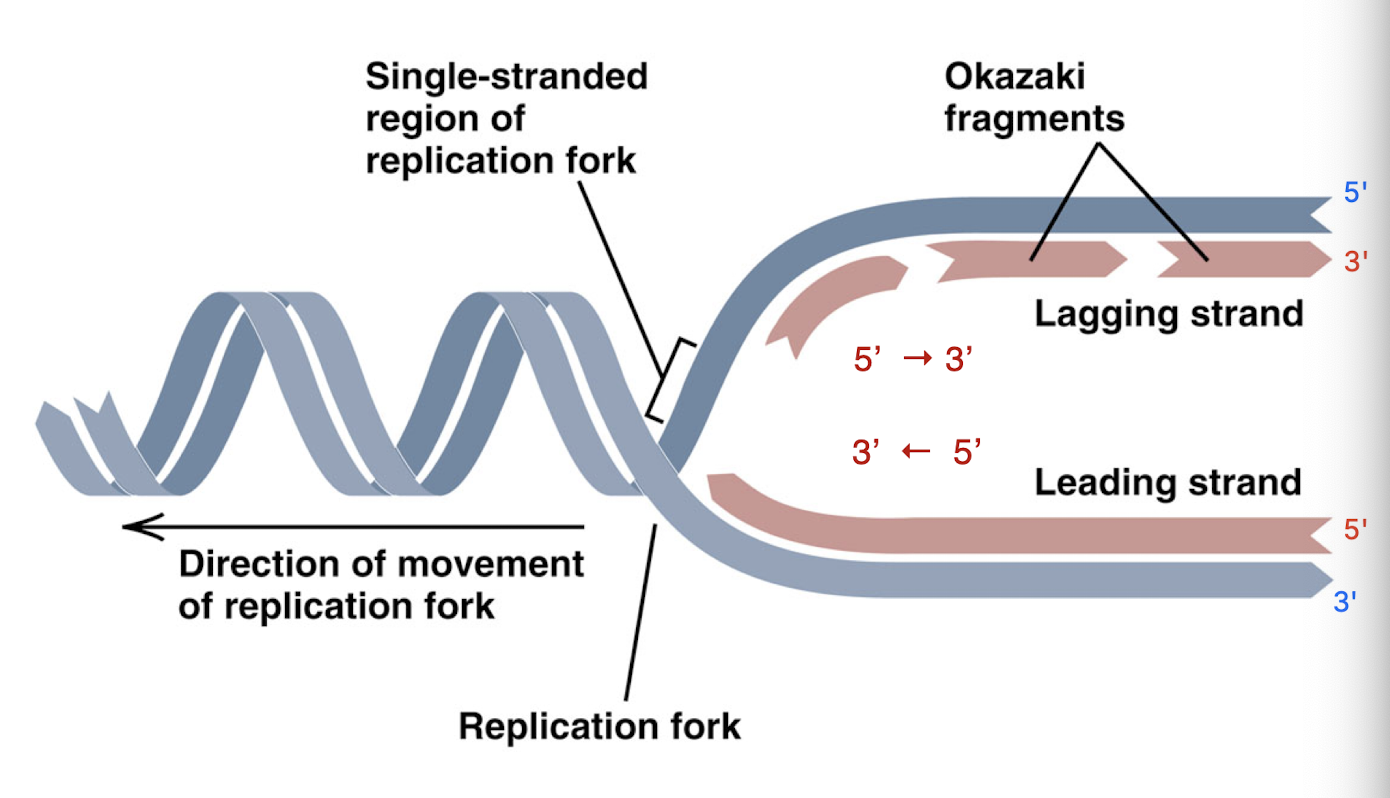
okazaki fragments
small chunks of the new DNA strand created as the lagging strand because of the direction DNA must build (onto the 3’ end on new strand)
because lagging strand builds away from replication bubble, must build in pieces as replication fork advances
replication of a linear eukaryotic chromosome
replication begins at replication origin and is bidirectional
multiple replication bubbles can exist and they will fuse once they meet
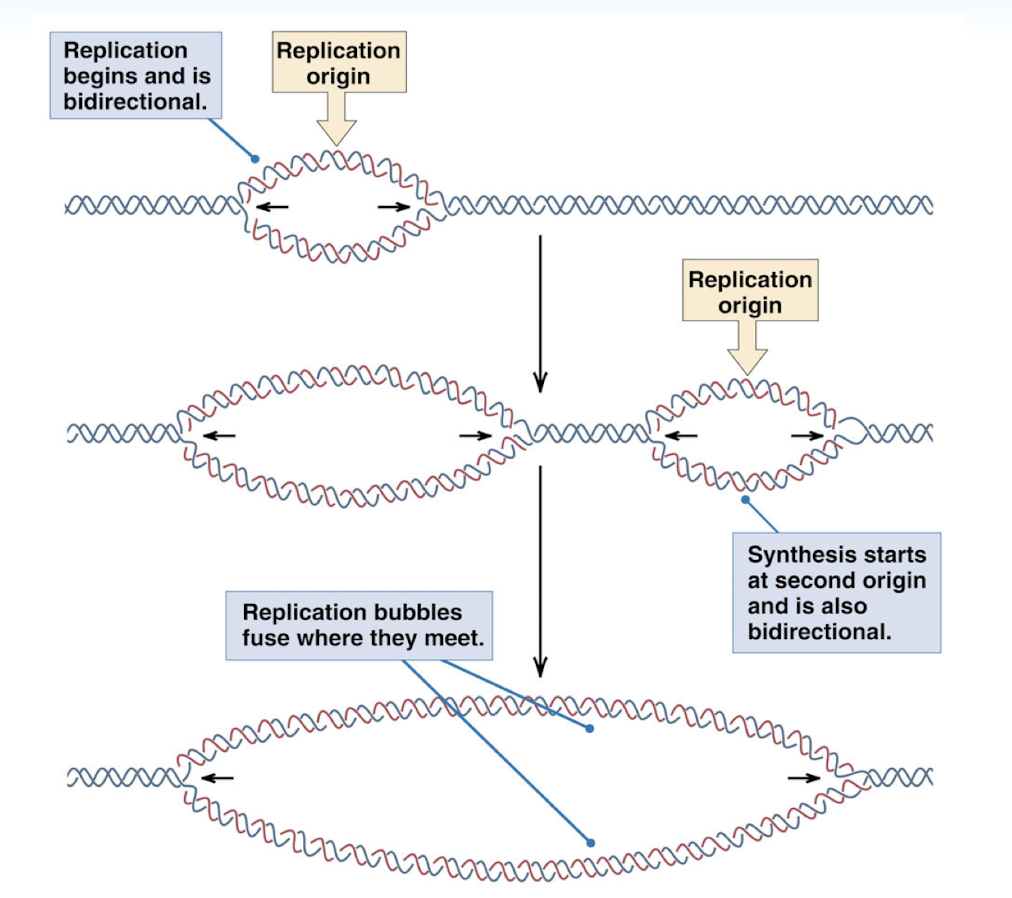
critical steps in DNA replication
unwinding, stabilization, and stress release
initiation my a Primosome Complex
chain elongation and proofreading
DNA replication proteins
helicase: enzyme, unwinding DNA double helix
SSB proteins: stabilizing the templates (single stranded DNA), DNA stressed when unwound and no longer double stranded
gyrase or topoisomerase: release mechanical stress of unwinding
primosome: initiate new strand synthesis
DNA polymerase complex: elongating new strand
DNA ligase: ligating (combining) Okazaki fragments
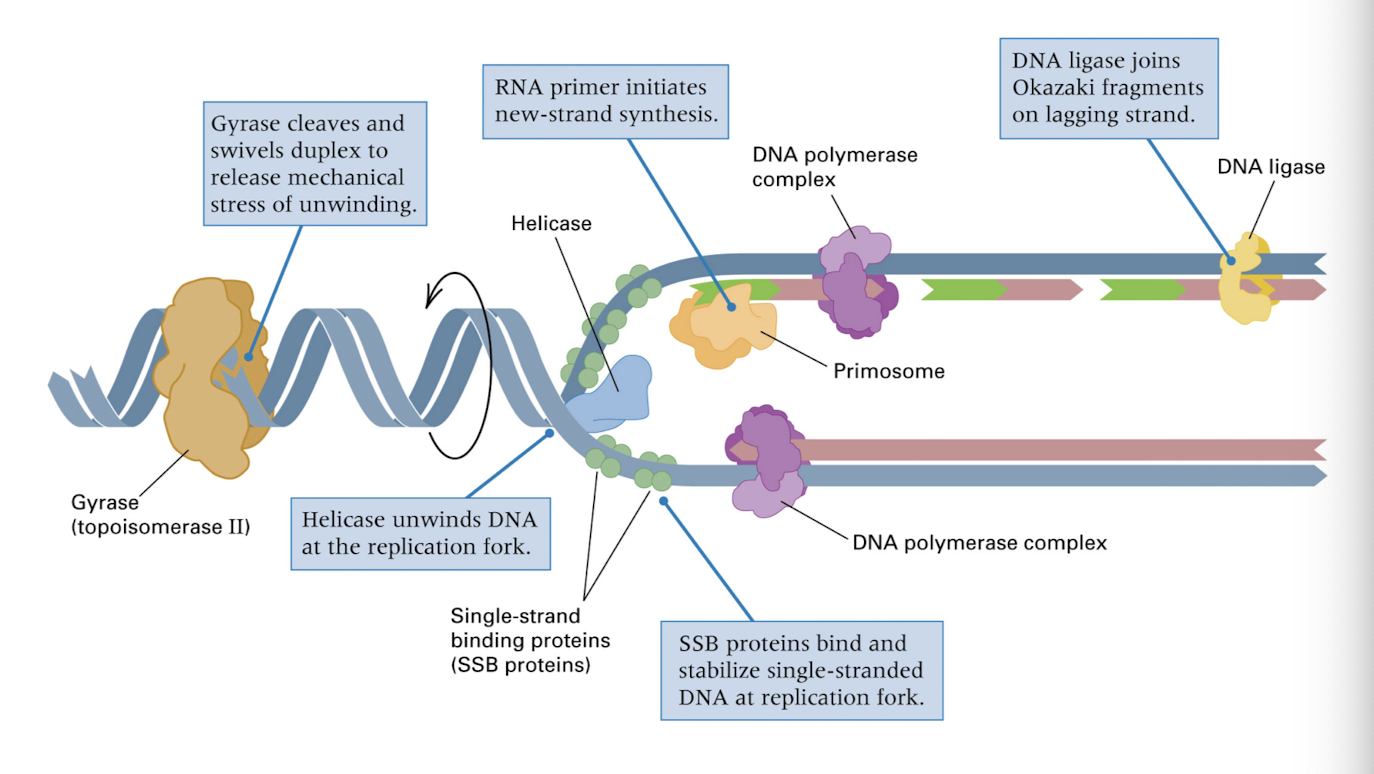
e.coli DNA replication fork
helicase unwinds DNA at replication fork
RNA primer initiates new strand synthesis
SSB proteins stablizie the single stranded DNA
DNA polymerase adds the complimentary nucleotides
DNA ligase joins fragments on lagging strand
gyrase cleaves DNA ahead of replication fork to release stress
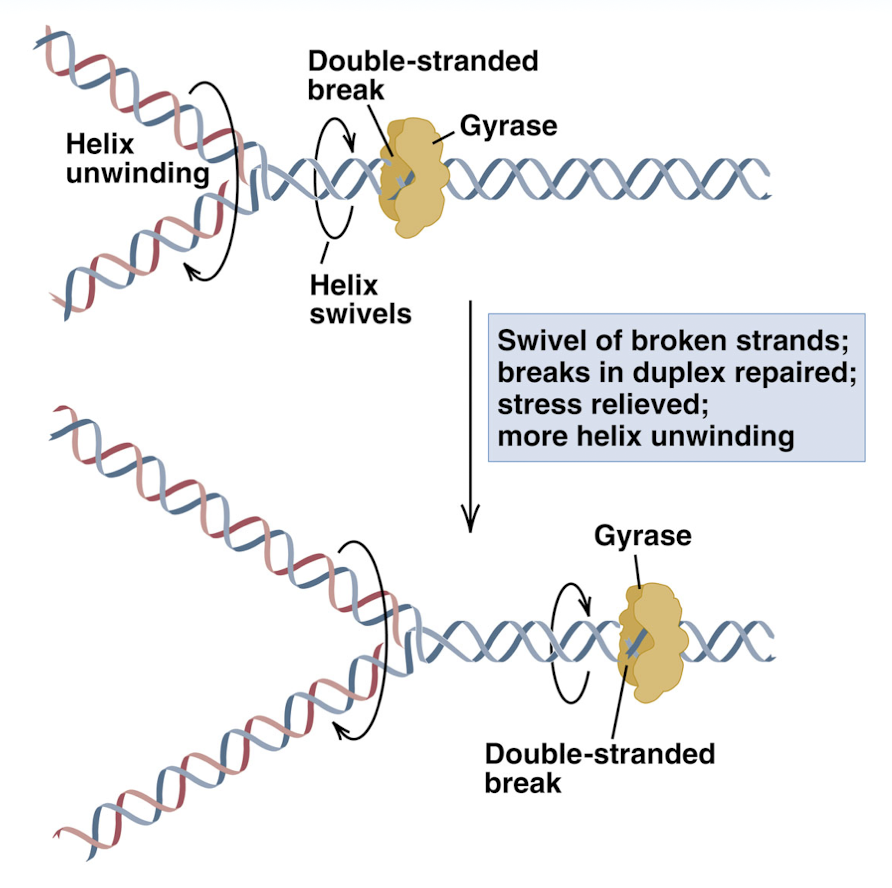
prevention of knotting of DNA by DNA gyrase
double strand broken and the helix can swivel to release stress before replication bubble reaches this section of DNA
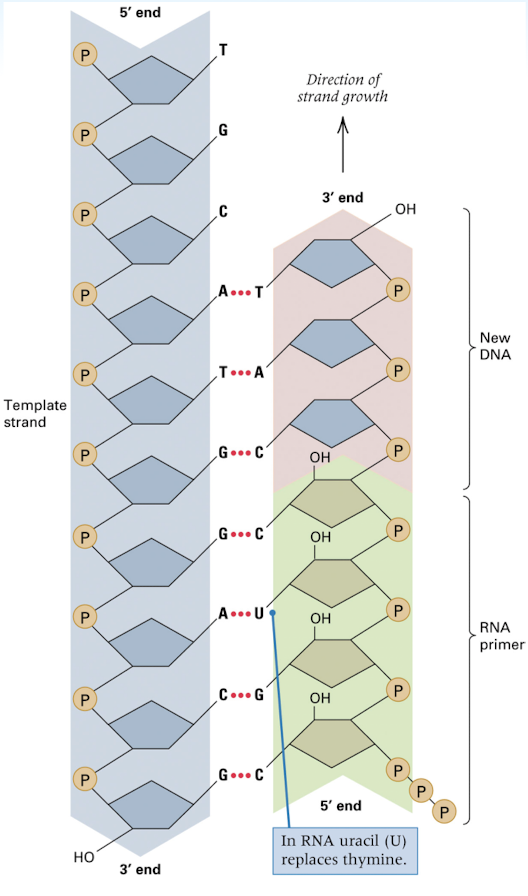
priming of DNA synthesis with a RNA segment
primer needed for DNA polymers (complimentary DNA bases to the template strand) to have a section of nucleotides to attach to to begins replication
new DNA chains are initiated by short RNA primers- RNA primer becomes 5’ end of new strand and is complimentary to the 3’ end of the template strand
