MCB 2210 L16: Mitochondria, Chloroplast and Peroxisome Targeting
1/22
Earn XP
Description and Tags
Slides 1-30
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
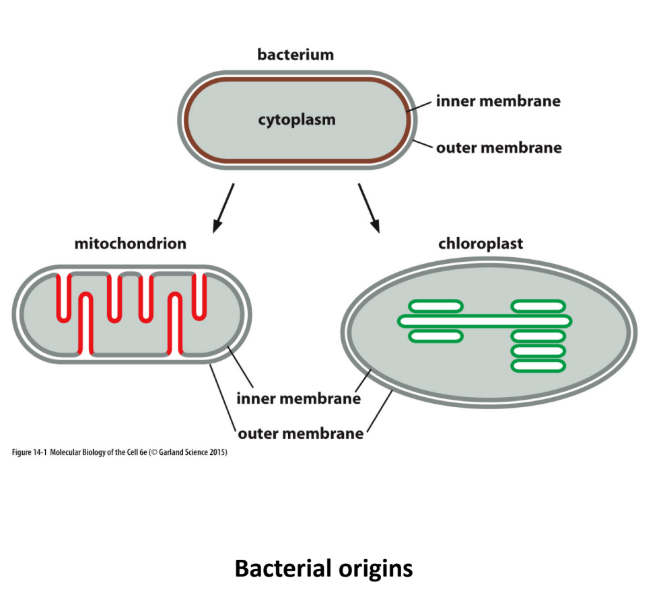
Explain the evolutionary ancestry of mitochondria and chloroplasts
Evolved from ancient bacterial eaten by eukaryotic cell
Inner + Outer membranes
Inner membrane = original bacterial membrane
Outer membrane = eukaryotic cell membrane (ate)
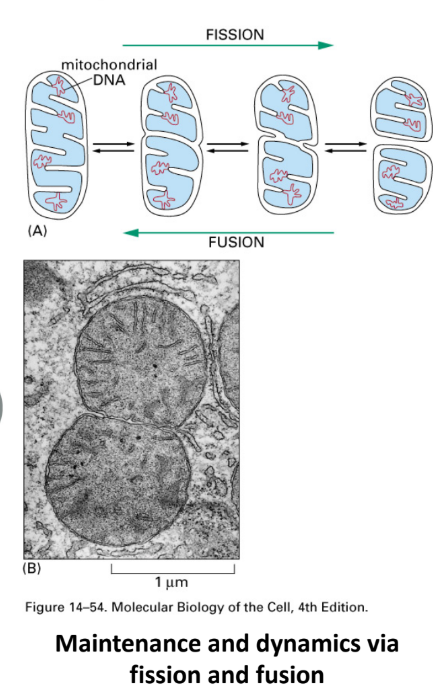
Describe fission and fusion and how they relate to mitochondria and chloroplasts
Mitochondria ≠ static → constantly Δshape through 2 processes
Fission = divide into 2 → cell division
Ensures 2 daughter cells have mitochondria
Remove damaged sections of mitochondrial network
Fusion = 2 fuse tg → long organelle
Share resources (proteins, healthy DNA) → repair/support underperforming sections of network
Note: chloroplasts undergo fission but NOT fusion
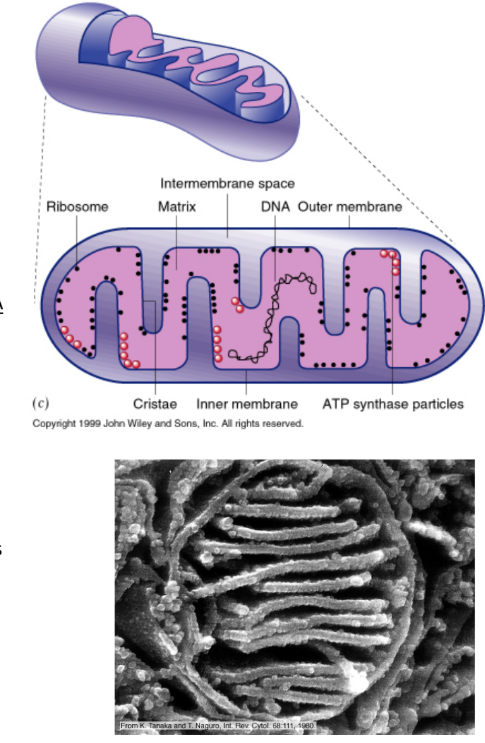
Describe structure, function & location of mitochondria
Structure
2 membranes
Outer membrane
Inner membrane = folded in → INCREASE surface area
Matrix = innermost space
Intermembrane space = space b/w 2 membranes
Shape = cylindrical + dynamic
Can fuse to tubes + Δ shape
Role of Δ morphology (shape) NOT understood
Function = produce ATP for cellular energy
Cytoplasm → Glycolysis = breaks down glucose → 2 ATP + pyruvate
Mitochondria = Oxidative metabolism (Citric acid/Krebs cycle) = pyruvate → mitochondria → broken into 30+ ATP
Location = throughout cytoplasm
Sometimes in areas where more energy is needed (muscle cells)
Associated w/ microtubules → move along microtubules through molecular motors
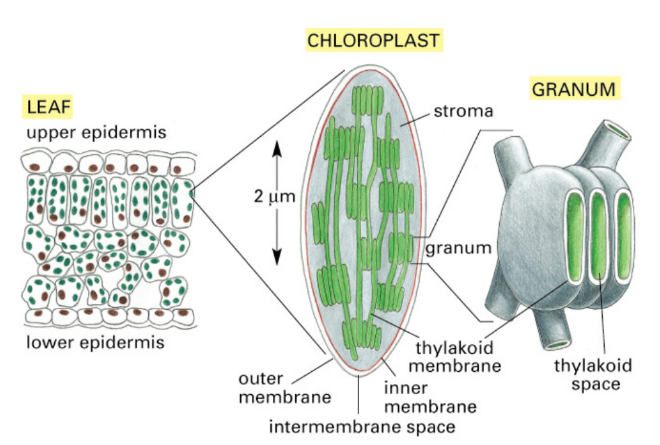
Describe structure + function + location of chloroplasts
Structure
2 membranes
3rd membrane internal layer in thylakoids
Thylakoid = tiny, sac-like membranes stacked inside plant chloroplasts
Inside = Thylakoid space/lumen
Outside = matrix = stroma (fluid in chloroplast surrounding thylakoids)
Location of machinery:
Thylakoid membrane → photosynthetic machinery
Stroma → carbohydrate synthesis machinery
Function
Use light energy to produce ATP
ATP → carbohydrate synthesis (glucose)
How are mitochondria involved in ATP production?
