Chapter 9: Gene Expression and Genomics
1/86
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
87 Terms
How are gene expression and enzyme activity controlled?
Cells don’t require all gene products at all times
Constitutive genes do need to be constantly “on.”
Inducible genes are only required at particular times.
Basic control of gene expression can take place during transcription, translation, or after translation.
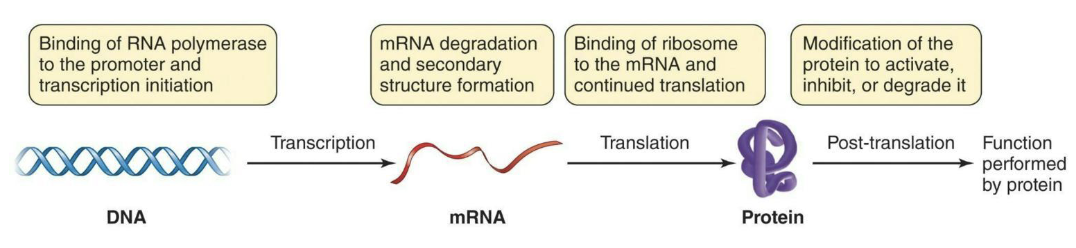
What is the activity of enzymes?
Inhibition of enzymes may result from binding of an inhibitor molecule.
Conformation is altered; substrate no longer binds.
This is known as “allosteric inhibition”
Covalent modification may also alter enzyme conformations, altering activity.
Enzyme covalent modification = Binding of molecules to specific sites on proteins can cause conformational changes that inhibit or enhance the activity of the protein
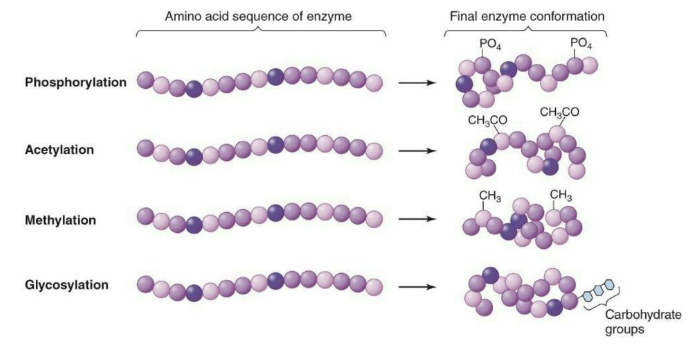
What is the production of enzymes?
While modifying enzyme activity can control a cell’s biochemistry, the energy input to make the enzyme is already expended.
A better means of energy conservation would be to only make enzymes as needed.
Much of the control mechanisms work to prevent transcription (and thus, translation) of genes when they aren’t required.
Often, this control involves regulatory molecules and DNA-binding proteins that affect the ability of RNA polymerase to bind to the promoter on DNA.
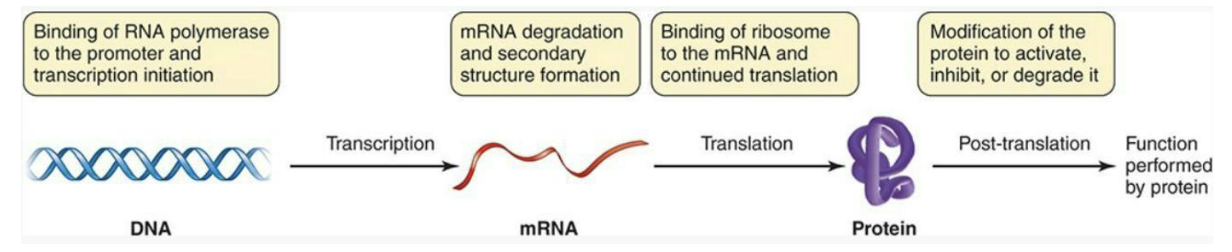
How do regulatory proteins control transcription?
Operon = Transcriptional unit of DNA with a series of structural genes under control of single promoter and their transcriptional regulatory elements
The lac operon of E.coli is a prime example of gene regulation
It includes an operator region where regulatory proteins bind
The lac operon = 3 structural genes (lacZ, lacY, and lacA) along with a single promoter and an operator
LacZ → β-Galactosidase
LacY → Permease
LacA → β-Galactoside transacetylase
A regulatory gene, lacI, is located upstream of the operon and encodes the LacI repressor protein, which controls the operator
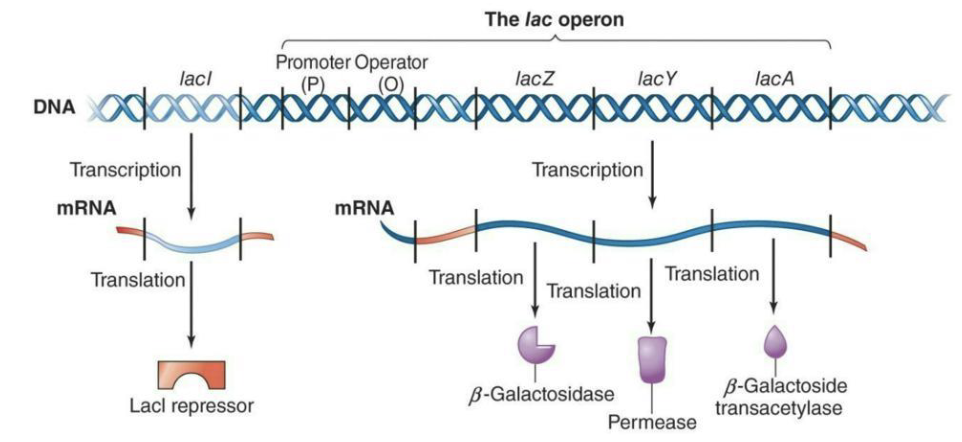
What is the lac operon?
Inducible expression
The system is only turned on (induced) when needed.
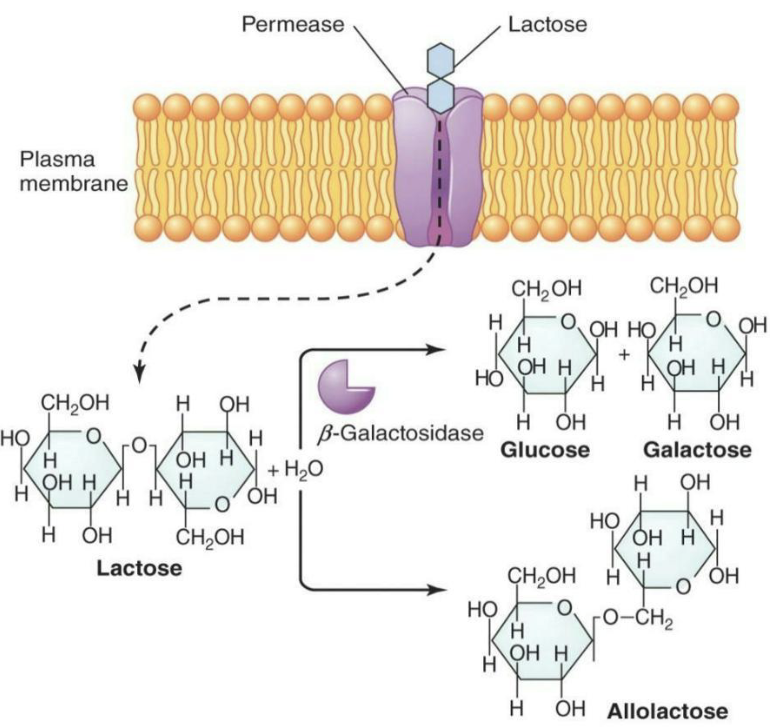
Permease (lacY) is a symporter that co-transports lactose into the bacterial cell along with a proton (H⁺)
Lactose can be cleaved by β-galactosidase (lacZ) into glucose and galactose or be converted to allolactose
Allolactose is an inducer that increases transcription by turning on the lac operon
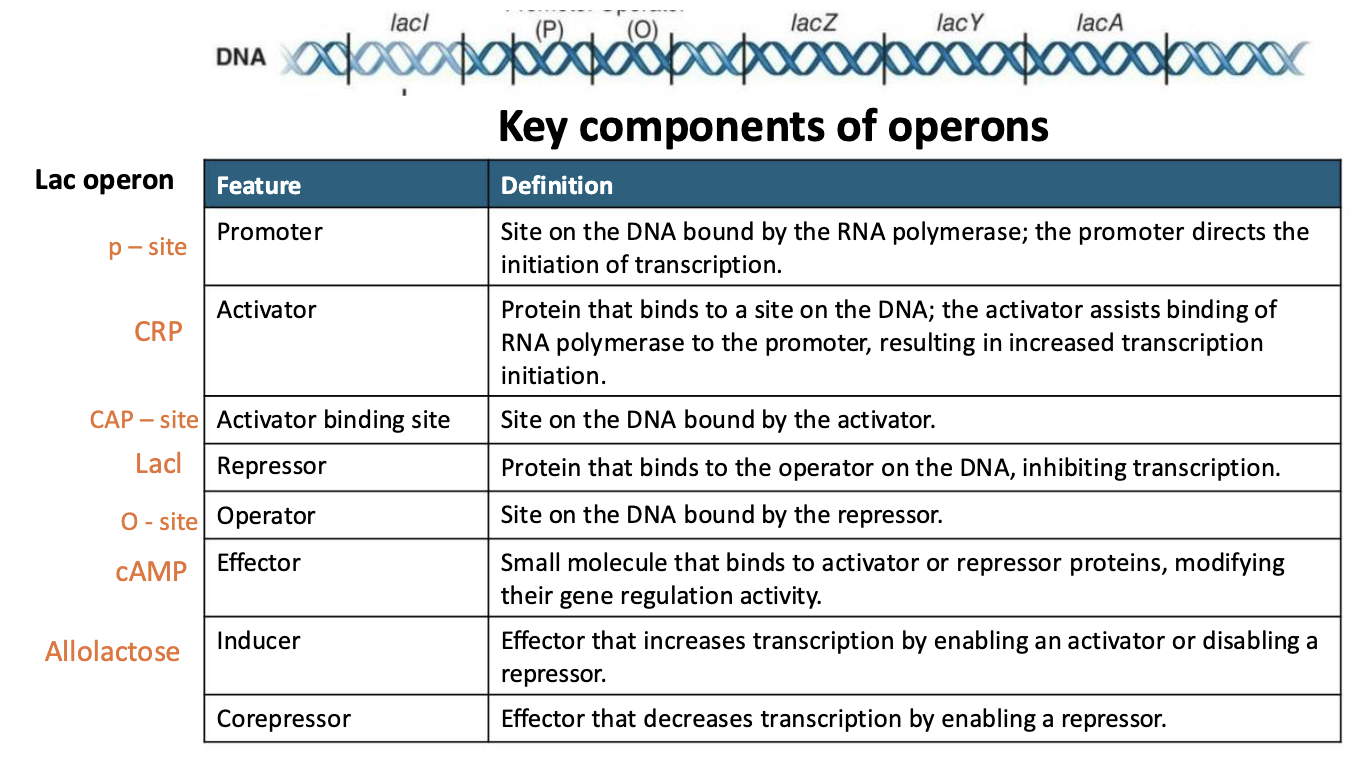
What are the key components of the operons?
Promoter (p-site): Site on the DNA bound by the RNA polymerase; the promoter directs the initiation of transcription.
Activator (CRP): Protein that binds to a site on the DNA; the activator assists binding of RNA polymerase to the promoter, resulting in increased transcription initiation.
Activator binding site (CAP-site): site on the DNA bound by the activator.
Repressor (LacI): Protein that binds to the operator on the DNA, inhibiting transcription.
Operator (O-site): Site on the DNA bound by the repressor.
Effector (cAMP): Small molecule that binds to activator or repressor proteins, modifying their gene regulation activity.
Inducer (Allolactose): Effector that increases transcription by enabling an activator or disabling a repressor.
Corepressor: Effector that decreases transcription by enabling a repressor.
What is the negative control of transcription (inducible expression)?
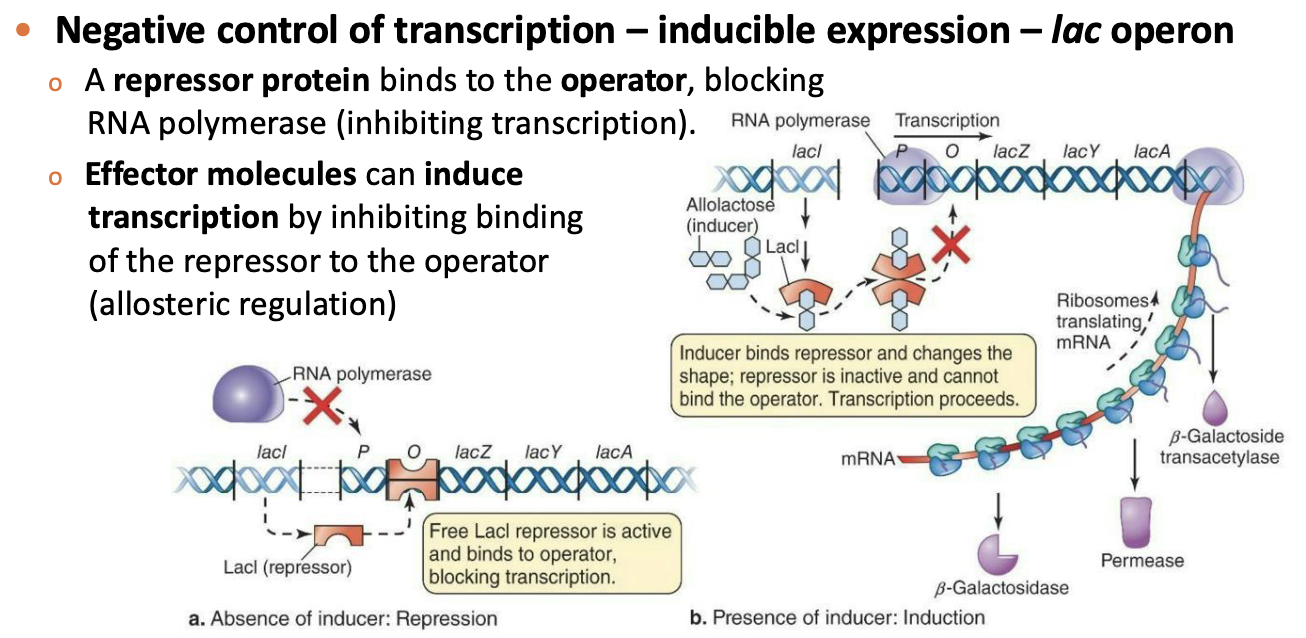
Negative control of transcription – inducible expression – lac operon
A repressor protein binds to the operator, blocking RNA polymerase (inhibiting transcription).
Effector molecules can induce transcription by inhibiting binding of the repressor to the operator (allosteric regulation)
What is the negative control of transcription (repressible expression)?
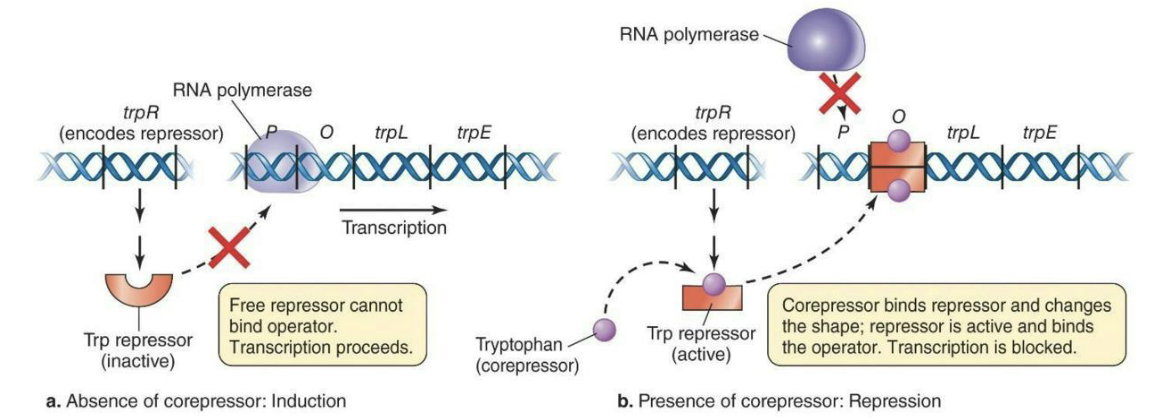
Negative control of transcription – repressible expression – trp operon
A repressor protein binds to the operator, blocking RNA polymerase (inhibiting transcription).
Effector molecules can also inhibit transcription by binding to the repressor and enhancing its ability to bind to the operator (e.g. trp operon).
Tryptophan = Corepressor (a type of effector molecule)
What is positive control of transcription?
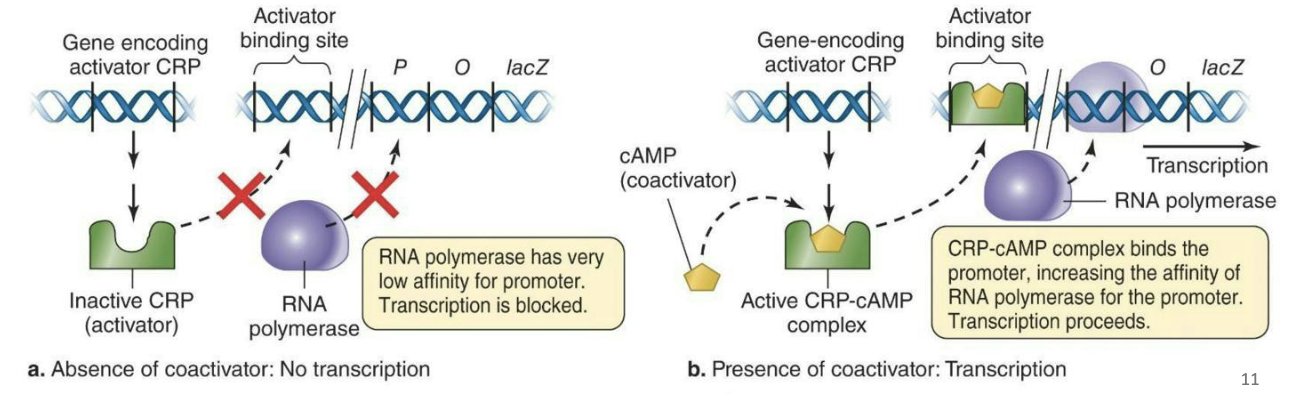
Positive control of transcription – lac operon
Regulatory molecules bind and increase transcription rates.
Usually, an activator protein (CRP) increases affinity of RNA polymerase for promoters
Effector molecule (cAMP) binds the activator protein, alters its conformation, makes it more likely to bind regulatory site on DNA.
Under low-glucose conditions, the amount of cAMP will increase resulting in transcription
How is the lac operon complex?
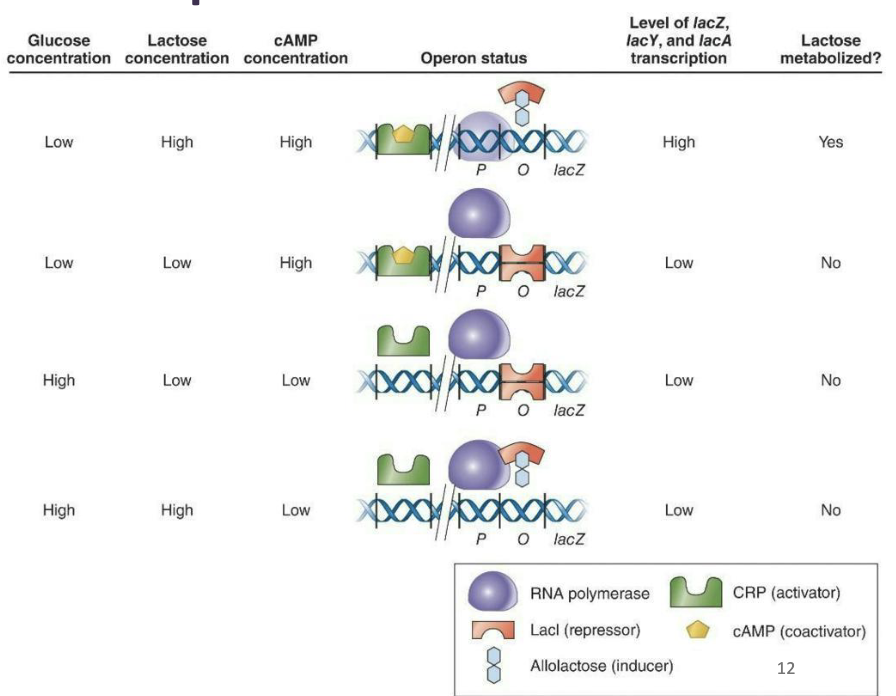
Positive control of transcription
The lac operon represents a composite of control methods.
This complexity allows for the best control if multiple sugar types are present.
The binding of Lacl by allolactose and CRP by cAMP regulate expression of the lac operon so that maximal expression occurs in the presence of lactose and absence of glucose
How is the lac operon investigated?
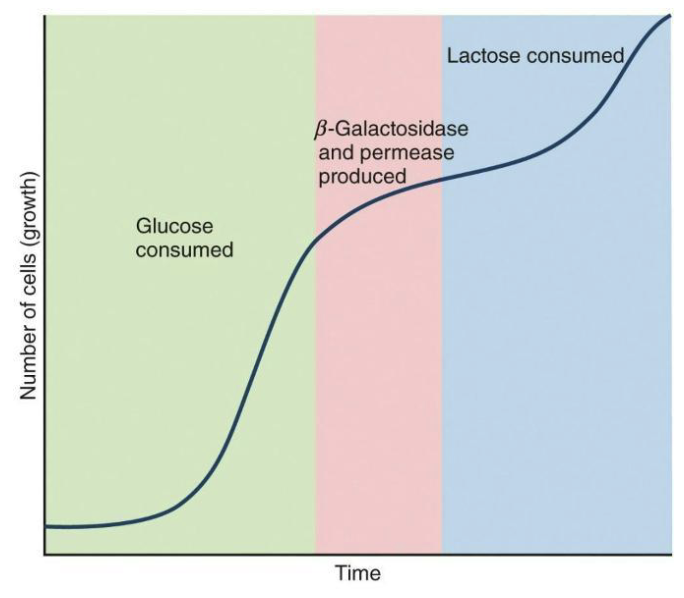
This complexity provides an opportunity to study how microbes best control gene expression.
