Biology of Cancer Exam 2 (Ch.6- and JC3-)
1/74
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
75 Terms
Cell Cycle
• The cell cycle divides the life cycle of all cells into stages
• M -Phase includes each stage of mitosis in which one cell divides into two
• Separation of replicated DNA
• Cytokinesis to separate two new cells
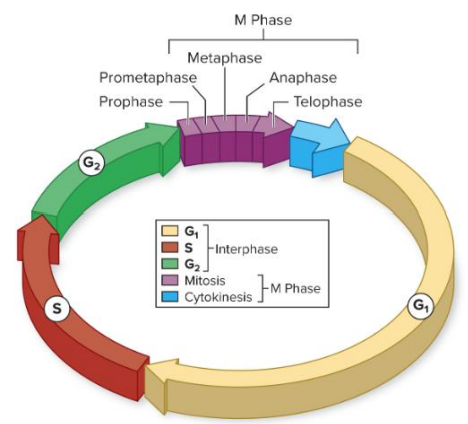
Interphase
*Interphase contains the remaining stages of the cell cycle
-Gap phase 1 (G1): Primary growth phase and the longest phase for most cell types
- Synthesis (S) phase: Phase of DNA synthesis
- Gap phase 2 (G2 ): Organelles undergo replication and cytoskeleton organization prior to mitosis
G1 ,G2 , G0 ,S, and M phase
• Gap phase 1 (G1): Primary growth phase and the longest phase for most cell types
• Synthesis (S) phase: Phase of DNA synthesis
• Gap phase 2 (G2): Organelles undergo replication and cytoskeleton organization prior to mitosis
• M -Phase includes each stage of mitosis in which one cell divides into two
>Separation of replicated DNA
>Cytokinesis to separate two new cells
• G0 phase • Temporary or permanent exit from the cell cycle
> Many of our body’s mature cells perform their normal cell functions during G0 phase
>Stimuli (e.g. growth factors) can stimulate a G0 cell to re-enter the cell cycle
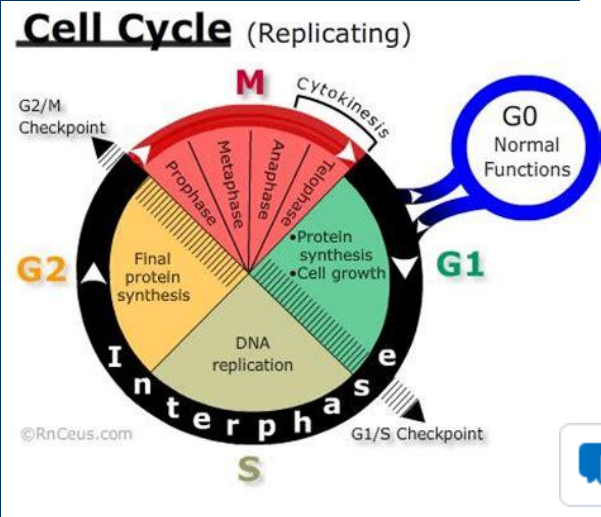
G1 /S Checkpoint
G1 /S Checkpoint
• Growth factors from other cells
• Nutritional state of the cell
G1 /S Checkpoint
• Commitment step to going through the cell cycle
• Influenced by growth factors from other cells
• Occurs just prior to DNA synthesis (and later cytokinesis)
• Assesses the availability of energy and raw materials
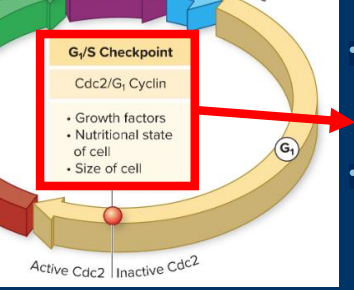
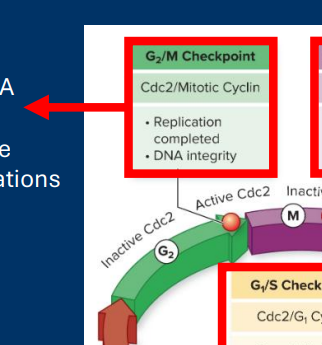
G2 /M Checkpoint
G2 /M Checkpoint
• DNA integrity
G2 /M Checkpoint
• Occurs after error-prone DNA synthesis
• Checks for DNA damage before “locking in”mutations via mitosis
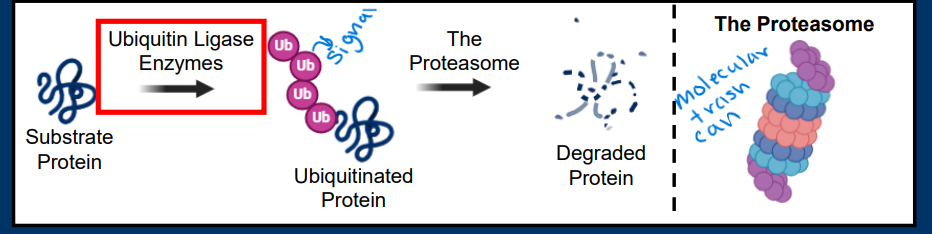
Spindle Checkpoint
• Spindle Checkpoint (Metaphase)
• Chromosome alignment
Spindle Checkpoint
• Occurs just prior to chromatid separation in anaphase
• Ensures that chromosomes are perfectly separated into new cells
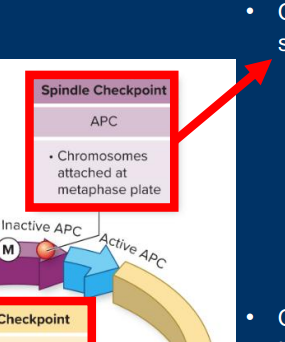
Cyclins (A, B, D, E)
• Each of the four cyclin proteins displays a characteristic pattern of expression throughout the cell cycle
• Cyclin D: Gradual up and downregulation through cell cycle
• Cyclin E: Peaked expression during G1 /S transition
• Cyclin A: Peaked expression in G2 phase
• Cyclin B: Peaked expression at G2 /M transition
*Be able to label a figure with each of the cyclins
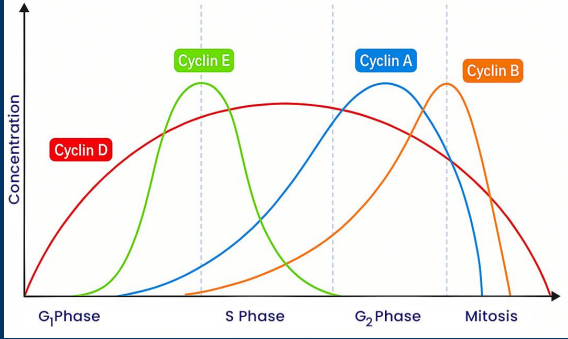
CDKs
• Progression through each checkpoint is regulated by one of many cyclin-dependent kinases (CDKs)
• CDKs enzymatic activity is inhibited unless they interact with a specific cyclin protein
- Examples = Cyclin A, Cyclin B, Cyclin D, and Cyclin E
• Different cyclins are expressed in unique times during the cell cycle. Therefore, different CDKs become active during the cell cycle
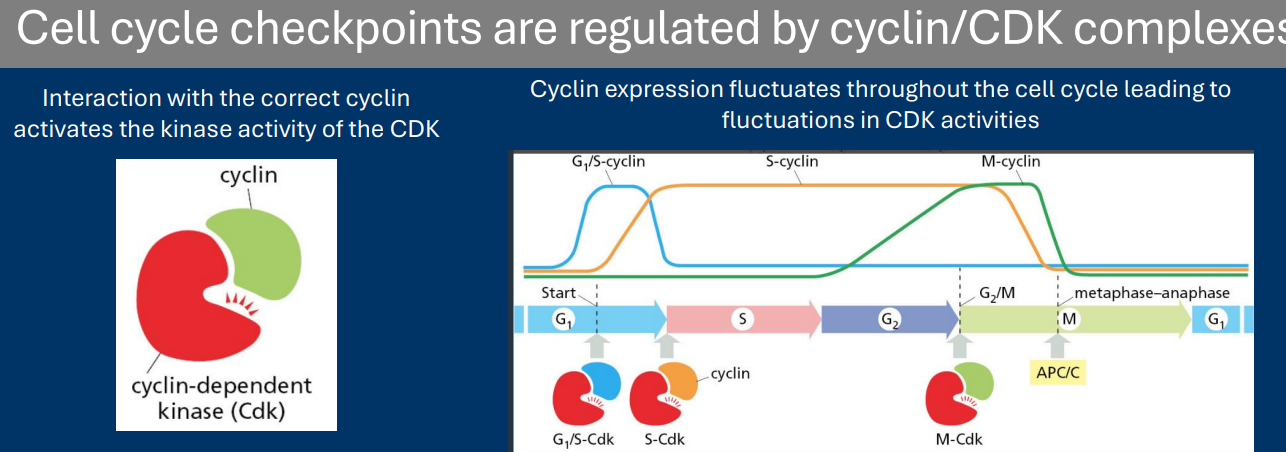
Ubiquitin/Proteasome System
• The ubiquitin/proteasome system (UPS) is a major pathway responsible for regulating protein degradation
• Two primary components:
- Ubiquitin ligases are enzymes that add poly-ubiquitin post-translational modifications to proteins
- Poly-ubiquitination of a protein leads to its transport to the proteasome, a cellular “trash can” that chews up the protein into free amino acids
• Ubiquitin ligases determine which proteins are targeted for degradation
(Protein degradation by the Ubiquitin/Proteasome System)