Convert pyruvate (product of glycolysis) → ATP
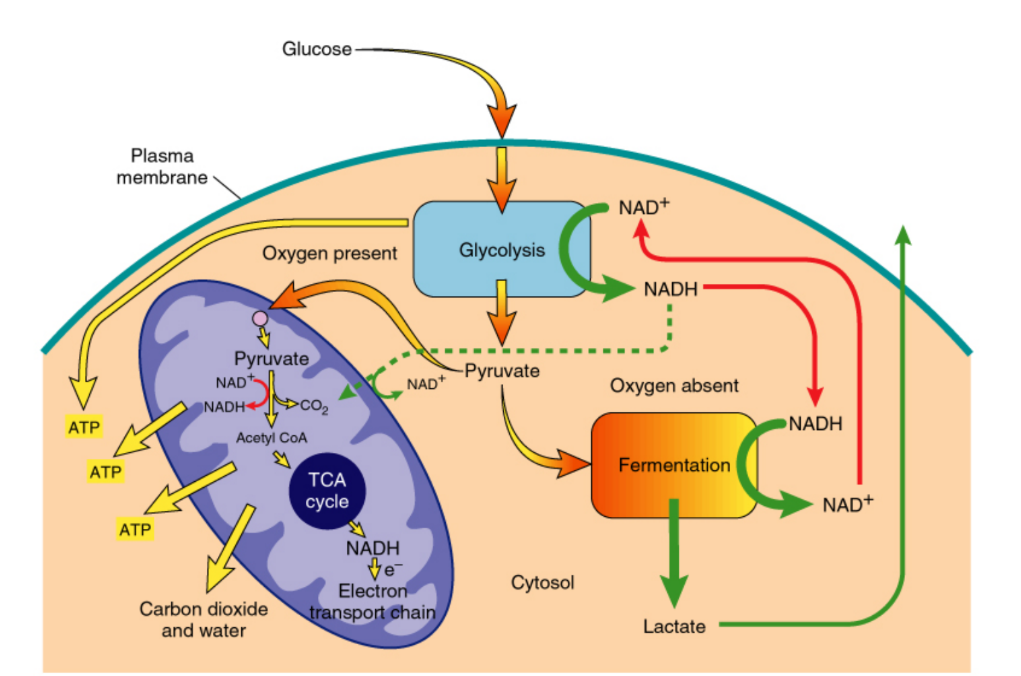
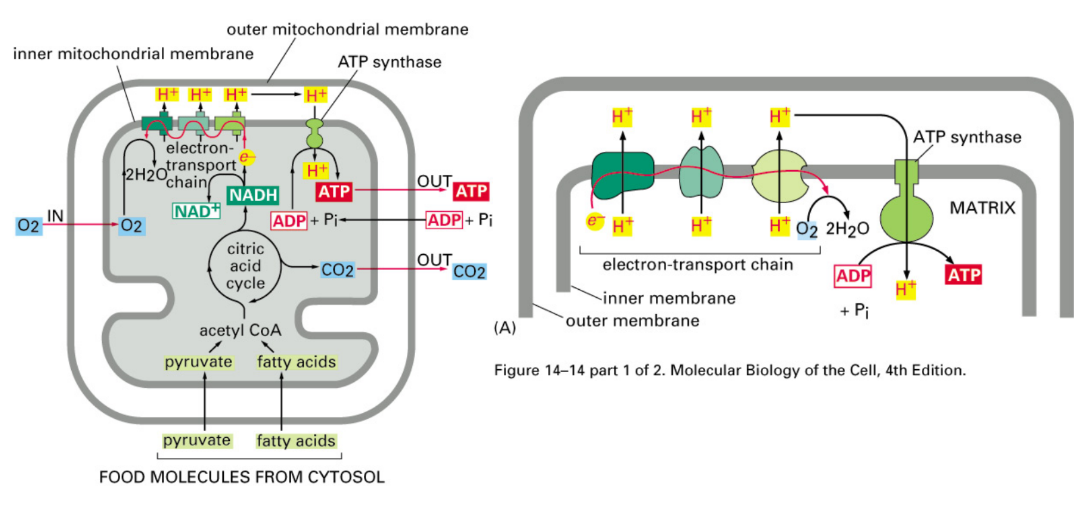
Describe the three steps required for ATP synthesis in mitochondria
Pyruvate → mitochondria → Citric acid cycle → NADH
NADH drops of e- @ Electron Transport Chain (ETC)
Pumps protons out → pressure gradient
Protons rush through ATP synthase → spin → ATP synthesis
What do chloroplasts use to generate proton gradient for ATP synthesis?