DNA polymerase can attach the new DNA nucleotides to the RNA primer that begins the synthesis of the new strand
addition of a deoxynucleotide (DNA nucleotide) to the 3’ OH end
replication occurs in the 5’ —> 3’ direction (new strand), bases are added to the 3’ end of the template strand
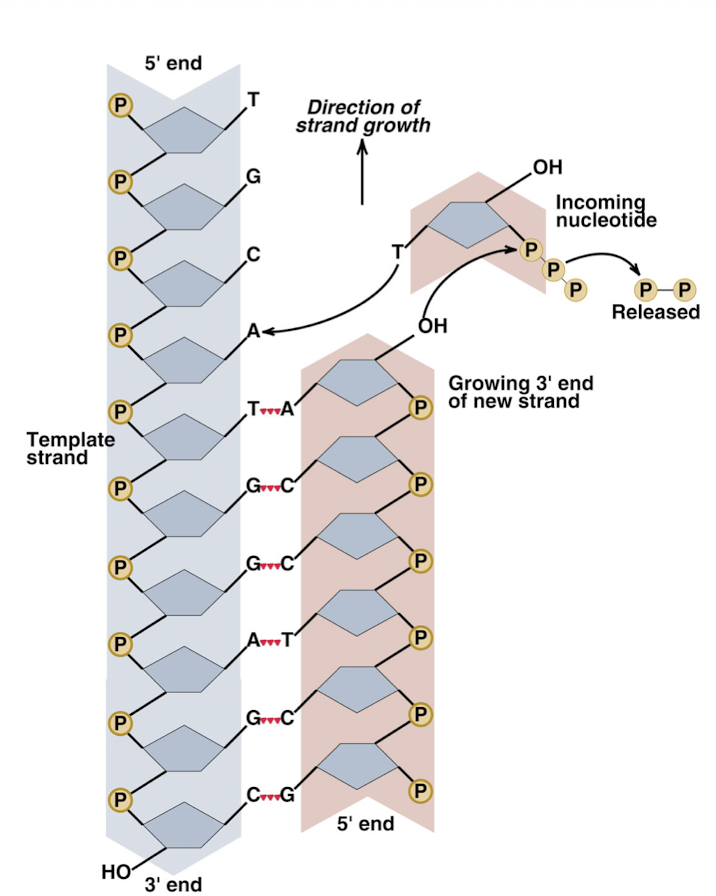
leading strand vs lagging strand
leading strand: moves towards the replication fork and is synthesized continuously in one large fragment
3’ strand of the template strand- DNA can just be added to the new strand easily
lagging strand: moves away from the replication fork and is synthesized in many, smaller Okazaki fragments (which are glued together by ligase)
must add to 3’ of template strand, which is the side of the strand the replication fork is moving towards
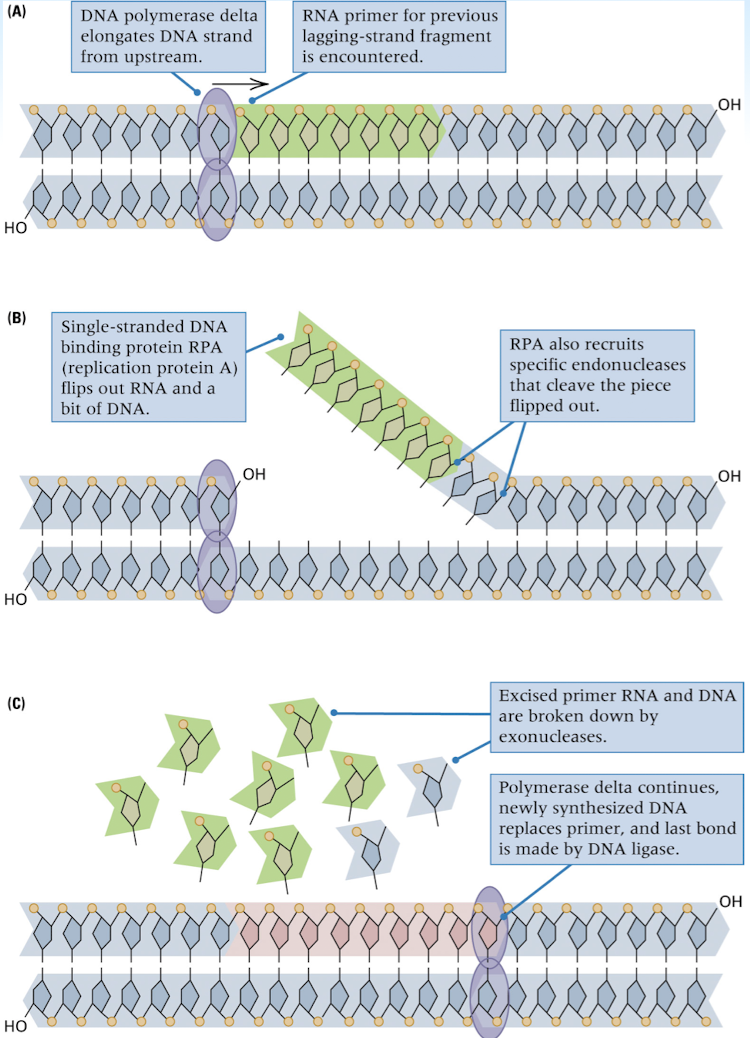
joining of Okazaki fragments
removal of RNA primer
replacement of the primer with the correct DNA sequence
joining where the adjacent DNA fragments come into contact
DNA polymerase in DNA replication
can add free nucleotides only to the 3’ end of the newly forming strand (so strand is built in the 5’-3’ direction)
can add a nucleotide only to a preexisting 3- OH group
correcrs mistakes in newly synthesized DNA (proofreading)
addition of a dideoxynucleotide to the 3’ OH end of a DNA chain…
terminates chain elongation
dideoxyribose: no 3’ OH group, so DNA replication cannot continue
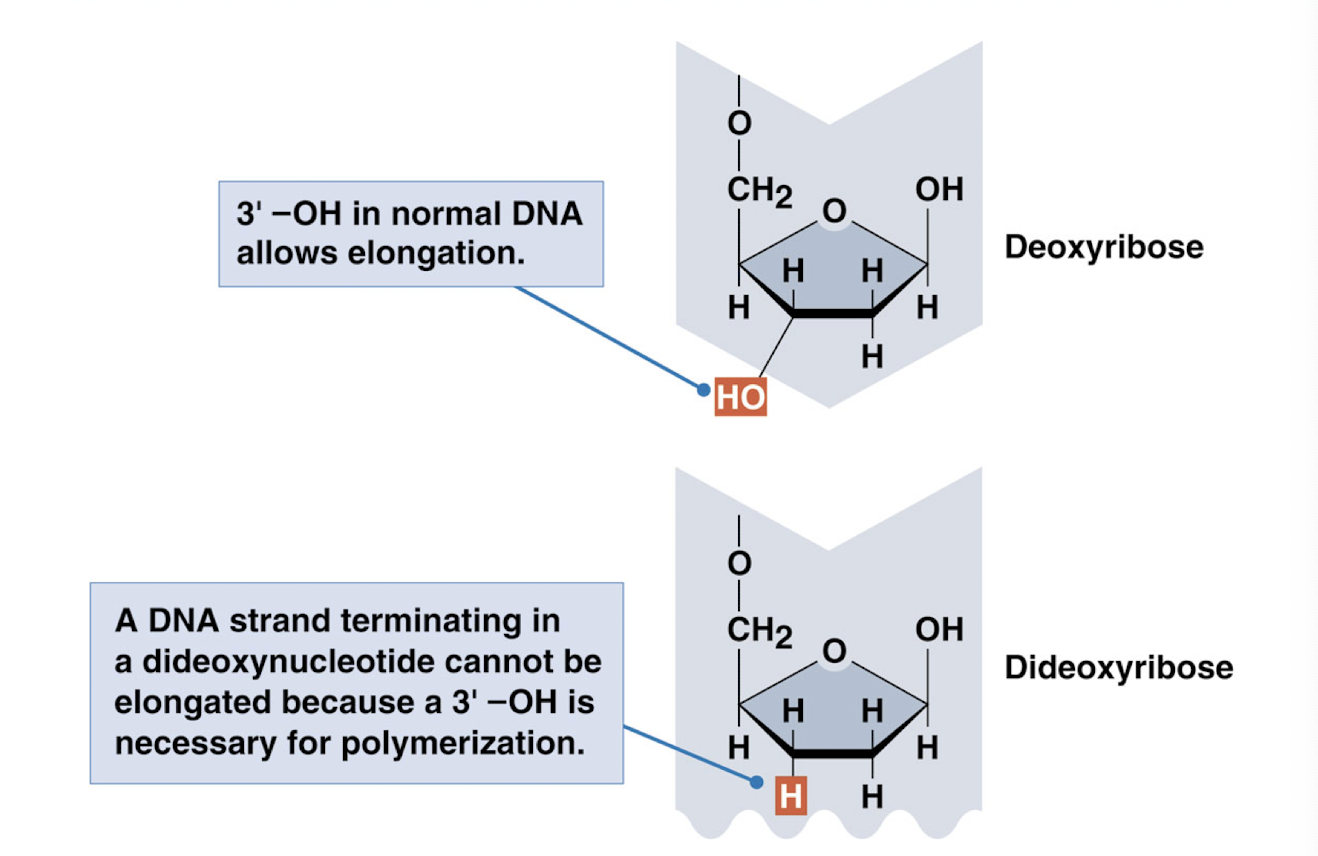
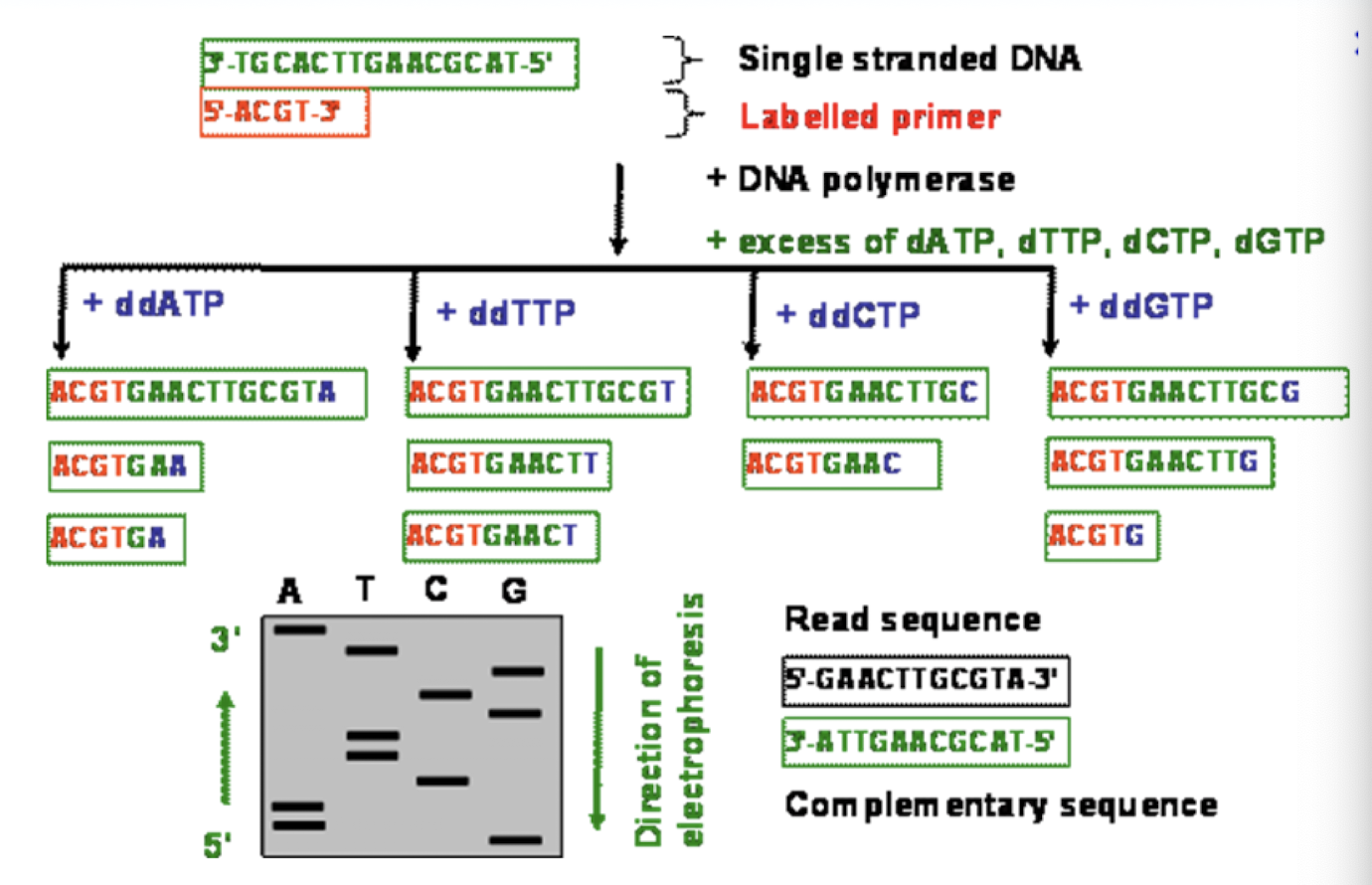
Sanger Sequencing
takes advantage of ddNTP chain terminators (N = some nucleotide, A, T, C, G)
DNA replication will be stopped where these are added- where whichever nucleotide the ddNTP is located
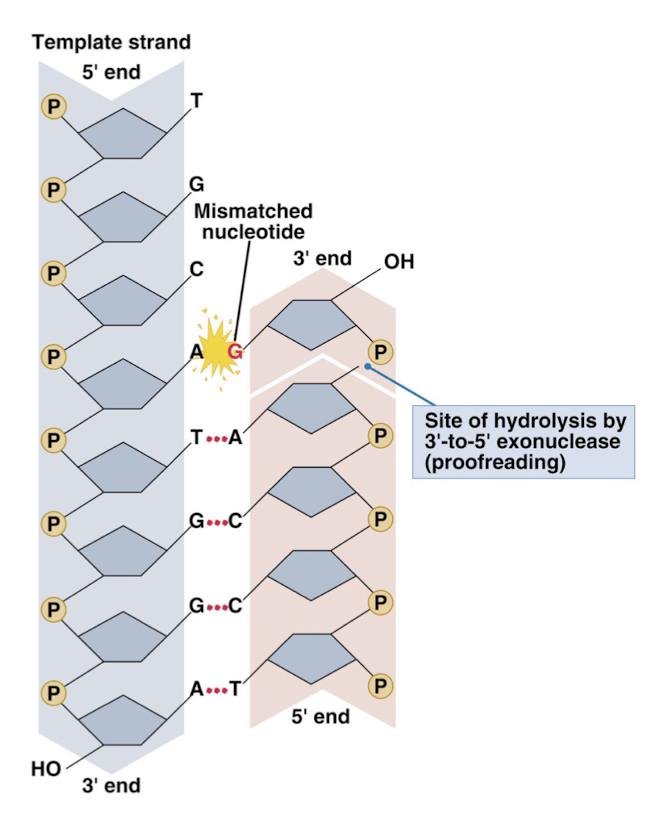
proofreading exonuclease function of DNA polymerase
when an incorrect base pair is formed, an enzyme cuts the DNA so that the incorrect nucleotide will be removed- ex: A-G removed because complimentary pairing should be A-T