Glucose is easier to use than lactose – faster growth occur.
Lag phase when lacZYA transcription and translation begin.
The lac operon isn’t expressed until all glucose is consumed (diauxic growth)―how is this controlled?
How were mutants expressed in the lac operon studied?
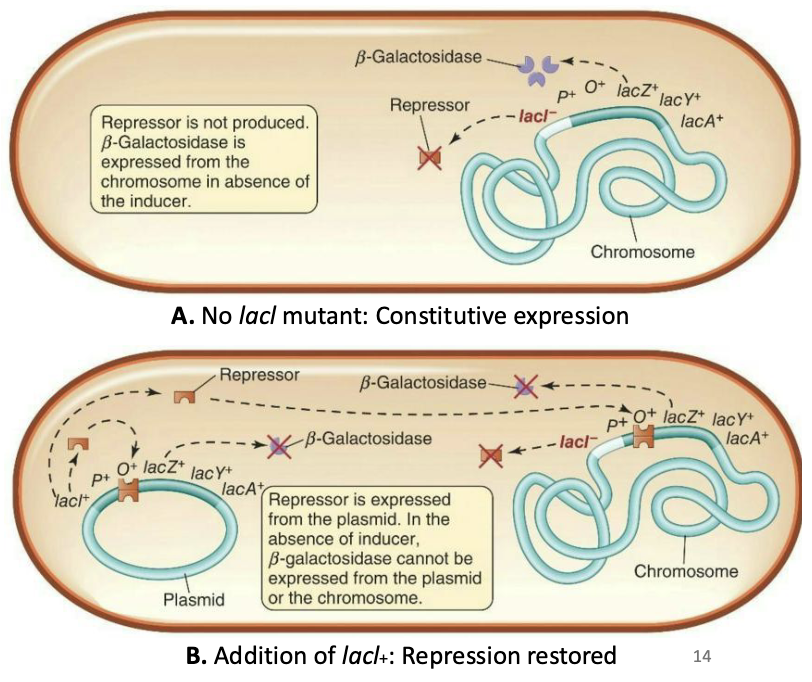
Mutants that constantly expressed the lac operon (even with no lactose present) were studied
Plasmid with the wild-type lacl can interact and repress (complement) a lacl mutant (l-)
Lacl is diffusible - this led to discovery of the LacI repressor protein.
Simple: Mutants with lacI⁻ constantly expressed the lac operon because they lacked the repressor. But adding a plasmid with lacI⁺ restored repression because the repressor is diffusible (can move through the cell and bind the operator on the chromosome).
How does the LacI repressor shut down expression of the structural genes?
Other mutant strains couldn’t shut down lac expression, even when “good” LacI was introduced.
The problem was in the DNA of the lac operon, not in the gene for production of the LacI protein – changes in the operator sequence.
Simple: Mutants with a changed operator sequence in the lac operon showed constitutive expression because the LacI repressor cannot bind the operator, so adding normal lacI⁺ does not restore repression.
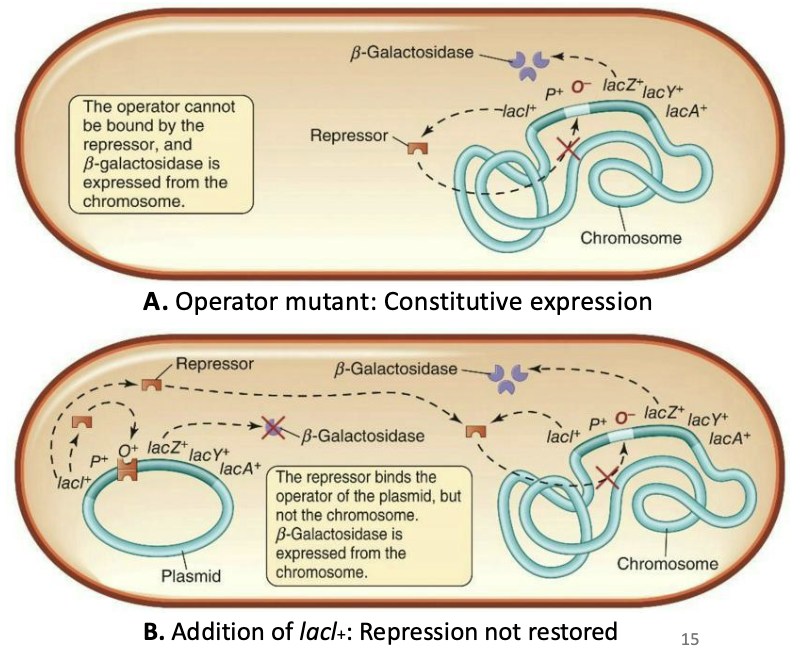
What did the previous studies studying the lac operon identify?
Several of these studies were very basic.
They laid the groundwork for doing similar work in many other operons.
This increased understanding of gene expression regulation in a variety of other areas.
This is also partially why we study the lac operon in so many biology courses!
How does a single environmental stimulus control multiple operons?
Larger sets of genes can also be coordinately regulated (global gene regulation).
Regulons are genes that are coordinated to respond to the same regulatory systems.
Catabolite repression = Shutdown of several systems that utilize various nutrients when glucose is present (cAMP-CRP complex)
The SOS response = A multigene system for wide-scale DNA repair
Pho regulon = Gene expression is regulated via the concentration of phosphate in the media
What are alternative sigmas factors (global gene regulation)?
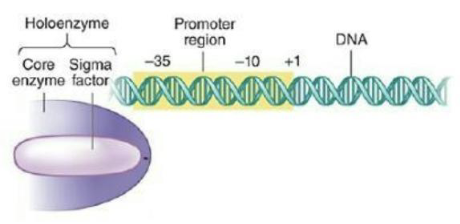
In bacteria, use of different sigma factors directs RNA polymerases to certain genes.
Alternative sigma factors are active only under specific conditions and are not required for cell viability.
Most E. coli promoters are recognized by sigma-70 (RpoD), but there are 6 alternative sigma-70 family members.
Sigma-54 (RpoN) = Regulating nitrogen utilization genes under nitrogen-limited conditions
Sigma-32 (RpoH) = Heat-shock protein gene regulator helping cell cope with protein-misfolding during heat stress
Sigma-38 (RpoS) = General stress-response gene regulator that activates genes needed for survival in stationary growth phase or in stress
What was the experimental evidence for the existence of regulons?
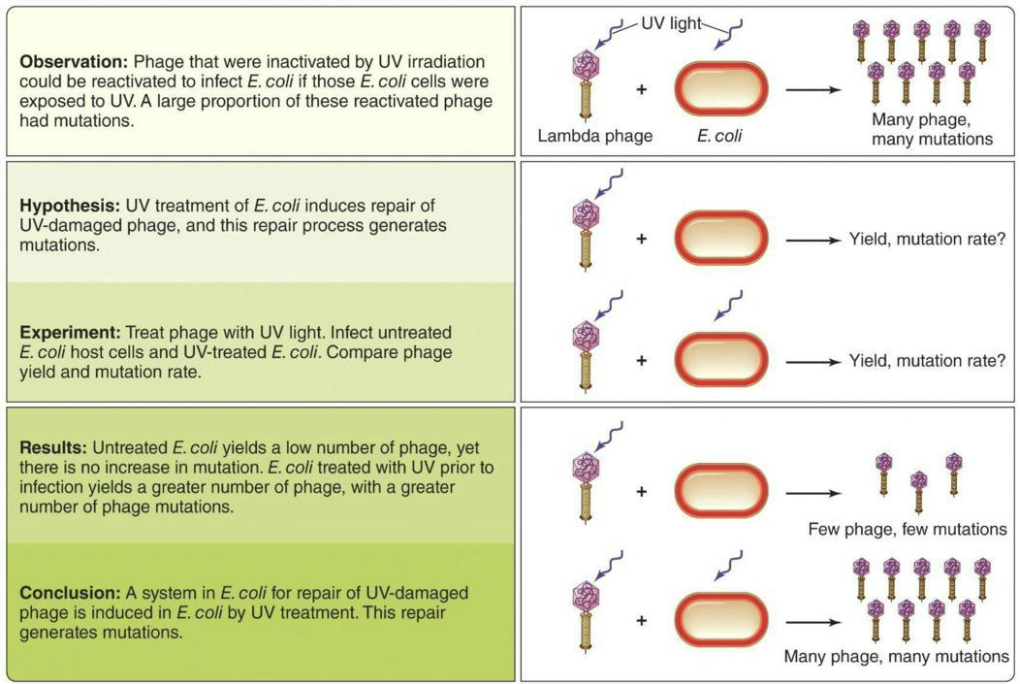
SOS response allows a cell to recognize and respond to serious DNA damage.
Early studies by Jean-Jacques Weigle showed UV exposure induces nonspecific repair mechanisms in bacteria.
Bacteria pre-exposed to UV light, then infected with damaged phages, repaired the phage DNA but also introduced more mutations.
Simplified: The SOS response fixes damaged DNA (bacteria and phage). It provided evidence for regulons because UV damage turned on many DNA repair genes at once, showing they are controlled together by a single system.
How did scientists discover which genes turned on when DNA was damaged?
Experimental evidence for the existence of regulons – SOS response
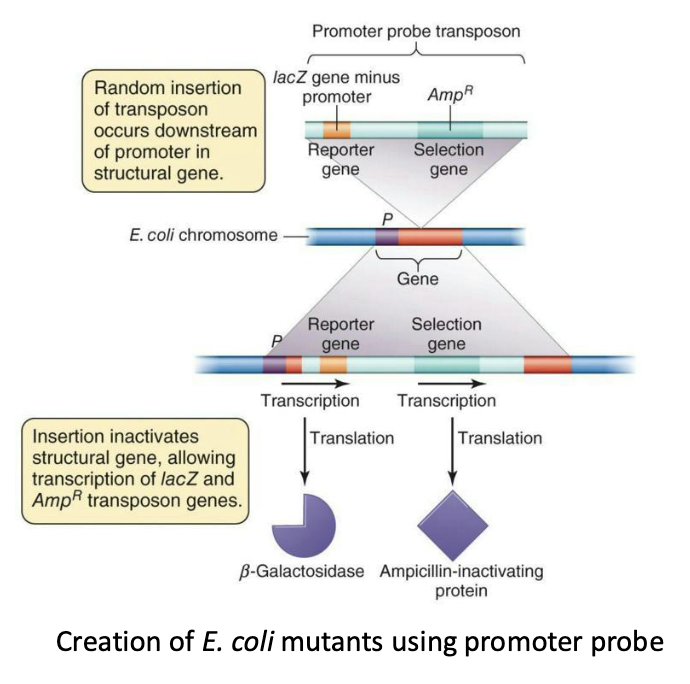
A promoter-probe transposon used to mutate E. coli.
It contains promoterless lacZ reporter gene that can only be expressed if inserted within actively transcribed gene.
When it inserted next to a gene that was expressed in response to DNA damage, it was also expressed (resulting in blue colonies).
This showed which genes were required for DNA damage repair
What happens after the introduction of the DNA-damaging agent mitomycin C?
Experimental evidence for the existence of regulons – SOS response
Insertions in din (damage inducible) genes are strongly induced immediately after the introduction of the DNA-damaging agent mitomycin C.
They are identified through a two-step screening process to determine which promoter-probe fusions are expressed only in the presence of mitomycin C and not expressed in the absence of mitomycin C.
Step 1: Find promoter fusions that turn on after mitomycin C treatment.
Step 2: Confirm they stay off when there’s no DNA damage.
Simple: When DNA is damaged by mitomycin C, multiple din genes are turned on at the same time. This shows they are co-regulated as a regulon, since they respond together only to DNA damage.
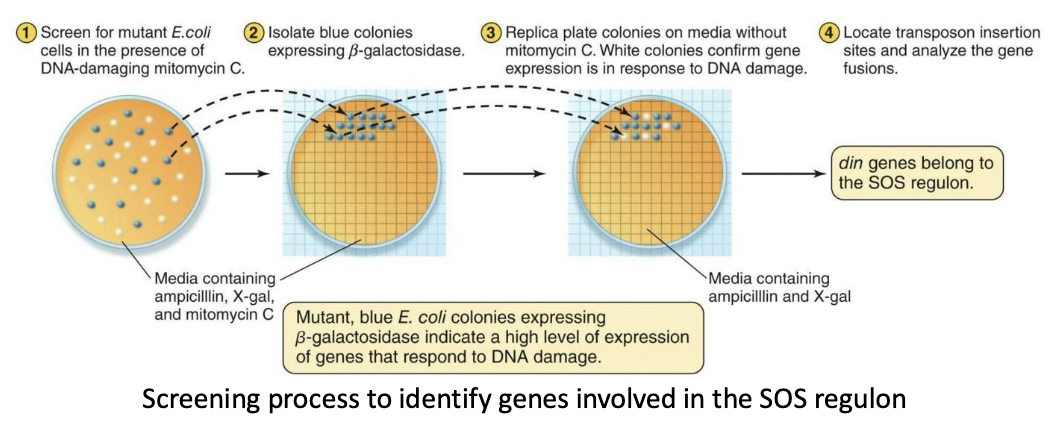
What happens once basic genes are identified?
Experimental evidence for the existence of regulons – SOS response
Once basic genes were identified, regulatory systems for the genes were studied.
Scientists damaged DNA and screened for blue (constitutively expressed) or white (unable to be expressed) genetic mutants from the previous strains.
Blue → DNA damage repair genes
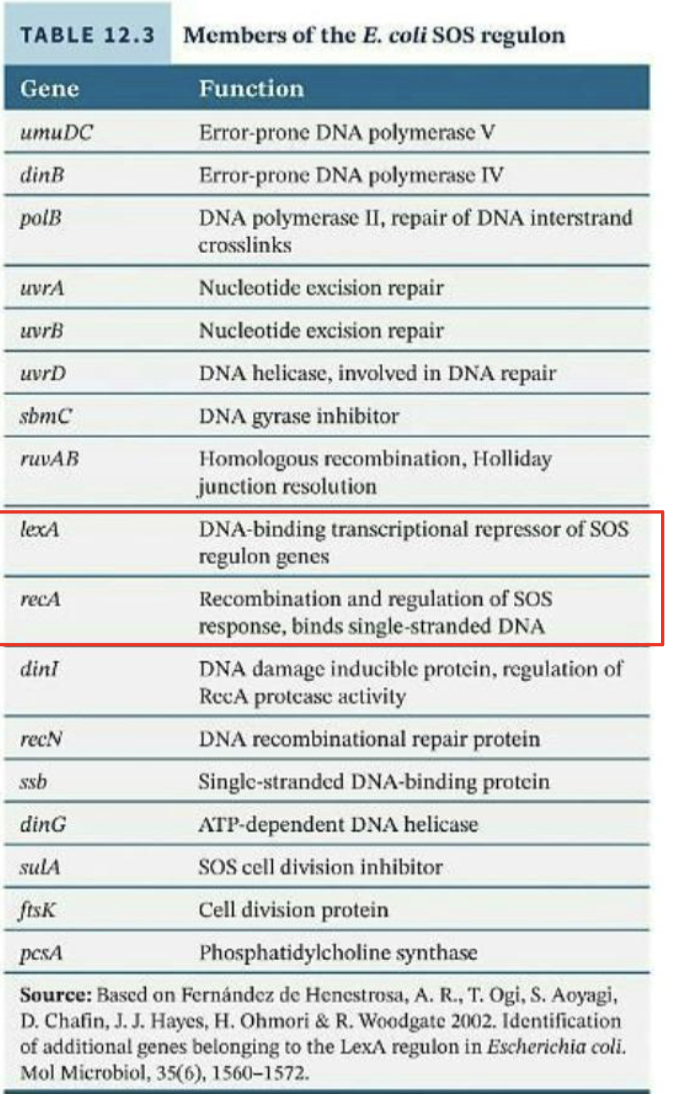
Using this process, numerous important genes and regulators were identified.
Two of the most important to the process were recA and lexA

How does the cell sense DNA damage?
It senses ssDNA
The LexA repressor inhibits the expression of genes under SOS control
When DNA damage occurs, the RecA protein interacts with ssDNA produced by damage to DNA, causing it to mediate cleavage of the LexA repressor
This action results in relief of the SOS genes from repression, activating their expression
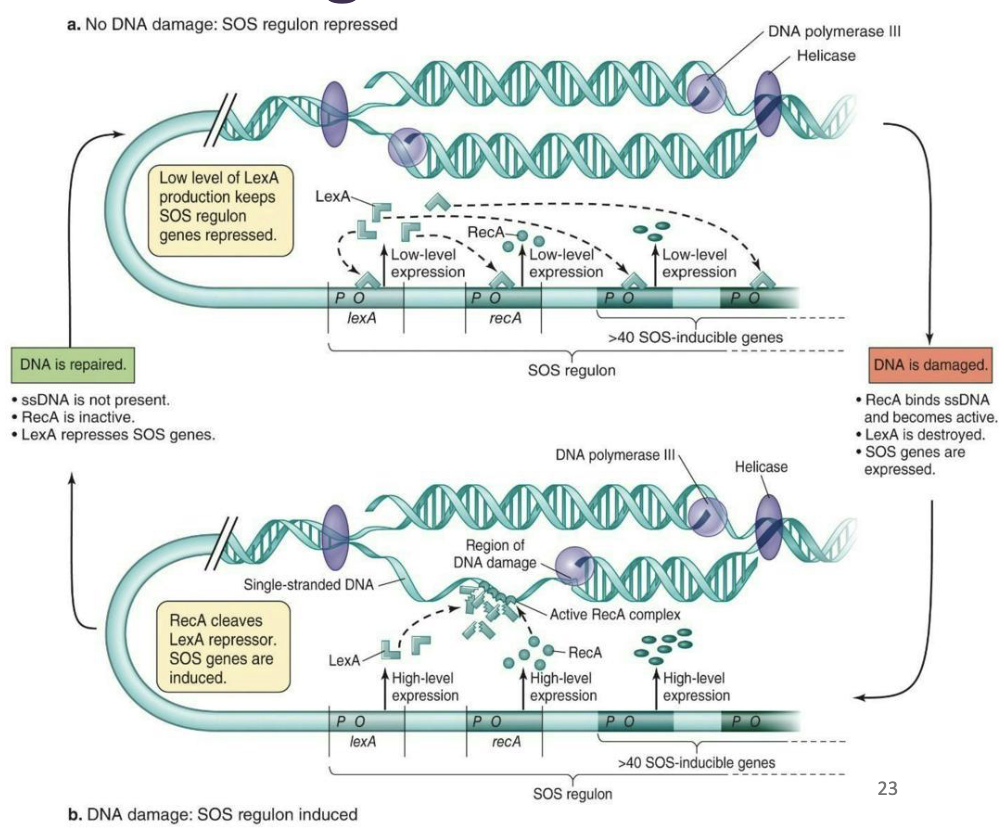
Summary
Constitutive expression refers to genes that always are expressed.
Inducible expression refers to genes that are expressed under certain conditions.
Gene expression regulation can occur at different levels (e.g., transcription initiation, protein activity).
The operon is the unit of transcription, whose expression is under the control of discrete regulatory elements, including promoters and operators.
In negative control, a repressor blocks transcription. Effectors can bind to repressors and modulate their activity.
These effectors can function as inducers or corepressors.
In positive control, an activator molecule may bind to an activator binding site, leading to increased transcription.
In the lac operon, activator protein (cAMP receptor protein or CRP) serves as an activator.
In the presence of glucose and lactose, E. coli exhibits a diauxic growth curve.
Alternative sigma factors bind to specialized promoters resulting in transcription initiation.
Regulons are coordinately regulated genes that respond to the same regulatory systems.
Catabolite repression and the SOS response represent well-studied regulon systems
How can mRNA be controlled?
Regulatory RNAs
All genomes carry regions of DNA coding for non-translated RNA.
rRNA/tRNA molecules
Small noncoding RNAs (sRNA) that can control gene expression at transcription or translation points
Antisense RNA, works by binding to complementary mRNA molecules in their leader sequences (5’ UTR), affecting gene expression
Ex: dsrA binds to rpoS transcript disrupting the inhibitory secondary structure, and allowing translation to proceed
Simple: Antisense RNAs like dsrA let the cell quickly turn on specific proteins, such as RpoS, by unblocking their mRNAs only when needed, like during stress.
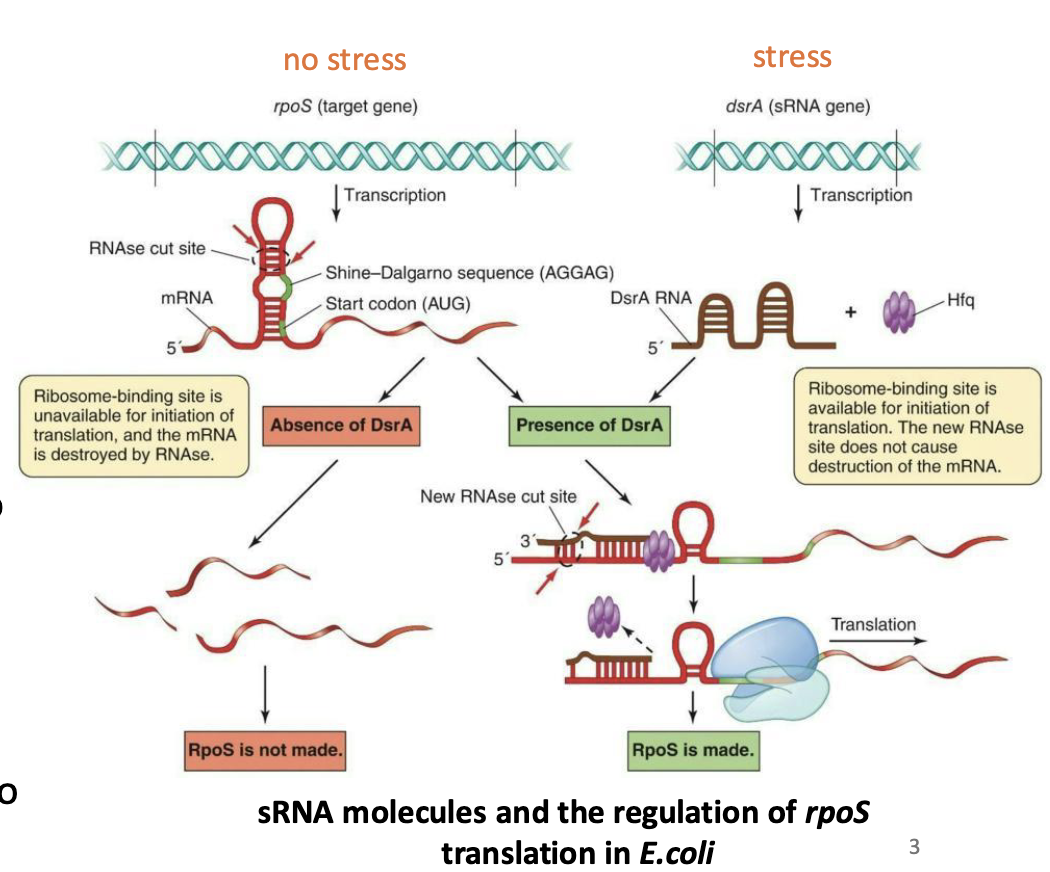
What is attenuation?