Use light energy
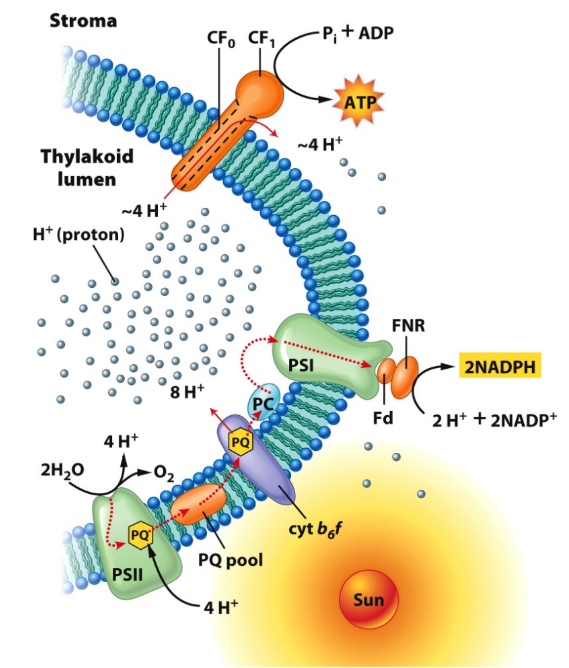
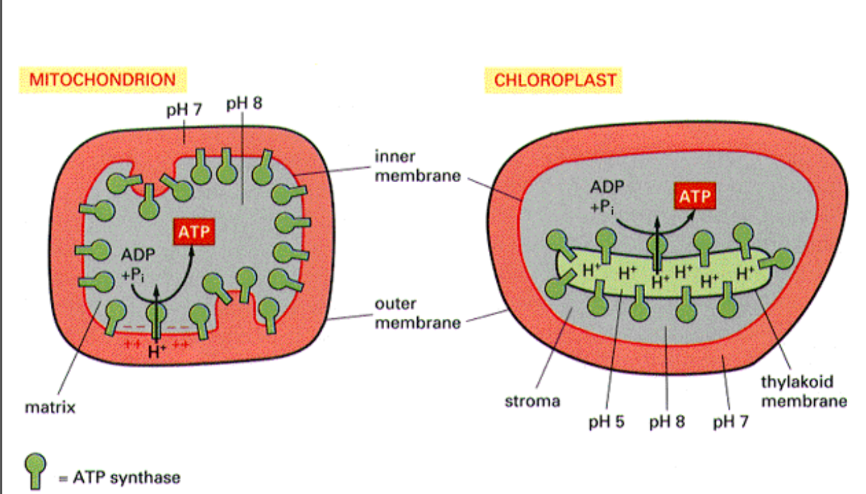
Where are proton gradients set up in mitochondria and chloroplasts?
Mitochondria → inner membrane
Chloroplast → thylakoid membrane
Both drive ATP synthesis in matrix/stromaZ
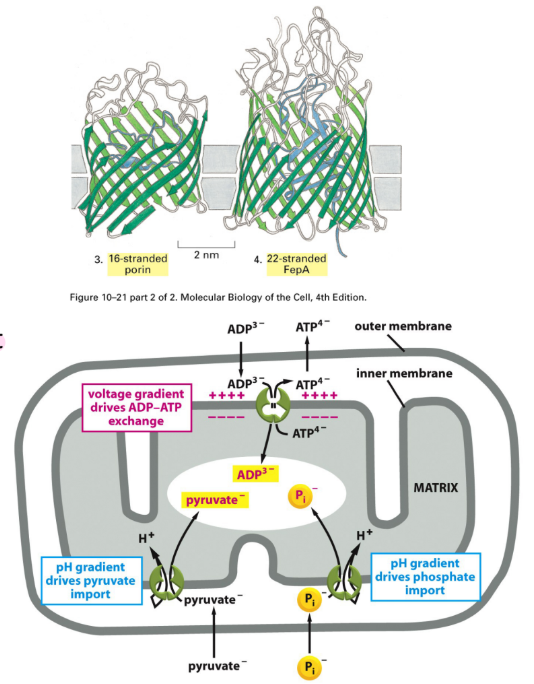
What are the 2 types of proteins found in inner and outer membranes of mitochondria? Describe how small molecule transport is achieved through these proteins.