endonucleases: enzymes that cleave the phosphodiester bond within a polynucleotide chain
exonucleases: enzymes that cleave phosphodiester bonds at the end of a polynucleotide chain
DNA polymerase can act as a 3’-to-5’ exonuclease
central dogma of biology
DNA —> RNA —> Protein
(transcription) (translation)
three major types of proteins
enzyme proteins- speeds up/facilitates reactions
regulatory proteins- regulate processes
structural proteins- make frame of cells
proteins
composed of one or more polypeptide chains
each polypeptide chain is a series of covalently joined amino acids
20 different amino acids commonly found in polypeptides, can be joined in any order and in any number (typically 100-1000)
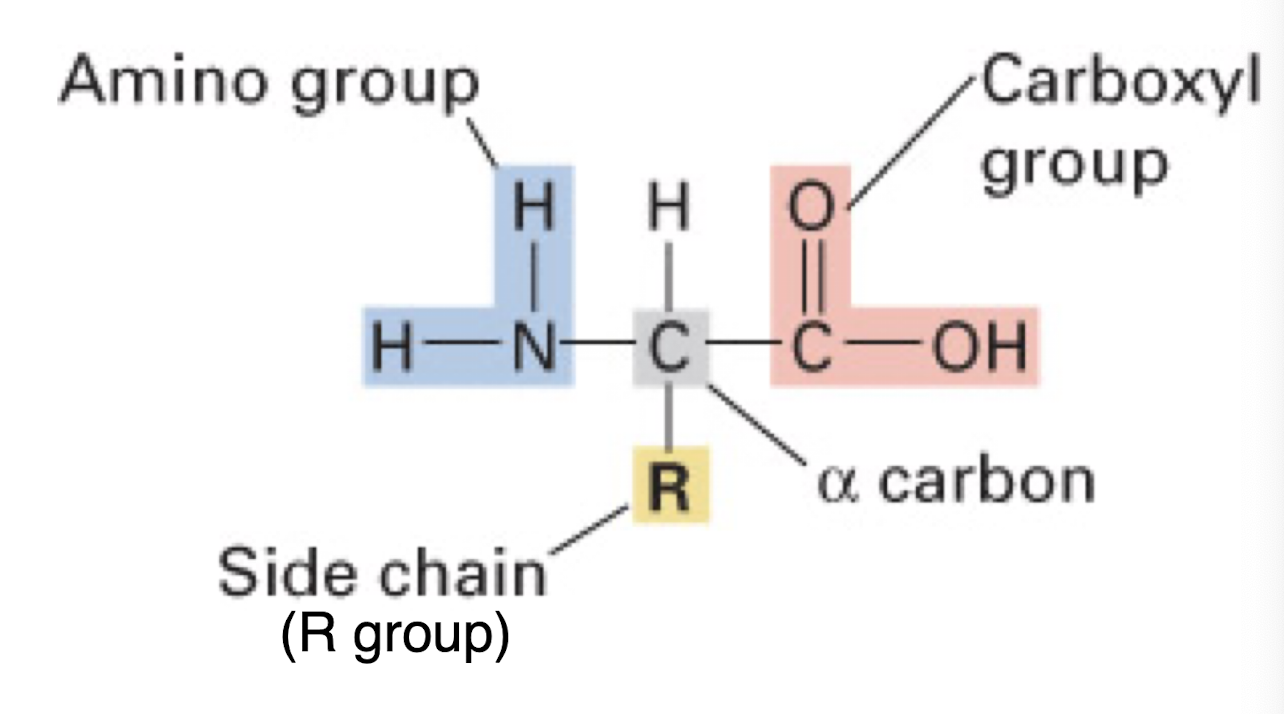
structure of an amino acid
Amino group (H2N), R group (various), Carboxyl group (COOH), and Hydrogen all attached to a central carbon (alpha carbon)
carbon backbone formed from central carbons
structure of amino acids depend on their R groups
some amino acids positively charged, some are negatively charged based on their R groups
positive: have extra amino group in R group
Alanine, Lysine
negative: have extra OH group in R group
Aspartic Acid, Glutamic Acid
charged: polarized molecules
hydrophilic (have charge) vs hydrophobic (have no charge)
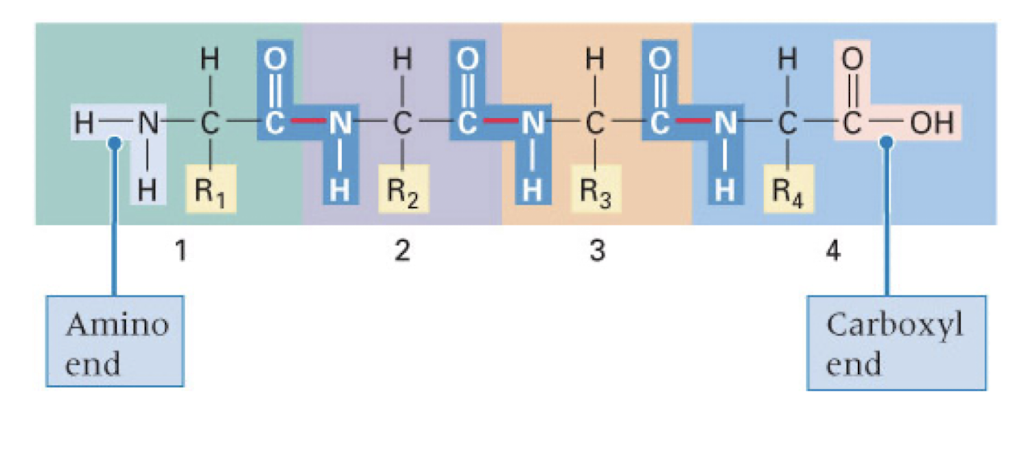
amino acids are joined by _____ bonds to create polypeptides
peptide bonds
R groups jut off the carbon backbone
amino acids are added to the carboxyl (-COOH)end of the polypeptide
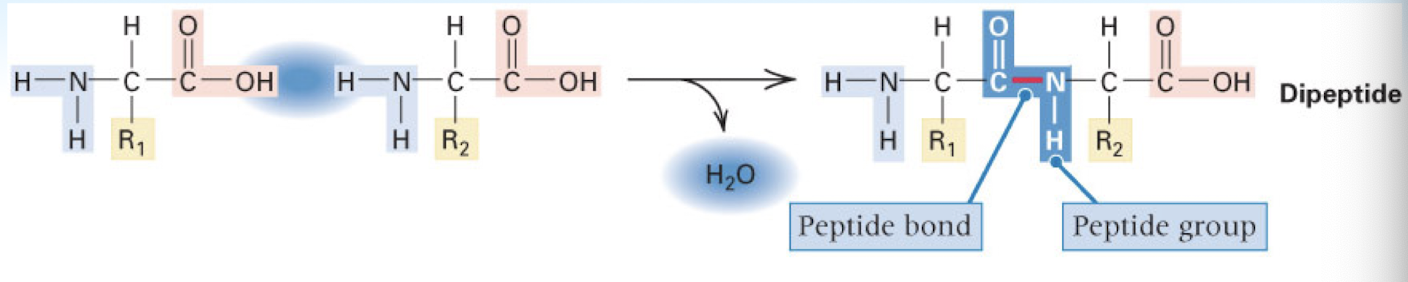
dehydration synthesis reaction
water removed when two amino acids combined in order to form a peptide bond —> forms dipeptide chain
as more amino acids added —> formed polypeptide chain
sickle cell anemia
normal red blood cells are smooth and round, move easily through blood vessels to carry oxygen to all parts of the body
sickle shaped cells are stiff and sticky, do not move easily through the body and tend to form clumps/get stuck in blood vessels