Control of transcription by mRNA secondary structure
Attenuation = Interaction between translation and transcription processes
If ribosome quickly follows RNA polymerase, terminator hairpin RNA loops are formed in the leader sequence and the polymerase detaches.
“Stalling out” of ribosome in mRNA leader sequence allows transcription to continue.
This process can’t occur in eukaryotes because translation and transcription isn’t coupled.
This occurs after initiation of transcription but prevents it from continuing.
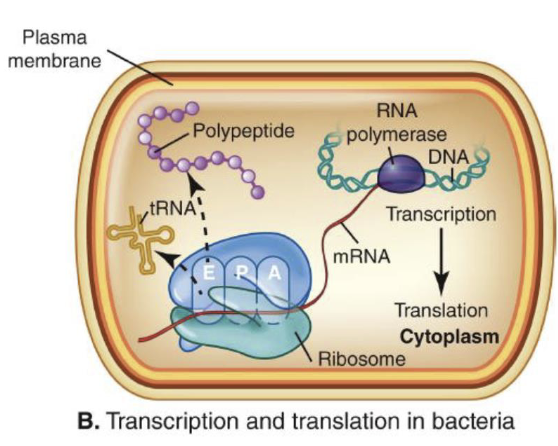
How is attenuation regulated after transcription initiation?
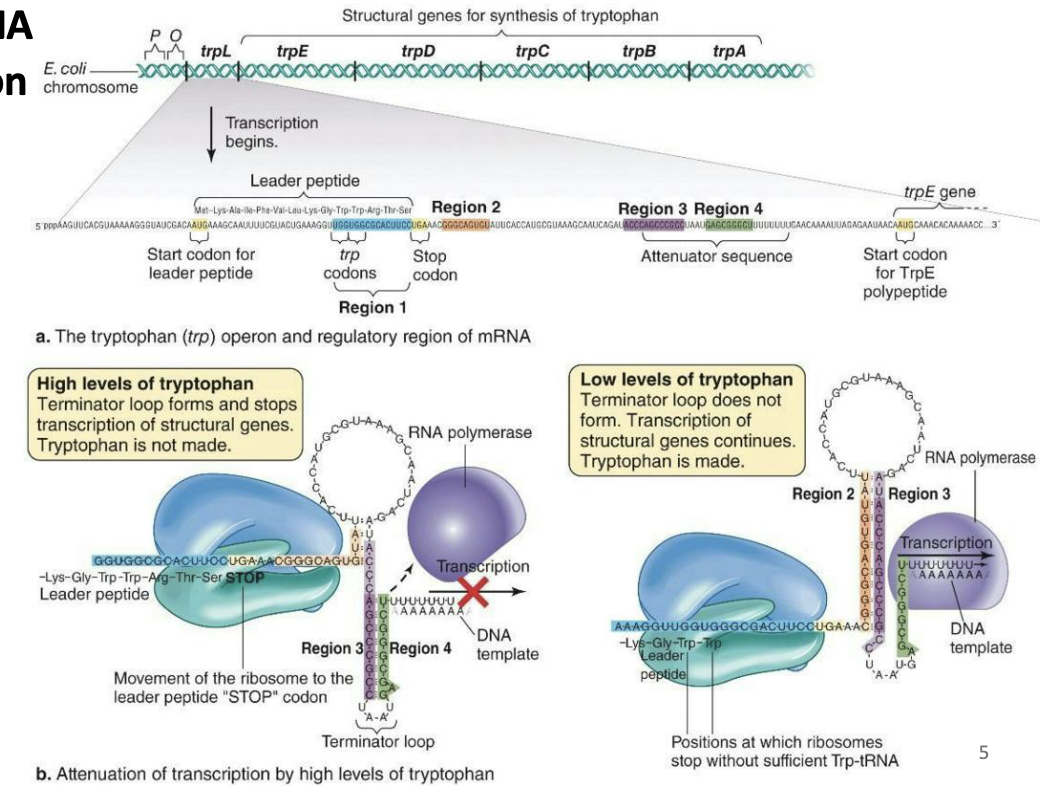
Control of transcription by mRNA secondary structure: Attenuation
Control of the trp operon
Regulation by attenuation occurs after transcription initiation
The trpL gene is the 1st gene in the operon, encodes a leader peptide, and contains an attenuator sequence
Under high levels of tryptophan, region 3 binds to region 4 to stop transcription (fast ribosome)
Under low levels of tryptophan, region 3 binds to region 2, allowing transcription to proceed (slow ribosome)
What are riboswitches?
Control of transcription by mRNA secondary structure
Riboswitches = Regulatory molecules that bind RNA and alter its shape
Changed shape in leader areas of mRNA can prevent continuation of transcription.
Changed shape around start codons in mRNA can prevent ribosomes from translating the mRNA.
These are capable of controlling either transcription or translation, depending on the conformation induced in the RNA molecule.
**Don’t memorize RNA-derived Compounds
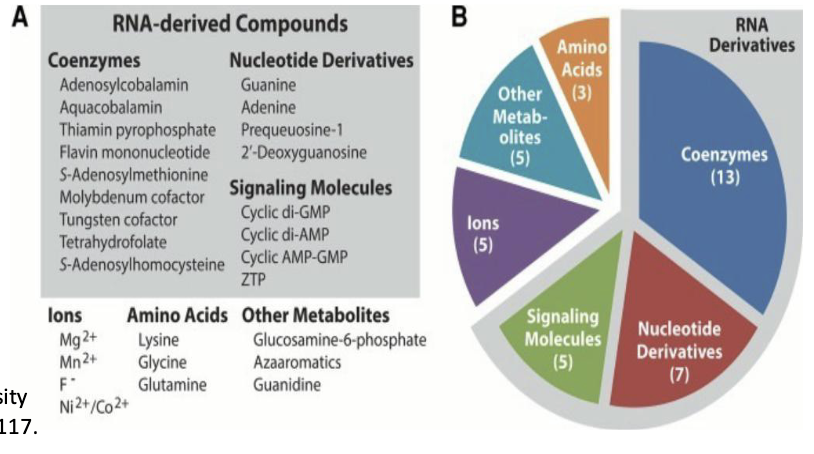

How do riboswitches work to regulate transcription vs. translation?
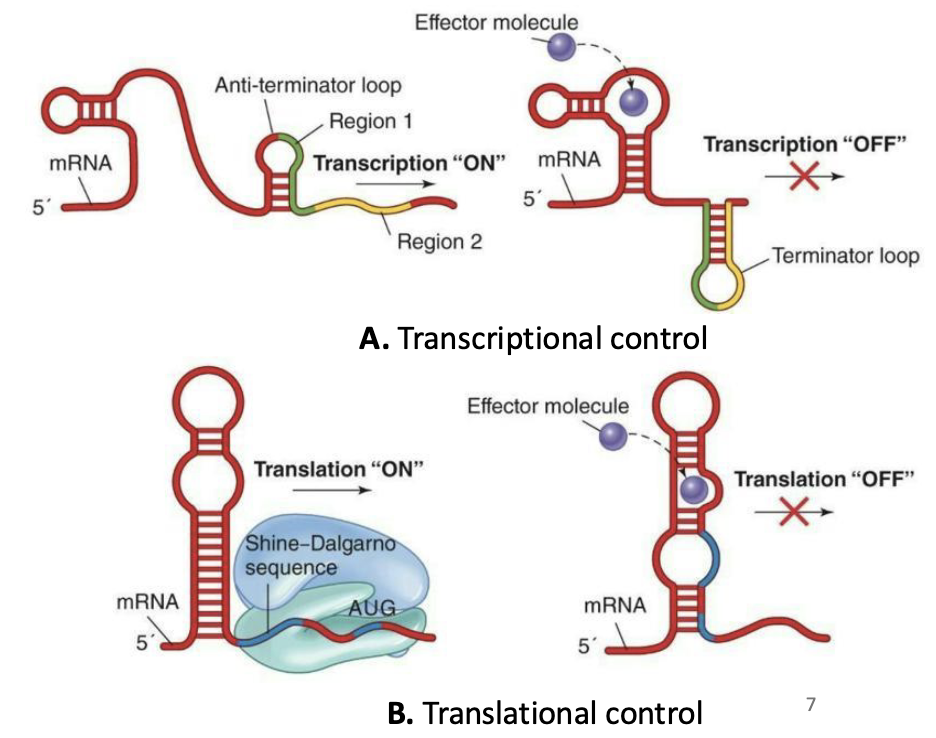
Control of transcription by mRNA secondary structure: Riboswitches
Messenger RNA can bind effector molecules such as vitamins or amino acids that regulate gene expression
Transcription riboswitches → affect RNA polymerase activity (whether mRNA is made).
Translation riboswitches → affect (inhibit) ribosome activity (whether mRNA is translated into protein).
How do bacteria communicate with their neighbors?
Gene expression can also be used as a means of communication between microbes.
This chemical signaling system is known as "quorum sensing."
The term refers to a number of members of a group that must be present in order to conduct business (a quorum).
Cells release autoinducer molecules into the environment as the population density increases.
Detecting changes in autoinducer levels causes regulation of gene expression.
What is Lux?
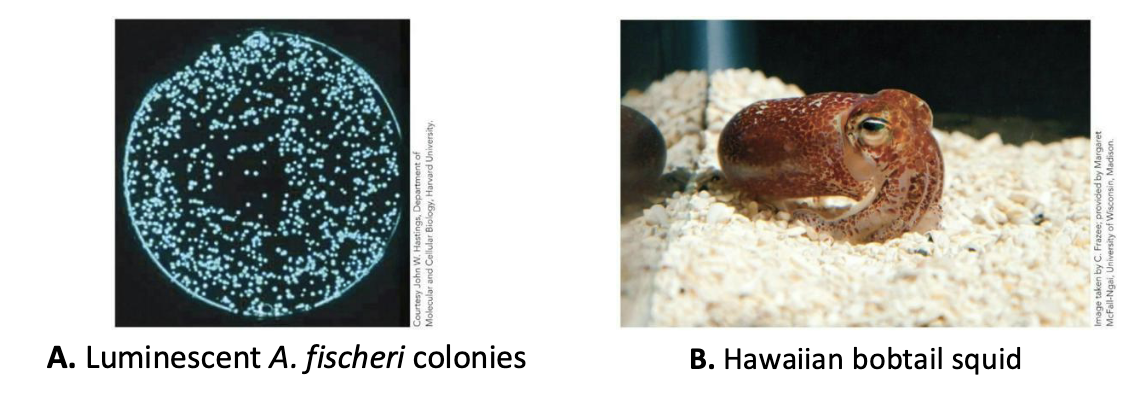
Lux, a prototypical quorum-sensing system
Found in Aliivibrio fischeri
Can live freely or in symbiosis with the Hawaiian bobtail squid.
The cells only emit light (via the enzyme luciferase) when in the light organ of the squid (or in a lab setting).

How do the bacteria know when to produce luciferase?
Lux, a prototypical quorum-sensing system
When grown to high density, the cells produce N-acyl-homoserine lactone (AHL).
This autoinducer stimulates luminescence.
The LuxI protein catalyzes AHL synthesis.
At low density, the cells don’t produce enough AHL to induce light emission.
Examination of how cells detect levels of AHL has been an area of active research.
LuxR regulator transcriptional activator interacts with AHL when it reaches a high enough concentration.
The complex binds the “lux box” DNA regulatory site.
This leads to transcription of luxA/luxB (coding for the luciferase protein) and luxI (positive feedback loop forming more AHL).
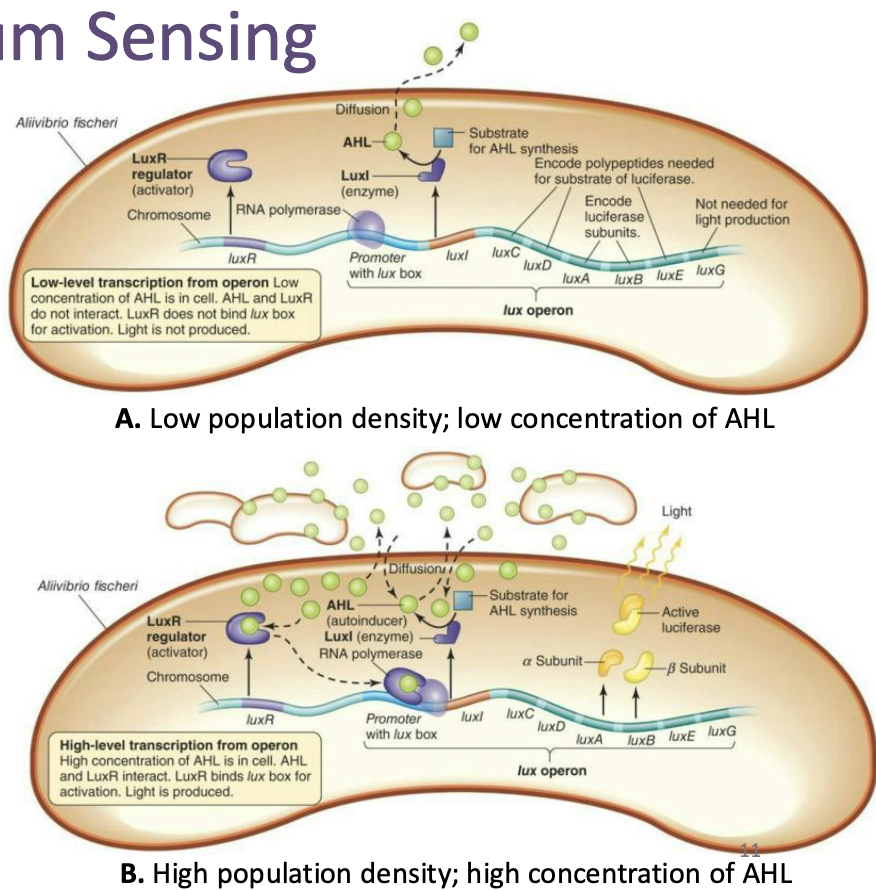
How is gene expression dependent on cell density?
Gene expression is dependent upon cell density
At low density, transcriptional regulation by quorum sensing involves the production of small amounts of AHL by enzyme Luxl encoded by the luxl gene
At high cell density, the concentration of AHL increases, and they can bind the LuxR transcriptional activator protein encoded by luxR
This results in increased affinity of the transcriptional activator protein for the lux box and increased transcription of the lux operon (positive feedback)
What is the widespread occurrence of quorum sensing?
A broad range of microbes possess quorum-sensing systems.
Mechanisms controlled include:
Motility
Conjugation (sex pili)
Biofilm formation
Pathogenesis (e.g., cholera toxin formation)
Autoinducers may even play a role in competition, interrupting or inhibiting a control pathway in other organisms in the environment.
How do environmental conditions affect gene expression?
Such systems can use one protein as a sensor and another to control transcription.
This allows for response to changes in the environment.
Signal transduction induced inside the cell alters it to respond appropriately.
These two-component regulatory systems are the most common regulatory systems in bacteria
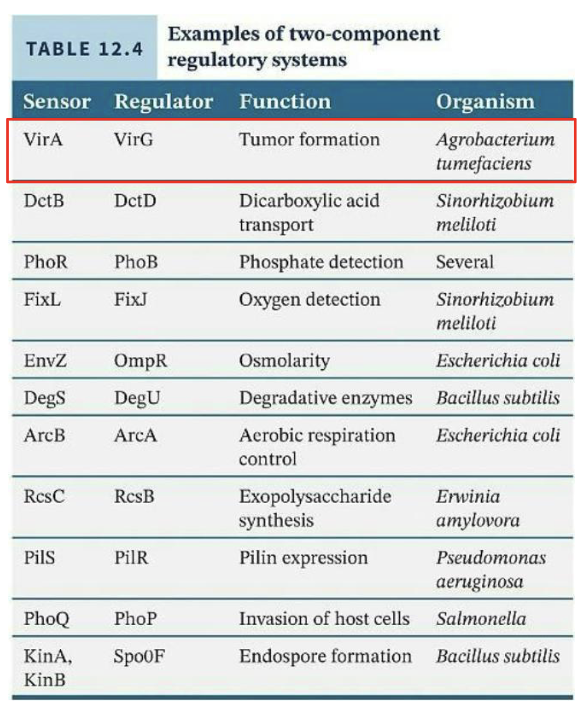
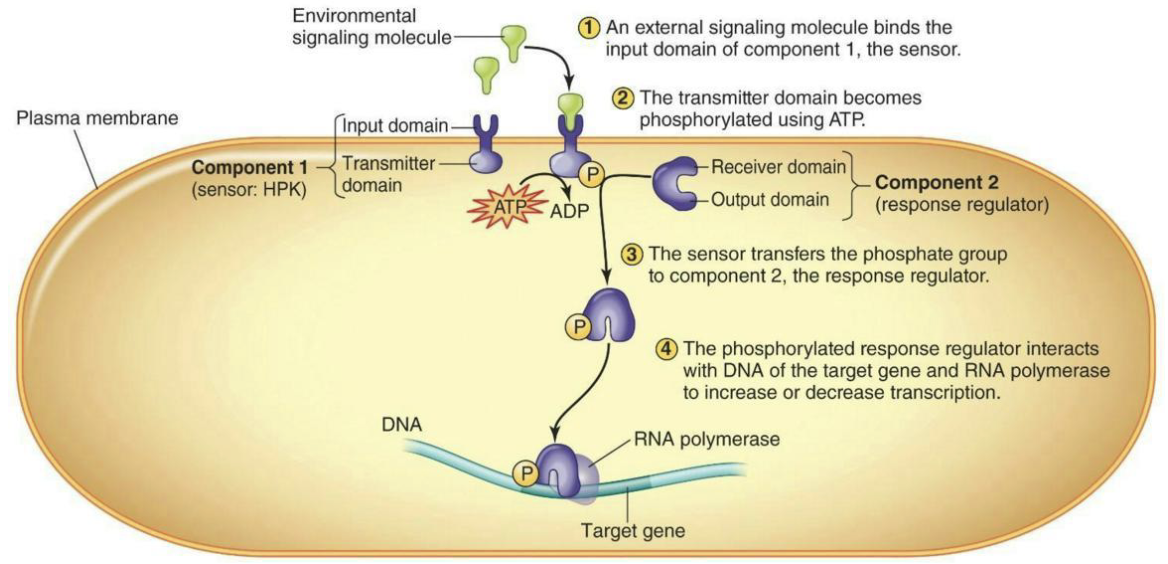
What does the two-component regulatory system involve?
Sensor kinase (often histidine protein kinase, or HPK) to detect the environmental stimulus
Input Domain → Outside cell
Response regulator (RR) to regulate transcription
Inside cell
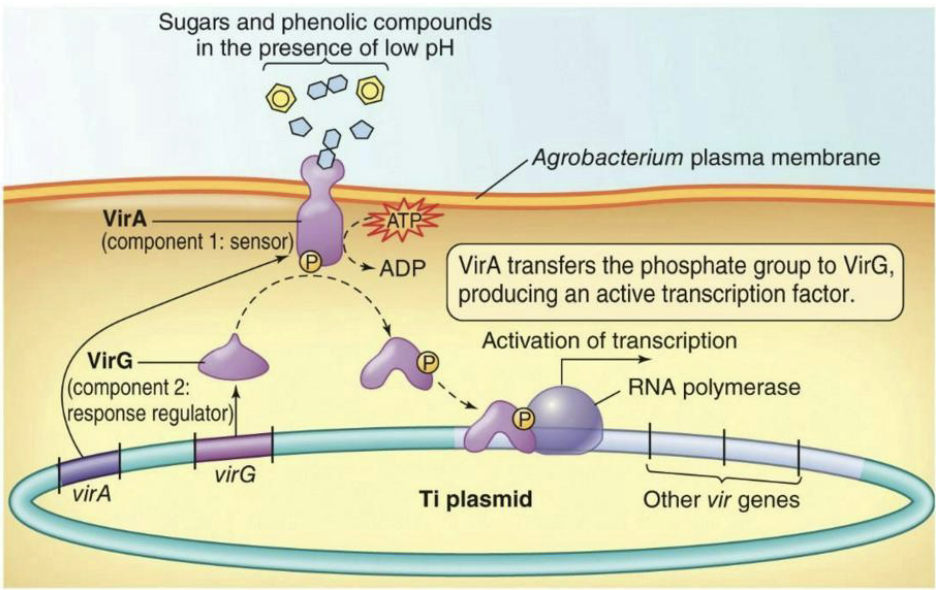
What is the virulence of Agrobacterium tumefaciens?
Causes tumours on plants, which led to a revolution in plant biotechnology
vir genes found on the Ti plasmid
Only expressed under conditions similar to a plant wound site (sugars and phenolic compounds in the presence of low pH)
virA/virG are required for expression of the other virulence genes.