Beta-barrel channels = type of pore-forming protein structure found in outer membranes of bacteria, mitochondria, & chloroplasts = barrel made of twisted beta-sheets
Porins = specialized, barrel-shaped transmembrane proteins → form water-filled channels in outer membranes of bacteria, mitochondria, & chloroplasts
Allows small molecules + ions to move b/w intermembrane space & cytoplasm (through outer membrane)
Transporters = specialized proteins (selective channels/carriers) located in cell membranes → selective
Located in inner membrane
Selectively moves small molecules across inner membrane in/out of matrix
Examples:
ADP/ATP exchanger = protein in inner mitochondrial membrane → ADP into mitochondria, ATP out mitochondria
Pyruvate/H+ cotransporter = protein complex in inner mitochondrial membrane
Pyruvate import to matrix
Final product of glycolysis → mitochondria for Citric Acid Cycle
Phosphate/H+ cotransporter (PiC) = protein in inner mitochondrial membrane
Phosphate (Pi) import
Pi → ATP synthase → ATP synthesis
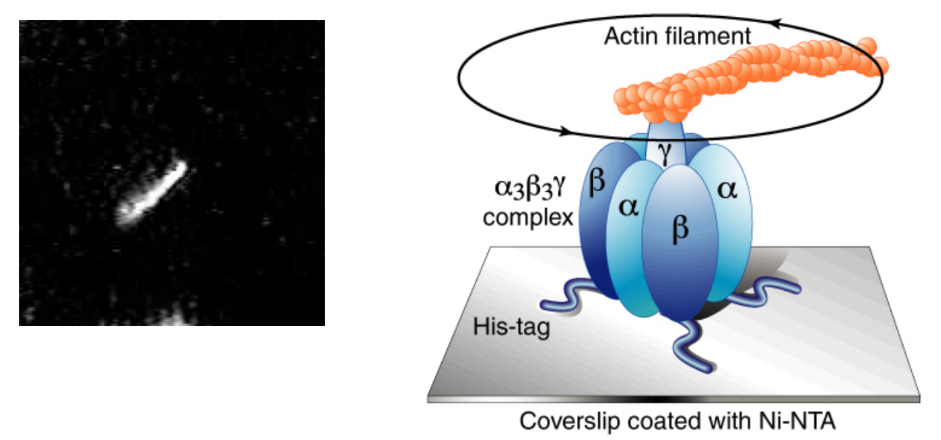
Describe the mechanism of ATP synthase + experiment that was used to prove it spins
Rotary mechanism = γ subunit (central shaft) of ATP synthase rotates → converts H+ gradient energy → chemical energy in ATP
Scientists attached fluorescent actin filament to γ subunit
Enzyme + energy → actin filament = spinning under microscope
Enzyme = anchored to glass using His-tag + Ni-NTA coating
Kept in place as enzyme spun
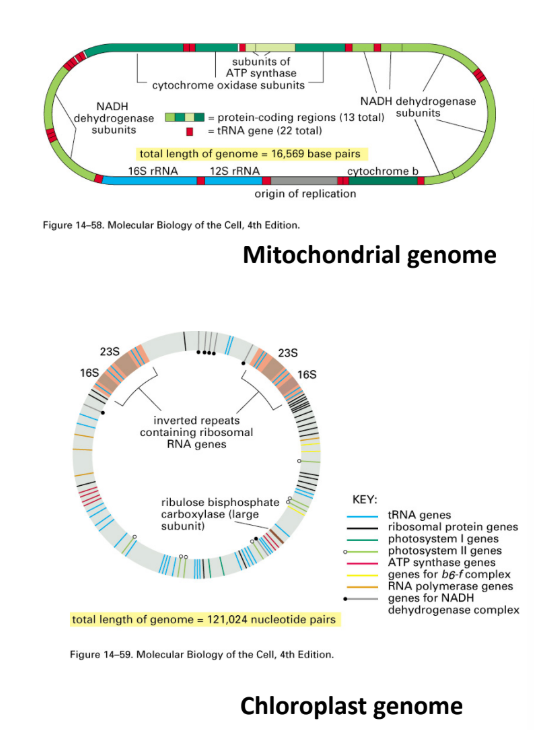
What is special about mitochondria and chloroplasts specifically?
Have their own genomes (circular)
>90% of mitochondria proteins encoded by nuclear DNA
Mitochondria = mixture of proteins encoded by nuclear + mitochondrial genes
Some enzymes = mixtures of nuclear + mitochondrial subunits
Δ tissue → Δ mitochondria
Mitochondrial genomes of different species = overlapping genes
Absent in mtDNA → nuclear DNA
Certain amount of genes → functioning mitochondria
Nuclear/mitochondrial genes depend on species
Chloroplast = mixture of proteins encoded by nuclear + chloroplast genes
ATP Synthase is described as a ____________
Chimera = describes something made of different origins
ATP synthase = protein subunits from 2 different places:
Mitochondrial DNA
Nuclear DNA
Describe mitochondrial protein synthesis pathways from mtDNA and nuclear DNA.
mtDNA:
Protein = encoded by mtDNA → synthesized in matrix w/ mitochondrial ribosomes
13 mtDNA-encoded proteins
Nuclear DNA:
>90% mitochondrial proteins = encoded by nuclear DNA → synthesized in cytoplasm
~1000 nuclear-encoded proteins
Nucleus → mRNA → protein by cytoplasmic ribosome
Pass through 2 membrane proteins to matrix
TOM = Translocase of the Outer Membrane
TIM = Translocase of the Inner Membrane
Describe protein localization to different mitochondrial compartments (matrix, inner membrane, outer membrane, intermembrane space). What signal is used?
