Handout 3B: Nucleic Acid Synthesis
1/25
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
26 Terms
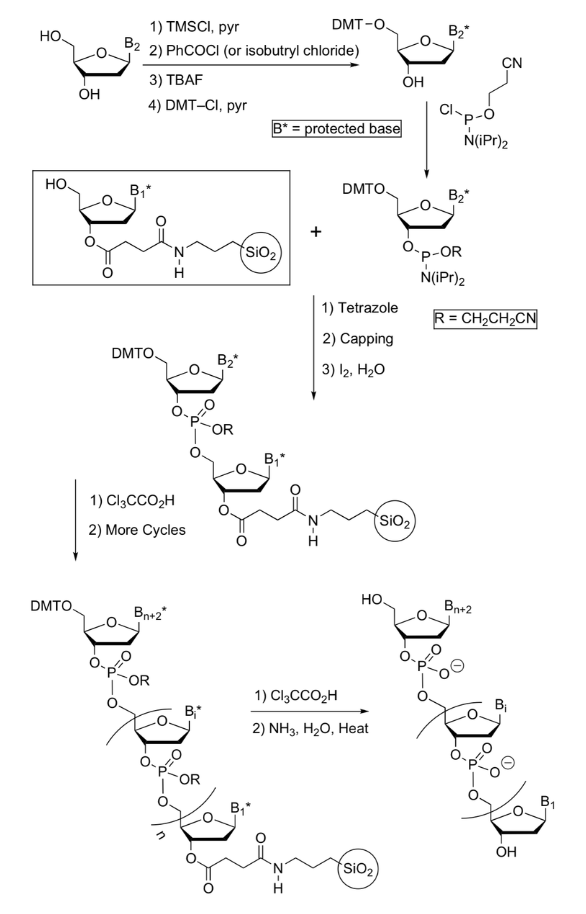
Protection of Nucleobases in DNA synthesis
protection a A, G, C are often done via transient protection using TMS protection to protect OH