VirA is a transmembrane HPK (histidine protein kinase) protein
VirG is a transcriptional activator RR protein.
How it works: VirA senses the signal, adds a phosphate to VirG → VirG becomes active → turns on the other vir genes needed for infection.
Overall, what is the function of the two-component regulatory system?
Collectively, different two-component regulatory systems can allow microbes to respond differently to environmental stimuli.
By specifically pairing particular HPKs and RRs, cells can better control which genes are expressed in response to cues from the environment.
How can bacterial cells regulate their behaviour?
The mechanism of chemotaxis - movement of motile bacteria toward favorable chemicals (nutrients) or away from harmful chemicals (toxins or poisons)
An example of a complex bacterial behavior modulated by shifts in protein activity
Controlled at the level of protein activity, rather than via changes in gene expression.
Chemotactic bacteria sense changes in chemical gradients over time.
Changes induce altered direction and duration of flagellar rotation, leading to directed movement over time.
How can you study chemotaxis using mutants?
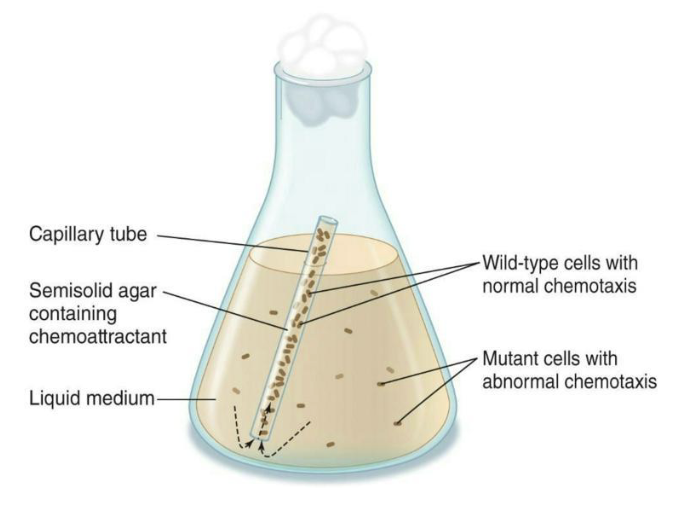
Chemotaxis mutants isolated using a capillary tube filled with nutrients
Microbes with normal chemotaxis will move into the tube.
Those with mutations in chemotactic proteins will remain outside the tube.
Multiple che gene mutant strains have been isolated and studied in this manner.
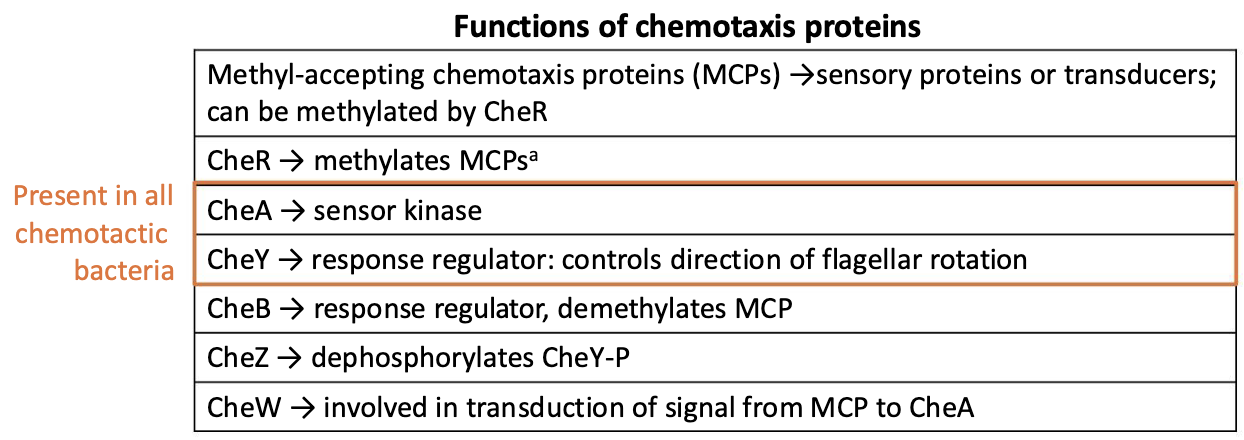
What are che proteins?
Che proteins = Two-component regulatory systems
CheA works as the sensor kinase, becoming phosphorylated.
CheA then phosphorylates CheY (the RR protein).
However, the phosphorylated RR proteins do not bind DNA; they bind to the flagellar motor, changing its activity
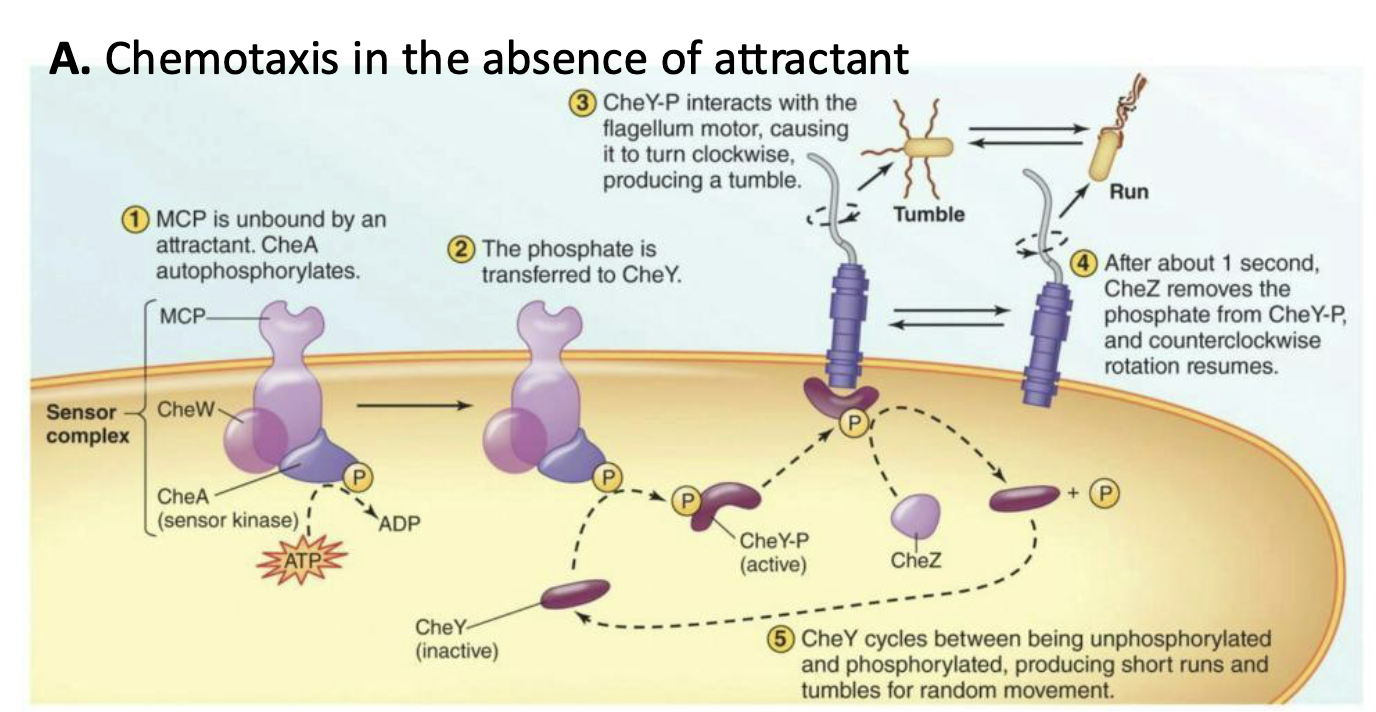
What happens to chemotaxis in the absence of attractant?
CheA is phosphorylated, which in turn phosphorylates CheY
The resulting CheY-P interacts with flagellum, signaling it to rotate clockwise (CW), causing a tumble
Removal of the phosphate from CheY-P disrupts interaction with flagella, resulting in counterclockwise (CCW) rotation, and the cell runs
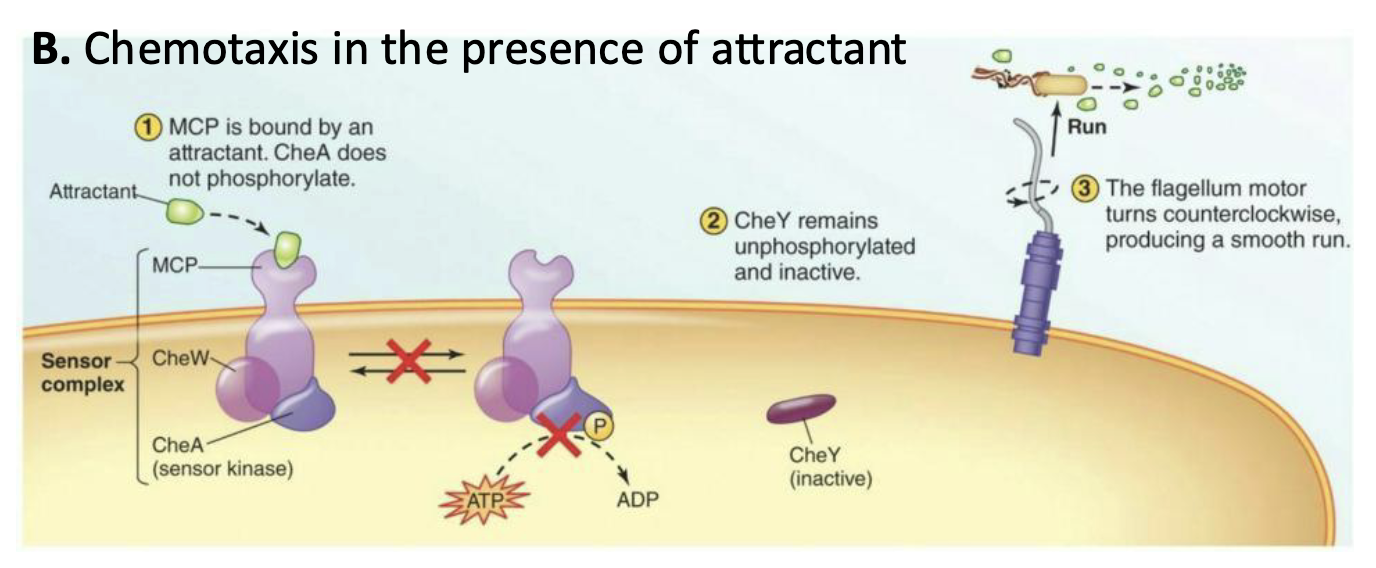
What happens to chemotaxis in the presence of attractant?
Phosphorylation of CheA is inhibited
The absence of CheY-P results in counterclockwise rotation of the flagella, causing a longer run
Summary
mRNA can be controlled by changes in its secondary structure (attenuation), riboswitches or sRNAs
Quorum sensing allows bacteria to sense population status
This signal is often the autoinducer molecule N-acyl-homoserine lactone
Bacteria use two-component regulatory systems to regulate gene expression in response to environmental signals
These consist of histidine kinases and response regulators
chemotaxis - movement of motile bacteria toward favorable chemicals (nutrients) or away from harmful chemicals (toxins or poisons), Controlled at the level of protein activity
What is some genome sequencing history?
1977 | Sanger develops a DNA sequencing technique which he and his team use to sequence the first full genome - an E. coli virus called X174 (5,375 nt). |
1983 | Mullis develops the polymerase chain reaction (PCR) -for amplifying DNA. |
1990 | Human Genome Project is launched. The project aims to sequence all 3 billion letters of a human genome in 15 years. |
1995 | Venter and colleagues completed the first bacterium genome sequence (Haemophilus influenza - 1.8 Mb) |
1996 | Venter and colleagues completed the first archaea genome sequence (Methanococcus jannaschii, - 1.66 Mb) |
1997 | Blattner and colleagues complete the Escherichia coli K-12 genome sequence |
2001 | First draft of the human genome sequence released. |
2007 | A new DNA sequencing technology is introduced that increases DNA sequencing output 70 fold, in one year! |
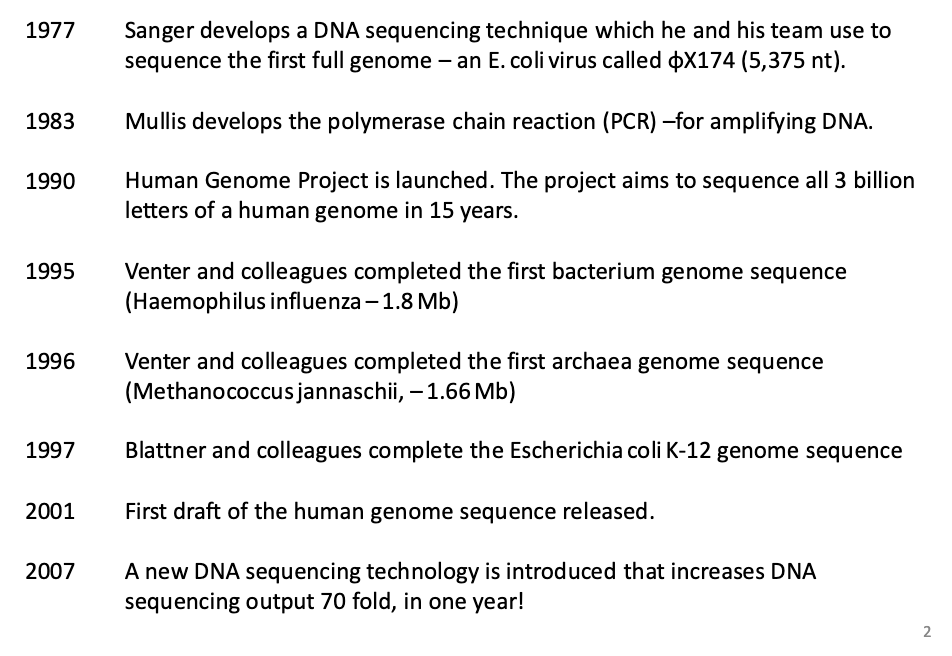

How are genome sequences determined?
DNA sequencing
Genomics = methods for studying the entire genome of a microbe
In 1995 team of scientists decoded the complete genome of Haemophilus Influenzae using “shotgun sequencing”
Genomics has been spurred by the development of recombinant DNA protocols.
However, this has created several new needs for
Improved DNA sequencing techniques
Formats for storage of very large data sets
Tools for analysis of large data sets generated
AI is revolutionizing the field of DNA sequencing by enhancing the speed, accuracy, and efficiency of genomic analysis
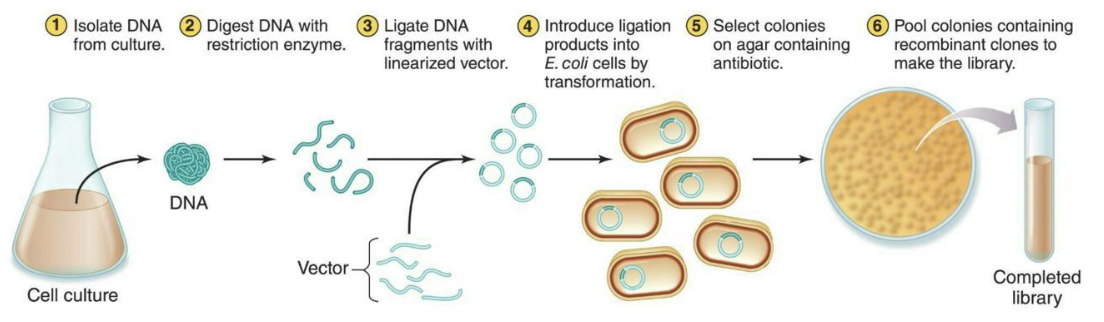
What are genomic libraries?
Made of the entire genome of an organism.
The specific fragments of DNA from the organism are cloned into plasmid vectors.
A genome of interest and a plasmid vector are digested with a restriction enzyme
The resulting fragments are mixed and ligated
The ligation products are introduced into suitable host cells, usually E. coli
Cells containing a plasmid are selected.
What is Sanger sequencing?
Sanger, or dideoxy, sequencing
In brief, requires three steps:
Cloning of the gene fragment to be sequenced
DNA synthesis
Electrophoresis
The method exploits primer specific DNA synthesis activity of DNA polymerase and the fact that DNA polymerases require a free 3' hydroxyl group.
By placing dideoxynucleotides (lacking that free 3' OH group) into the DNA synthesis mixture, the process is terminated with a distinct, labeled endpoint nucleotide.
Gel electrophoresis can separate the fragments of different lengths and detect which labeled nucleotide is on the end of each fragment, providing a sequence
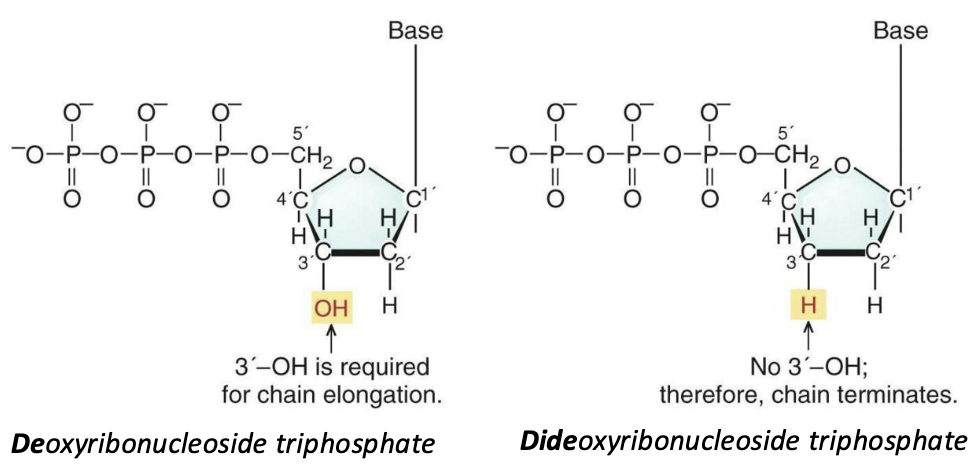
What does Sanger sequencing rely on?
The Sanger sequencing method relies on the use of special dideoxyribonucleotides (ddNTP) that can be inserted into a growing DNA strand by DNA polymerase but block the addition of further nucleotide
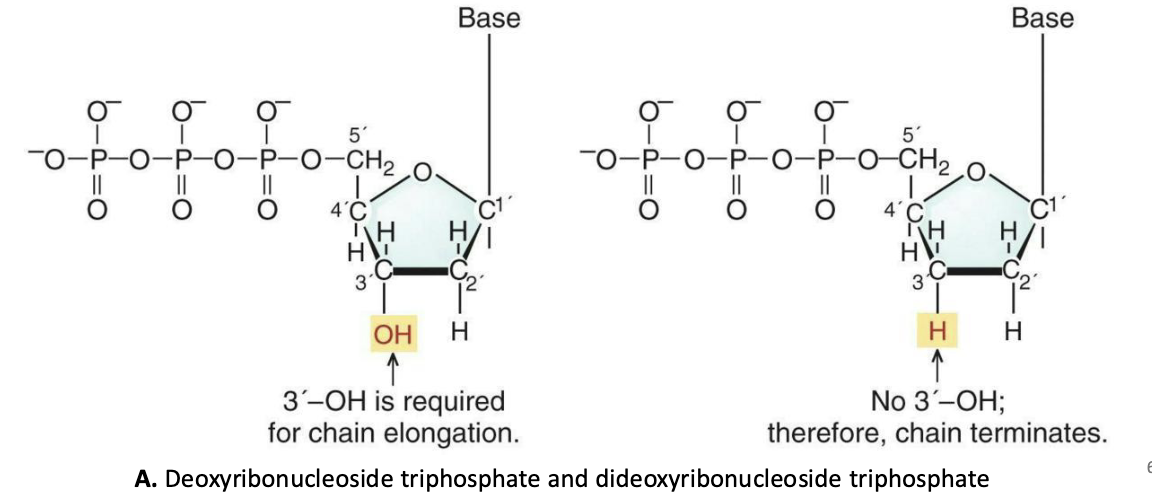
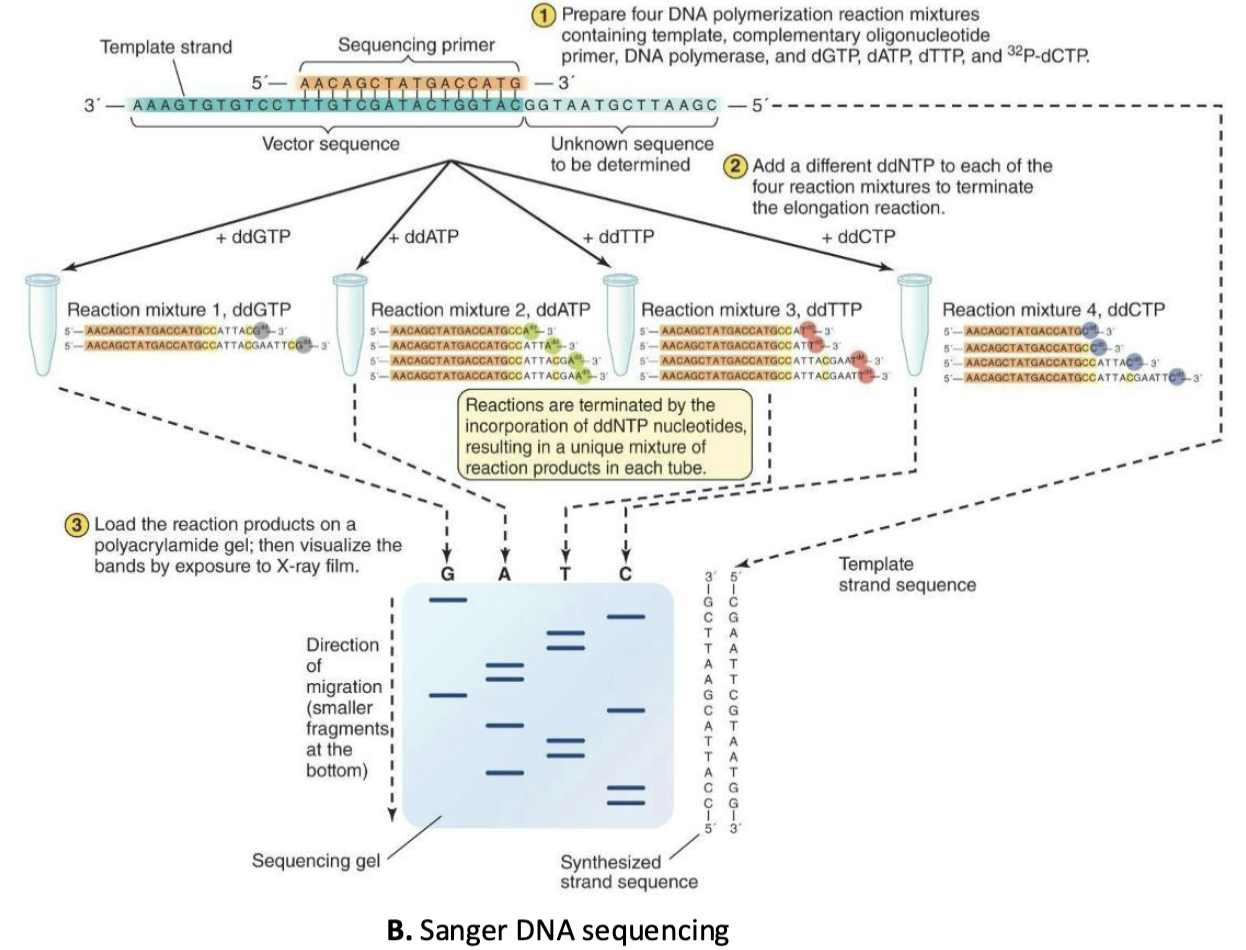
What are the steps of Sanger sequencing?