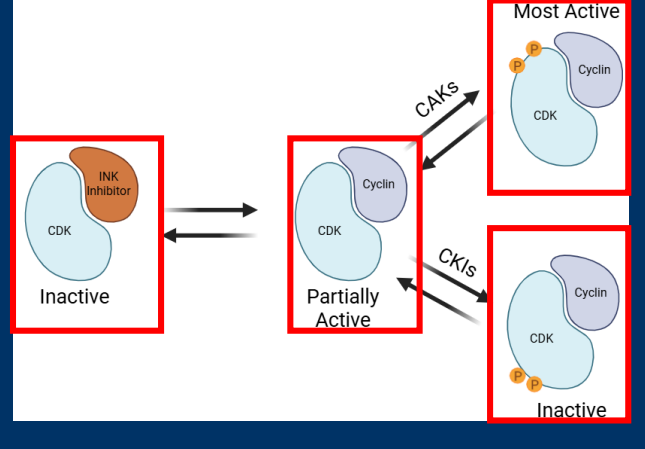
CAKs/CKIs
• Post-translational modifications alter CDK enzyme activity
• Post-translational modifications with activating phosphorylation by CDK activating kinases (CAKs)
• Post-translational modifications with inhibitory phosphorylation by CDK inhibitor kinases (CKIs)
-CDK enzyme activity (kinase) is regulated in various ways
-Regulation of CDK activity by interaction and phosphorylation
Retinoblastoma
• Cyclin D/CDK regulates cell cycle progression via the Retinoblastoma Protein (RB)
- RB = Important tumor suppressor gene
• In the absence of Cyclin D/CDK, RB binds to and inhibits E2F family transcription factors
- RB also recruits HDACs to silence gene expression
• Cyclin D/CDK phosphorylate RB and this causes a conformational change the releases E2F and HDAC
• E2F regulates the expression of many genes including Cyclin E and gene needed for DNA Synthesis
(Cyclin D (and E) regulate the retinoblastoma (Rb) transcription factor)
(Retinoblastoma suppresses the expression of E2Fregulated genes until phosphorylated by cyclin D/CDK)
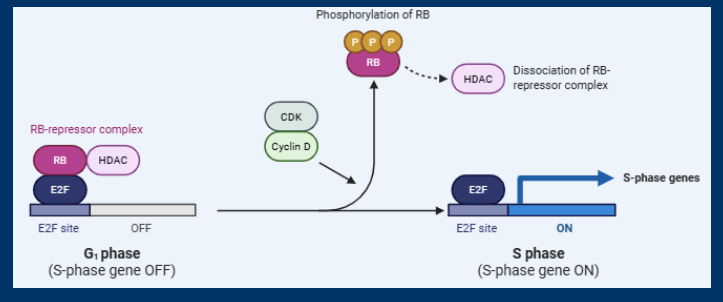
E2F Transcription Factors
• In the absence of Cyclin D/CDK, RB binds to and inhibits E2F family transcription factors
• Cyclin D/CDK phosphorylate RB and this causes a conformational change the releases E2F and HDAC
• E2F regulates the expression of many genes including Cyclin E and gene needed for DNA Synthesis
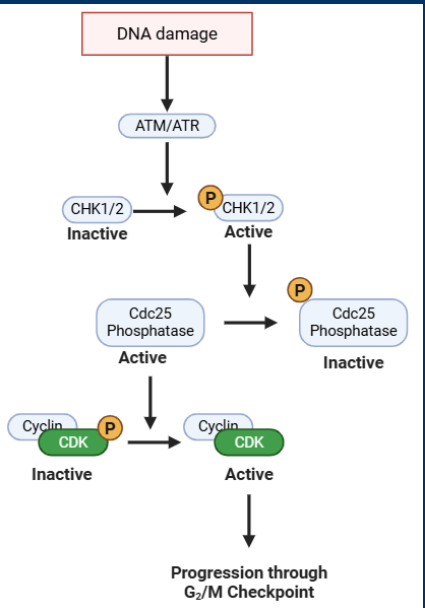
ATM/ATR
• The G2 /M checkpoint is regulated by two kinases ATM/ATR
> Kinase function activated by DNA damage
• Activated ATM/ATR phosphorylate the kinases CHK1/2
> CHK1/2 phosphorylation = activation
• CHK1/2 phosphorylate a phosphatase (Cdc25)
> Cdc25 phosphorylation = inactivation
• Cdc25 activates a cyclin B/CDK complex by removing and inhibitory phosphorylation
• End result: DNA damage = no progression through G2 /M
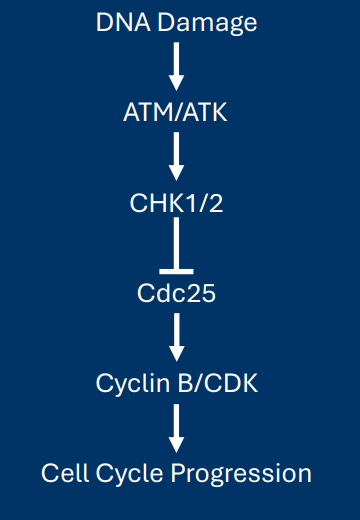
Cause→ Mechanism→ Effect ATM and ATR
Cause – DNA damage causes enzymatic activation of two kinases: ATM and ATR
Mechanism – ATM/ATR activation sets off a signal cascade:
• ATM/ATR phosphorylate/activate the kinases CHK1/2
• CHK1/2/ phosphorylate/inactivate the phosphatase Cdc25
• Inactive Cdc25 can no longer activate cyclin B/CDK
Effect – Cell cycle progression stops when DNA damage is present
Critical thinking questions
1.) How can the context of each cell cycle checkpoint tell us about the factors by which it is regulated?
2.) What are the different modes of regulation that are used for cyclins verses cyclin dependent kinases?
3.) What proteins discussed today will act like a tumor suppressor? Which will act like a proto-oncogene?
4.) How can the expression patterns for cyclins tell us about which cell cycle checkpoints they regulate?
1.) What is necessary as in for example G2/M checkpoint is the last checkpoint before mitosis which means there must be a DNA damage check or it will “lock” in mutations via mitosis
2.) Cyclin expression is regulated. Cells produce/degrade the protein to control how much is present (regulated during the cell cycle → Cell pathways make more or less of the protein at key points in the cycle)
CDK activation is regulated. Cells continually express the same amount of protein. Cyclin interactions and post-translational modification affect CDK activity
(The total amount of CDK protein remains the same. Activity is regulated by the presence of cyclins or post-translational modifications)
3.) Retinoblastoma Protein (RB)→ Tumor suppressor gene
Potentially cdks as protooncogenes?