Amines need to be protected, as they would interfere with the synthesis w/ their nucleophilicity
Thymine does not need a protecting group as it doesn’t contain an amine
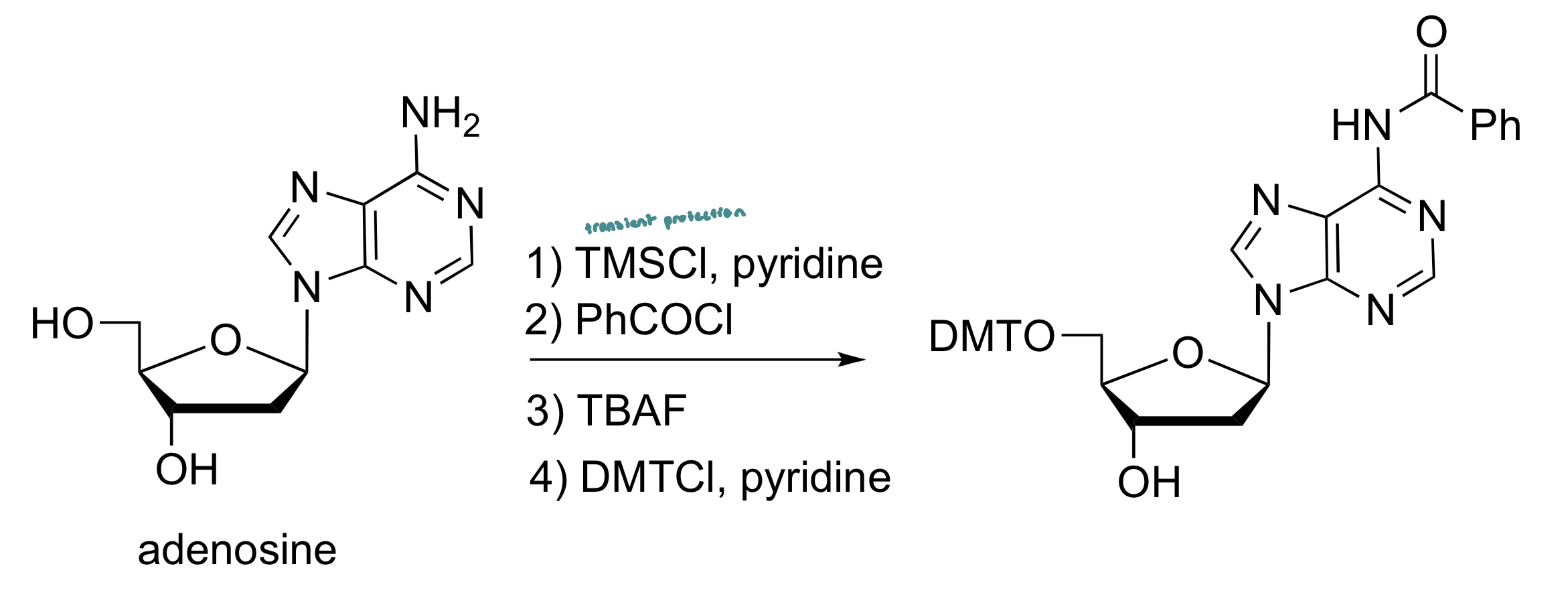
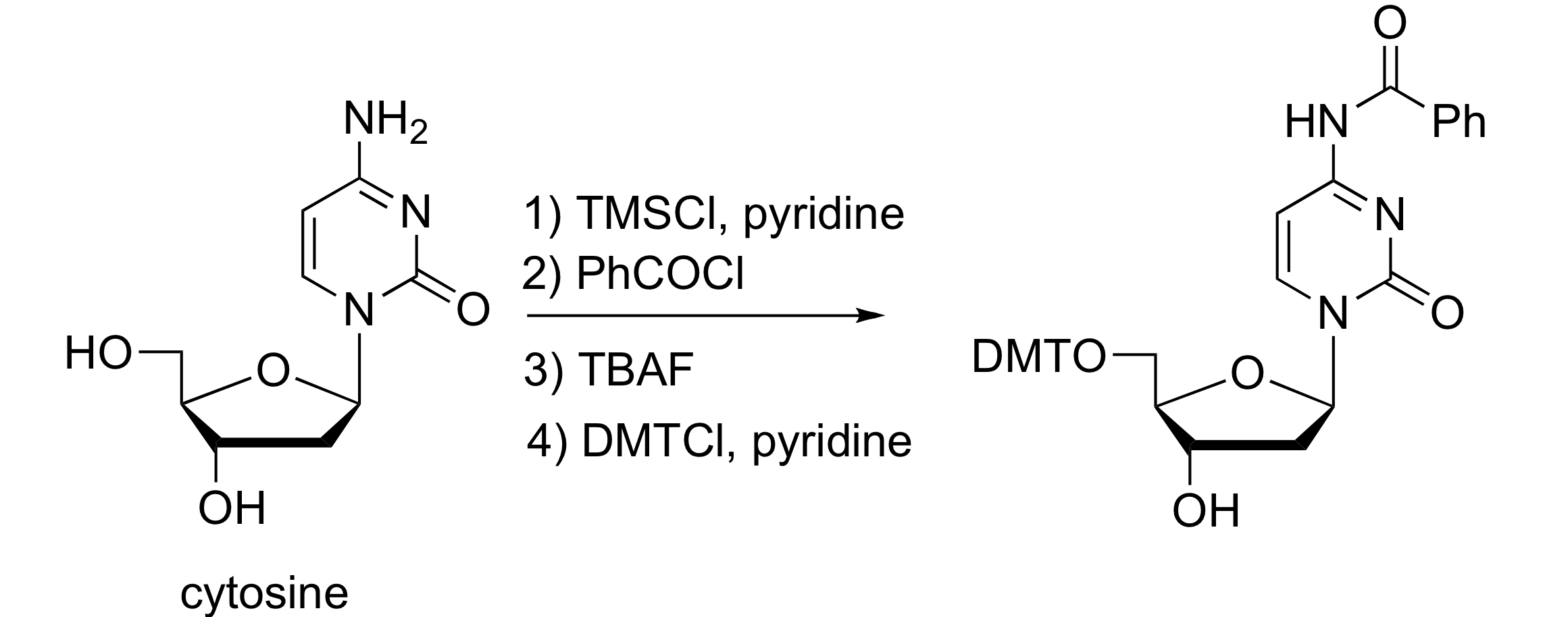
Adenine and cytosine are protected at their amino group as a benzamide