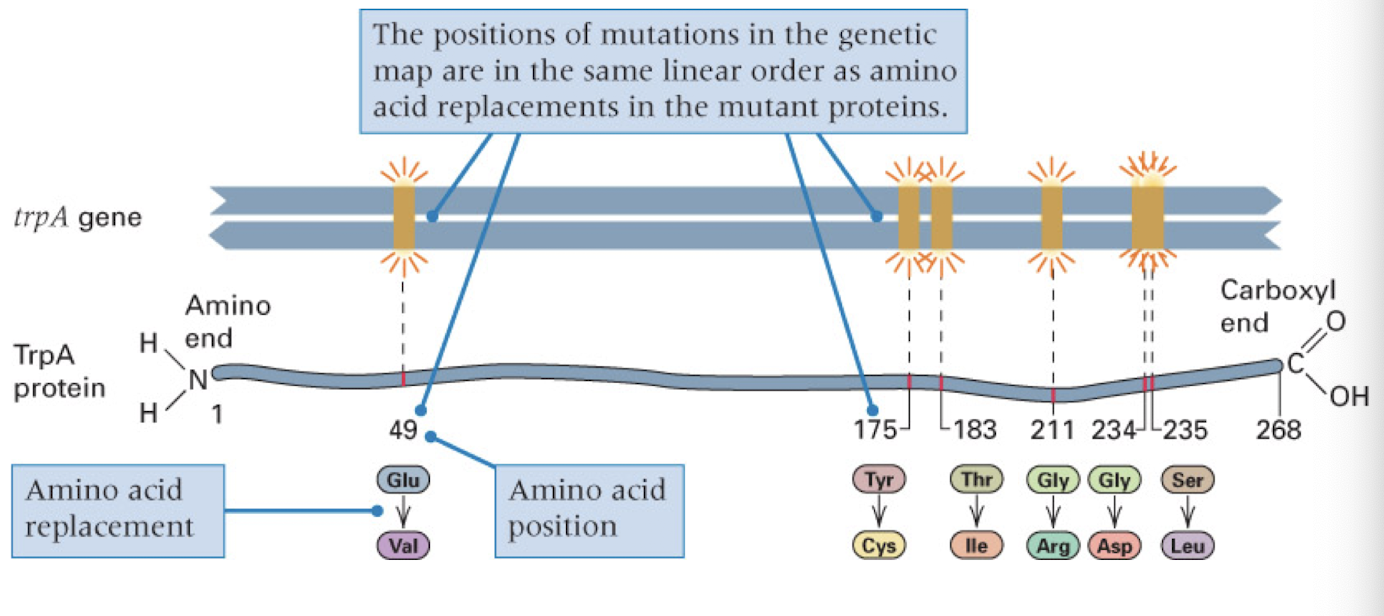
DNA and protein are colinear
DNA sequence determines amino acid sequence in a point-to-point manner
mutation in DNA can be traced back to a change in the AA in the DNA
basis for genetic disease
transcription
process by which the information contained in DNA is copied into a single strand RNA molecule of complementary base sequence
RNA polymerases
large, multiple subunit complexes who active form is called the RNA polymerase holoenzyme
bacteria: one RNA polymerase transcribes all genes
eukaryotes:
RNA polymerase I- rRNA
RNA polymerase II- mRNA and certain small RNAs
RNA polymerase II- tRNA, 5S rRNA
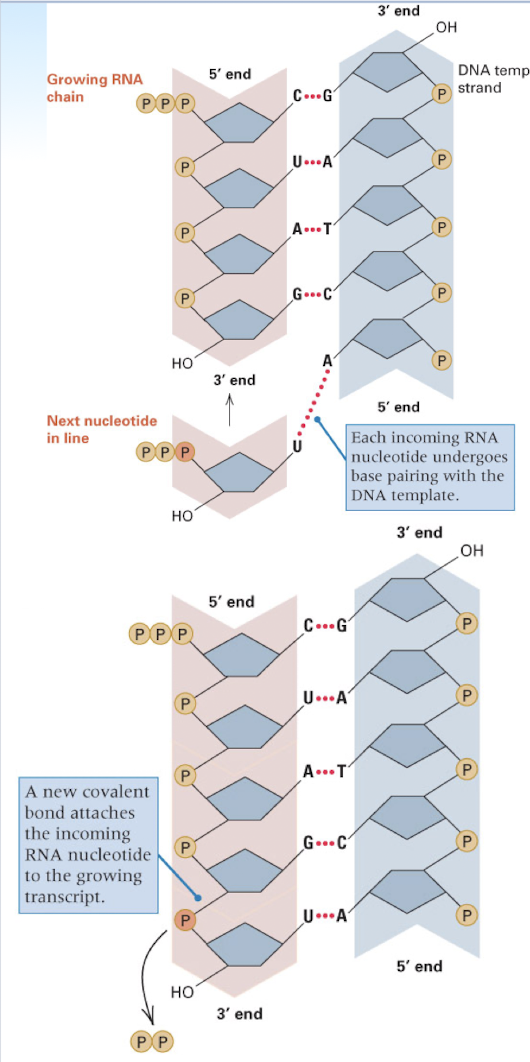
RNA synthesis
DNA serves as template
U base pairs with A in RNA synthesis (rather than T)- complementary base pairing with template strand
still builds onto the 3’ end of the growing new strand
RNA polymerase does not require a primer- can initiate instantly
can recognize unique region of DNA to begin RNA synthesis: promoter sequence
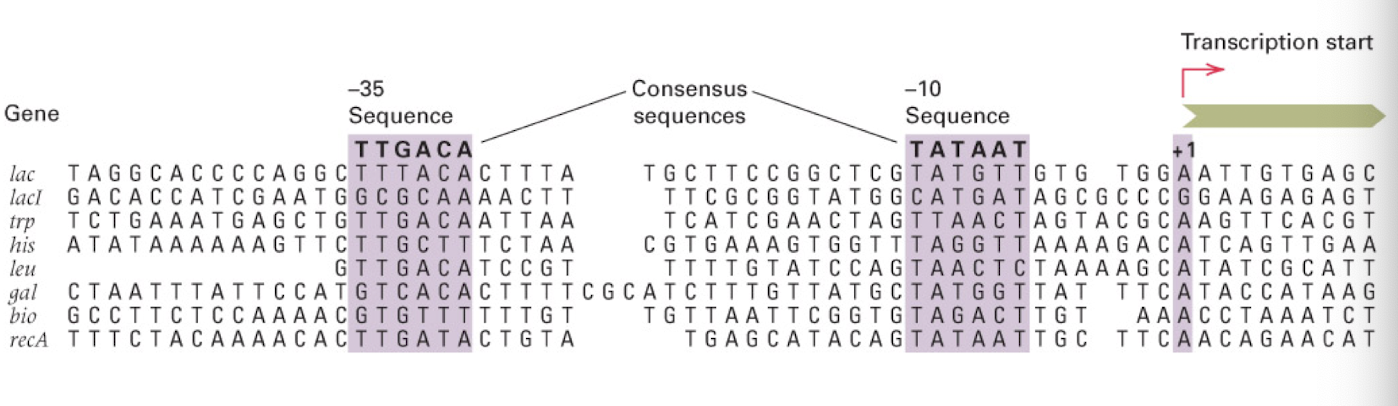
e. coli promoters have conserved sequences
-35 and -10 nt regions determine promoter strength
consensus sequences
share sequence homology, but not identical
sigma factor protein: help to recruit RNA polymerase to promoter region
the more closely a sequence resembles these sequences, the stronger the promoter (???)
eukaryotic RNA polymerase II promoters contain a…
TATA box- much further from from transcription start (25-30 nucleotides) than in e. coli
farther so there is more control
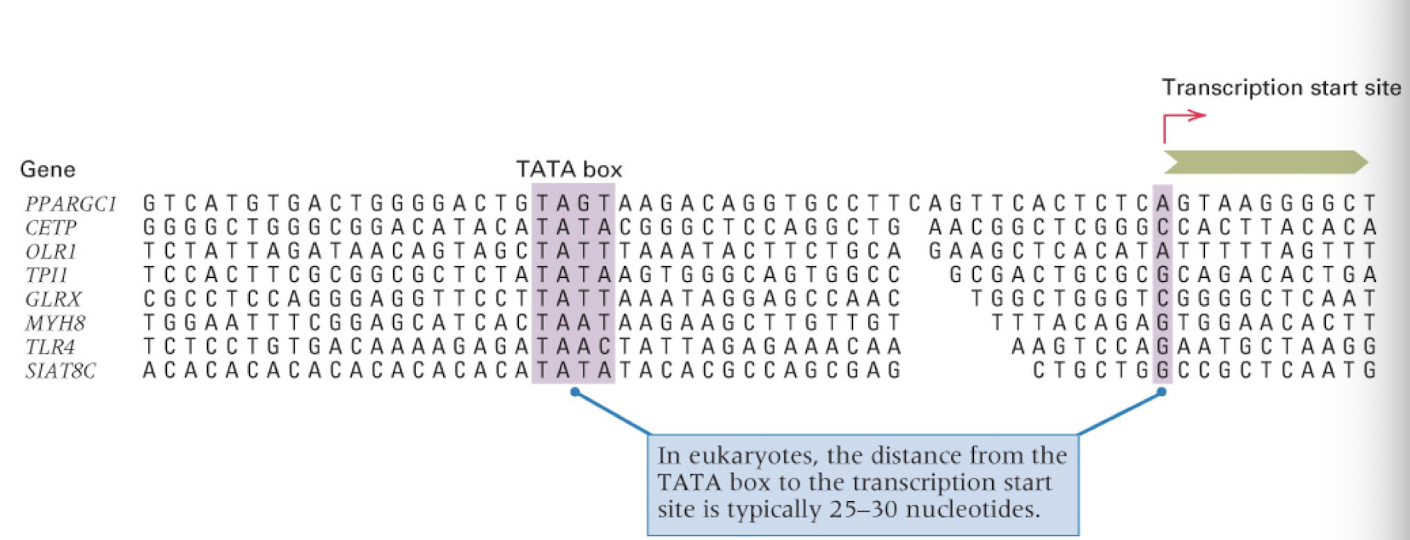
stages of transcription
chain initiation: RNA polymerase binds to initiate transcription
chain elongation: only the template strand is transcribed, transcription bubble is ~15 nt long with 8-9 nt paired with the 3’ end of the RNA
nucleotides added to the RNA strand are complimentary to the template strand (but code for the non template strand)