4.)Cyclin expression patterns act as temporal markers for cell cycle checkpoints because their concentrations rise and fall, peaking only when specific phases or transitions are required. By mapping when a particular cyclin-CDK complex is synthesized and subsequently degraded, researchers can identify which checkpoint that specific complex regulates.
Critical thinking questions
5.) What series of events connects RTK activation to the transcription of genes needed for DNA synthesis?
6.) During the G2M checkpoint, what instances of phosphorylation lead to enzyme activation? What instances of phosphorylation lead to enzyme inhibition?
7.)How does the cell “recognize” the presence of unattached chromosomes?
8.) What are two ways in which the activation of APC leads to cell cycle progression?
5.) RTK autophosphorylation sets off a signal cascade of:
• MAPK-dependent AP-1 activation
• Cyclin D expression leading to CDK activation
• Phosphorylation of RB leading to release of E2F inhibition
• Transcription of DNA Synthesis genes
6.)
• CHK1/2 phosphorylation = activation
• Cdc25 phosphorylation = inactivation
7.)Unattached kinetochores are “recognized” by the presence of Mad/Bub proteins (Mad/Bub bound to unattached kinetochores)
8.)APC causes the ubiquitination of two important things:
• Degradation of securin activates the enzyme separase which separates sister chromatids
• Degradation of cyclin A and cyclin B
If were talking about”Targeted disruption of Skp2 results in accumulation of cyclin E” what cell cycle checkpoint is relevant?
G1S
The _________ of cyclins is controlled by transcriptional mechanisms
The upregulation of cyclins is controlled by transcriptional mechanisms
*Cyclin expression is regulated by transcription and degradation
E2F transcription factors promote Cyclin ____ expression
E2F transcription factors promote Cyclin E expression
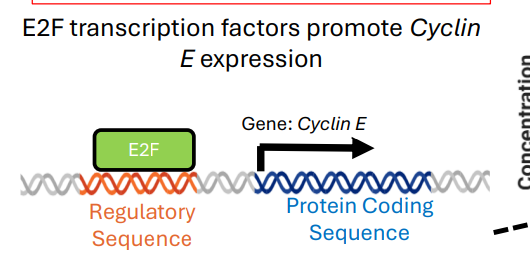
The __________of cyclins is controlled by proteolytic mechanisms
The downregulation of cyclins is controlled by proteolytic mechanisms
Cyclin E expression is controlled by two sequential pathways of transcriptional regulation
1) Growth factor-dependent MAPK activation leading to AP-1 transcriptional activation
• Cyclin D is a transcriptional target of AP-1
2) Cyclin D/CDK-dependent inactivation of RB leading to transcriptional activation of E2F transcription factors
• Cyclin E is a transcriptional target of E2F
Which cyclin is upregulated first? Cyclin D or E?
Cyclin D (as it is upregulated it phosphorylates RB which upregulates cyclin E)
Protein degradation by the UPS regulates the cell cycle in two ways. Both are exemplified by the APC ubiquitin ligase
• Degradation of cyclins
• Degradation of proteins with functions needed for checkpoints (e.g. Securin/Separase)
The SCF ubiquitin ligase regulates cell cycle progression
Made up of three primary components
• Skp1: An adaptor protein that connects the other two components
• Cullin: A protein that recruits ubiquitin to the complex
• F-box proteins: One of very many proteins that are responsible for SCF substrate specificity
The human genome contains many F-box proteins
• “F-box” domains interact with Skp1
• Other domains interact with specific cellular proteins
* Which F-box protein is bound to the SCF complex determines which proteins will be degraded