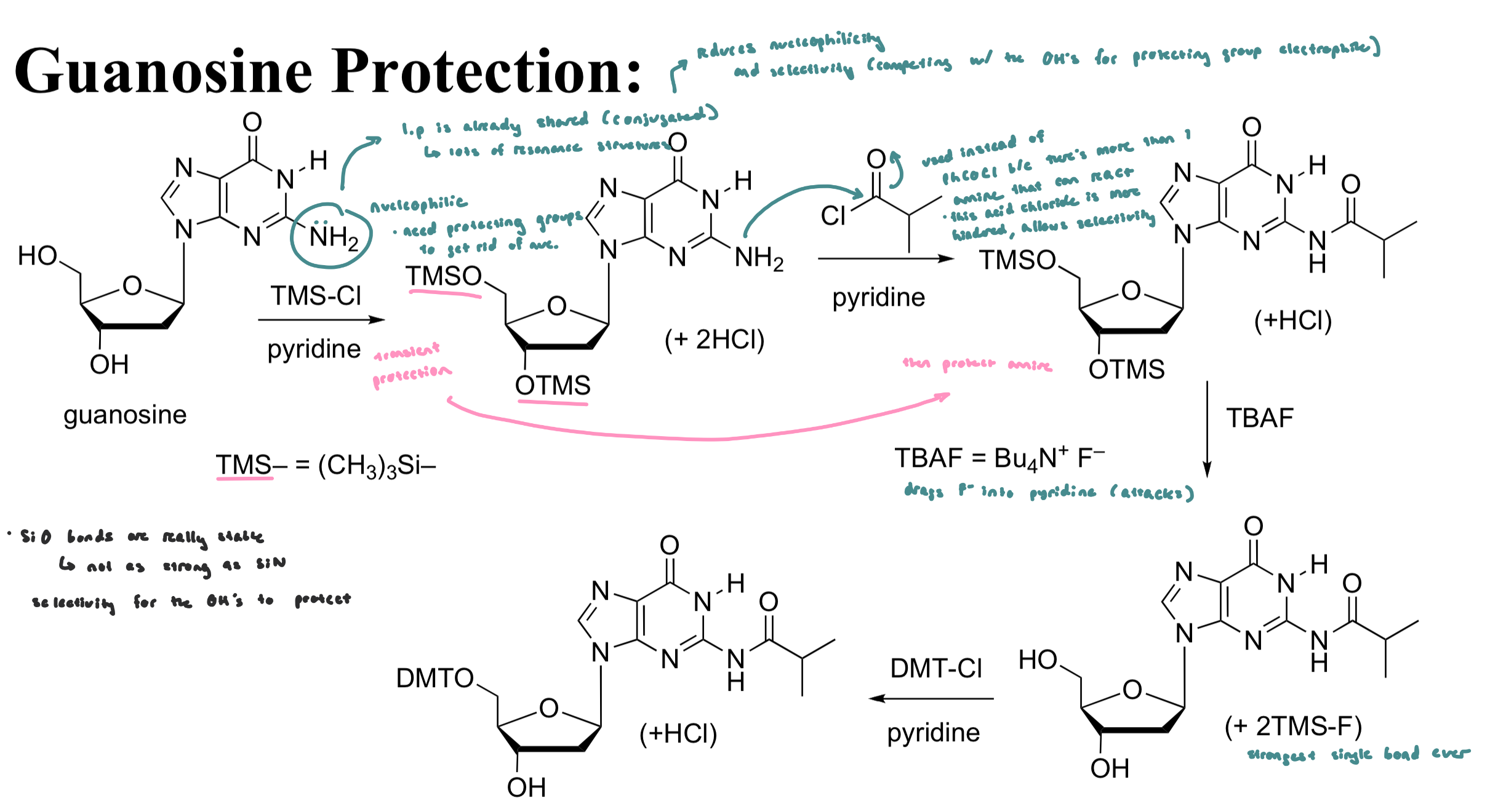
Guanine is protected at its amine as an isobutrylamide (benzamide is difficult to remove from a guanine)
TMS
TMS = trimethylsilyl (Me3Si)
Reaction of an alcohol with TMS-Cl goes via an addition/elimination mechanism, via a pentavalent intermediate