The Sanger sequencing method is based on the synthesis of a DNA strand, using a template strand to guide the insertions of nucleotides.
Plasmid with cloned DNA is denatured to provide ss template
Four DNA polymerization reactions prepared
32P-dCTP (highlighted in yellow) is radioactively labeled
ddNTP is added to each reaction – polymerization reaction is terminated
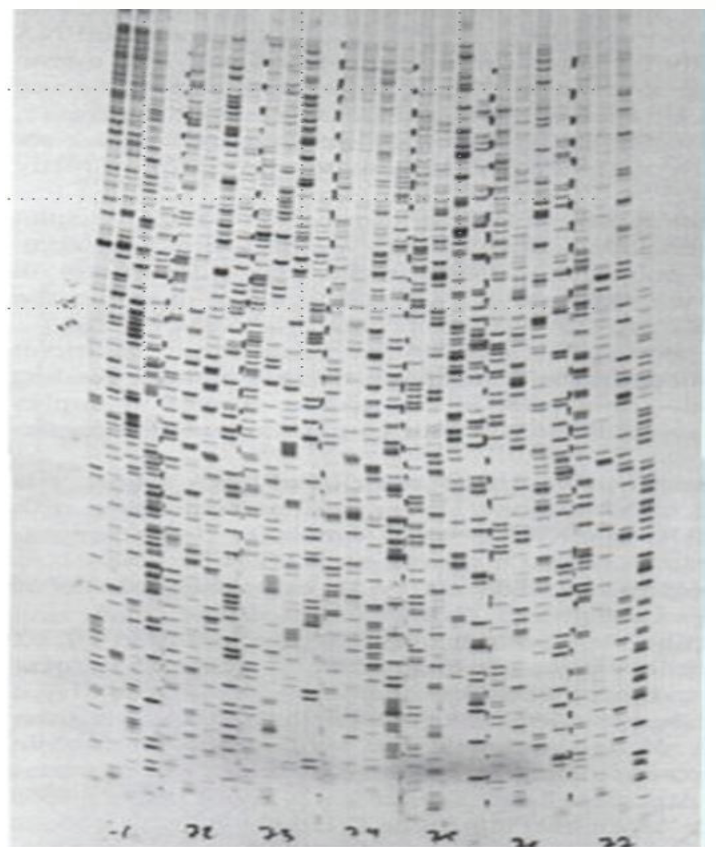
Products of each reaction are detected by exposure to X-ray film after electrophoresis
Simple: Each band represents where DNA synthesis stopped, and the lane tells you which base caused the stop. Reading fragments from smallest to largest reveals the sequence order.
What are method improvements to Sanger sequencing?
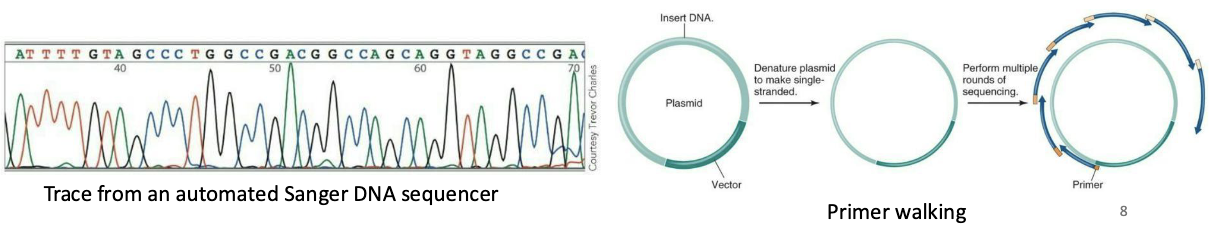
Sanger, or dideoxy, sequencing – method improvements
The use of thermostable polymerases, allows multiple rounds of synthesis from a single template strand
Automated methods using fluorescent labels instead of radioactive labels are safer, cheaper, and easier. Each ddNTP can labeled with a different fluorescent dye.
Sequences of 700 to 1,000 bases obtained cheaply in hours.
Longer sequences obtained by “primer walking,” using repeated rounds of sequencing with primers complementary to the end of the last segment sequenced (not possible for LARGE genomes like human genome)
What is high-throughput sequencing?
Also known as next-generation sequencing (NSG) methods
For large scale sequencing, NGS replaced Sanger sequencing
Are far cheaper than Sanger sequencing partially since they don’t require gel electrophoresis.
Far more efficient because it allows the collection of sequences from many molecules simultaneously.
short read, massively parallel sequencing technique is a fundamentally different approach that revolutionized sequencing capabilities
What are examples of NSGs?
Pyrosequencing
454 sequencing
Ion Torrent platform
Illumina sequencing
What is Pyrosequencing (NSG)?
No need in chain-terminating ddNTP, only dNTP used
Detects addition of a nucleotide to the end of a synthesized strand of DNA by production of light
Faster and cheaper than Sanger method
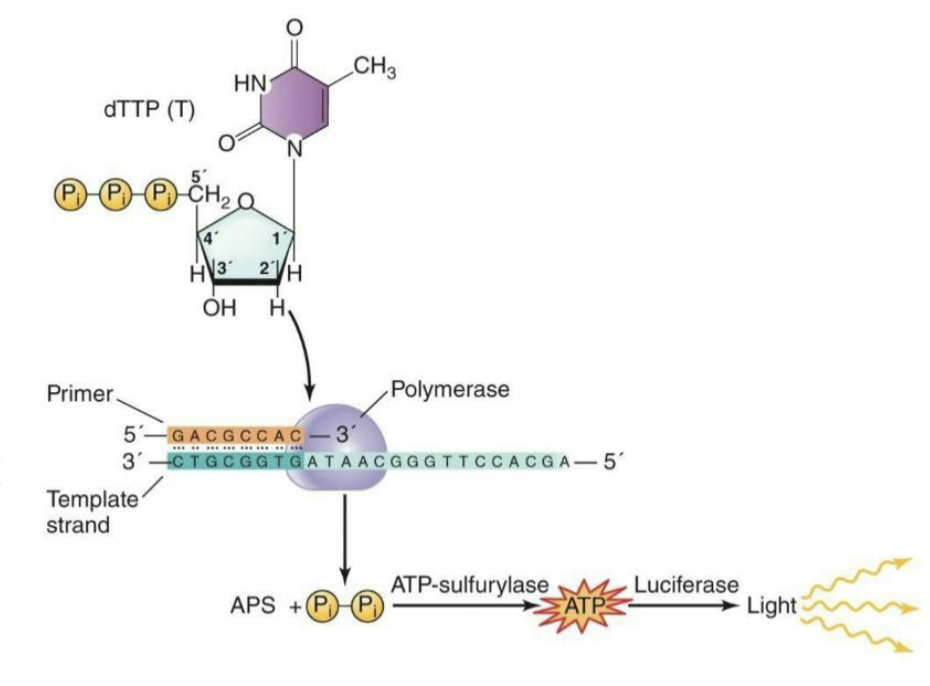
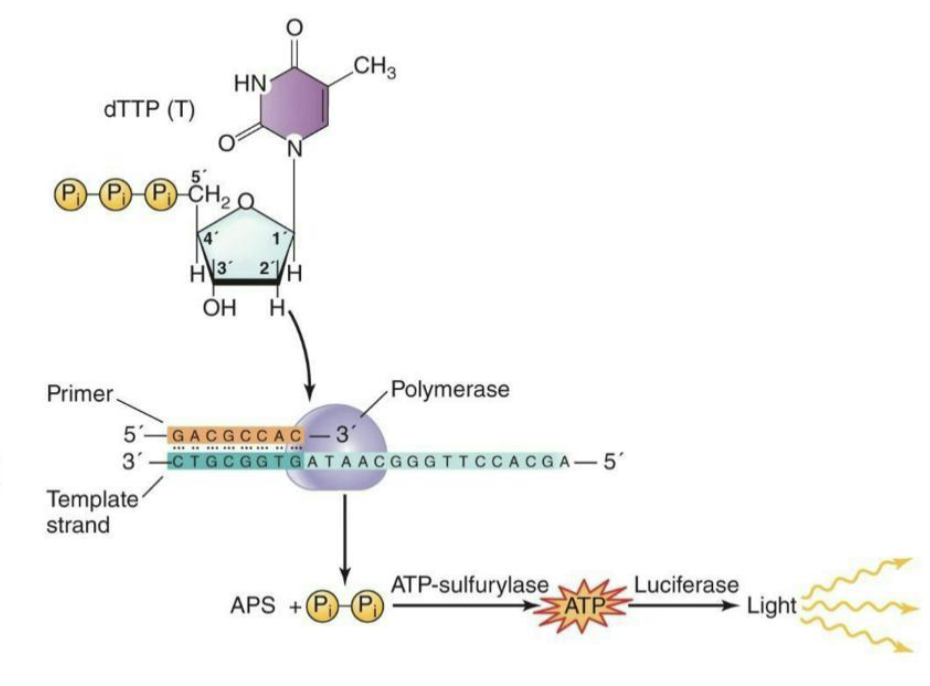
What are the steps of Pyrosequencing?
In the strand shown, DNA polymerase adds dTTP to the growing DNA strand
ATP-sulfurylase reacts with the pyrophosphate that is released upon incorporation with adenosine phosphosulphate (APS), generating ATP
This ATP interacts with luciferase and releases a burst of light
The automated detection of this light by a camera indicates that the nucleotide was added to the growing DNA strand
What is 454 sequencing (NSG)?
DNA doesn’t have to be cloned first, only fragmented.
Technically, many sequencing reactions carried out at once in a well format.
Software analysis makes sense of the many reaction results generated at the same time.
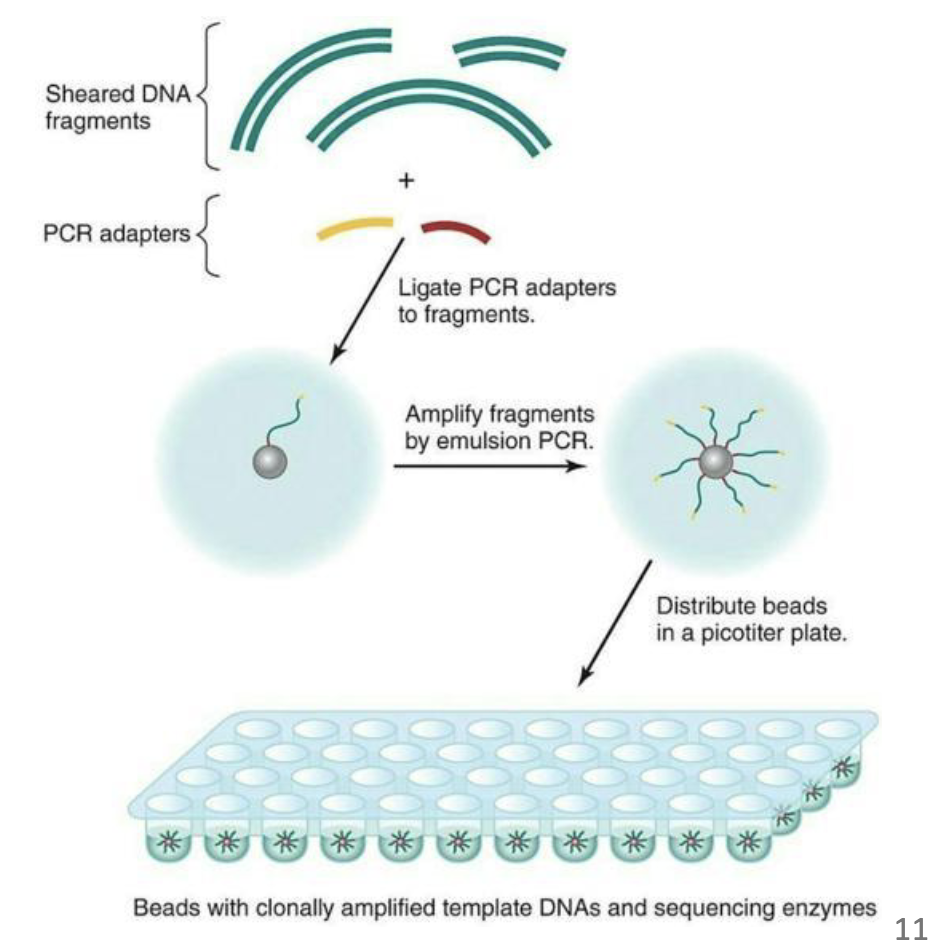
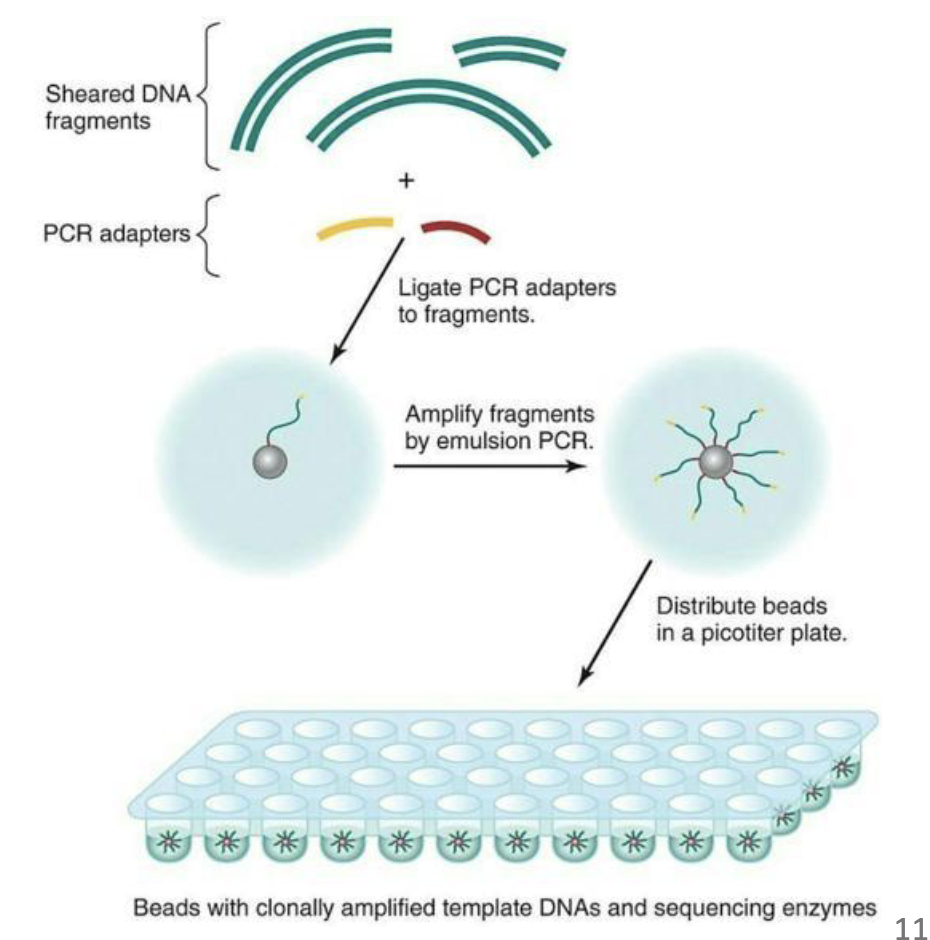
What are steps of of 454 sequencing?
This method starts with shearing of the DNA into fragments.
Fragments are ligated with nucleotide adapters that facilitate the trapping of these fragments on tiny beads
Then amplified by PCR within water droplets, a process called emulsion PCR.
Individual beads with the amplification products are distributed in a flow cell, where repeated pyrosequencing reactions are carried out alternating between each of the four dNTP bases.
ATP produced by the DNA synthesis results in light production by luciferase, which can be measured by a CCD camera.
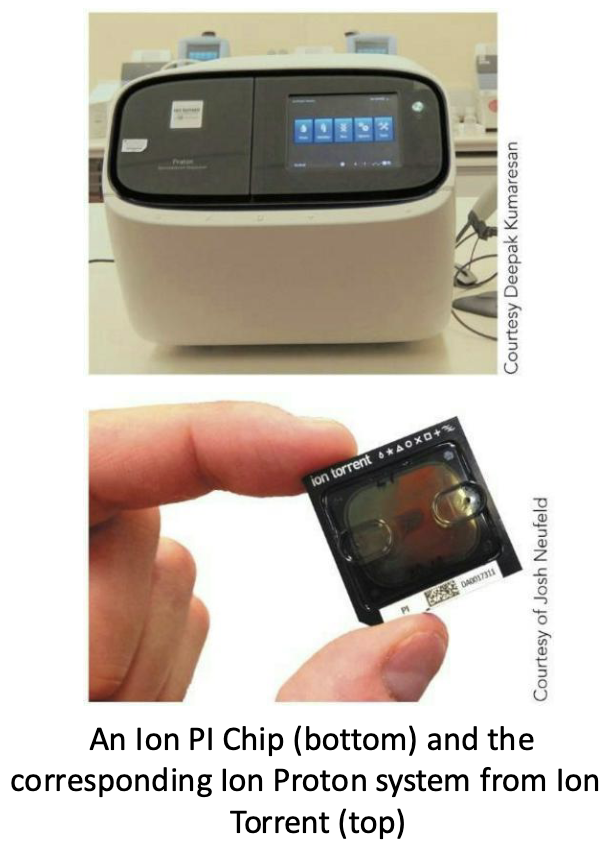
What is the Ion Torrent platform (NSG)?
Same principle as 454- pyrosequencing, but uses a sensor that detects hydrogen ions released each time a nucleotide is incorporated – the world’s smallest pH meter
Reads up to 200 bp

What is Illumina sequencing (NSG)?
DNA polymerase adds fluorescently labeled nucleotides to fragments of DNA. Images are taken, the fluorescent label is removed, more fluorescent labels are added, and so on until millions of DNA fragments are sequenced for hundreds of bases.
Finally, the fragments are analyzed and assembled by a computer.

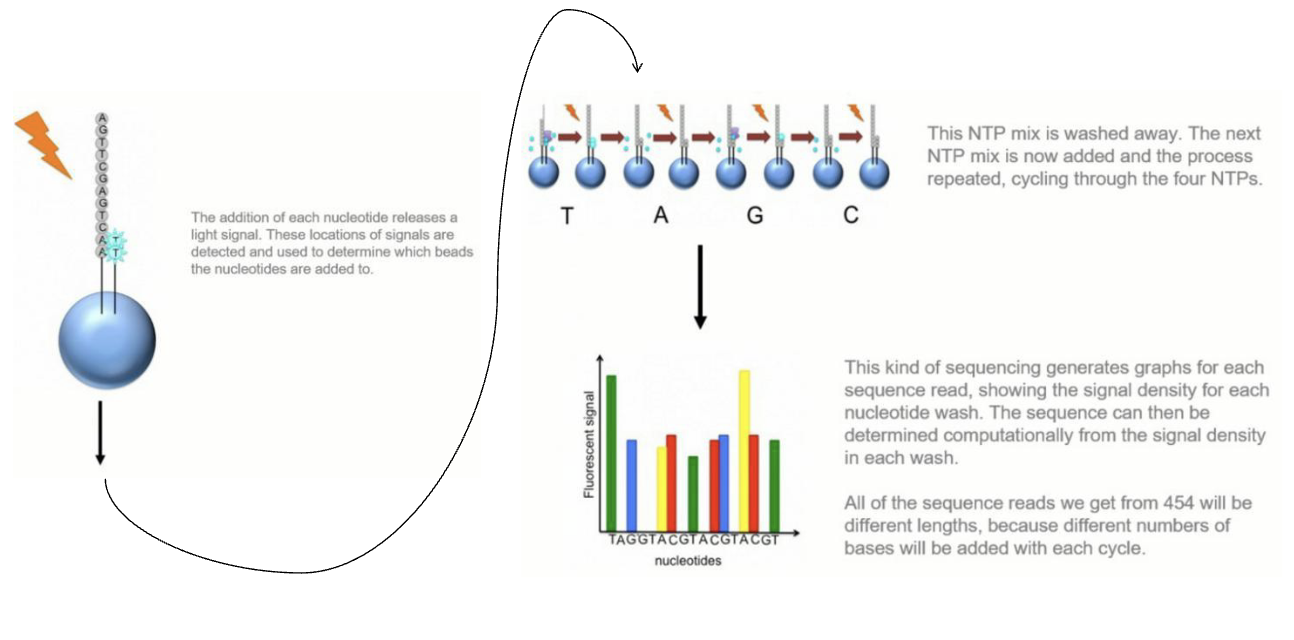
What are the steps of 454 Sequencing?