Nuclear encoded proteins = targeting signal
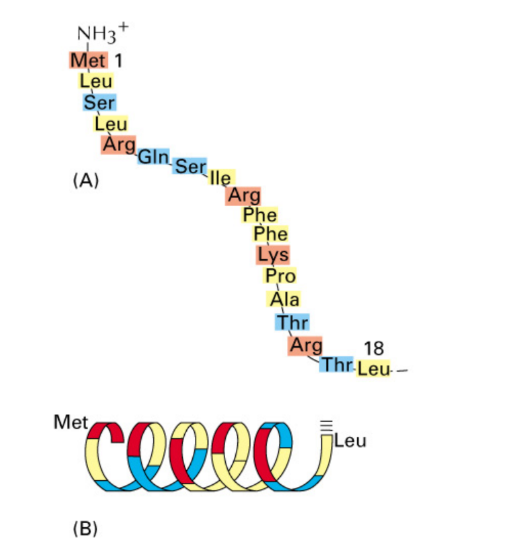
N-terminal amphipathic α-helix + (+) charged amino acids= helix w/ hydrophobic + hydrophilic sides
Necessary & sufficient → import → mitochondria
Signal = bound by receptor protein of outer mitochondrial membrane
Post-translational transmembrane transport = process where protein synthesis → cytosol BEFORE transport across mitochondrial membrane
Mitochondrial-encoded proteins ≠ targeting signal
Signal NOT NEEDED
Synthesized in matrix
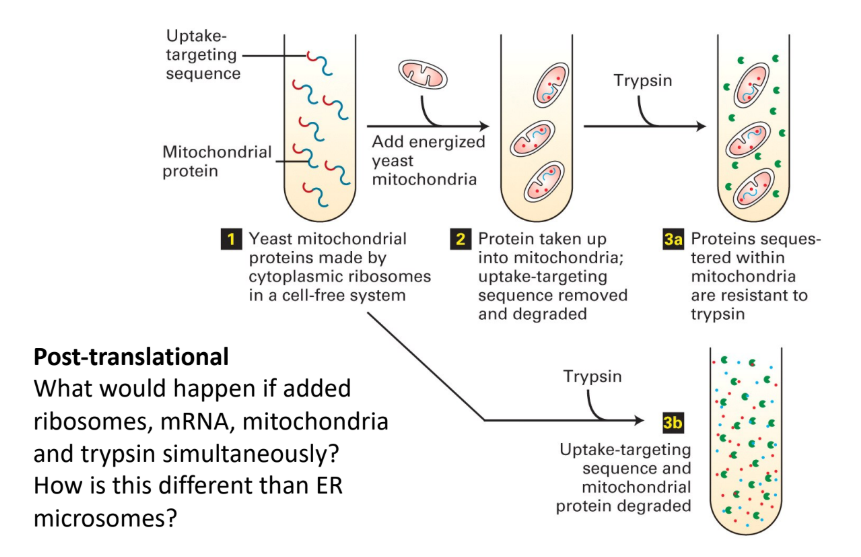
Describe the experimental paradigm used to prove protein transport mechanism to mitochondria.
Cytoplasmic ribosomes → synthesize yeast mitochondrial proteins (test tube) + N-terminal amphipathic α helix (targeting sequence)
Yeast mitochondria → test tube
Proteins = taken up into mitochondria
Targeting sequence = removed + degraded
Trypsin (protease) → test tube
Proteins = intact bc sequestered/hidden inside mitochondria
Trypsin → test tube W/O mitochondria
Proteins = degraded (no protection)
What happens if you add ribosomes, mRNA, mitochondria, and trypsin simulatenously?
How is this different than ER microsomes?
Trypsin → digest protein as soon as protein = synthesized
No proteins imported safely
Difference in when import occurs
Mitochondrial import = post-translational
Mitochondria → entire protein digested
ER import = co-translational
ER → part of protein safe inside ER
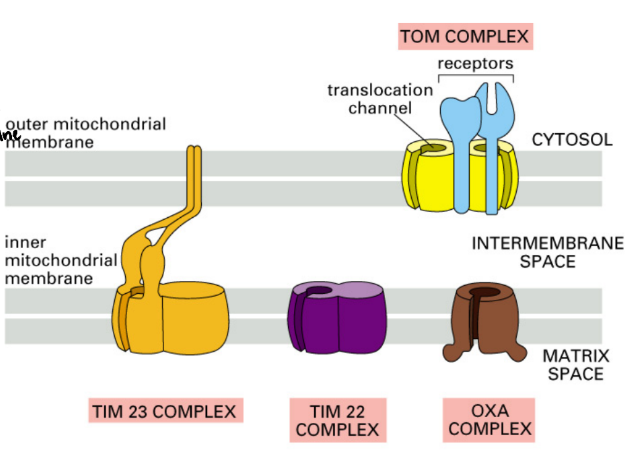
Describe transport of nuclear-encoded proteins across the inner & outer membranes of mitochondria
TOM (Transporter Outer-Membrane mitochondria) = outer membrane pore complex + receptor for signal
Protein transport → intermembrane space
Protein = all possible protein destinations in mitochondria (outer membrane, inner membrane, intermembrane space, matrix)
TIM (Transporter Inner-Membrane mitochondria) = inner membrane pore complex