termination: RNA polymerase reaches a terminator sequence and the RNA and polymerase are released
basal factors required to form the class II pre-initiation complex
6 factors and RNA polymerase II = preinitiation complex
factors:
TFIIA, TFIIB, TFIID, TFIIE, TFIIF, TFIIH
many are multi subunit
factors and RNAP must also bind in a specific order (in vitro)
transcription chain initiation by RNA polymerase II
transcription chain initiation by RNA polymerase II
TBP- TATA box binding protein
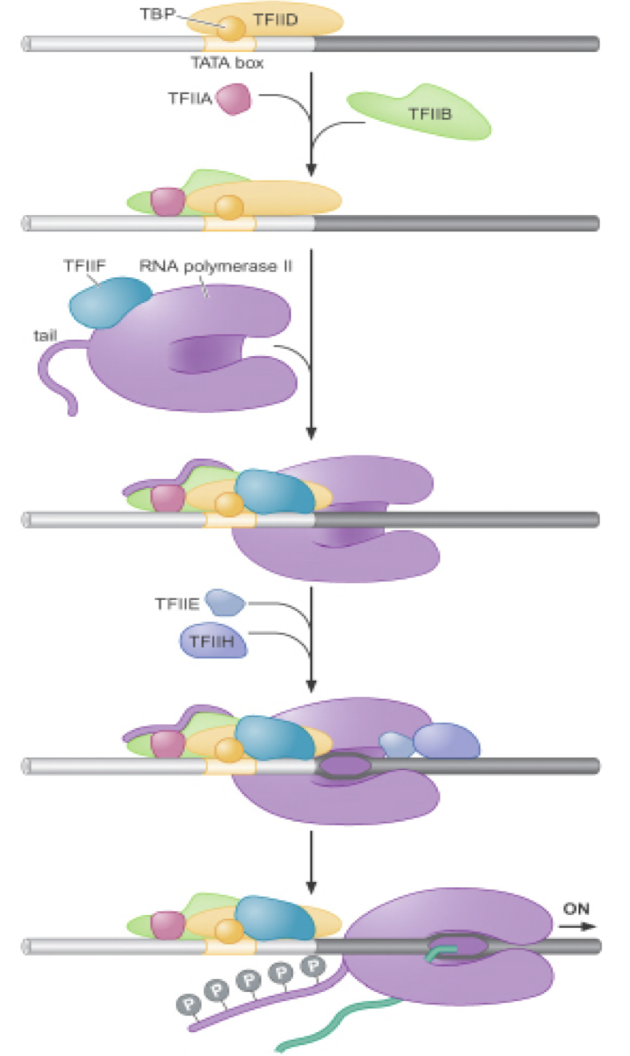
TBP (TATA box binding protein) binds to and distorts DNA using a…
a beta sheet inserted into the minor groove- bends DNA ~90 degrees and forces open the minor groove
TBP-DNA complex
the TBP is in purple, the DNA TATA sequence is in gray
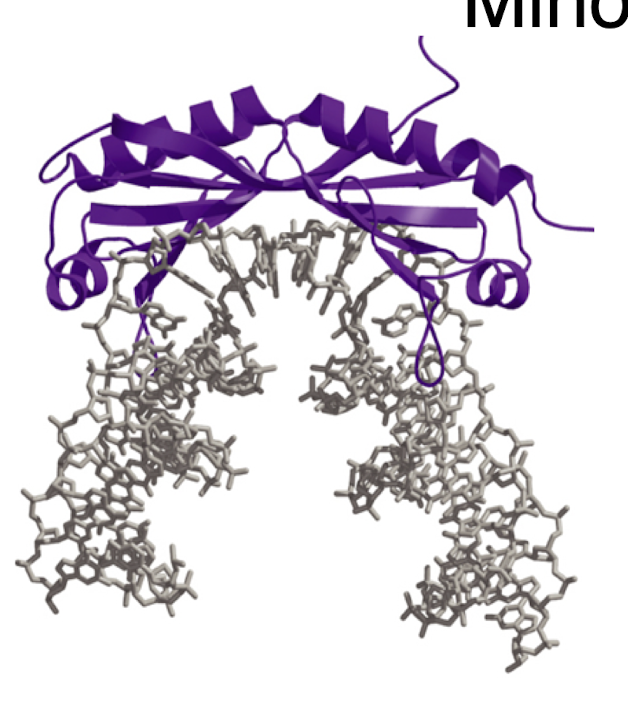
functions of TFIIA, TFFIIB, TFIID, and TFIIF, TFIIE, TFIIH
TFIIA: binds to TBP and could be considered a TAFII (TBP associated factor)
TFIIB: needed for the polymerase/TYIIF complex to bind to TFIID- linker between the two
TFIIF: binds to the RNAP and reduces non specific binding of the RNAP to DNA
TFIIE: binds after polymerase/TFIIF binds and stimulates and recruits TFIIH
TFIIH: required for promoter clearance, has DNA helicase activity for melting DNA at transcription bubble, has Kinase activity for phosphorylation of the CTD of the large subunit of RNAP
promoter clearance
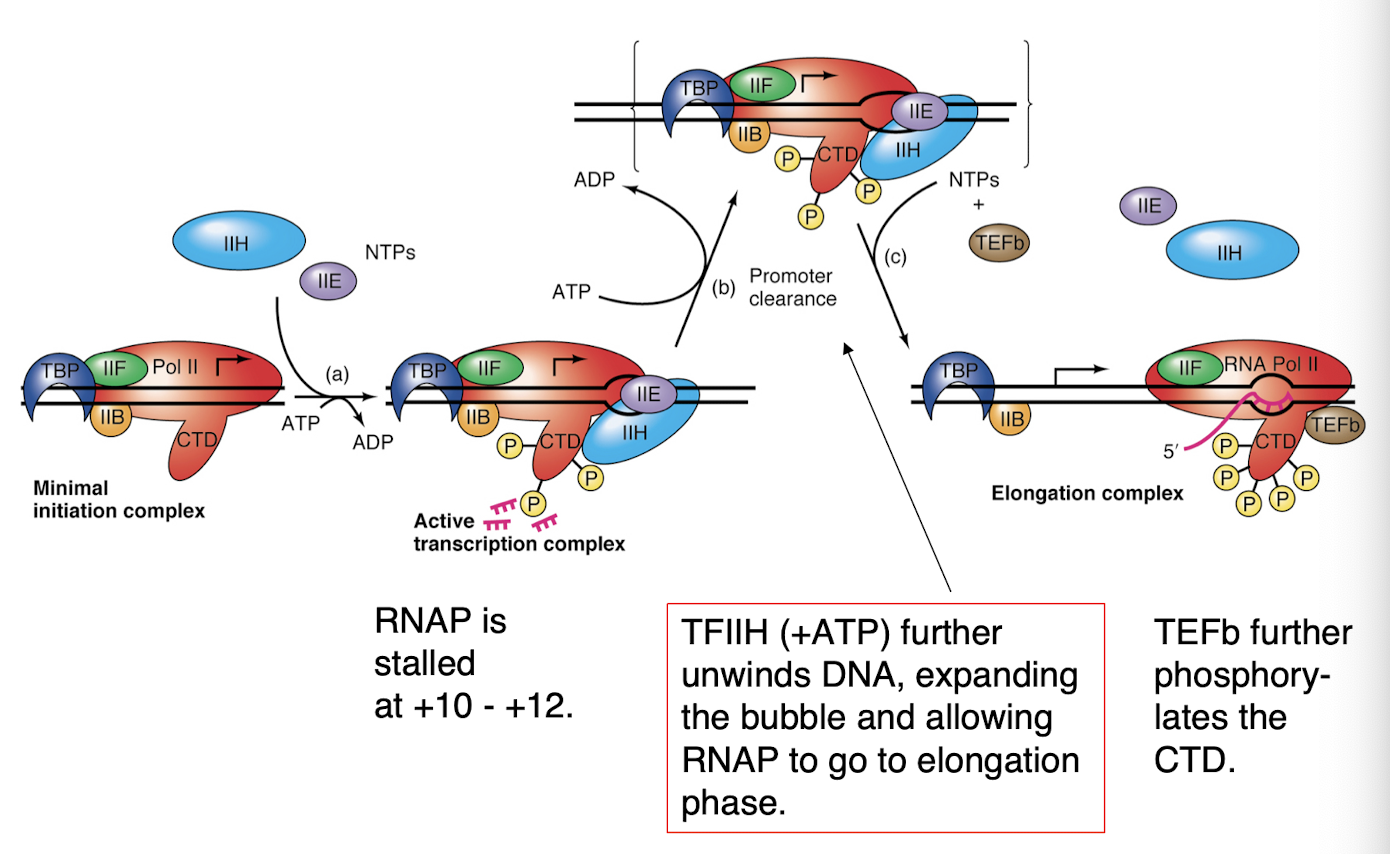
chain termination
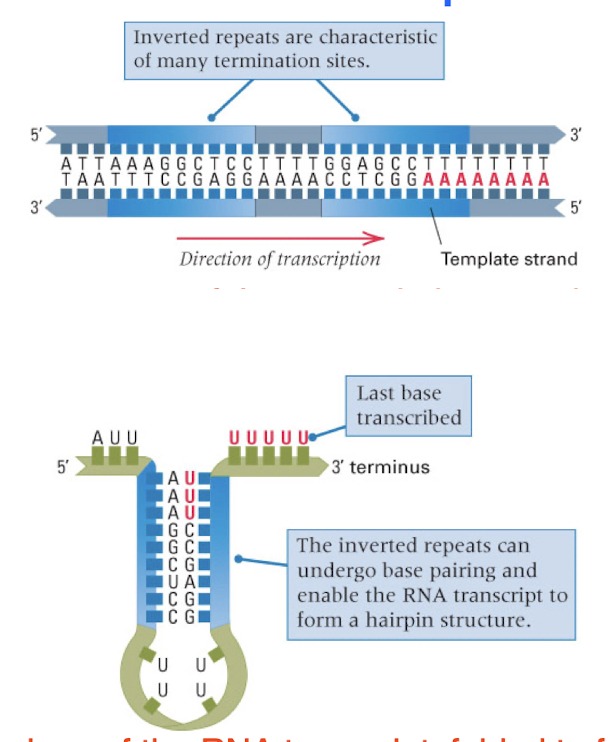
self termination: most common in bacteria, transcription stops when the polymerase encounters a particular sequence of nucleotides
termination requires the presence of a termination protein
self termination sequences contain…
a hairpin and U-rich region
some terminators also require a terminator protein
initiation of a second rounds of transcription…
does not need to wait for the completion of the first
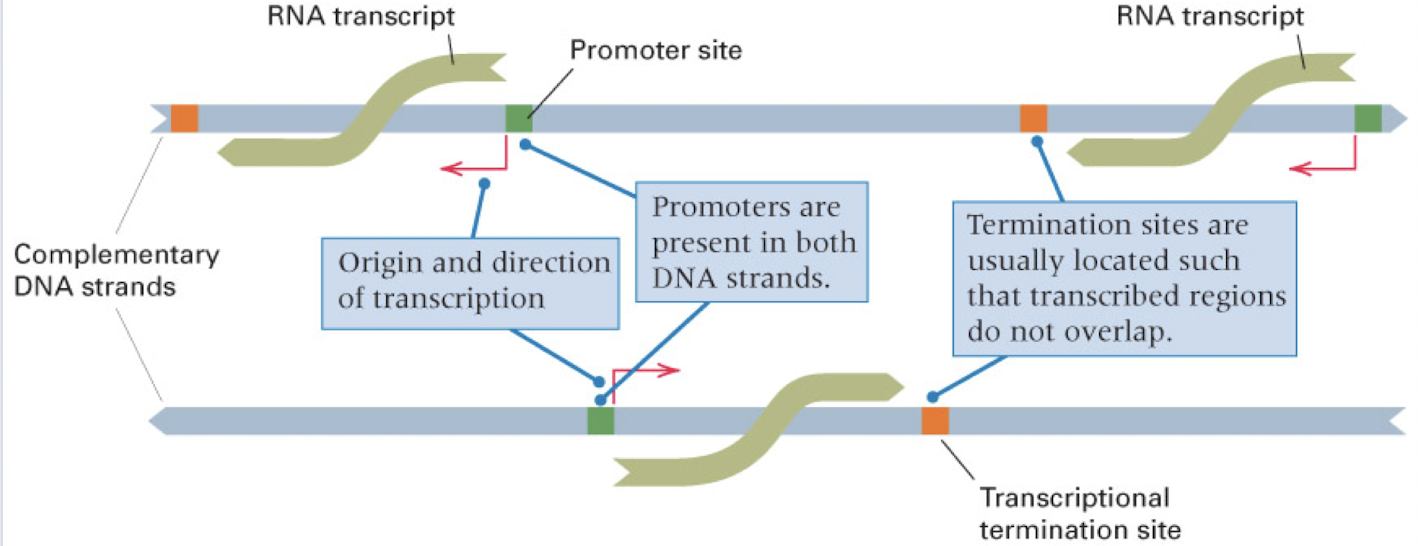
how many DNA strands are transcribed for any one gene?