Immunoprecipitation
Immunoprecipitation is a common method for determining if two proteins physically interact in the cell
Western blot is used to determine which other proteins coimmunoprecipitated with the target protein
End of Journal club #3
End of Journal club #3
PI3K
PI3K signaling regulates cellular metabolism
• Activation of membrane RTKs leads to the activation of the kinase PI3K (phospho-inositol3-kinase)
• PI3K is a lipid kinase
- Phosphorylates PIP2 to PIP3
• In this system, PI3K is acting as an proto-oncogene (Activation leads to pathways necessary for cell division)
• The phosphatase PTEN counteracts PI3K by removing phosphate from PIP3
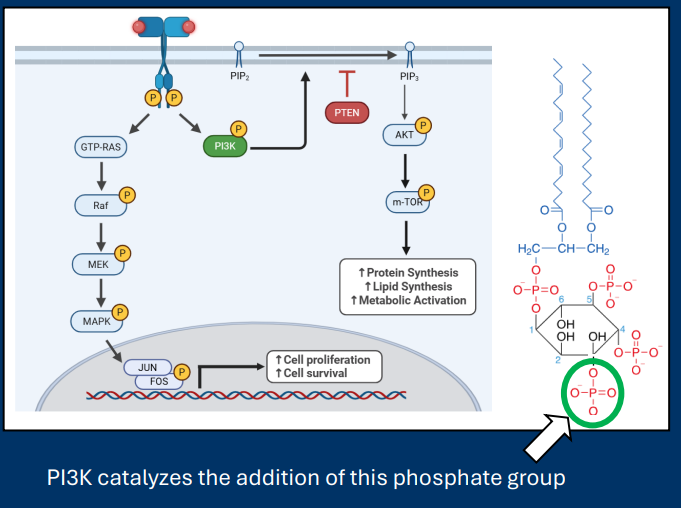
PIP2 /PIP3
• PIP3 leads to the activation of m-TOR
>Promotes anabolic metabolism to produce components for cell division
• PIP2 /PIP3 is phosphorylated by PI3K
• The phosphatase PTEN counteracts PI3K by removing phosphate from PIP3
PTEN
• The phosphatase PTEN counteracts PI3K by removing phosphate from PIP3
>PTEN functions as a tumor suppressor gene
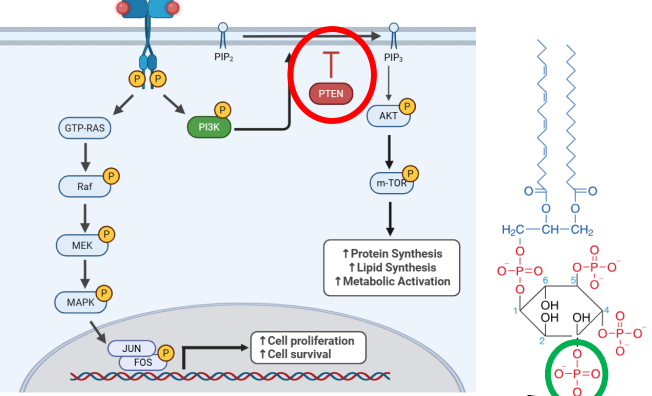
m-TOR
• PIP3 leads to the activation of m-TOR
• Promotes anabolic metabolism to produce components for cell division
*Master Regulator of metabolism
-Says make more
p53
• p53 is one of the most well studied tumor suppressor genes
-Altered in almost all human cancers
• p53 expression is regulated by the ubiquitin-proteasome system
• MDM2 is the ubiquitin ligase responsible for p53 ubiquitination and degradation
• Activators of p53 interfere with p53/MDM2 interaction
- This prevents p53 degradation and leads to more p53 expression
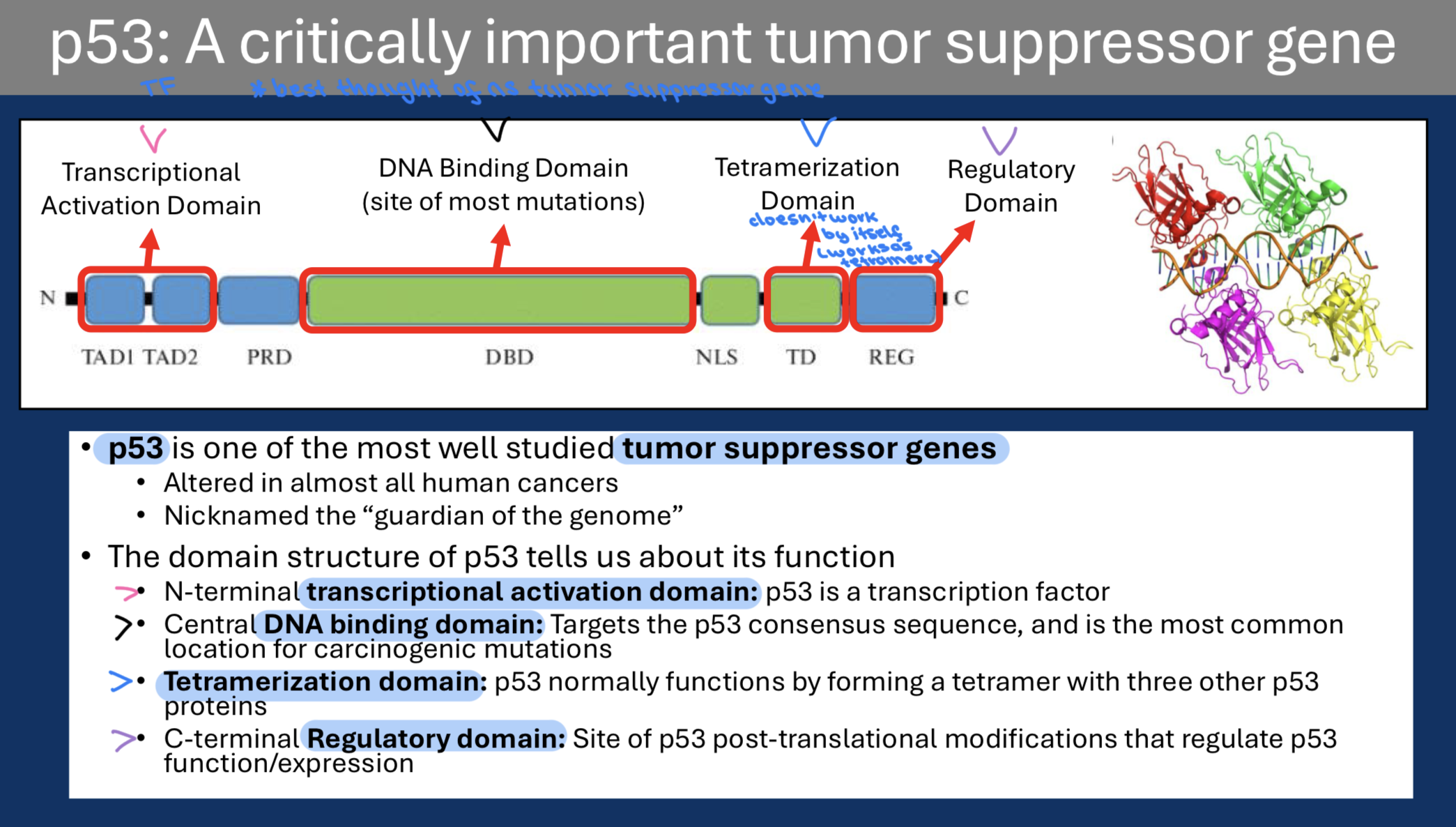
Two examples of p53 activation:
• DNA Damage: ATM and CHK2 kinases phosphorylate p53 and this modification prevents interaction with MDM2
• Oncogene activation: Constitutive RAS activation leads to modifications of MDM2 that prevent interaction with p53
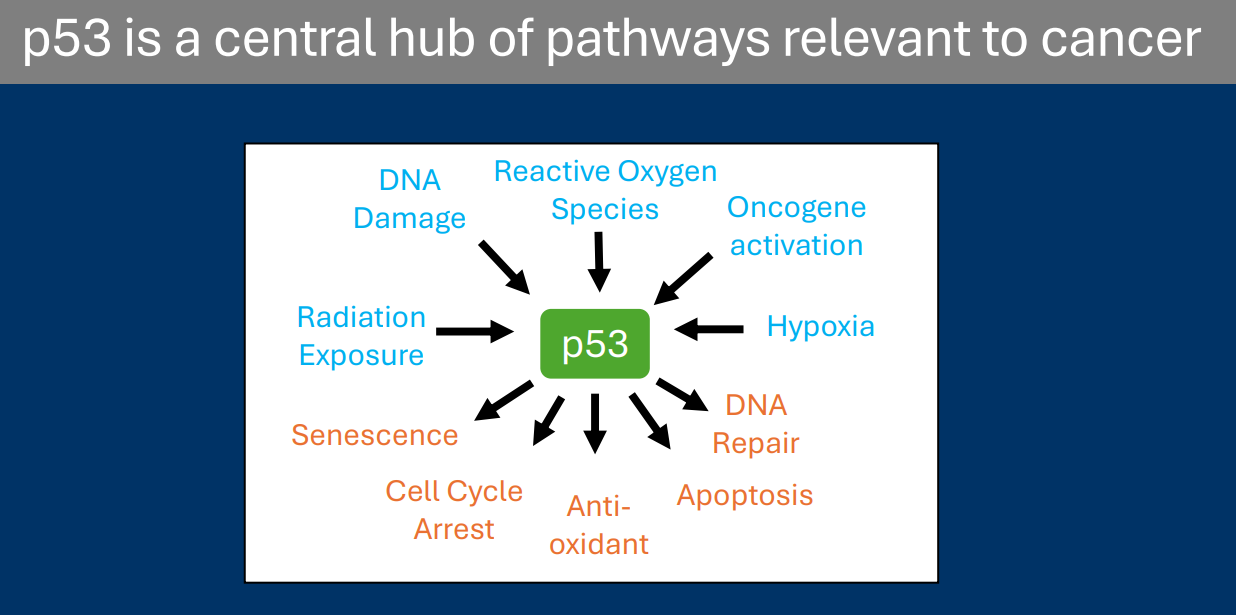
The domain structure of p53 tells us about its function
• N-terminal transcriptional activation domain: p53 is a transcription factor
• Central DNA binding domain: Targets the p53 consensus sequence, and is the most common location for carcinogenic mutations
• Tetramerization domain: p53 normally functions by forming a tetramer with three other p53 proteins
• C-terminal Regulatory domain: Site of p53 post-translational modifications that regulate p53 function/expression
MDM2
Regulation of p53 expression by MDM2
• MDM2 is the ubiquitin ligase responsible for p53 ubiquitination and degradation
p53 and MDM2 are part of an auto-regulatory feedback loop
• p53 promotes the expression of the MDM2 gene
• MDM2 promotes p53 degradation
(Regulators of p53 disrupt the p53/MDM2 interaction)
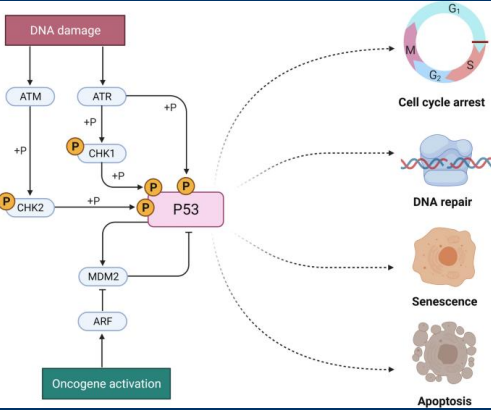
Reactive Oxygen Species
• Metabolic pathways generate dangerous intermediates known as reactive oxygen species (ROS)
• If not removed, ROS will go on to cause cellular damage including damaging DNA
• p53 promotes the expression of antioxidant genes that scavenge ROS
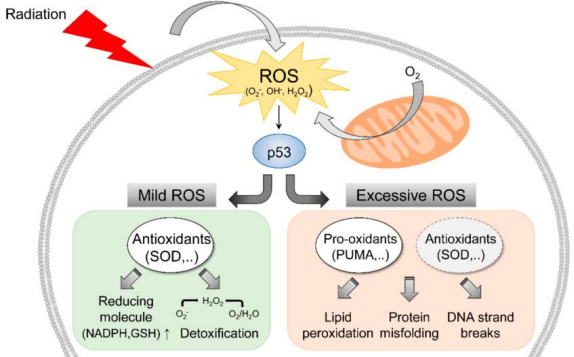
Tumor Suppressor Gene
Context matters for defining tumor suppressor genes
• Healthy cells: Tumor suppressors play essential roles to regulate cell division, DNA repair, apoptosis, etc.