Protein transport → matrix
OXA Complex = inner membrane protein that inserts protein into matrix
Protein transport → matrix
Protein exposed to matrix → targeting signal cleaved off
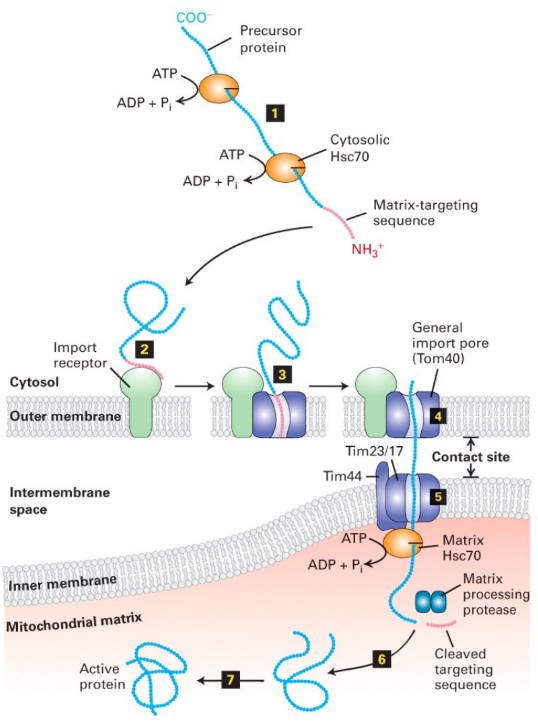
Describe transport of proteins into matrix
Occurs across both membranes @ once
Cytosolic chaperones keep protein unfolded
Import receptor + TOM40 Complex = recognize signal sequence
Protein transported through TOM40 complex → TIM44-TIM23/17 complex @ contact site
Contact site = specialized region where outer + inner membranes are tightly tethered to facilitate transfer of precursor proteins
Matrix chaperone = binds protein + pull unfolded proteins across inner membrane → matrix
Matrix protease = cleaves signal sequence
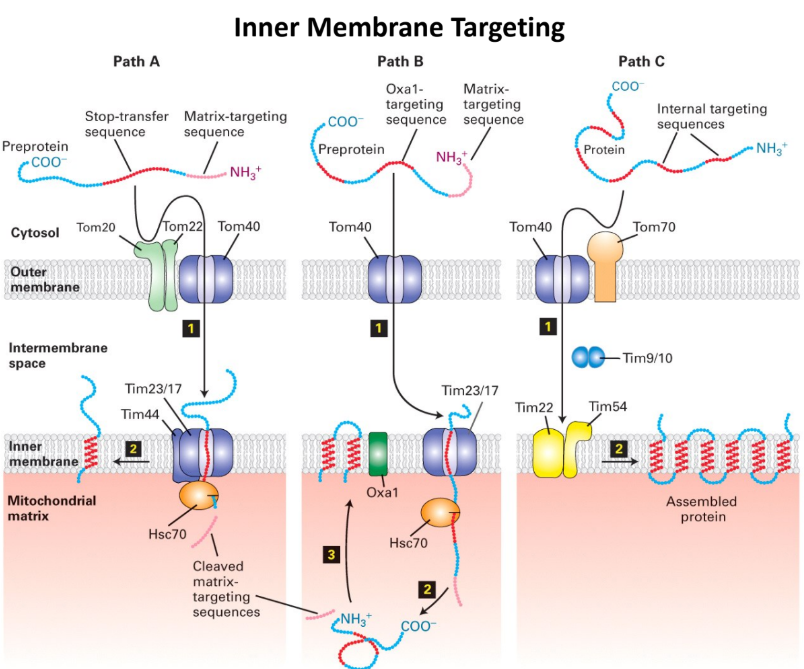
What are the 3 different pathways to the Mitochondrial Inner Membrane?
Path A = Stop-Transfer Route (most common for single transmembrane protein)
N-terminal matrix-targeting sequence + stop-transfer anchor sequence
Protein → TIM23 complex → stop-transfer sequence = “stuck” in inner membrane
Complex = opens laterally → protein embedded in membrane
Path B = Oxa1-Mediated Route (used by proteins → “re-inserted” into membrane from matrix side)
TOM/TIM23 pathway: Protein imported → matrix
Oxa1 = protein in inner membrane = inserts proteins into mitochondrial membrane
Recognizes protein & inserts back into inner membrane
Pathway for (nuclear + mitochondrial)-encoded proteins
Path C = Multi-transmembrane Route (Multipass proteins) → Ex. ATP/ADP antiporter
NO N-terminal targeting sequence + HAS internal targeting sequences
Internal targeting sequences = acts as signal sequence + anchor in membranes
Bound by small chaperones in intermembrane (Tim9+10) → prevent folding/clumping
Enter matrix through TIM22 Complex (NOT TIM 23)
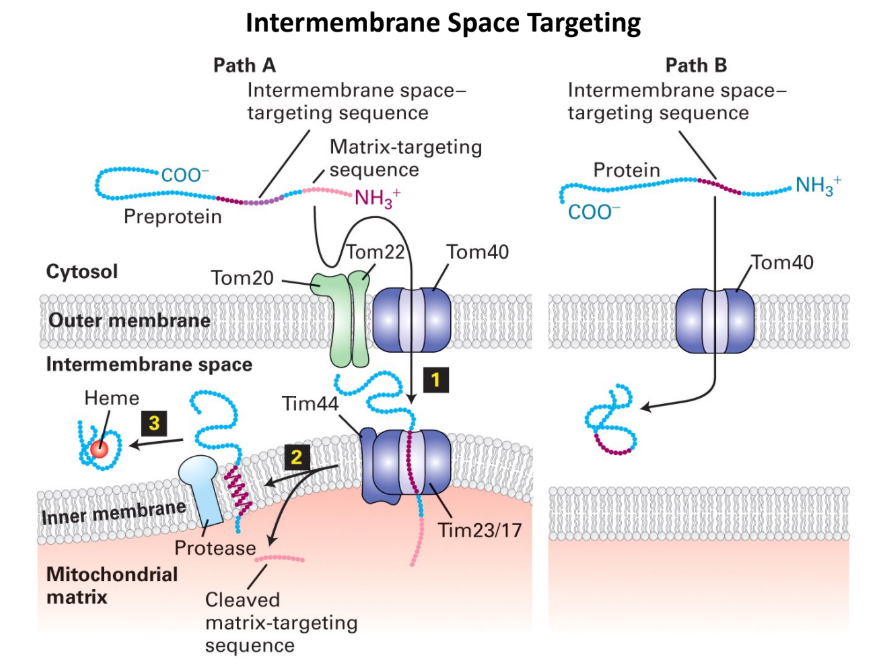
What are 2 pathways that proteins are targeted to intermembrane space?