only one
though, different genes can be transcribed on different strands
in bacteria, transcription and translation are…
coupled
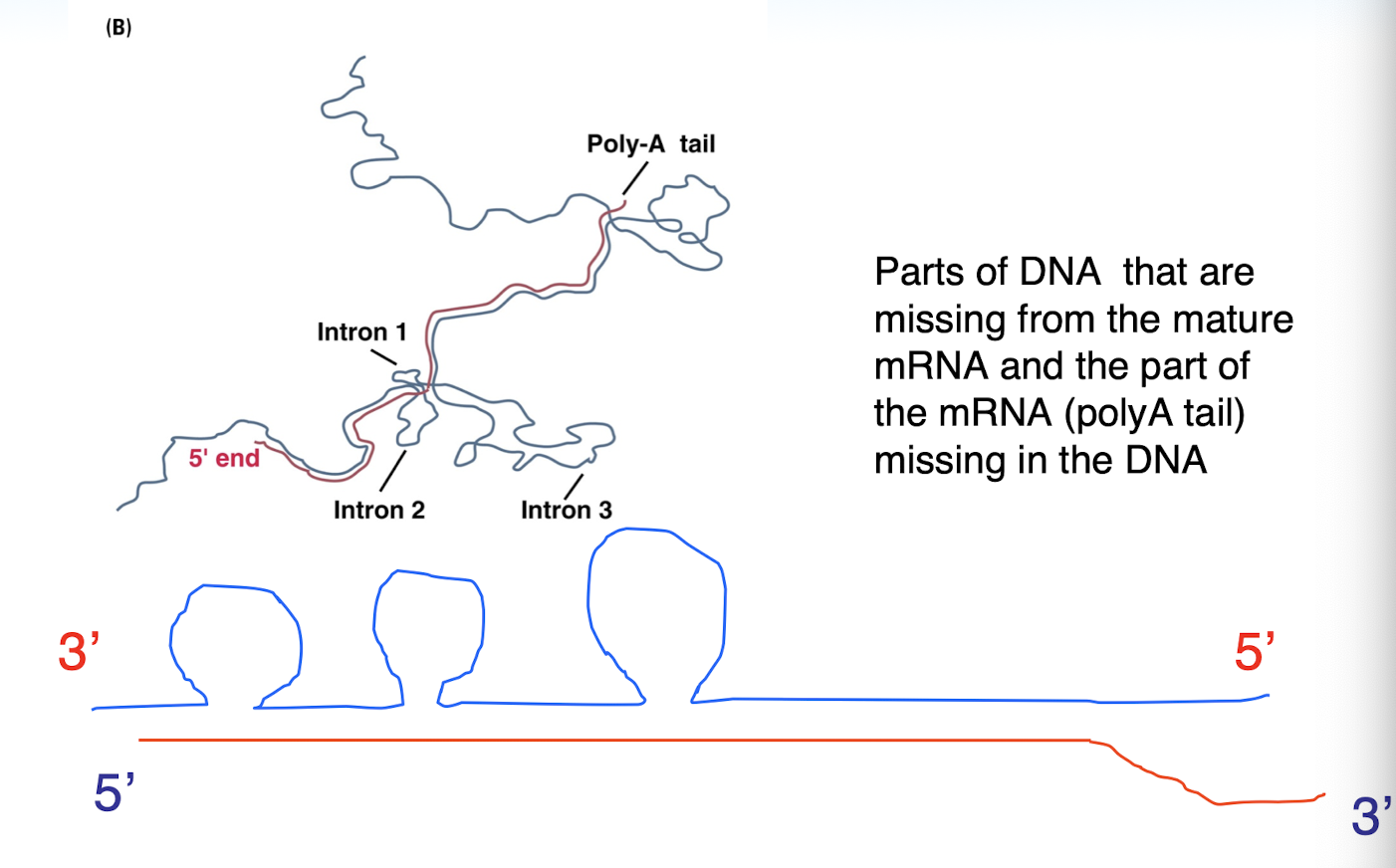
structure of messenger RNA
RNA processing is aided by RNA polymerase itself
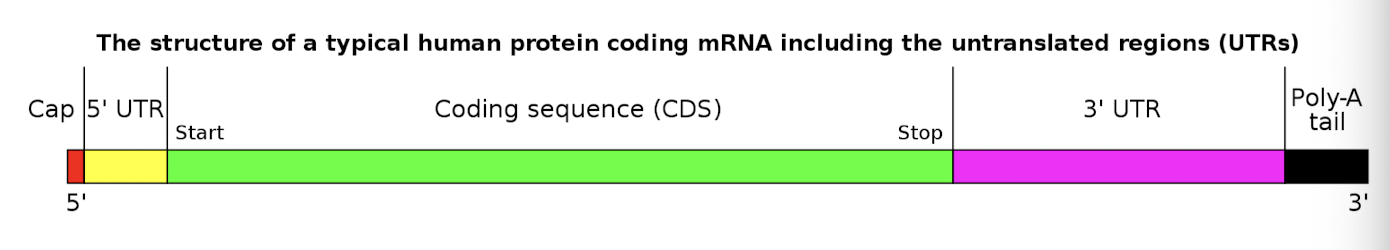

RNA processing
the conversion of a primary transcript into an mRNA or tRNA molecule
includes splicing, cleavage, modification of termini, and (in tRNA) modification of internal bases
prokaryotes: primary transcript is mRNA
eukaryotes: primary transcript must be processed
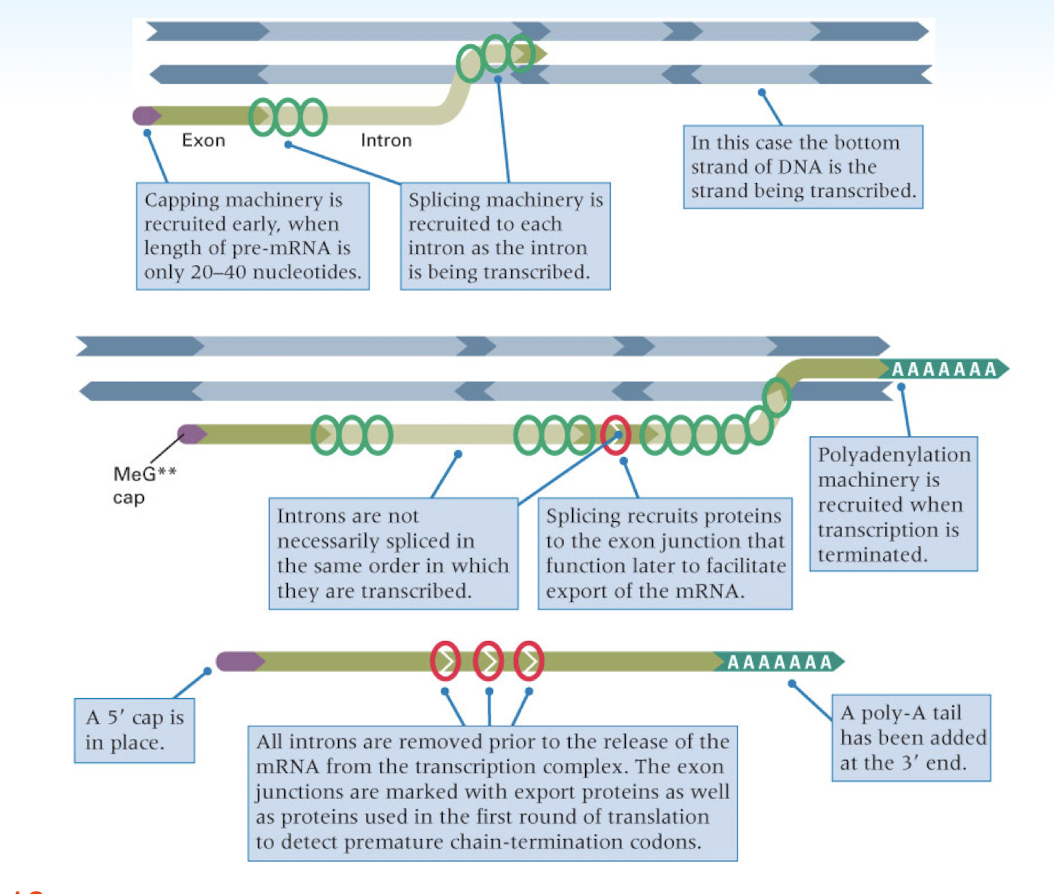
steps of RNA processing
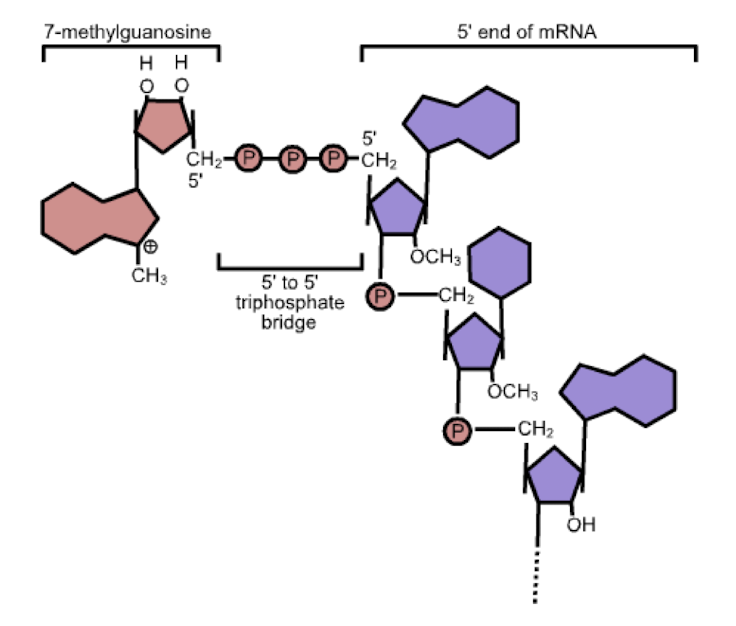
5’ guanine cap- addition of a 7-methylguanosine to the 5’ end of the RNA strand
3’ polyadenylation- addition of ~200 nucleotide polyA tail (stops enzymes from destroying strand in cytoplasm)
splicing- removal of intron and joining of exons
5’ capping functions
regulation of nuclear export
prevention of degradation by exonucleases
promotion of translation (attachment to ribosome)
promotion of 5’ proximal intron excision
snRNPs
small nucleoprotein particles that mediate splicing
RNA splicing takes place in nuclear particles known as spicesomes
snRNPs are composed of proteins and several specialized small nuclear RNA (snRNA) molecules
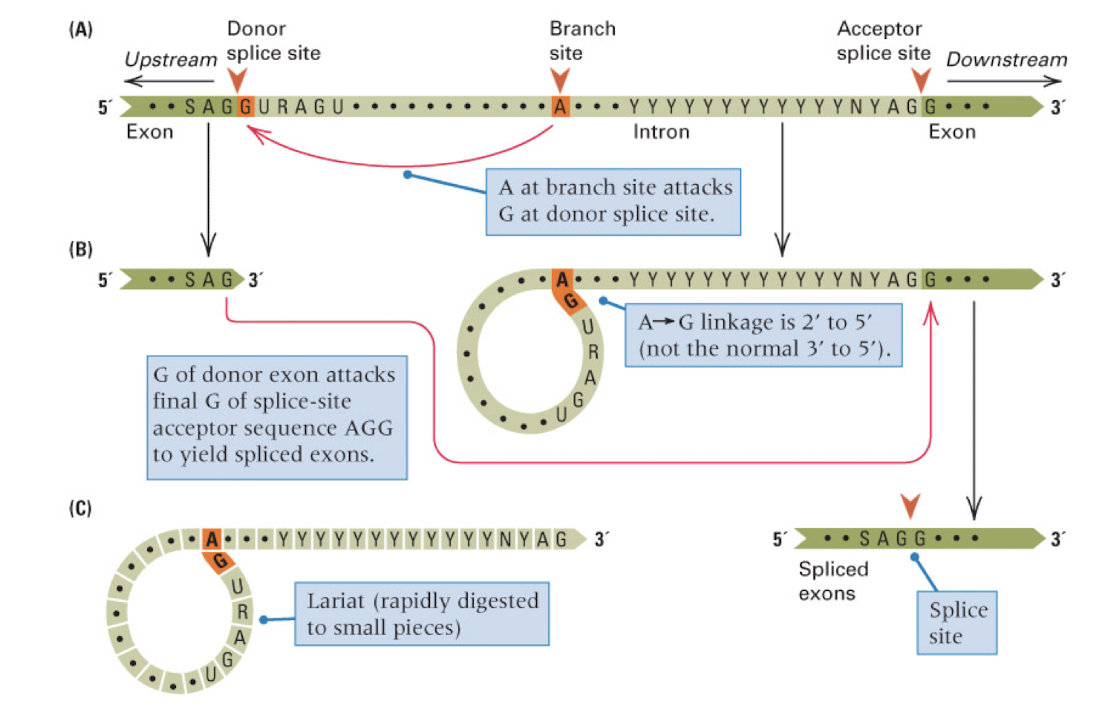
mechanism of RNA splicing
introns have 5’ GU and 3’ AG splice sites and an internal branch site A
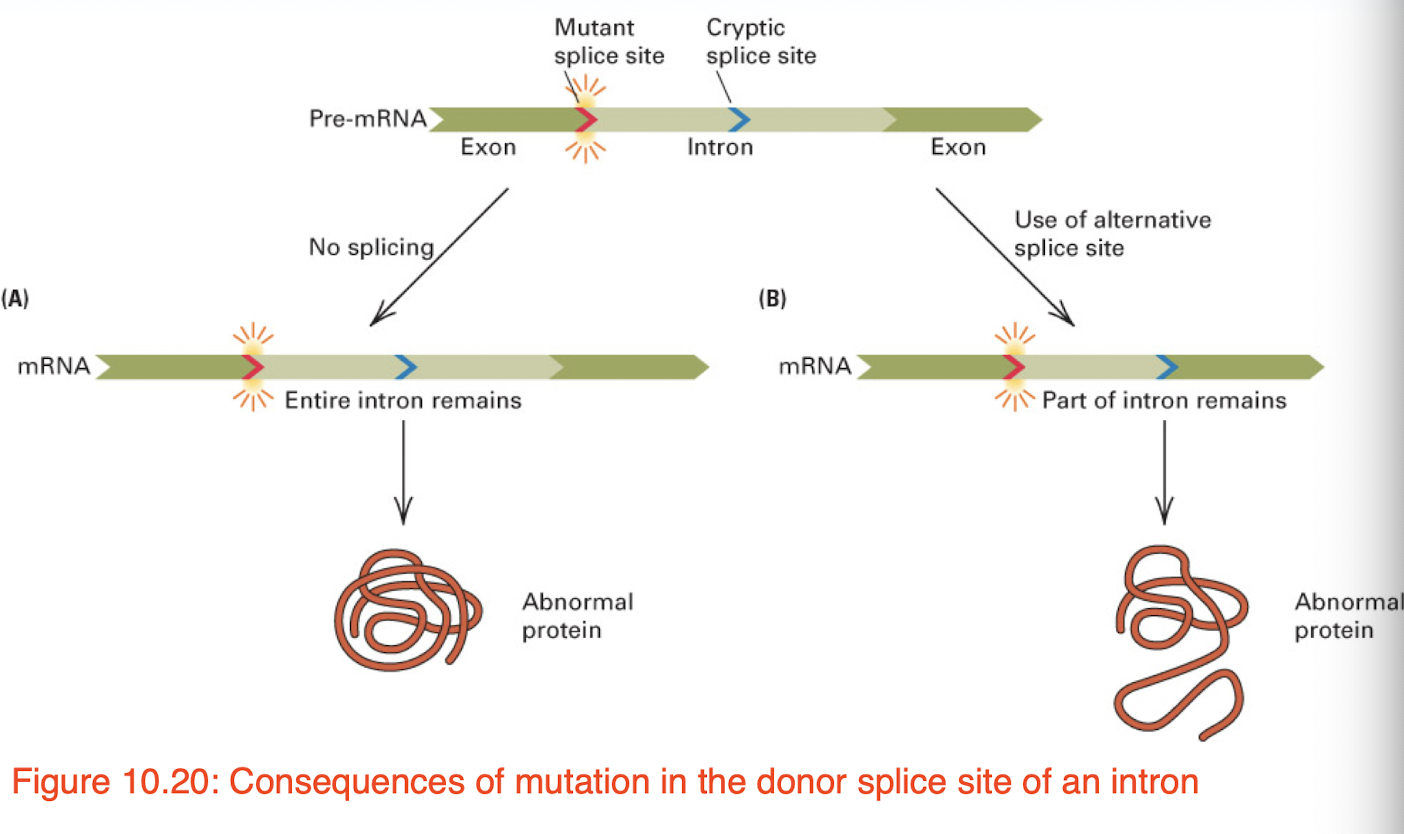
splice site mutations may result in…
the retention of an intron
ribozyme (RNA enzyme or catalytic RNA)