• Cancer cells: Mutation to tumor suppressor genes impairs gene/protein function leading to one of many aspects of carcinogenesis
Tumor suppressor genes are often considered in relation to the cell cycle
• Non-mutated tumor suppressors reduce progression through the cell cycle
• Mutated tumor suppressors fail to reduce cell cycle progression
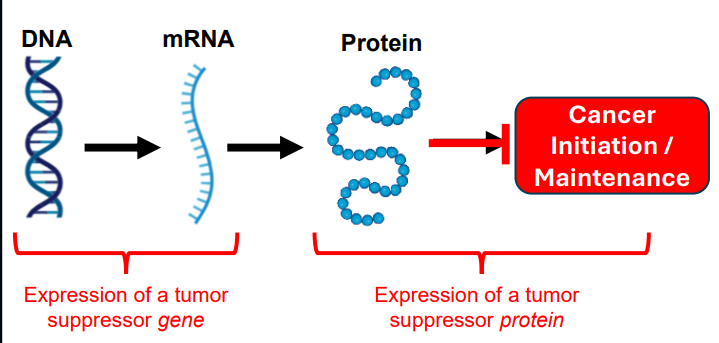
Sporadic Retinoblastoma
• Two normal copies of Retinoblastoma gene at birth
• Sporadic mutations must occur twice to delete both copies
• Odds of two mutations in the same cell are very low (~ 108 retinoblasts per eye)
• Sporadic retinoblastoma is almost always restricted to one eye
• Sporadic ~60% of cases
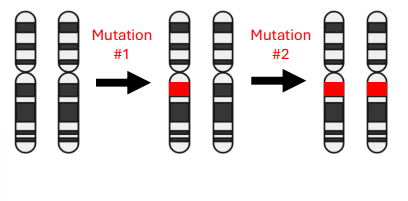
Familial Retinoblastoma
• Born with one mutated copy of Retinoblastoma gene
• Only one additional mutation is required to develop the disease
• Odds of developing mutation twice in two different cells is much higher
• Familial retinoblastoma is generally bilateral (both eyes)
• Familial ~40% of cases
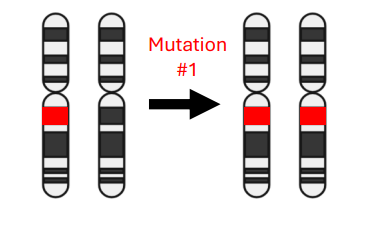
Two-Hit Hypothesis
• Two-Hit Hypothesis: Tumor suppressor genes typically require both alleles to be inactivated for cancer to develop
• Cancer susceptibility genes make carriers heterozygous for tumor suppressor genes
• Two important considerations:
1) Germline mutations mean an individual only needs one sporadic event for carcinogenesis
2) Susceptibility genes can be in DNA repair pathways, so the loss of one allele can make it easier to develop secondary mutations
• The Two-Hit Hypothesis is a helpful learning tool, but many exceptions can be found
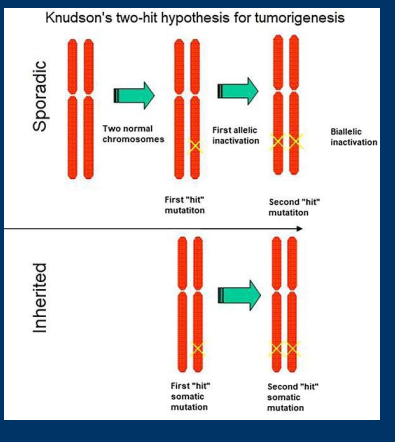
Haploinsufficiency
Haploinsufficiency: Genes for which loss of a single allele is sufficient to cause disease
Dominant negative effects
Dominant negative effects: Mutation of one allele creates a protein product that interferes with the remaining wild type allele
Gain of function mutations
Gain of function mutations: Mutations that give a protein novel functions that can cause disease
Critical Thinking Questions
1.) How can things like upstream causes and downstream effects identify a protein as a tumor suppressor gene?
2.) What are common upstream causes of tumor suppressor gene activation?
3.) What are common downstream effects of tumor suppressor gene activation?
4.) How does the “Two-Hit Hypothesis” explain why familial retinoblastoma is generally bilateral, but not sporadic retinoblastoma?
5.) How do the concepts of haploinsufficiency, dominant negative effects, and gain of function mutations disagree with the “Two-Hit Hypothesis”?
1.) Upstream analysis identifies a protein as a tumor suppressor if its activity increases in response to signals that normally trigger cell cycle arrest or apoptosis. Downstream analysis identifies a protein as a tumor suppressor if its presence or activation leads to the inhibition of pathways that drive tumor growth, metastasis, or cell survival.
Upstream Cause->Identify signals that activate the protein (e.g., DNA damage, stress).
Downstream Effect->Identify cellular processes inhibited by the protein (e.g., cell cycle, survival
2.) DNA damage, Hypoxia, Oncogene activation, replication stress
3.) DNA repair, cell cycle arrest, apoptosis, senescence
4.) Familial retinoblastoma every cell inherits one mutated RB1 gene (the first hit), making it highly likely that multiple retinal cells in both eyes will undergo a second, spontaneous somatic mutation. This is in contrast to sporadic where there are two normal copies so two mutations have to occur in the same cell and this is less likely it is even less likely it would occur in both eyes
5.) Haploinsufficiency, dominant negative effects, and gain-of-function mutations disagree with the traditional “Two-Hit Hypothesis” by allowing for tumor development after only one genetic alteration (or a single hit leading to a dysfunctional protein product) rather than requiring the inactivation of both alleles
Critical Thinking Questions
6.) How does the phosphatase PTEN behave like a tumor suppressor gene?
7.) What are the four p53 domains discussed in class? How does each domain contribute to p53 function?
8.) What is the logic behind the regulatory feedback loop in which p53 regulates MDM2 expression?
9.) What is the difference between cell cycle arrest, senescence, and apoptosis? Which is most likely to be caused by severe DNA damage?
10.)In what ways could a p53 mutation behave more like an oncogenic mutation?
6.)PTEN acts as a vital tumor suppressor by functioning as a dual-specificity phosphatase that inactivates the PI3K/AKT/mTOR signaling pathway, curbing cell growth, survival, and proliferation.
7.)
• N-terminal transcriptional activation domain: p53 is a transcription factor
• Central DNA binding domain: Targets the p53 consensus sequence, and is the most common location for carcinogenic mutations
• Tetramerization domain: p53 normally functions by forming a tetramer with three other p53 proteins
• C-terminal Regulatory domain: Site of p53 post-translational modifications that regulate p53 function/expression
8.)p53 acts as a transcription factor that induces the expression of MDM2, which in turn acts as an ubiquitin ligase to degrade p53, preventing runaway activity and unnecessary apoptosis.
9.)
• Cell cycle arrest: A temporary stop to cell division while the cell repairs DNA damage
• Senescence: A permanent withdrawal from cell division (but the cell survives)
• Apoptosis: Programmed cell death
Severe DNA damage is most likely to cause apoptosis.
10.)