The addition of each nucleotide releases a light signal. These locations of signals are detected and used to determine which beads the nucleotides are added to.
This NTP mix is washed away. The next NTP mix is now added and the process repeated, cycling through the four NTPs.
This kind of sequencing generates graphs for each sequence read, showing the signal density for each nucleotide wash. The sequence can then be determined computationally from the signal density in each wash.
All of the sequence reads we get from 454 will be different lengths, because different numbers of bases will be added with each cycle.
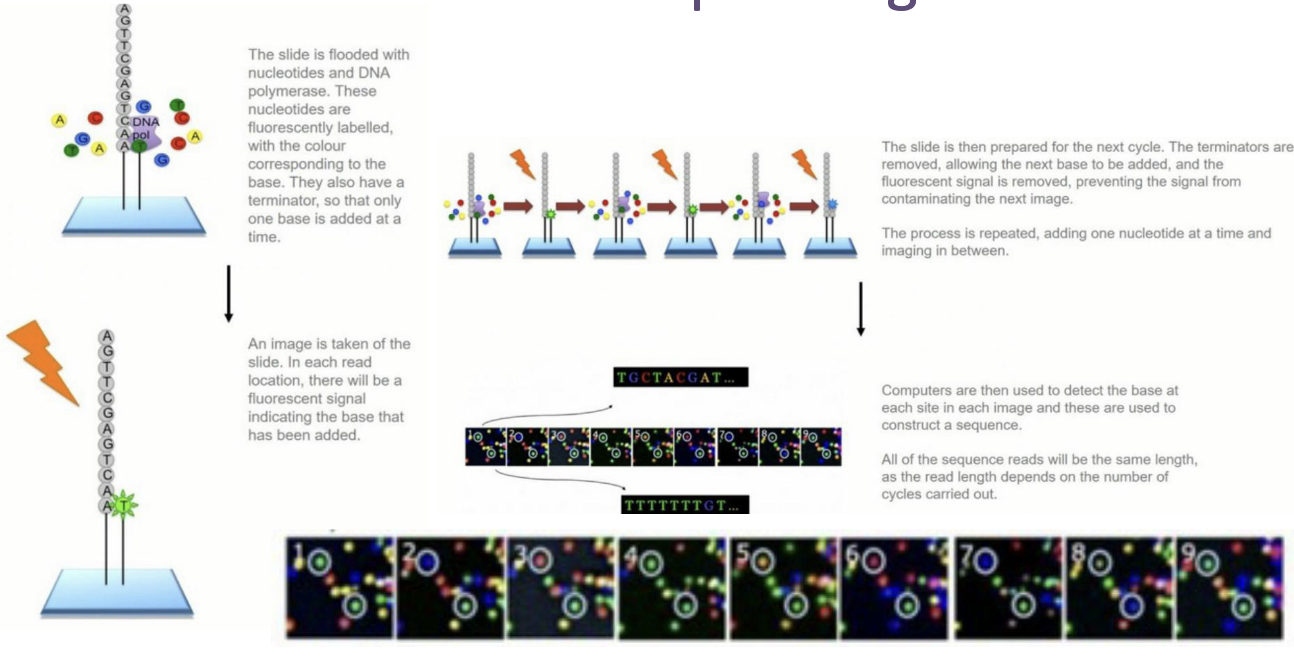
What are the steps of Illumina Sequencing?
The slide is flooded with nucleotides and DNA polymerase. These nucleotides are fluorescently labelled, with the colour corresponding to the base. They also have a terminator, so that only one base is added at a time.
An image is taken of the slide. In each read location, there will be a fluorescent signal indicating the base that has been added.
The slide is then prepared for the next cycle. The terminators are removed, allowing the next base to be added, and the fluorescent signal is removed, preventing the signal from contaminating the next image. The process is repeated, adding one nucleotide at a time and imaging in between.
Computers are then used to detect the base at each site in each image, and these are used to construct a sequence. All of the sequence reads will be the same length, as the read length depends on the number of cycles carried out.
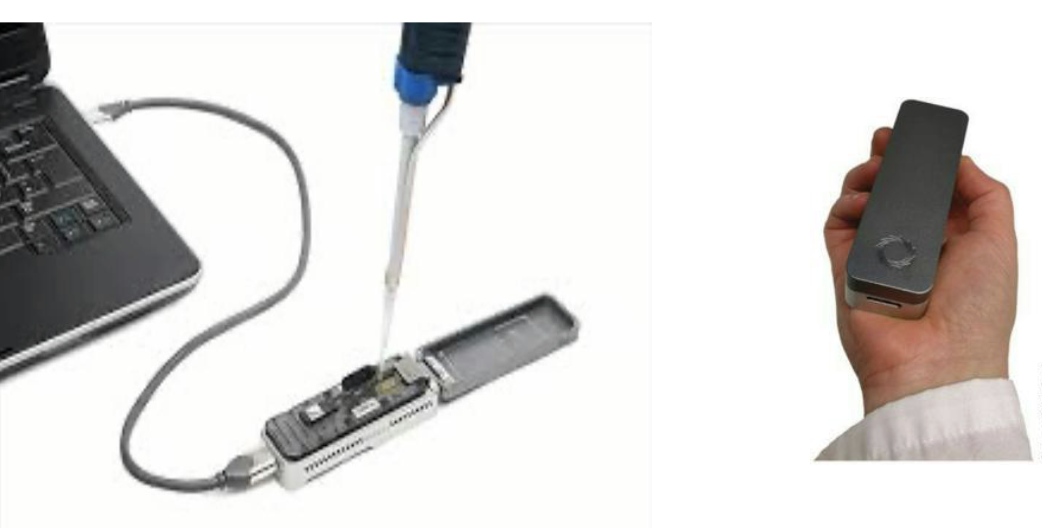
What is Nanopore sequencing?
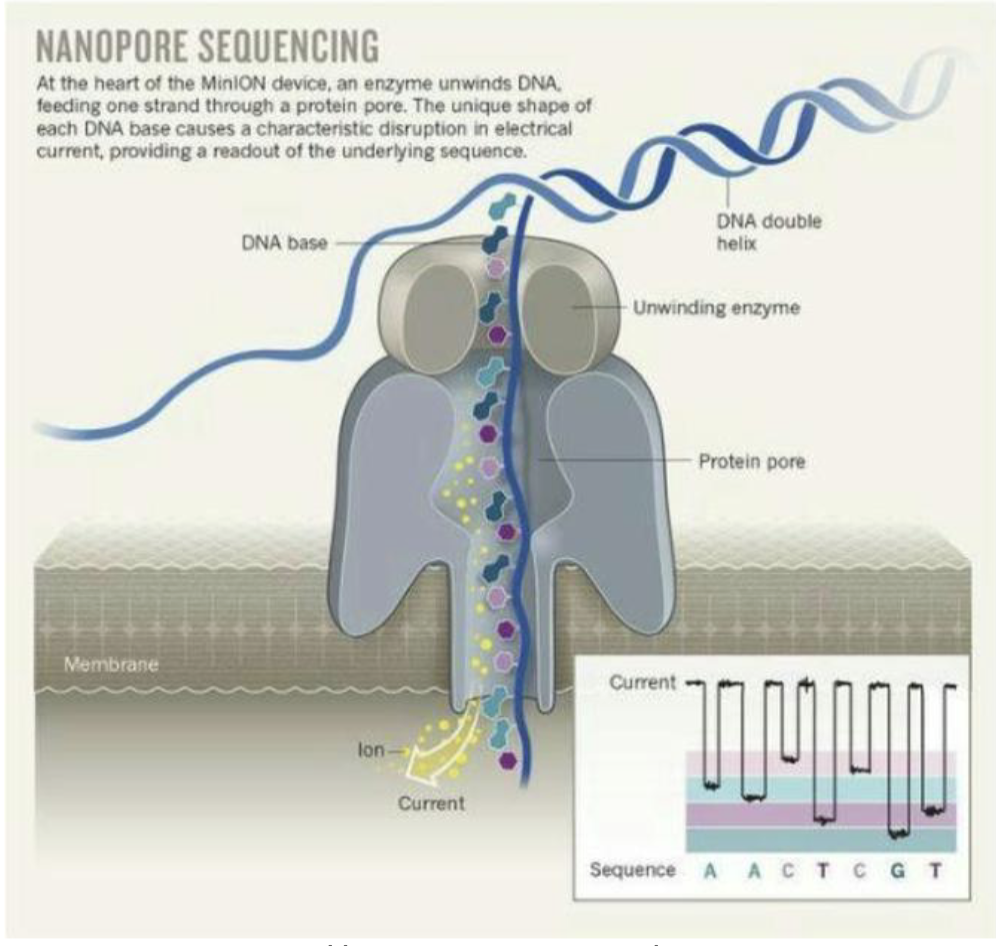
Nanopore sequencing — MinION
New polymerase-independent method that guides ssDNA through protein pore and reads bases as they exit
Enormously long read lengths (over 100 kb!)
Protein nanopores are embedded into a synthetic membrane bathed in an electrophysiological solution and an ionic current is passed through the nanopores
As molecules such as DNA or RNA move through the nanopores, they cause disruption in the current
This signal can be analyzed in real-time to determine the sequence of bases in the strands of DNA or RNA passing through the pore
What is the size of nanopore sequencers?
Sequencers the size of USB sticks for only 1000$!
Nanopore sequencing is less ideal for SNPs (single-nucleotide variants) because it doesn’t guarantee accuracy in that position; it’s better for structural variants (large DNA changes)

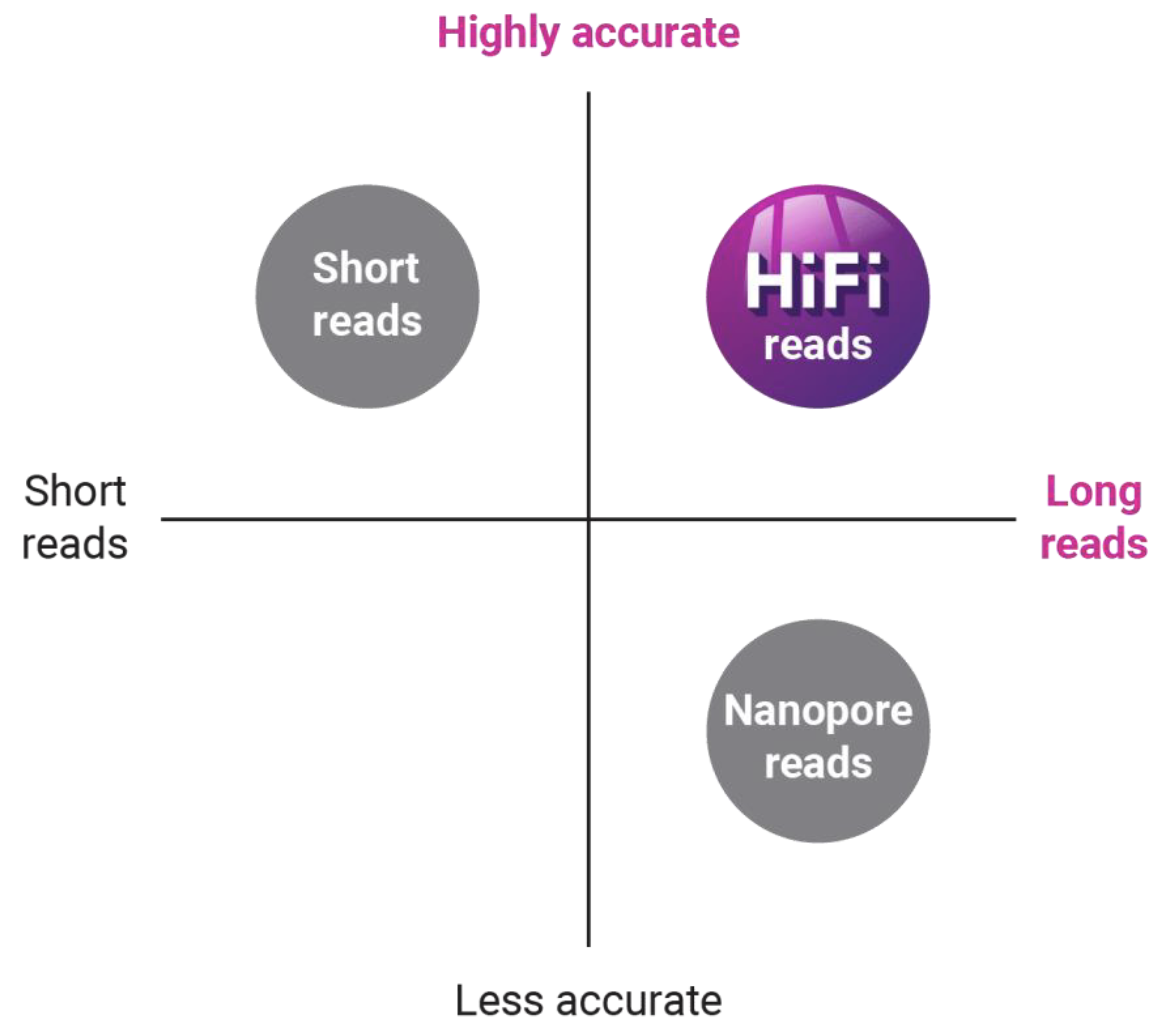
What is PacBio sequencing - long reads…?
Long reads allow you to readily assemble complete genomes and sequence full-length transcripts
High accuracy provides over 99.99% accurate sequencing results
Uniform coverage enables sequencing through regions inaccessible to other technologies
Single-molecule resolution lets you capture sequence data with over 99% single-molecule accuracy
Epigenetics that can be explored through direct detection of base modifications during sequencing
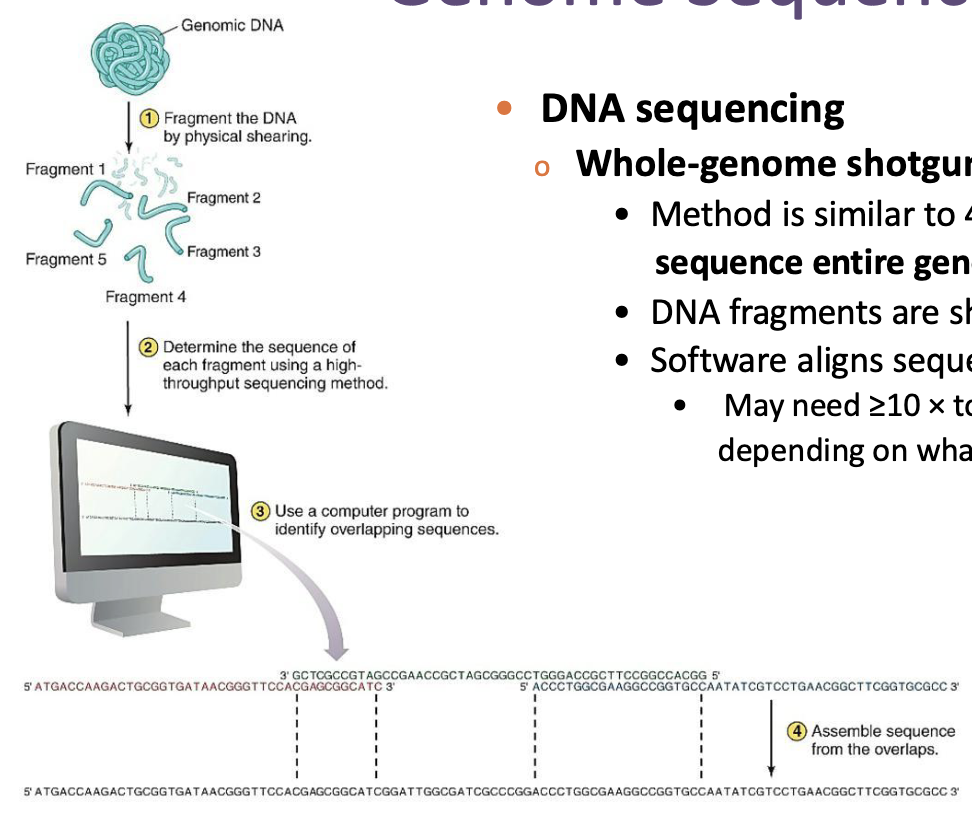
What is whole-genome shotgun sequencing?
Method is similar to 454 sequencing but attempts to sequence entire genome in one setup.
DNA fragments are sheared and sequenced.
Software aligns sequences.
May need ≥10 × total genome length to do so depending on what you are trying to analyze
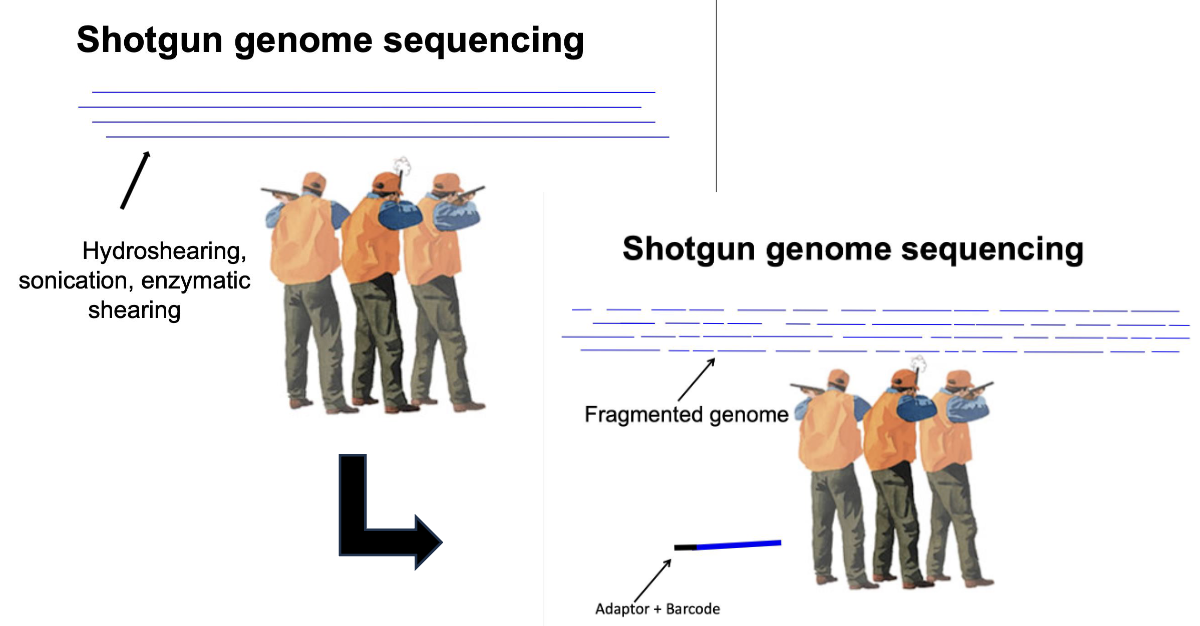
What is an analogy for shotgun sequencing?
Shotgun genome sequencing works by randomly breaking the genome into many small DNA fragments (“shooting”) using methods like hydroshearing, sonication, or enzymatic shearing.
Adaptors and barcodes are added to the fragments so they can be recognized by the sequencing machine and different samples can be tracked, and then each fragment is sequenced separately and assembled by computers using overlapping regions to reconstruct the original genome.
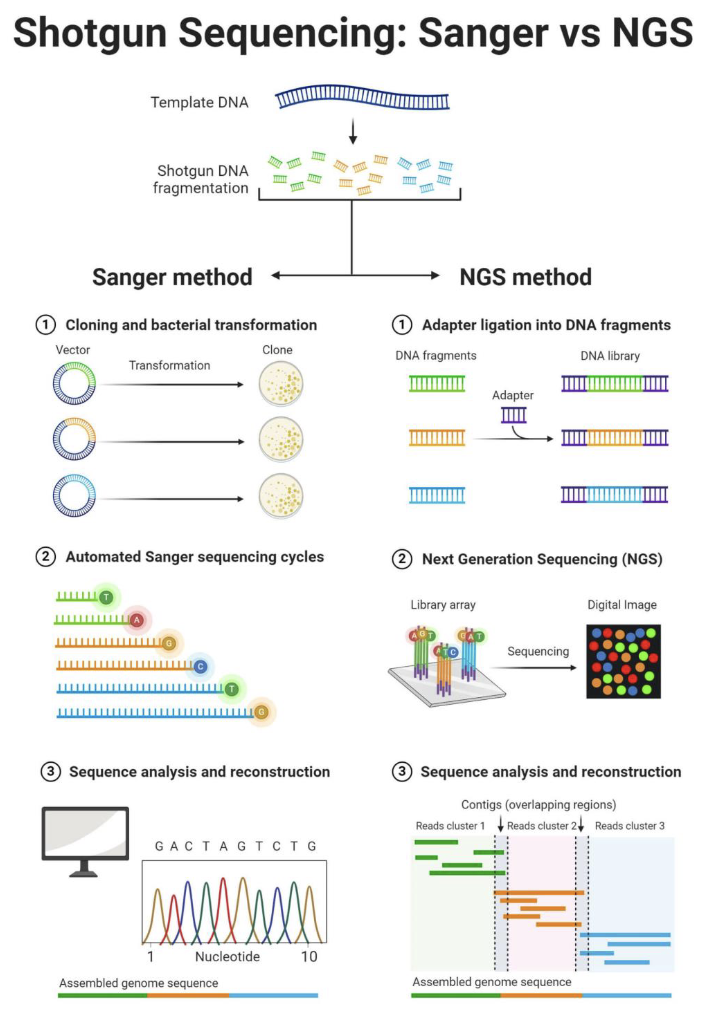
Shotgun Sequencing: Sanger vs NGS?
Shotgun sequencing breaks the genome into many random DNA fragments.
Sanger method: fragments are cloned into bacteria and sequenced one at a time using ddNTPs.
Benefit: Produces long, accurate reads in the correct order, so you don’t need to computationally assemble short fragments like in NGS.
NGS method: adaptors are added and millions of fragments are sequenced simultaneously on a sequencing chip.
In both methods, computers assemble overlapping sequences to reconstruct the genome.
How can gene expression be measured using genomic tools?