Si-O bonds are very strong and drive the reaction selectively over formation of Si-N bonds (Si-F bonds are even stronger than Si-F bonds)
TBAF (tetrabutylammonium fluoride) is used to remove TMS from the R-O-TMS via an addition/elimination mechanism
Guanosine Protection
Adenosine Protection
Cytosine Protection
Thymine Protection
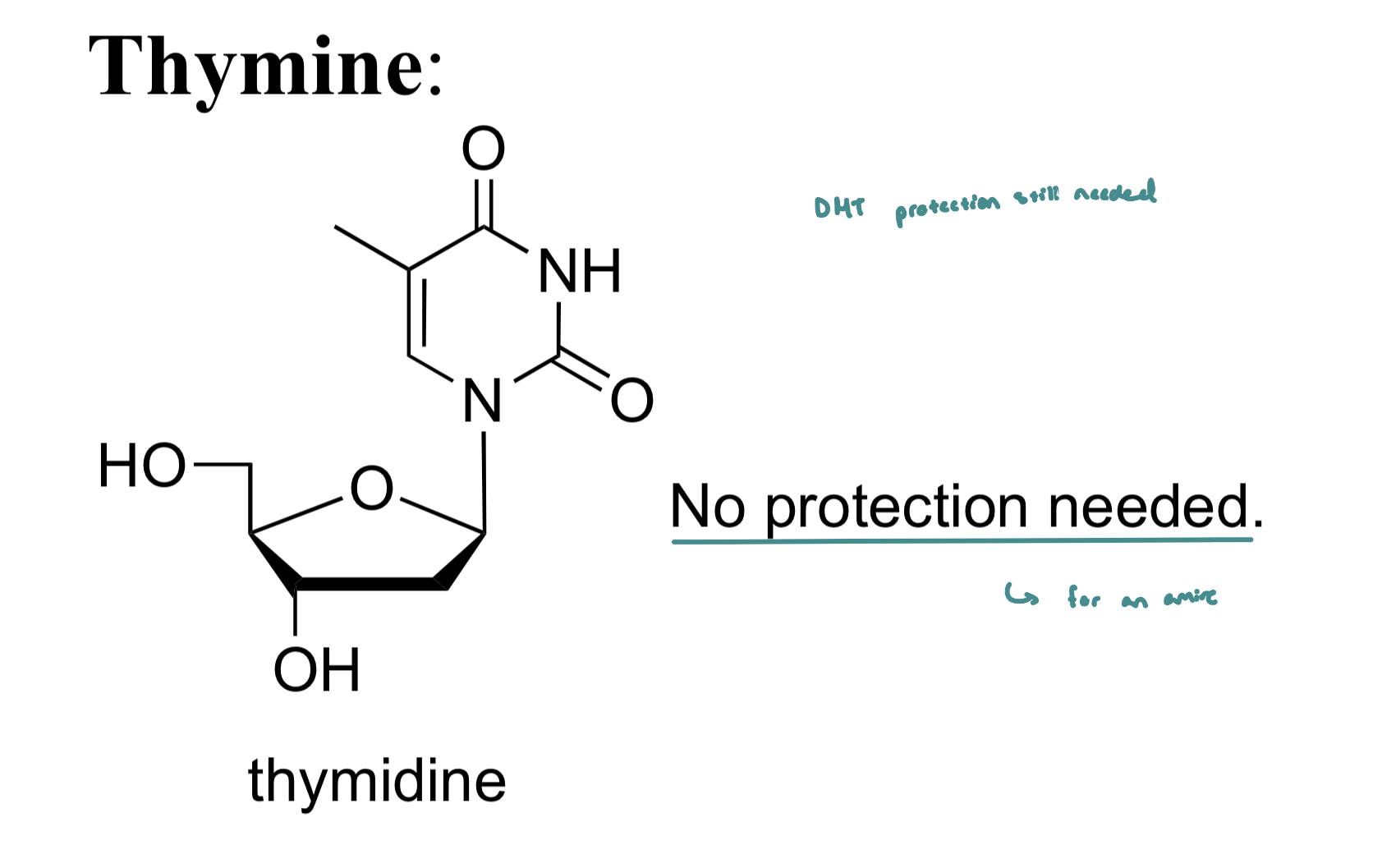
DMT Protection
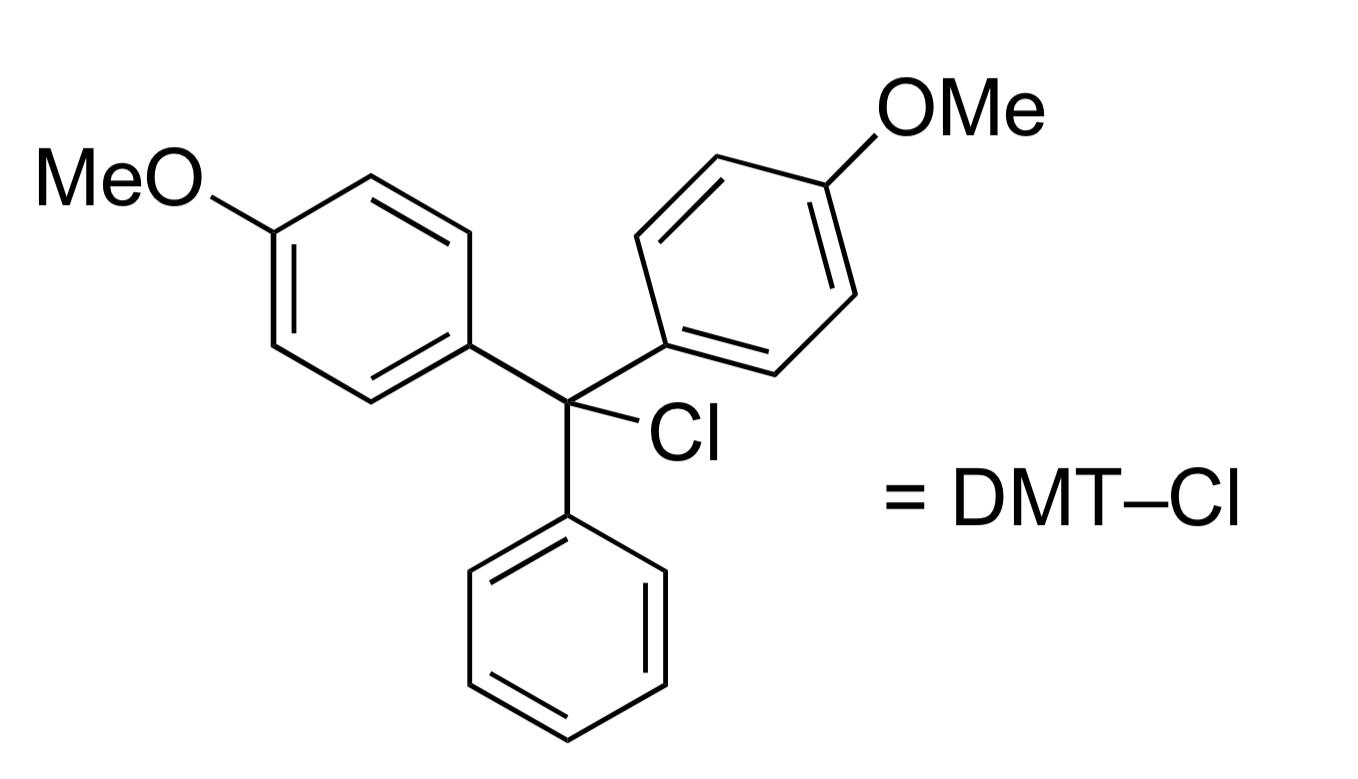
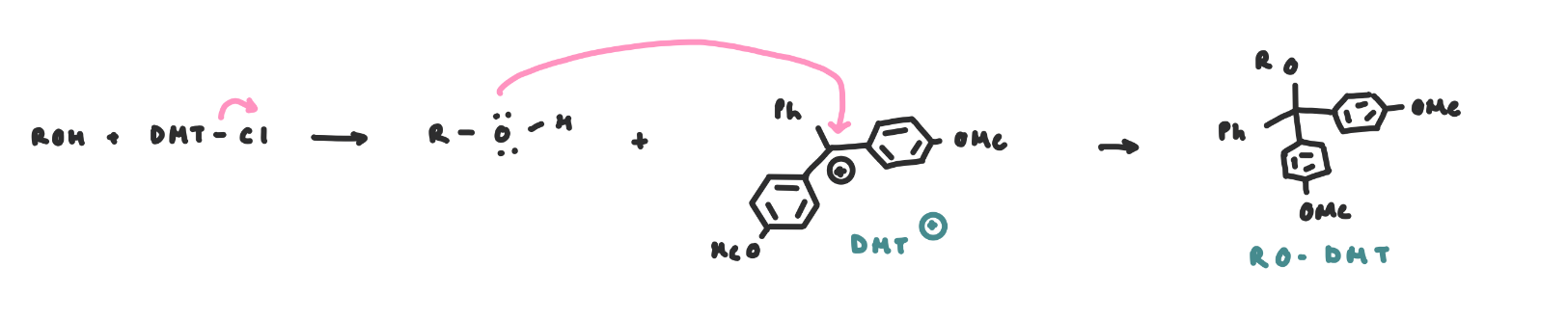
Once the nucleobases are protected, the 5’OH can be protected as RO-DMT (4,4’-dimethoxyltrityl)
DMT is similar to trityl, but is easier to remove and is colored, which is used to monitor reaction progress
Selective to primary over secondary alcohols (due to steric bulk)
DMT Incorporation
Resin Incorporation
Next, the nucleoside w/ protection at its nucleobase and DMT at its 5’OH is attached to resin.