RNA molecules possessing a well defined tertiary structure that enables it to catalyze the chemical reaction in RNA splicing
many natural ribozymes catalyze either the hydrolysis of one of their own phosphodiester bonds or the hydrolysis of bonds in other RNAs
3’ polyadenylation functions
nuclear export
translation
stability of mRNA
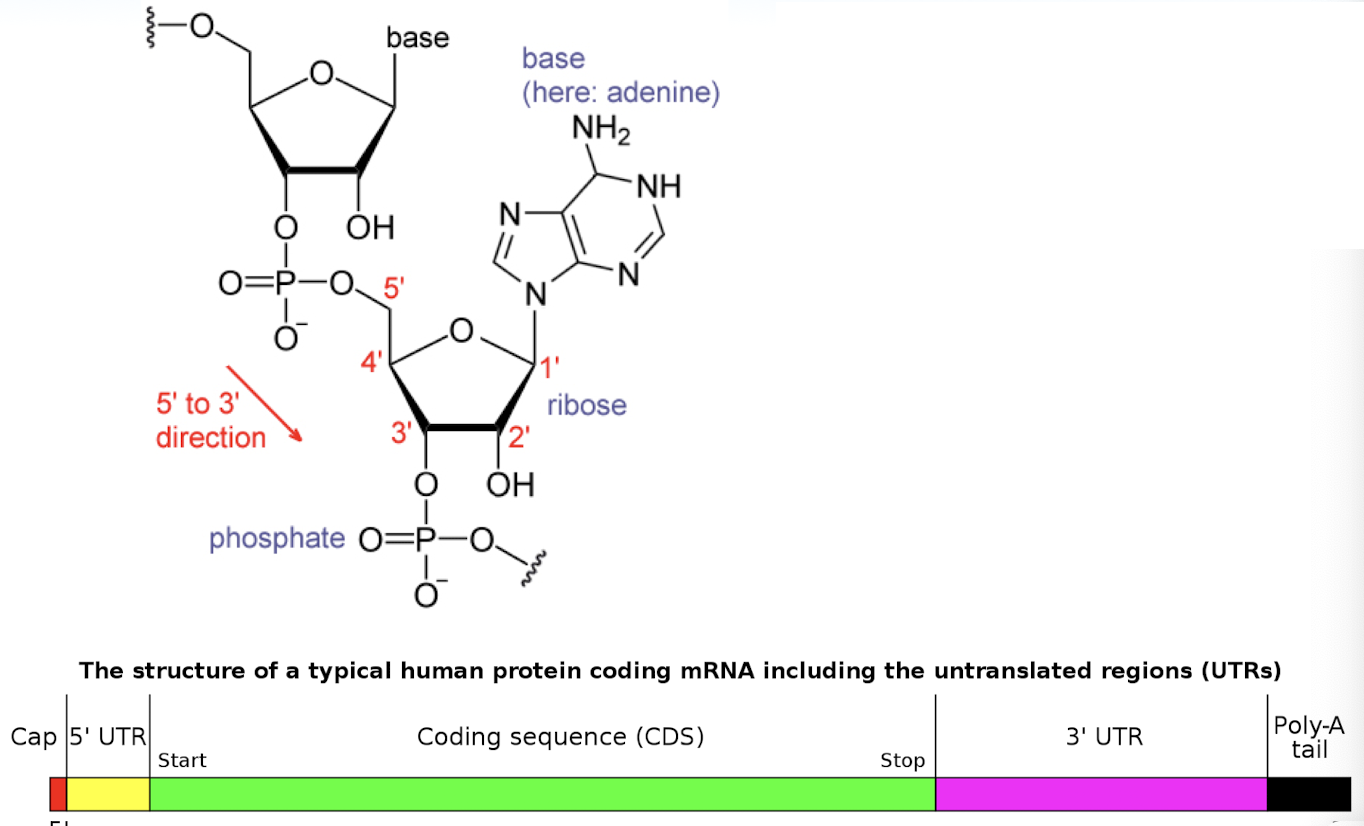
mRNA
provides the nucleotide coding sequence that determines the amino acid sequence
ribosomes
responsible for protein synthesis
tRNA
adaptor molecules that decode the nucleotide sequence to amino acid sequence
aminoacyl tRNA synthetase
attaches an AA to tRNA (based on complimentary anti codon to the codon on mRNA)
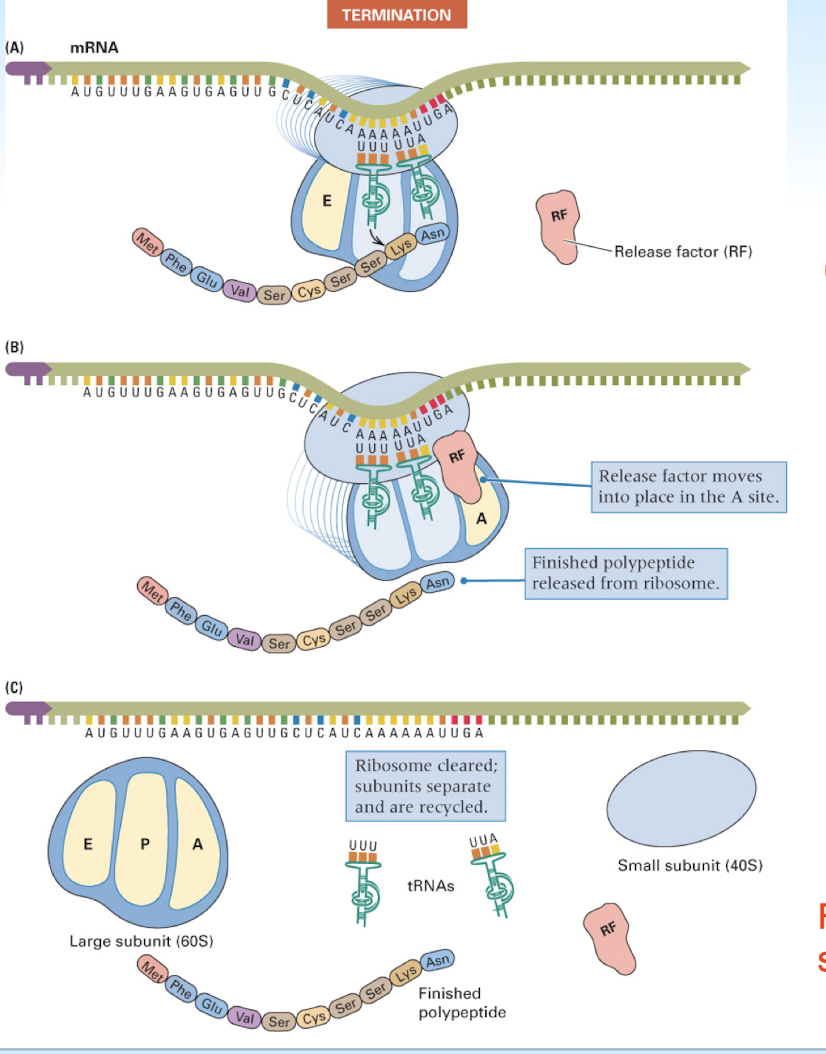
initiation, elongation, and release factors
specialized proteins that aid in each stage of translation
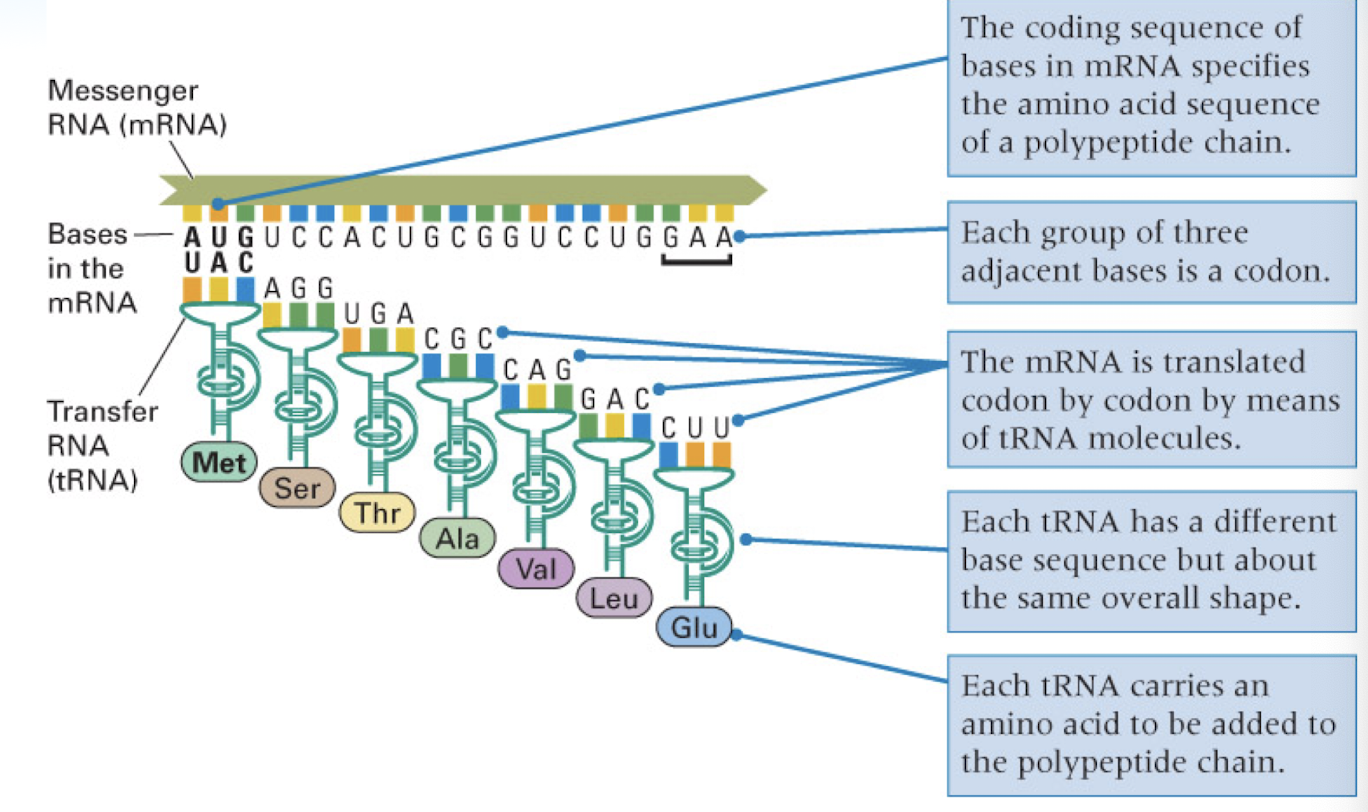
mRNA is translated into protein
structure features of tRNA
the 5’-terminal monophosphate group
the acceptor stem is a 7-base pair stem made by the base pairing of the 5’-terminal nucleotide with the 3’-terminal nucleotide
the CCA tail is important for the recognition of tRNA by enzymes critical in translation
the anticodon arm is a 5-bp stem whose loop contains the anticodon
the D and T arm secondary structures are important for the function of tRNA
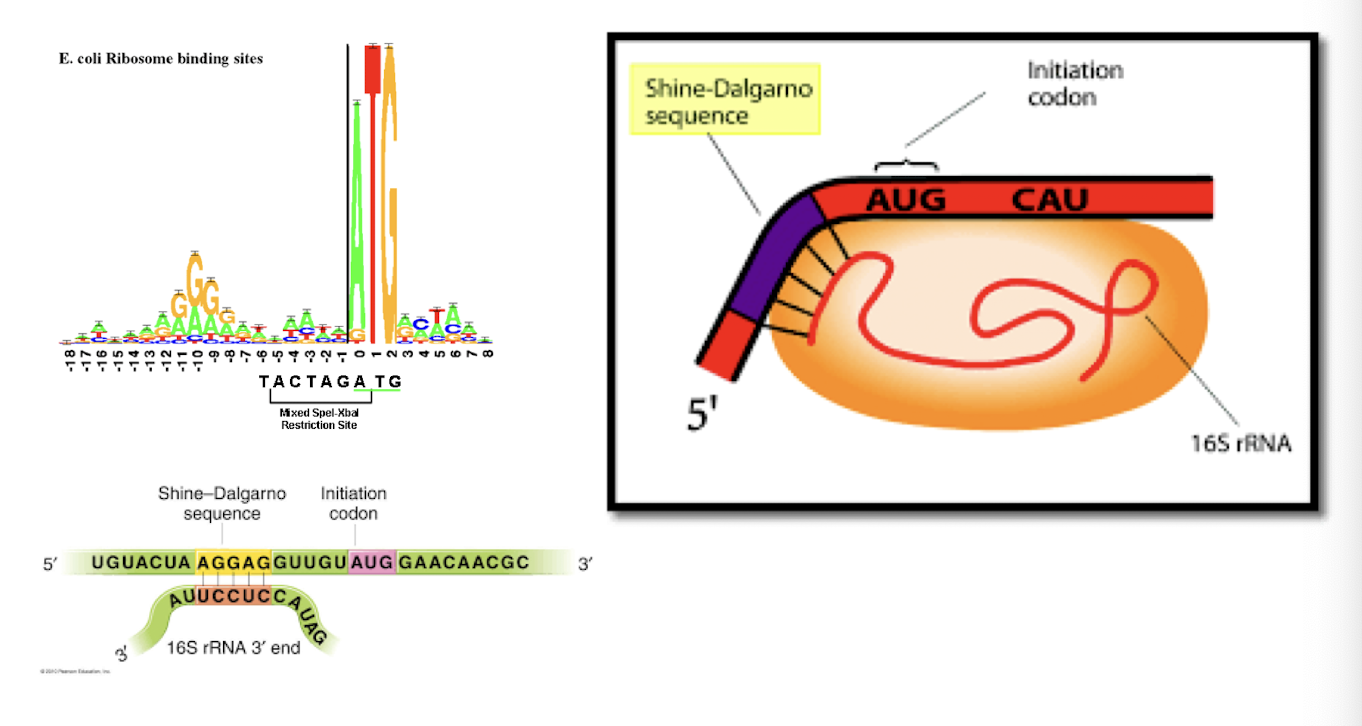
translation initiation in prokaryotes
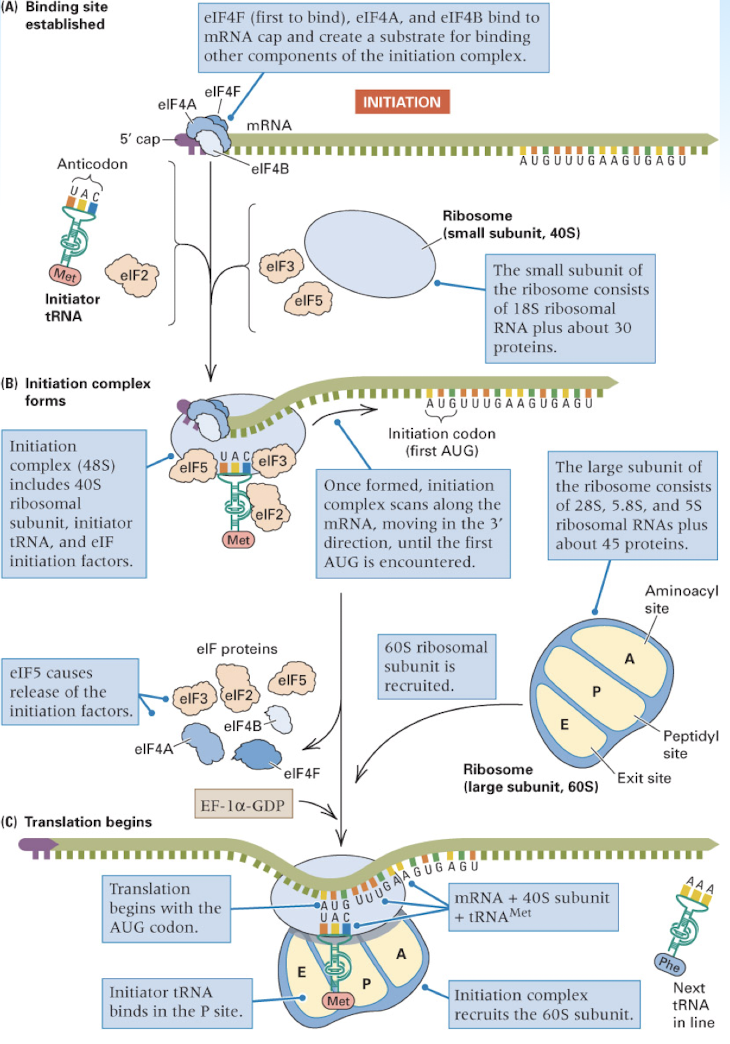
initiation of protein synthesis in eukaryotes
A Site: the ribosomal site which binds the incoming aminoacyl-tRNA (acceptor site)
P Site: the site which holds the peptide tRNA, that is the tRNA which is covalently linked to the growing polypeptide chain (Peptidyl site)