Cells regulate gene expression in response to environmental and metabolic conditions.
Examining the mRNA transcripts and proteins in a cell under specific environmental conditions helps researchers understand the adaptations of microbes.
Transcriptomes = Collection of transcribed mRNA molecules in a cell
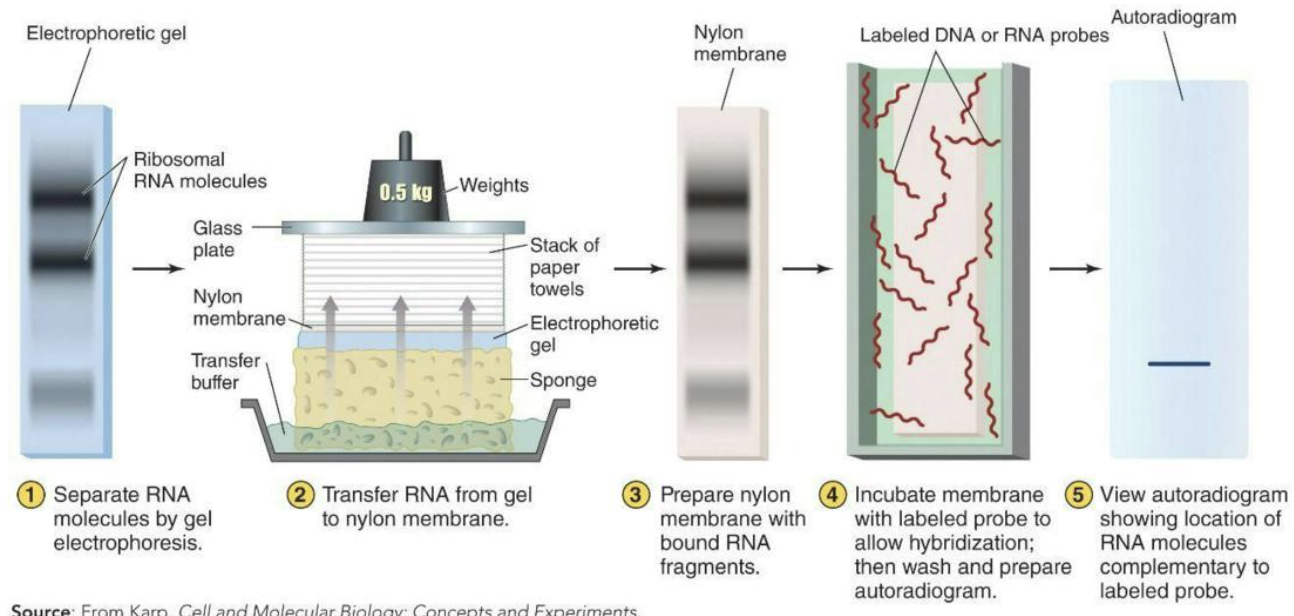
Northern blots – a technique that helps to measure transcriptional expression of individual genes
Separation of RNA fragments by electrophoresis, followed by blot transfer and probing with labeled DNA fragments
What are microarrays (transcriptomes)?
Method for examining transcriptional activity of all genes in a cell simultaneously.
The technique is a miniatured, automated reverse of Northern blots.
Probe DNA fragments are amplified by PCR and placed on a glass slide in a known pattern.
Total cell mRNA is converted to complementary DNA (cDNA) by reverse) transcriptase, labeled with a fluorescent molecule, and passed over the microarray slide.
The more intensely a “spot” on the microarray lights up, the more cDNA (ergo, the more mRNA) is present.
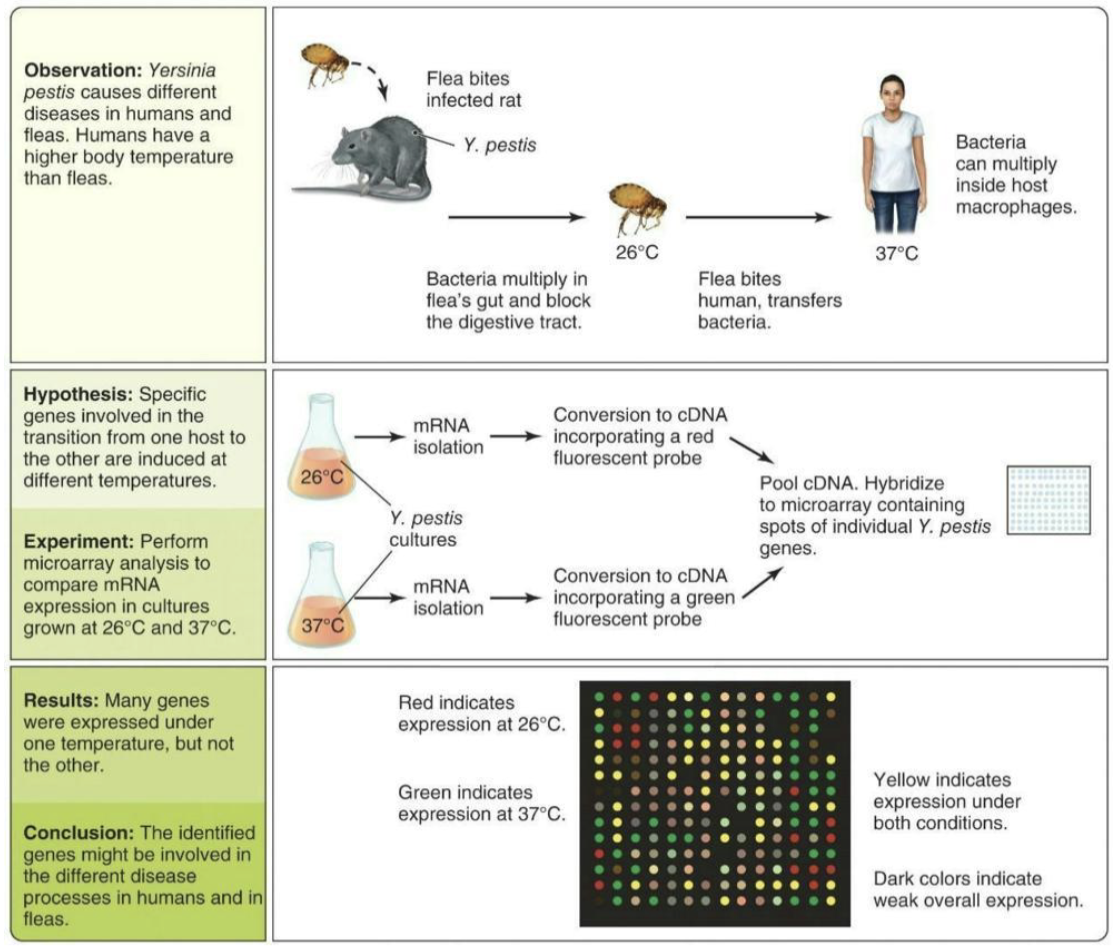
What is an example of microarrays?
An example of this method in use is comparison of mRNA expression profiles in Y. pestis microbes growing at flea or human body temperatures.
The identified genes are candidates for future research because they may be important for causing disease.
What is RNA-seq technology (transcriptomes)?
RNA sequencing performed by conversion of mRNA into cDNA by reverse transcriptase.
cDNA is then sequenced using rapid automated methods.
Bioinformatic tools then help compare all sequences to known RNA sequences
Has been used successfully to study Saccharomyces cerevisiae and other organisms.
Will continue to evolve in speed and benefit in tandem with DNA sequencing and analysis methods.
What is proteomes?
The collection of expressed proteins in a cell
Differences in protein types and abundance reflect changes in gene expression and/or protein stability
Can be studied by multiple methods, including
2D-polyacrylamide gel electrophoresis (PAGE)
Mass spectrometry
Liquid chromatography-mass spectrometry (LC-MS)
X-ray crystallography
Nuclear magnetic resonance (NMR)
What is 2D-PAGE (Proteomes)?
Allows separation of proteins on a gel based on
Isoelectric point (pH where protein has no charge) of protein, which is a function of amino acid composition
Mass (separation on the gel, smaller fragments move faster)
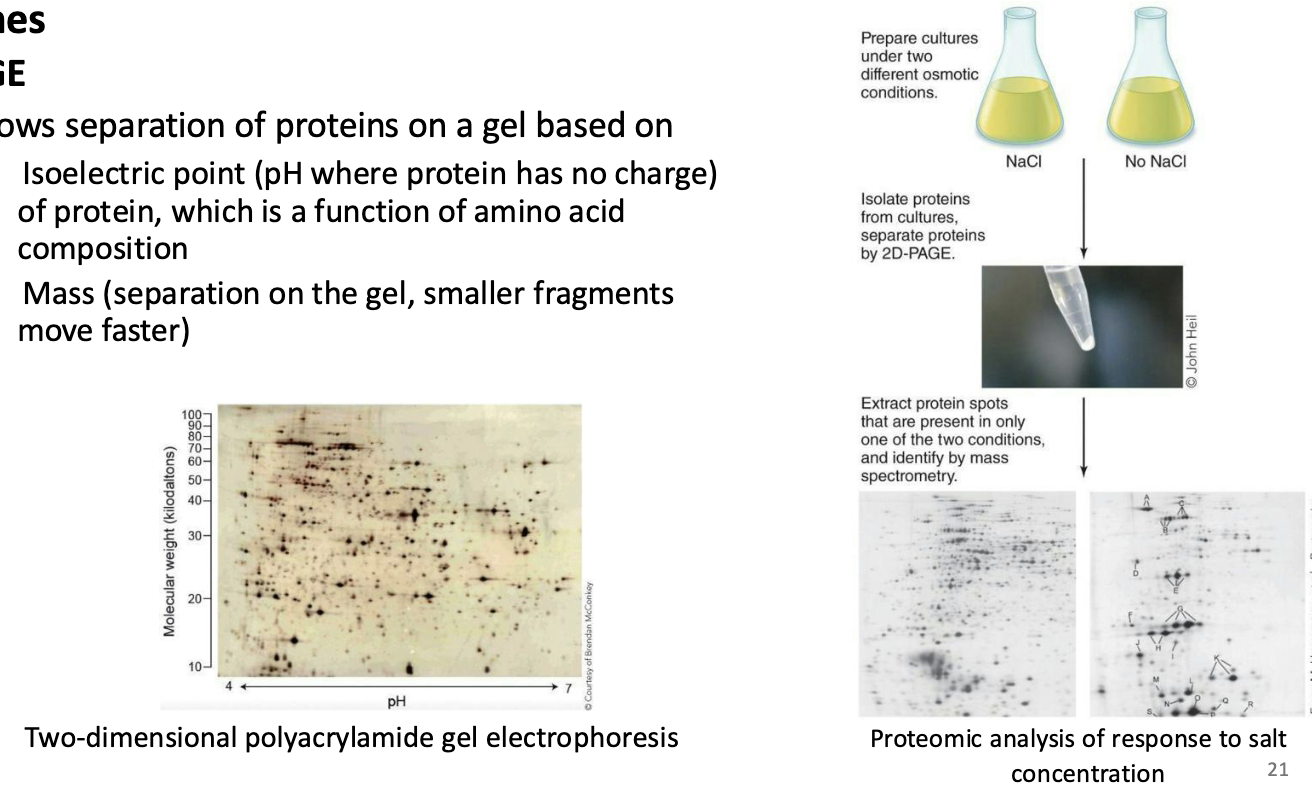

What is mass spectrometry (proteomes)?
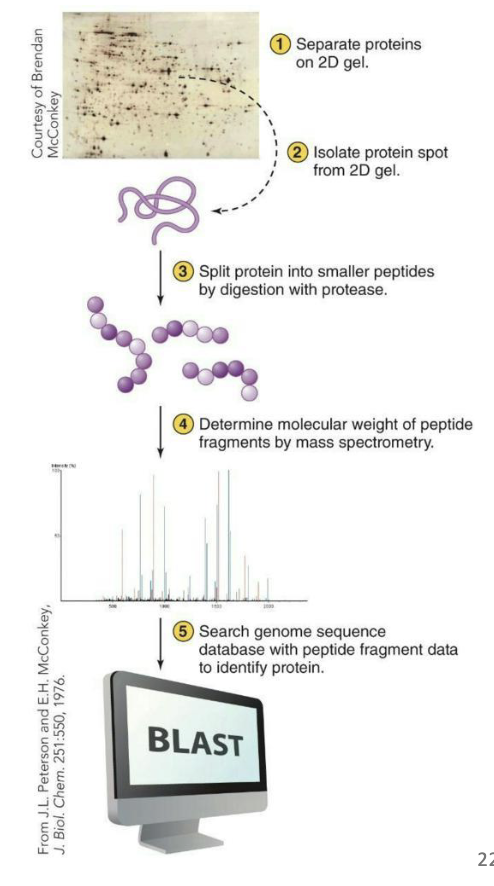
Mass spectrometry can be used to determine amino acid sequences of portions of the polypeptides from 2D-PAGE
Spots are first extracted from the gel and then digested into smaller fragments using proteases
The fragments are analyzed by mass spectrometry to determine the amino acid sequence based on individual mass-to-charge ratios
Comparison of sequence to known protein sequences can help determine identity
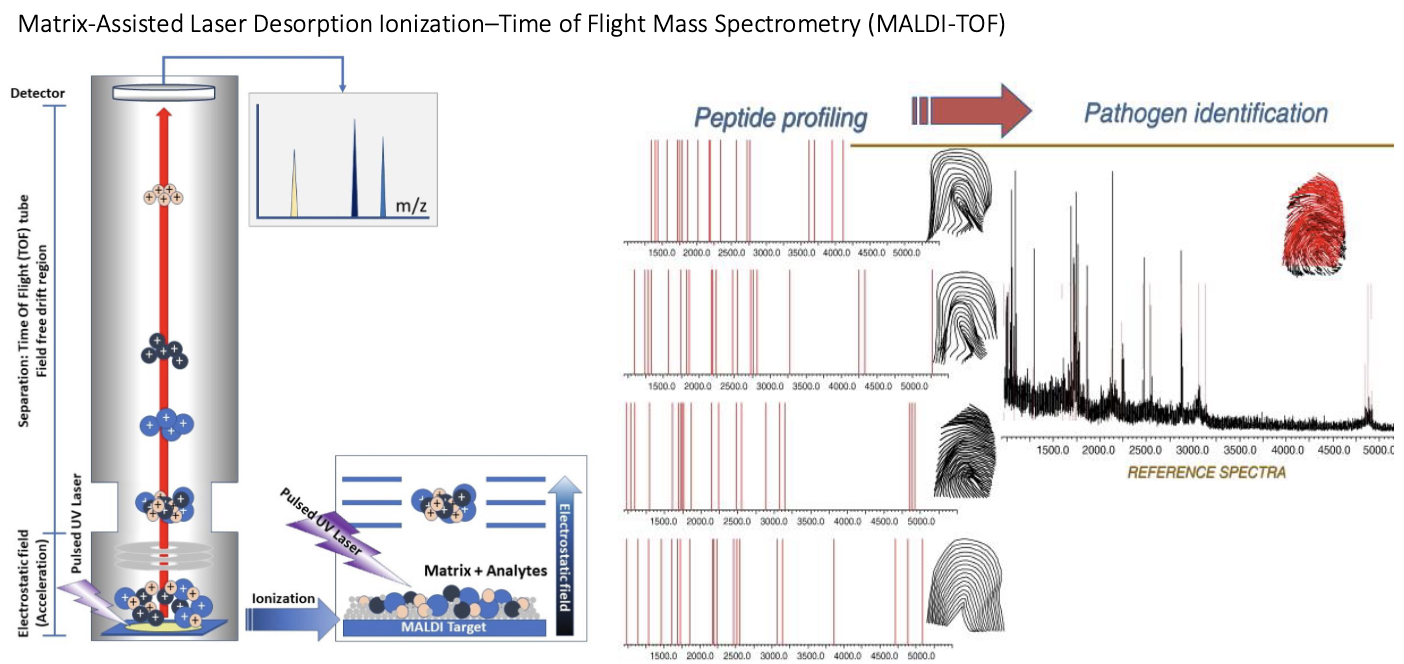
What is Matrix-Assisted Laser Desorption/Ionization Time-of-Flight (MALDI-TOF) MS?
Identifies bacteria and some fungi from medical cultures.
How it works:
Measures mass-to-charge ratio (m/z) of full polypeptides.
Produces a unique peptide/protein fingerprint for each microorganism.
Compares sample fingerprint to a database of known spectra to identify the organism.
Rapid and accurate microbial identification based on protein patterns.
What are methods for determining protein 3D shape?
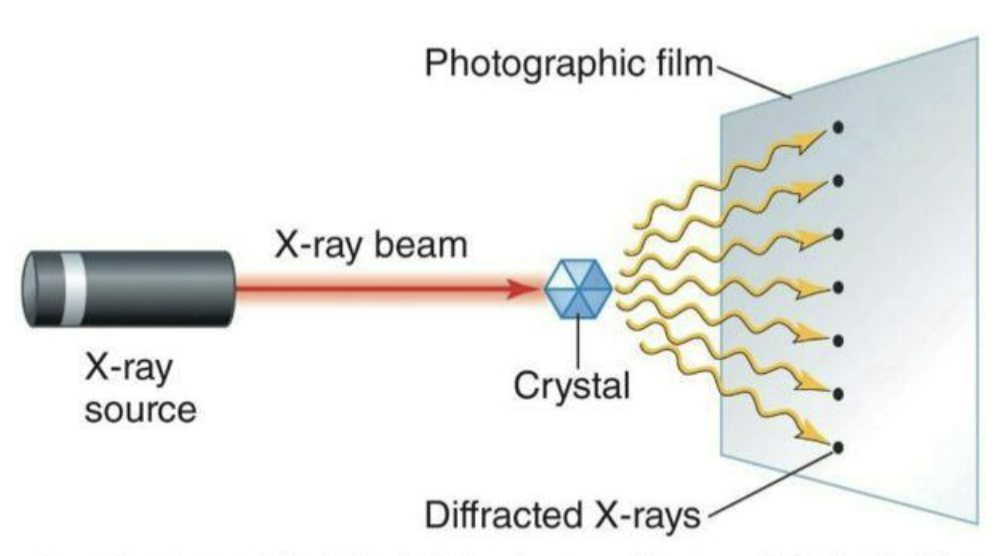
X-ray crystallography
X-ray beam shot at crystallized protein.
Diffraction pattern can be used to discern protein shape.
Nuclear magnetic resonance
Measures distances between atomic nuclei to discern protein shape.
Can measure proteins in solution.
Limited to proteins of about 30 kDa.
AI systems like AlphaFold (DeepMind)
Made over 200 million protein structure predictions freely available
Use deep learning algorithms to predict protein structures based on their amino acid sequences.
Can predict protein structures in a fraction of the time with near- experimental accuracy
What information can be obtained from comparing genomes?
Comparative genomics is the study of evolutionary processes using the tools of genomics.
Provides us with information about the relationships between different species
Genetic variability – result of mutations/changes in DNA sequence
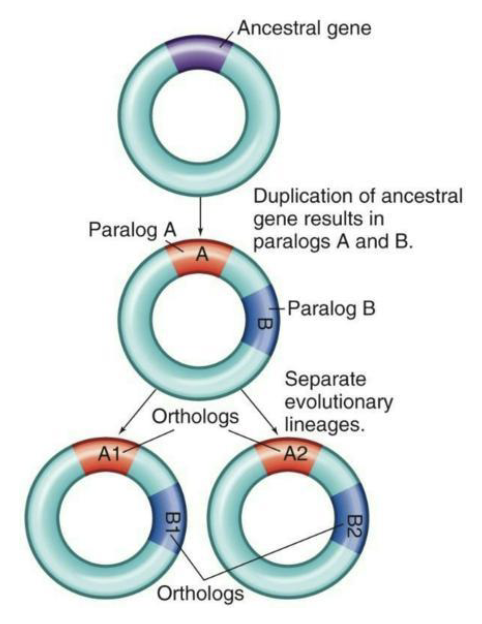
Different genes may have arisen from duplication events.
Paralogs = Genes that are different from each other, yet arose from a duplication event
Orthologs = Genes that have evolved from the same ancestor with the same function in different organisms
Paralog families can become very large
Studying them can provide evolutionary insight
What is horizontal gene transfer?
Sharing genetic material by microbes
Genomes tend to have characteristic G+C nucleotide content.
E. coli = 50% G+C
Streptomyces coelicolor = 72% G+C
Saccharomyces cerevisiae = 38% G+C
Areas of genomes with significantly lower or higher G+C content than the rest of the genome are likely areas where horizontal transfer has occurred.
Identifying and studying these areas can lend further insight into comparative genomic studies.
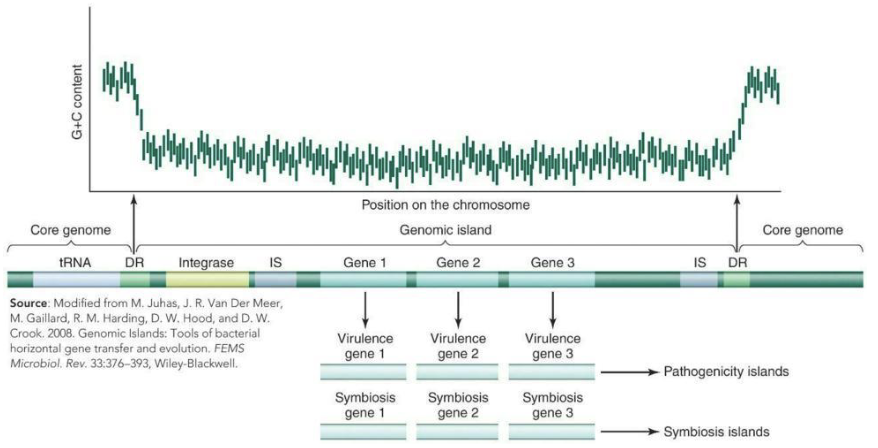
What are genomic islands?
Large segments of DNA exist in one genome but not in another genome that is otherwise closely related area
DNA segments of 10‒200 kb associated with tRNA genes, transposable elements, plasmids, or bacteriophages
Genomic islands often integrate insertion sequences (IS) and direct repeat sequences (DR)
Pathogenicity islands contain virulence genes, whereas symbiosis islands contain genes for symbiosis.