• p53 oncogene functions
• Dominant negative p53: Mutant p53 in heterozygous individual interacts with wildtype p53 and prevents the function of the healthy allele’s protein
• Gain of function p53: Mutations that gain a new function such as binding to a new protein that incorrectly targets p53’s transcriptional function
“Tumor Suppression in the Absence of p53-Mediated Cell-Cycle Arrest, Apoptosis, and Senescence” JC #4 Major findings
It demonstrates that p53-mediated tumor suppression can occur independently of its canonical, acute roles in cell-cycle arrest, apoptosis, and senescence. Researchers found that mice with mutant p53 (3KR) that were defective in these three processes still did not succumb to early-onset tumors, indicating that other, non-canonical p53 functions, such as metabolism regulation and antioxidant defense, are sufficient to suppress tumor initiation
Apoptosis
Cell death by apoptosis
• Highly regulated –“Programmed cell death”
• Membrane “blebs” enclose cell components for phagocytosis
• Organized to prevent inflammation
• During apoptosis cells shrink and cell membranes undergo “blebbing”
*Apoptosis is regulated by pro- and anti-apoptotic factors
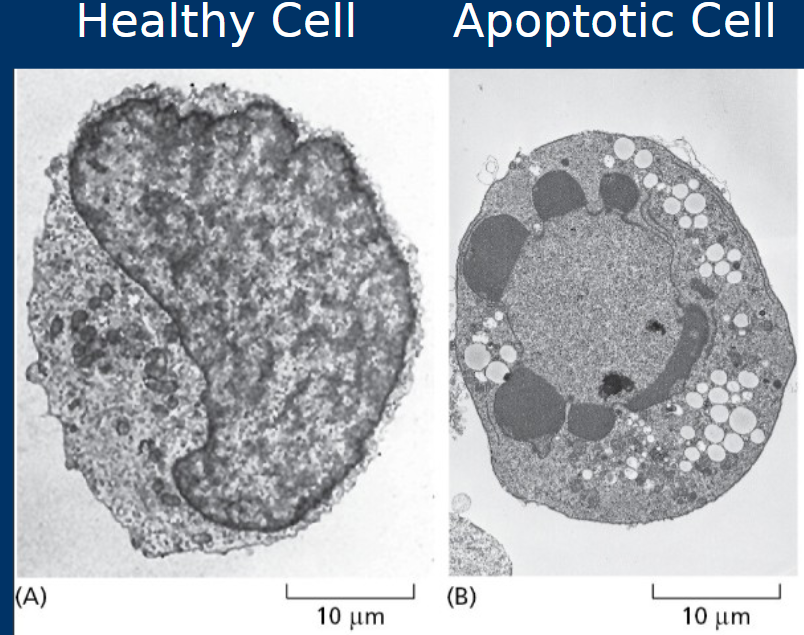
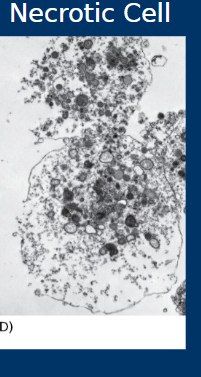
Necrosis
• Cell death by necrosis
• Less organized
• Cell rupture spills cell contents into tissue
• Generally causes inflammation and associated with negative outcomes
Phagocytosis
Phagocytosis is a vital process where specialized immune cells, primarily macrophages, detect, engulf, and digest (things like dead cells or bacteria)
• Membrane blebs become apoptotic bodies which can then be phagocytosed by immune
cells such as macrophages
Intrinsic pathway
Intrinsic: Internal cellular mechanisms that recognize cell damage (i.e. DNA damage,
oxidative damage, etc.)
• The intrinsic pathway is called “cell
autonomous”. It is a cell recognizing that it must
eliminate itself usually due to irreparable damage
(e.g. DNA damage)
Extrinsic pathway
• Extrinsic: External mechanisms in which immune cells identify carcinogenic cells and induce
them to undergo apoptosis
• The extrinsic pathway is induced by other cells which recognize unhealthy cells and use cell
signaling to induce apoptosis
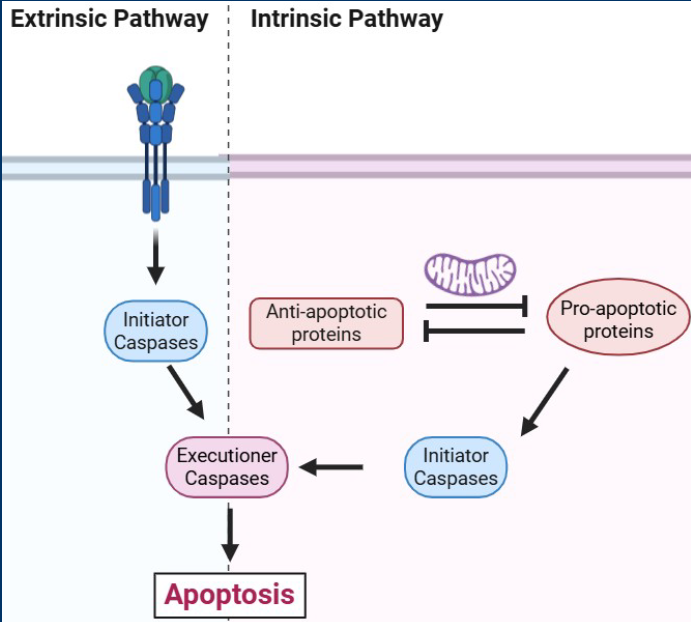
Initiator caspases
Initiator caspases are activated early in apoptotic pathways
-Responsible for cleaving and activating other caspase enzymes
- Form of enzymatic signal amplification
- Examples = Casp8 (Extrinsic) and Casp9 (Intrinsic)
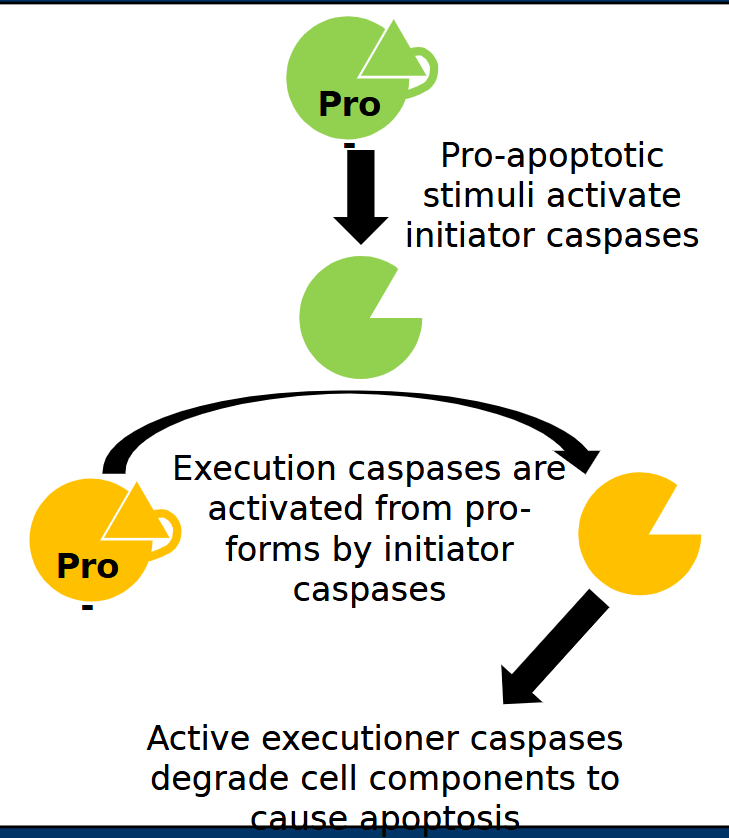
Executioner caspases
Executioner caspases cleave cellular components to cause apoptosis
- Cleavage of cytoskeleton to cause membrane disruption
- Cleavage of PARP
- Cleavage/activation of DNase enzymes to degrade nuclear DNA
- Examples = Casp3 and Casp7 (Intrinsic and extrinsic)
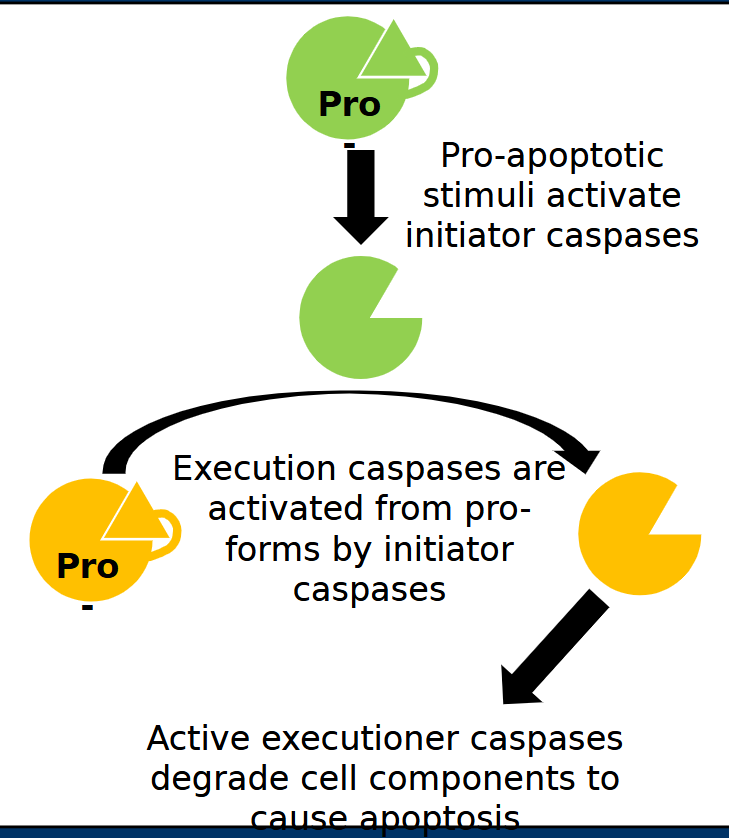
Casp8
Casp8 is an initiator caspases which means it is activated early in apoptotic pathways
• Responsible for cleaving and activating other caspase enzymes
• Form of enzymatic signal amplification