Path A = Protease cleavage
Protein enters inner membrane through Tim23/17 complex
Protease cleaves protein → released into intermembrane
Path B = Direct entry
Protein passes through outer membrane
Protein stays in intermembrane space
Does NOT try to enter inner membrane
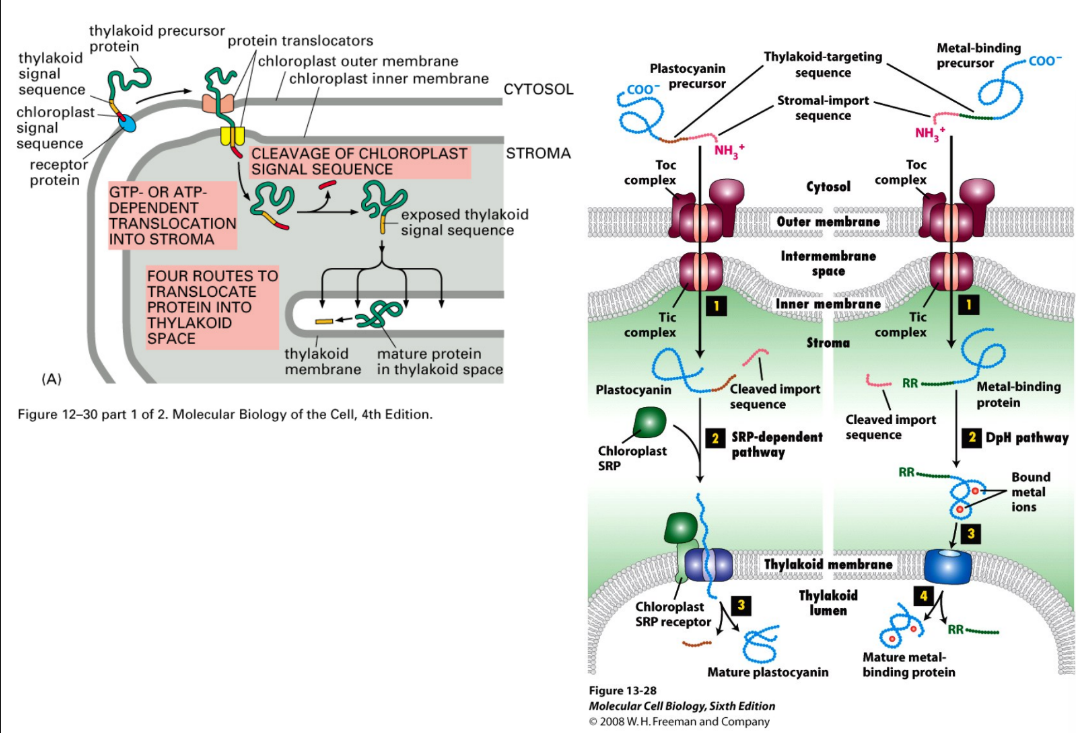
Explain protein targeting to chloroplast
Similar to mitochondria
Chaperones = assist post-translation import (unfolded)
Stroma targeting signal = N-terminal amphipathic helix
Protein → outer membrane → bind to receptor
Transport occurs through TOC + TIC (similar to TOM + TIM)
Directed to different areas through secondary signals + complexes
Signal sequence = cleaved once protein → stroma
Thylakoid → 4 routes, use thylakoid targeting sequence
Plant cells = mitochondria + chloroplasts
Membrane receptors MUST be able to tell signals apart
Same signal → transported to both
Different signals → each organelle
Poorly understood
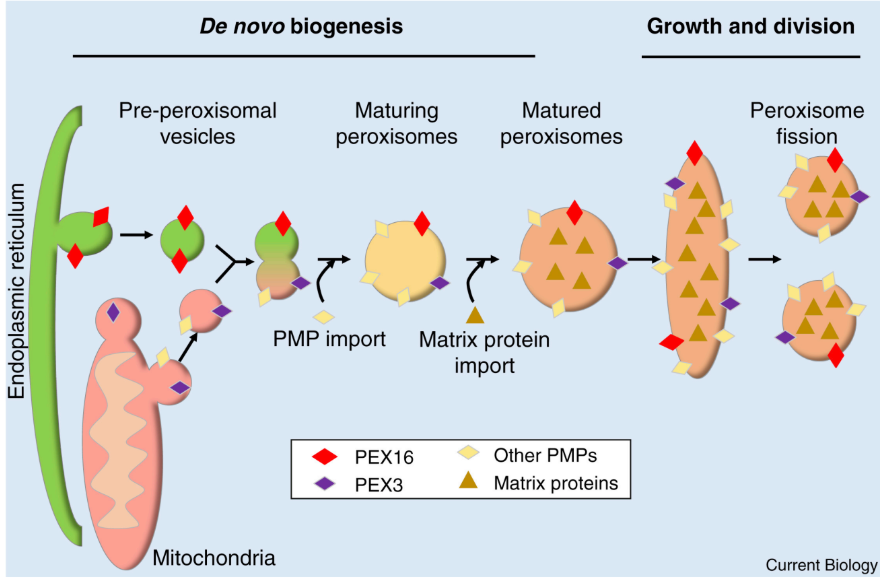
Peroxisomes + 2 mechanisms to multiple
Peroxisomes = organelles that specialize in oxidative reactions = degrade long fatty acid chains through β-oxidation = chimeras of ER + Mitochondria → bounded by single membrane bilayer