Can we apply genomics to the study of uncultivated microbes?
If only a small number of microbes can be cultured, how can you study the rest?
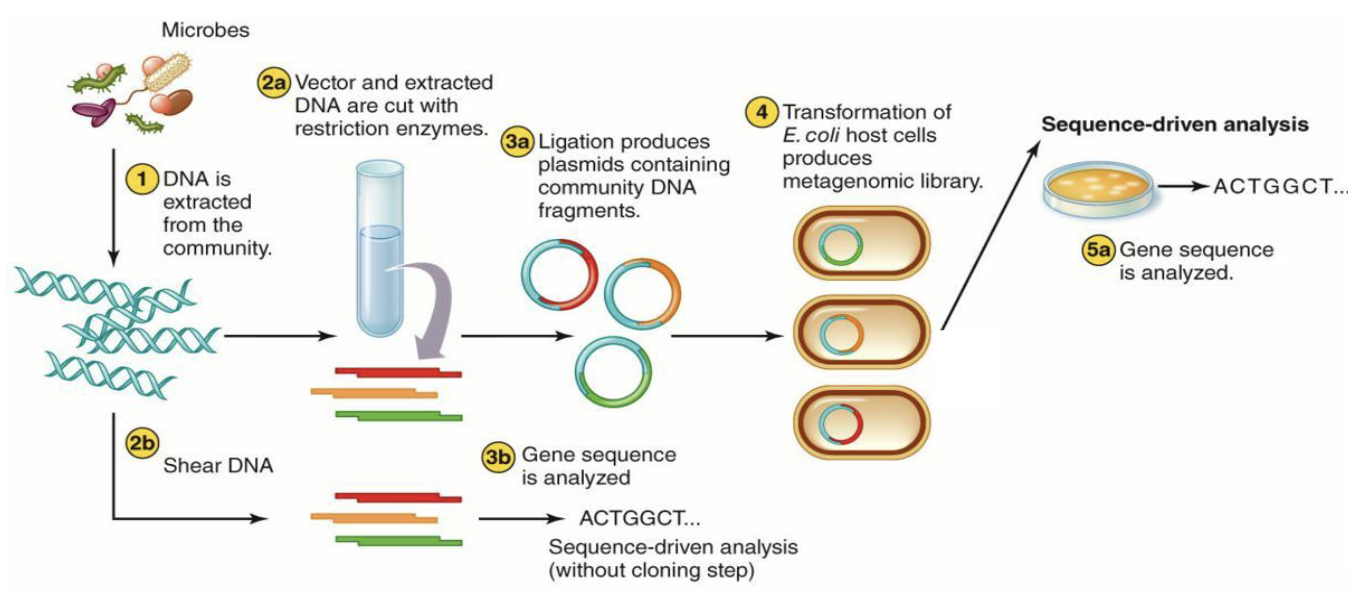
Metagenomics involves the construction and analysis of gene libraries from DNA extracted directly from complex microbial communities.
This field is changing our understanding of life on Earth, finding evidence for newly discovered organisms in very diverse and challenging locations.
Acid mine drainage
Deep-sea thermal vents
Wastewater treatment areas
Some of the genes discovered have possible applications in biotechnology and nanotechnology.
How is DNA extracted for metagenomics?
Next gen sequencing allows DNA to be extract directly from microbial communities and sequenced
Very useful for studying unculturable microorganisms
Analysis of sequences is the challenging part.
The trick is to eliminate sequences that we already know about to find new genes, potentially from new organisms.
What are universal genes?
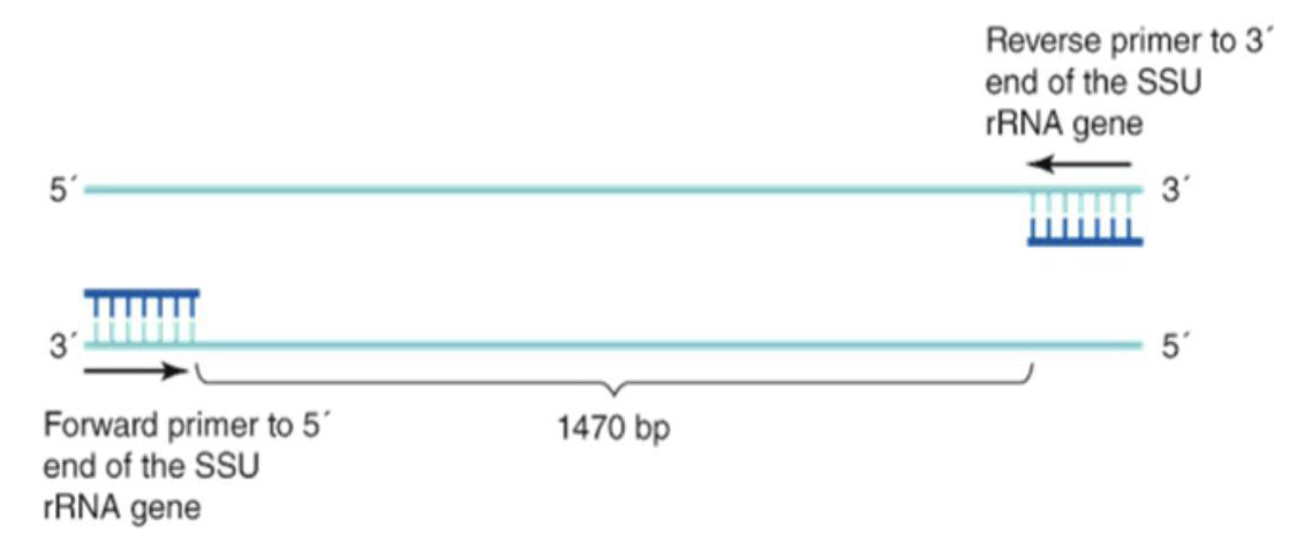
Can determine phylogenetic relationships between organisms by using universal genes like SSU rRNA
Universal primers can be used that bind to conserved sequences in many organisms
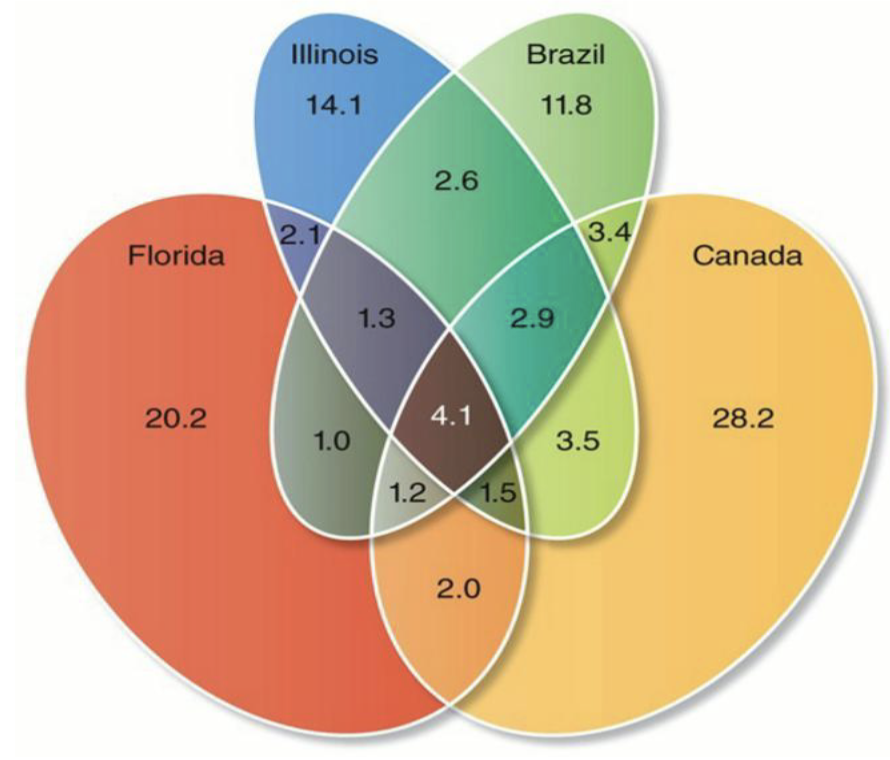
What is the comparison of microbiota in different soils?
Closely related microbes often classified into operational taxonanomic units (OTUs) based on similarity of a marker gene (often >97% similarity of SSU rRNA)
Caveats - bacteria species naming can be not consistent so sometimes >97% similarity can include 2 different species such as in the case of E. coli and Shigella which has >99% rRNA similarity
What else can you use to sequence genomic information?
Can also sequence all genomic information from an environmental sample rather than a specific marker gene amplified by PCR
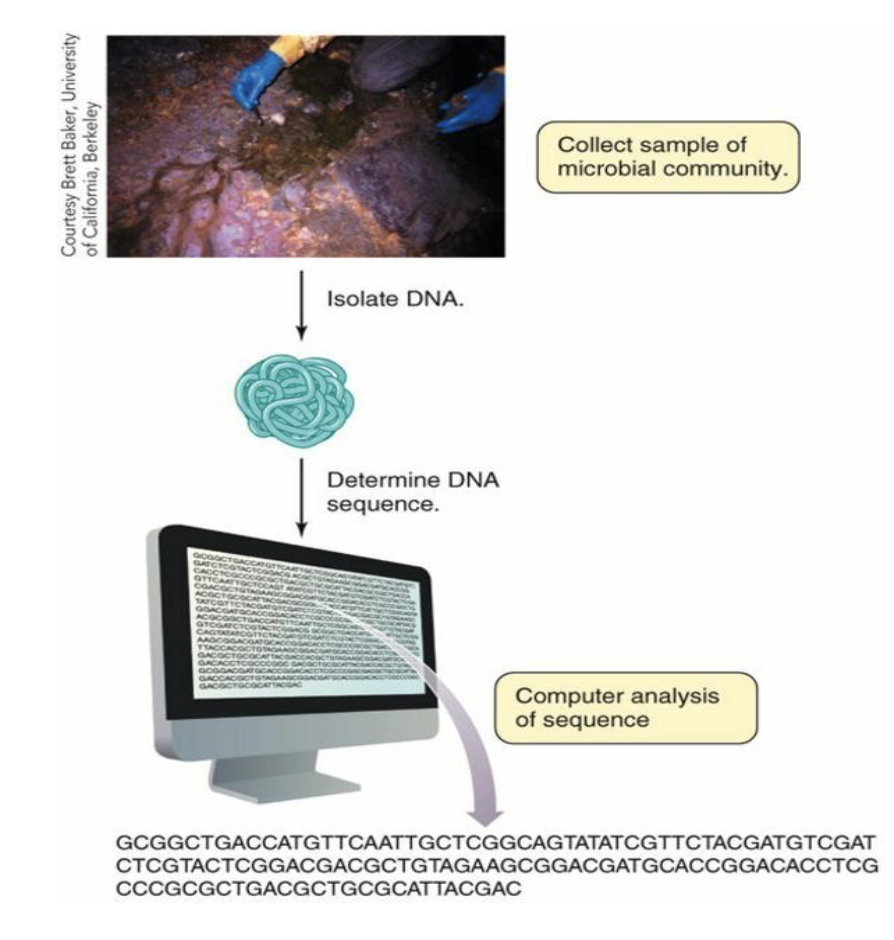
What are cultivation-independent techniques?
Purpose: To determine “who’s there” in a microbial community without needing to grow microbes in culture.
Direct sequencing (without growth or cloning)
Extraction of DNA from an environmental sample
Followed by PCR (often for SSU rRNA genes) and sequencing
Compare to databases of known sequences for identification
Example: PCR amplification of a 16S rRNA gene region from bacteria isolated from yogurt
Forward primer (27F): 5’-AGAGTTTGATCMTGGCTCAG-3’
Reverse primer (519R): 5’-GWATTACCGCGGCKGCTG-3’
These primers amplify a specific region of the 16S rRNA gene to see which bacteria are present.
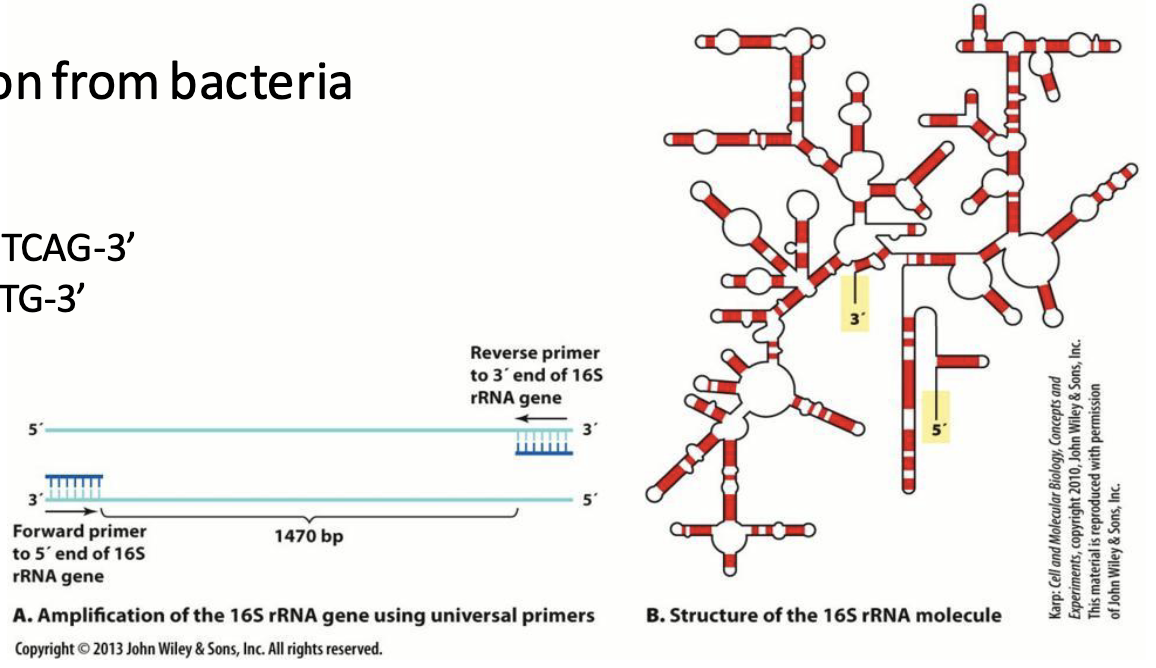
How can this be used in forensics (fungi)?
Fungi identify the geographic origin of dust samples
Analyses identified ~40,000 fungal taxa (~700 per sample)
Using the unique fungal composition of dust samples, can predict where a dust sample originated geographically
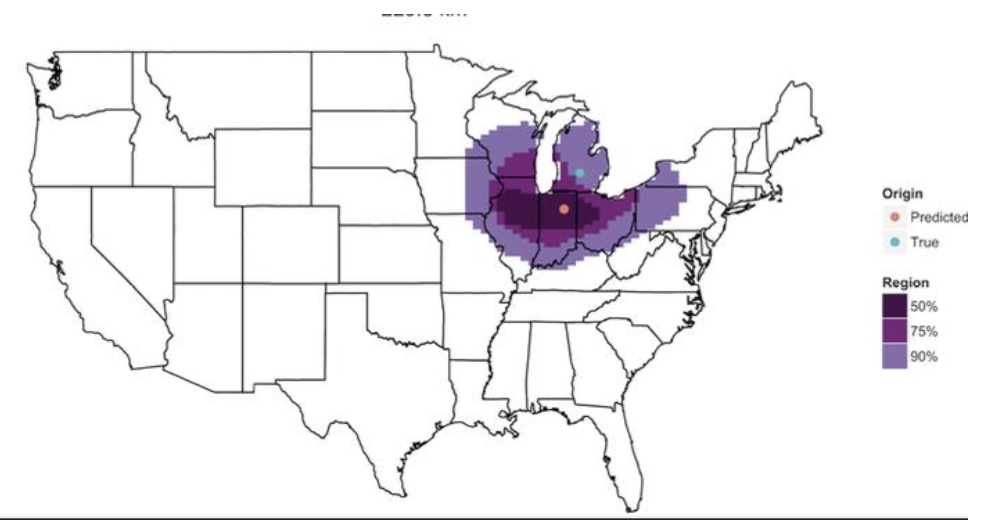
How can this be used in forensics (trace soil samples)?
Case study: A soil microbial community DNA profile was obtained from the small sample of soil recovered from the sole of a shoe
Determine the composition of the bacterial communities in both evidence and other places of interest (crime scene shoe print, alibi location, suspect house, etc)
The tread soil community had a high similarity to the footprint soil community (>0.9) and the soil samples from the suspects home also clustered more closely to the crime scene soil than the alibi soil
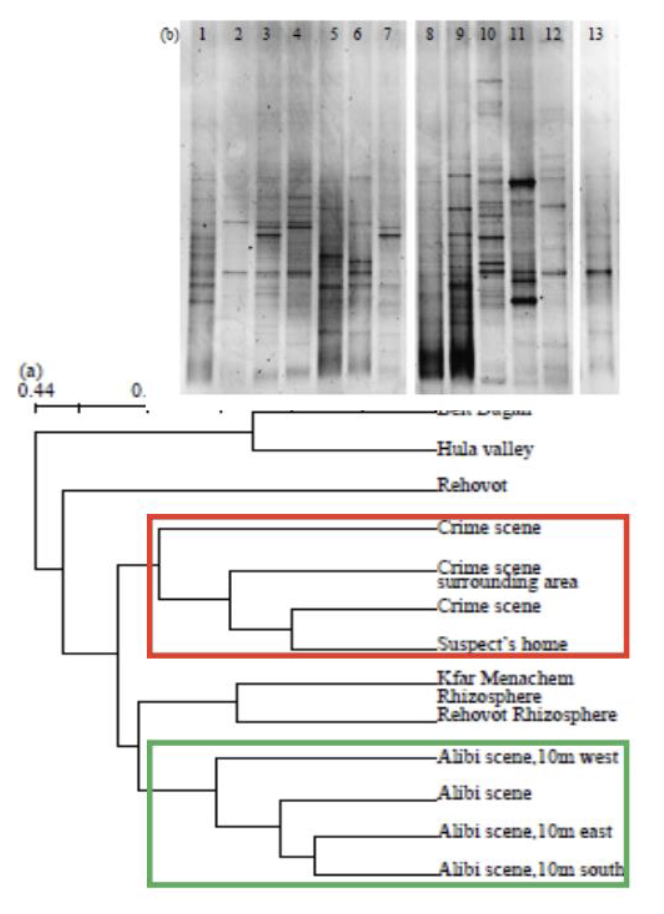

What are the stats of the cost per human genome?
Sequencing cost dropped from >$100M (2001) to < $1,000 today.
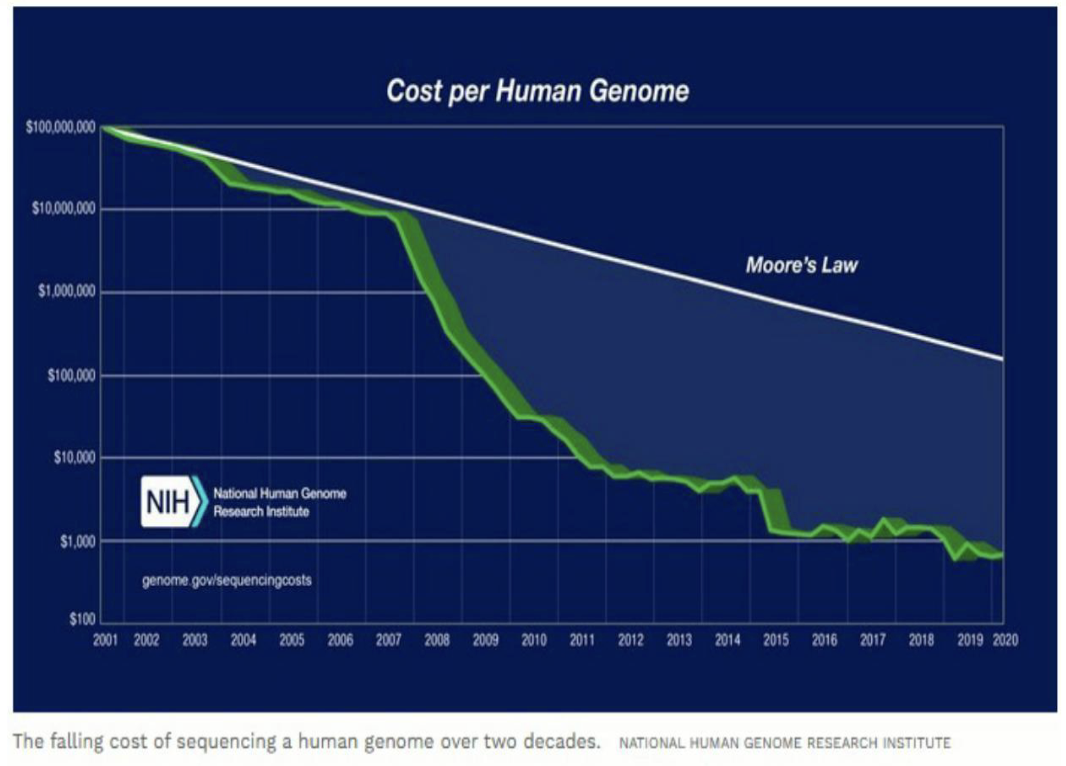
Genomics Summary
Sanger, dideoxysequencing and primer walking allowed sequencing of large pieces of DNA
Next gen sequencing methods have further increased the speed at which DNA can be sequenced
Nanopore sequencing couples rapid speed and throughput with large fragments sequencing
Comparative genomics yields new insight into evolutionary processes
Horizontal gene transfer can be detected by comparison of %G+C content across the genome
Genomic islands are large stretches of DNA that can move by horizontal gene transfer and encode traits like pathogenicity, antibiotic resistance or symbiosis
Metagenomics involves sequencing DNA from complex microbial communities
Can be used to determine phylogenetic relationships by amplifying certain genes (ex. SSU rRNA), or for discovering new gene sequences by sequencing all genomic information