• Examples = Casp8 (Extrinsic) and Casp9 (Intrinsic)
TNFα Receptor and • FAS Receptor pathways
• Both pathways converge on the activation of the initiating caspase called Casp8
• Casp8 activates the executioner caspases Casp3 and Casp7
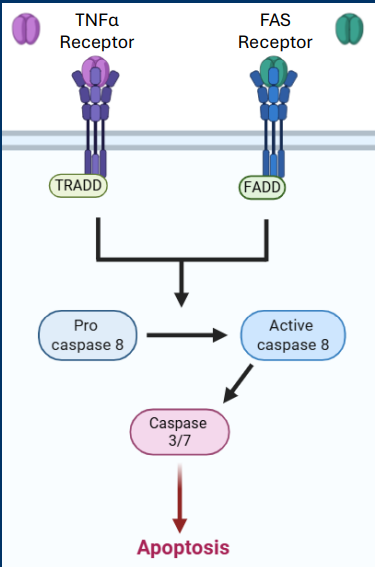
Casp9
Casp9 is an initiator caspase which are activated early in apoptotic pathways
• Responsible for cleaving and activating other caspase enzymes
• Form of enzymatic signal amplification
• Examples = Casp8 (Extrinsic) and Casp9 (Intrinsic)
• The apoptosome has main functions, but the main function is the activation of the initiator caspase (Casp9)
• Casp9 cleaves/activates the executioner caspases Casp3 and Casp7
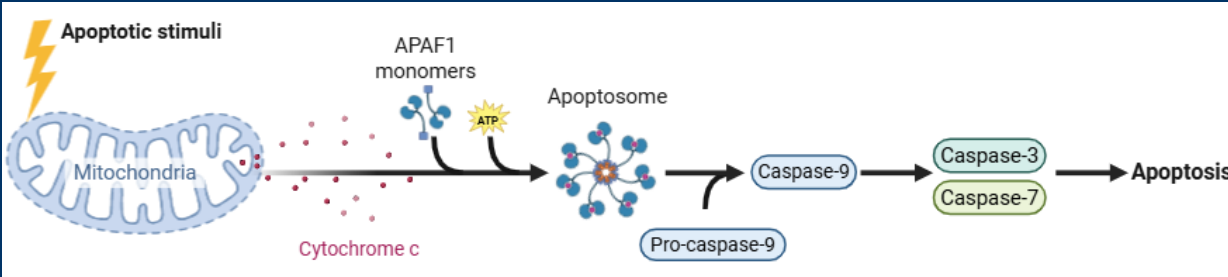
Casp3
Casp3 is an Executioner caspases cleave cellular components to cause apoptosis
• Cleavage of cytoskeleton to cause membrane disruption
• Cleavage of PARP
• Cleavage/activation of DNase enzymes to degrade nuclear DNA
• Examples = Casp3 and Casp7 (Intrinsic and extrinsic)
>Some executioner caspases degrade specific proteins with important cellular functions
> For example, Casp3 cleaves and inactivates PARP as a mechanisms to prevent further DNA repair as the cell commits to death
• Casp8 and Casp 9 activates the executioner caspases Casp3 and Casp7
Casp7
Casp7 is an is an Executioner caspases cleave cellular components to cause apoptosis
• Cleavage of cytoskeleton to cause membrane disruption
• Cleavage of PARP
• Cleavage/activation of DNase enzymes to degrade nuclear DNA
• Examples = Casp3 and Casp7 (Intrinsic and extrinsic)
-• Casp8 and Casp 9 activates the executioner caspases Casp3 and Casp7
TNFa
• TNFα Receptor
• Ligand = TNFα (inflammatory cytokine)
• Intracellular effector = TRADD (TNF receptor-associated death domain
(extrinsic)
• Both pathways converge on the activation of the initiating caspase called Casp8
• Casp8 activates the executioner caspases Casp3 and Casp7
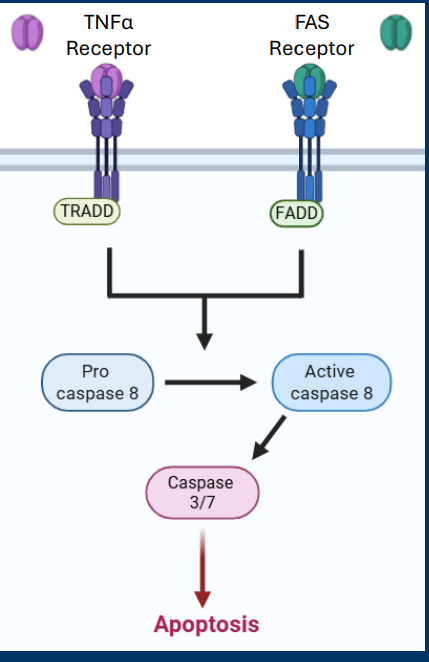
TNFR
TRADD
TRADD (TNF receptor-associated death domain)
Part of the TNFα Receptor pathway ( it is the intracellular effector)
FAS receptor
* FAS Receptor
• Ligand = FASL (FAS ligand)
• Intracellular effector = FADD (Fas-associated death domain)
• Both pathways converge on the activation of the initiating caspase called Casp8
• Casp8 activates the executioner caspases Casp3 and Casp7
(extrinsic)
FASL
FASL (FAS ligand)
part of FAS Receptor pathway
FADD
FADD (Fas-associated death domain)
it is the intercellular effector in the FAS receptor pathway
BH3 domain
• BH3 = BCL2 Homology
> Intrinsic apoptosis relies on a family of proteins that contain one to four BH domains
-Classified based on number of BH3 domains (one, three, or four domains)
-Classified as either pro- or anti-apoptotic
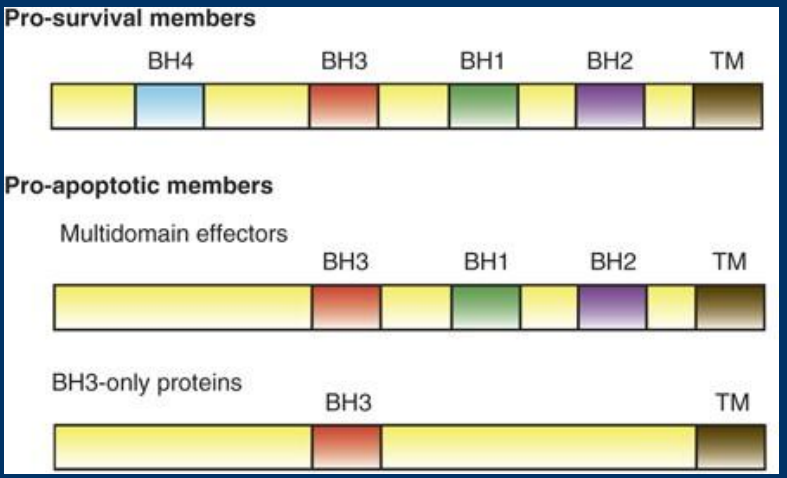
BCL2
BCL2 = B – Cell Lymphoma 2
• BCL2 is the most highly mutated gene in germinal B-cell lymphoma
• Cancer patients generally have mutations that lead to BCL2 overexpression
• Based on this alone, can you determine if BCL2 is a pro-apoptotic or antiapoptotic gene?