β-oxidation = metabolic process of breaking down fatty acids inside mitochondria to generate ATP
Major byproduct = H2O2
De Novo Biogenesis = creating new peroxisomes
ER + Mitochondria → “Pre-peroxisomal vesicles”
Pre-peroxisomal vesicles + PEX16 + PEX3 fuse → “maturing peroxisome”
Imports Peroxisomal Membrane Proteins (PMPs) + Matrix proteins (enzymes for β-oxidation) from cytosol
Growth & division
Peroxisome imports more proteins + lipids → elongation
Fission → 2 smaller peroxisomes
Import = post-translational
N-term/C-term peroxisomal targeting sequences (PTSs) → Unique cytoplasmic receptors
Signals NOT cleaved after protein = inside
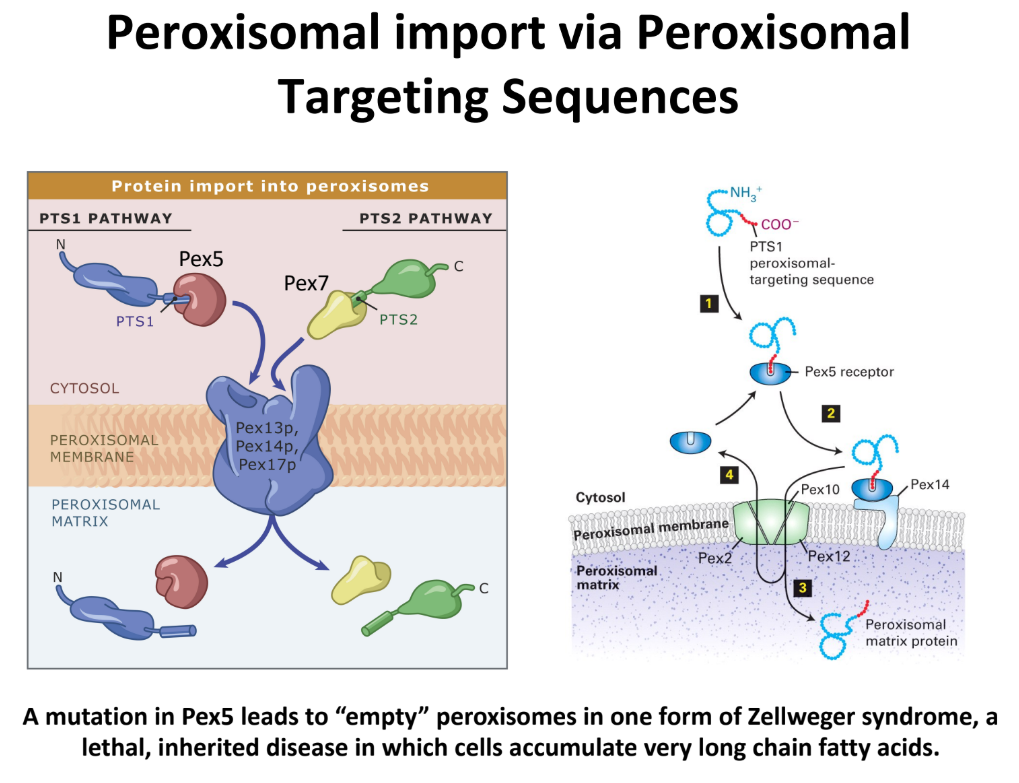
What are the 2 main pathways that bring proteins into peroxisome matrix? Define Zellweger Syndrome
PTS1 Pathway = uses Pex5 receptor (cytosolic)
Pex5 binds → protein in cytosol → translocation complex on membrane → releases protein into matrix
PTS2 Pathway = uses Pex7 receptor (cytosolic)
Transport proteins w/ different type of signal sequence
Zellweger Syndrome = mutation in Pex5 → empty peroxisomes
Lethal, inherited disease
Leads to cells w/ accumulated long fatty acid chains