E Site: a site which transiently binds to the outgoing, deacylated tRNA (exit site)
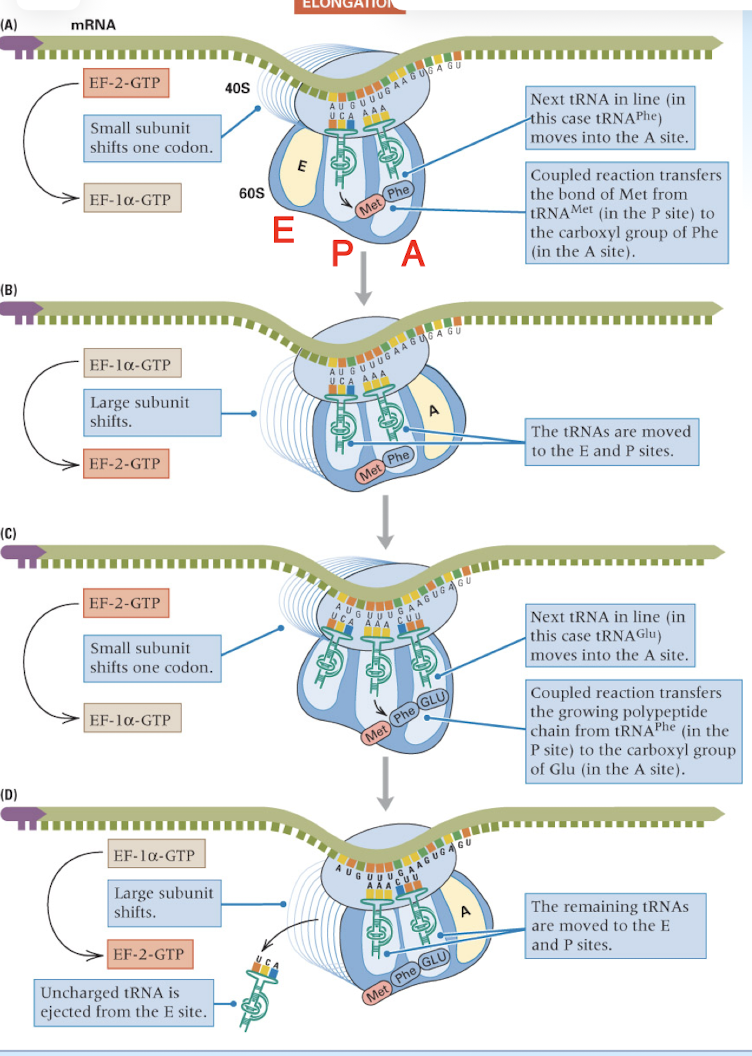
elongation is a repeated cycle of three processes
bringing each aminoacylated tRNA into line
forming the new peptide bond to elongate the polypeptide
moving the ribosome to the next codon along the mRNA
translation termination occurs when…
a stop codon is reached
transcription and translation are ______ in bacteria
coupled
in eukaryotes, transcription takes place in the nucleus and translation occurs in the cytoplasm
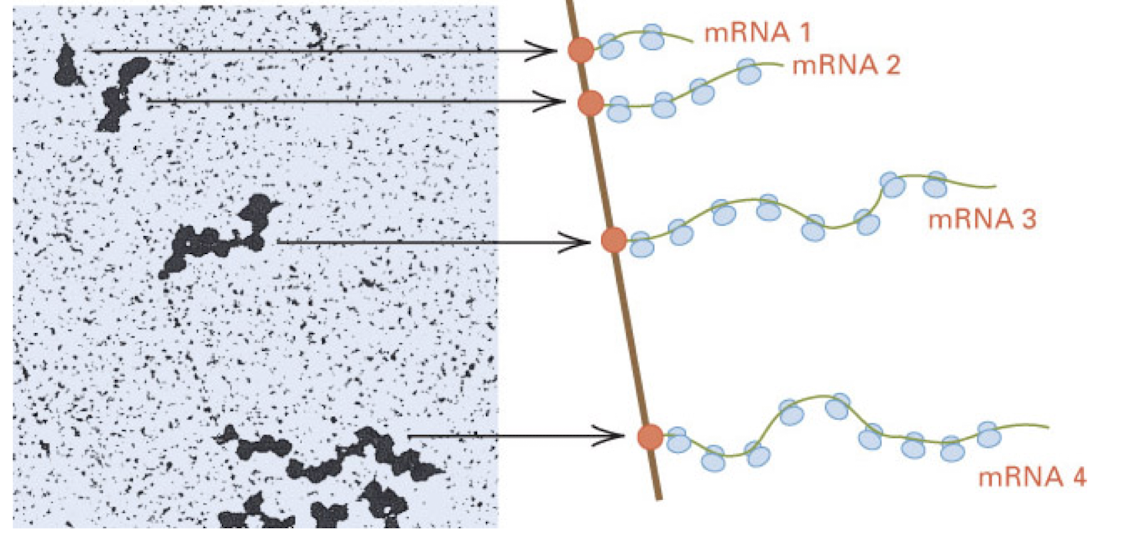
prokaryotic ribosomes can initiate at ____________ in a mRNA
internal sites
prokaryotic mRNAs- can be polycistronic
ribosomes can initiate translation within an mRNA, therefore making multiple polypeptides
eukaryotic mRNAs- are only monocistronic
ribosomes must initiate translation at the 5’ end, therefore only one polypeptide can be made at a time
translation in prokaryotes vs eukaryotes
transcription and translation
prokaryotes: simultaneous and colocalized
eukaryotes: separate (nucleus and cytoplasm)
initiation
prokaryotes: f-met
eukaryotes: Met
ribosome binding
prokaryotes: SD dequence
eukaryotes: 7 me-G Cap
polycistronic mRNA
prokaryotes: common
eukaryotes: rare or nonexistent
RNA processing
prokaryotes: no
eukaryotes: yes (5’ G Capping, 3’ PolyA tail, splicing)
primary protein structure
sequence of residues making up the protein (chain of amino acids)
secondary protein structure
refers to local sub-structures: alpha helix and beta strand or beta sheets (initial folding of AA chain)