Anti-aprotic (protooncogene)
BCL-xL
Anti-apoptotic members of the BH3 family include: BCL2, BCL-xL, Mcl-1
• Generally have four BH domains
• Anti-apoptotic BCL2/BCL-xL counteract this( Proto-apoptotic BH domains making pores) by binding to Bak/Bax to “distract” them from forming homooligomer pores
-Other pro-apoptic factors (e.g. Bid) interact with BCL2/BCL-xL to prevent Bak/Bax interaction
Bak
• Pro-apoptotic members of the BHC family include:
• Bak and Bax (three BH domains)
• Bid, Bim, Bad, Puma, Noxa, Bik (one BH domain)
• Pro-apoptotic Bak/Bax form homooligomer structures that form mitochondrial membrane pores
**
• Too high expression of Bak/Bax or too low expression of BCL2/BCL-xL allows Bak/Bax to oligomerize the mitochondrial membrane
> Forms pores leading to MOMP
• If pro-apoptotic Bak/Bax succeed in forming pores, MOMP leads to the release of cytochrome c into the cytosol
Bax
• Pro-apoptotic members of the BHC family include:
• Bak and Bax (three BH domains)
• Pro-apoptotic Bak/Bax form homooligomer structures that form mitochondrial membrane pores
*
• Too high expression of Bak/Bax or too low expression of BCL2/BCL-xL allows Bak/Bax to oligomerize the mitochondrial membrane
> Forms pores leading to MOMP
• If pro-apoptotic Bak/Bax succeed in forming pores, MOMP leads to the release of cytochrome c into the cytosol
Bid
Pro-apoptotic members of the BHC family include:
• Bid, Bim, Bad, Puma, Noxa, Bik (one BH domain)
>Other pro-apoptic factors (e.g. Bid) interact with BCL2/BCL-xL to prevent Bak/Bax interaction
• They also have upstream crosstalk mechanisms
• BH3-only proteins (e.g. BID) are regulated by both pathways (Intrinsic and extrinsic)
MOMP
• Intrinsic apoptosis revolves around a fight to form large pores in the mitochondrial outer membrane, known as mitochondrial outer membrane permeabilization (MOMP)
-MOMP Releases Cytochrome C into the Cytoplasm
-
• Too high expression of Bak/Bax or too low expression of BCL2/BCL-xL allows Bak/Bax to oligomerize the mitochondrial membrane
-Forms pores leading to MOMP
• If pro-apoptotic Bak/Bax succeed in forming pores, MOMP leads to the release of cytochrome c into the cytosol
Cytochrome C
• If pro-apoptotic Bak/Bax succeed in forming pores, MOMP leads to the release of cytochrome c into the cytosol
• This is the same cytochrome c that acts as an electron carrier during the mitochondrial electron transport chain
-Cytosolic Cytochrome C forms the Apoptosome
• Once in the cytosol, cytochrome c interacts with the protein APAF-1 and causes it oligermize into a structure known as the apoptosome
APAF-1
• Once in the cytosol, cytochrome c interacts with the protein APAF-1 and causes it oligermize into a structure known as the apoptosome
(Apoptotic protease activating factor 1 (APAF-1) is a key cytosolic protein that initiates the intrinsic (mitochondrial) pathway of apoptosis by forming the apoptosome)
Apoptosome
>Cytosolic Cytochrome C forms the Apoptosome
• Once in the cytosol, cytochrome c interacts with the protein APAF-1 and causes it oligermize into a structure known as the apoptosome
• The apoptosome has main functions, but the main function is the activation of the initiator caspase (Casp9)
• Casp9 cleaves/activates the executioner caspases Casp3 and Casp7
Critical thinking questions
1.)What are some differences between apoptotic and necrotic mechanisms of cell
death?
2.)What are the differences between the upstream causes of the extrinsic and
intrinsic apoptotic pathways?
3.)In what ways are the downstream effects of the extrinsic and intrinsic
apoptotic pathways similar?
4.)How does caspase activation work? What is the difference between an initiator
and an executioner caspase?
1.) Apoptosis is highly regulated and it prevents inflammation. Necrosis is not very organized, causes inflammation and the cell contents spill into tissue (in apoptosis “blebs” keep cell enclosed for phagocytosis)
2.) Intrinsic: Internal cellular mechanisms that recognize cell damage (i.e. DNA damage, oxidative damage, etc.) vs Extrinsic: External mechanisms in which immune cells identify carcinogenic cells and induce them to undergo apoptosis
The differences between the upstream causes of the intrinsic and extrinsic apoptotic pathways lie in whether the death signal originates from internal cellular stress (intrinsic) or external extracellular signals (extrinsic)
3.)Both pathways rely on the activation of caspase enzymes (Executioner and initiator ) to achieve apoptosis in terms of down stream effects.
4.)Initiator caspases are activated early in apoptotic pathways they are responsible for cleaving and activating other caspase enzymes (Casp8/9).Executioner caspases cleave cellular components to cause apoptosis.
• Specific cleavages remove inhibitory segments and activate proteolytic activity of caspase enzymes
• Activated caspases have proteolytic activity towards many protein components of the cell
Critical thinking questions
5.) What are some proteolytic targets of executioner caspases?
6.) How does an immune cell recognize a cancerous cells to induce the extrinsic
pathway?
7.) How do the interactions of BH3 domain proteins contribute to mitochondrial
outer membrane permeabilization during intrinsic apoptosis?
8.) What is the significance of cytochrome c during intrinsic apoptosis?
9.) How would you expect each of the proteins discussed today to be mutated
during cancer? Would their function be activated or inactivated?
5.) Cytoskeleton, PARP, DNase enzymes
6.) Cytotoxic T-cells use membrane receptors to determine if a cell is expressing “healthy” proteins. Tumor cells present abnormal proteins marking themselves as carcinogenic.
7.) Bak/Bax interact to create pores which leads to MOMP being able to be released ( this only happens when BCL2/BCL-xL are blocked by Bid from interfering with Bak/Bax interaction)
8.) Cytochrome C interacts with APAF-1 to form the apoptosome which activates Casp9→ casps3/7
9.)
Methods of detecting apoptosis
Western blot (caspase 8 (extrinsic) caspase 9 (intrinsic)
qRT-PCR (Quantitative real-time PCR) Elevated mRNA expression of a gene indicates the activation of
transcriptional pathways that express the gene ( Intrinsic→ gene expression, p53 → Bak/bax-| BCL2/BCL-xL
DNA Isolation and Visualization on Gels (DNAfase)a )gene indicates the activation of
A TUNEL (Terminal deoxynucleotidyl transferase dUTP nick end labeling)
• DNA from non-apoptotic cell has very few free DNA ends
• DNA from apoptotic cell has many free DNA ends
• Flow cytometry is a method for visualizing lipids/proteins expressed on the outer cell surface
• Cells do not have to be lysed or killed -(flippase moves PS outside the cell so a fluorescent marker could bind to this)