Resin = CPG (Controlled pore glass)
Silica gel, glass, CPG, etc. are (SiO2)n, a solid polymeric matrix of silicondioxide
attaches nucleosides (protected sugar base units) covalently to a solid support to make purification easy and allows for stepwise synthesis
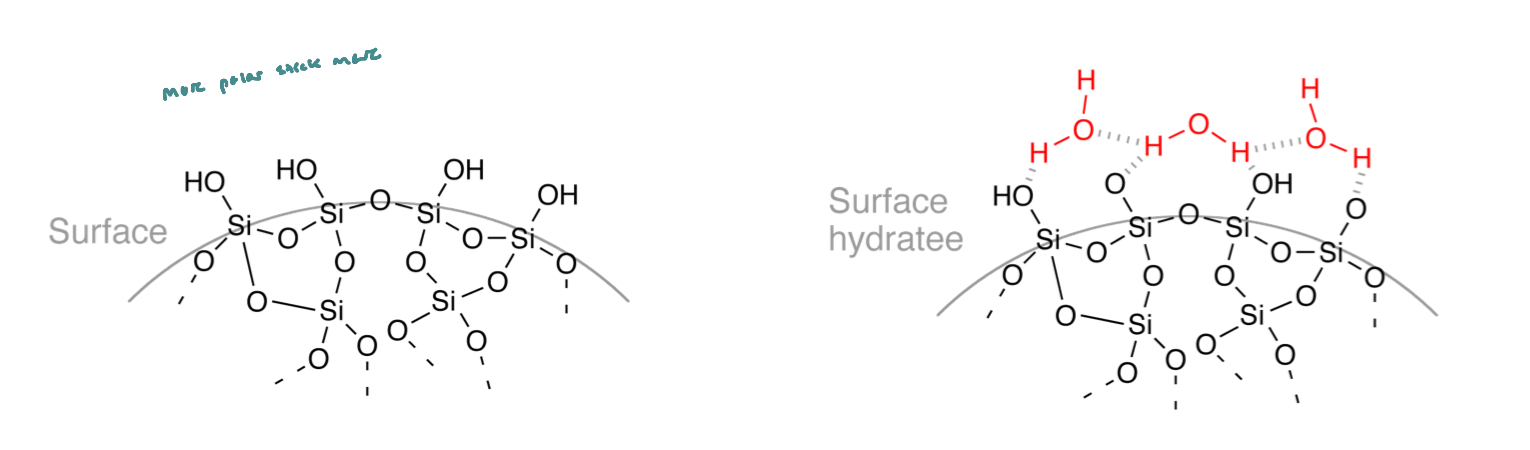
Resin Incorporation Mech
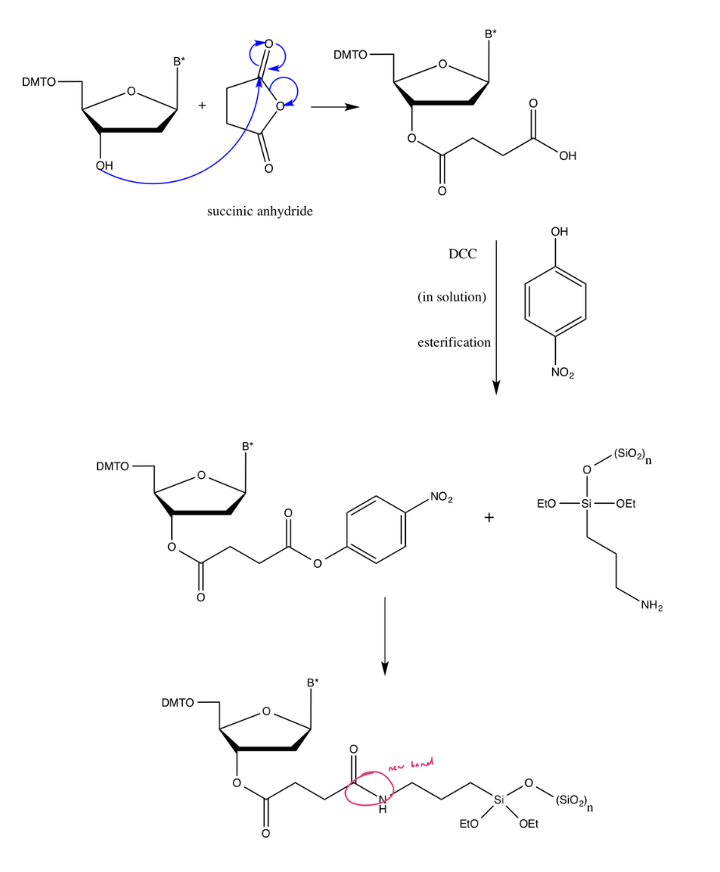
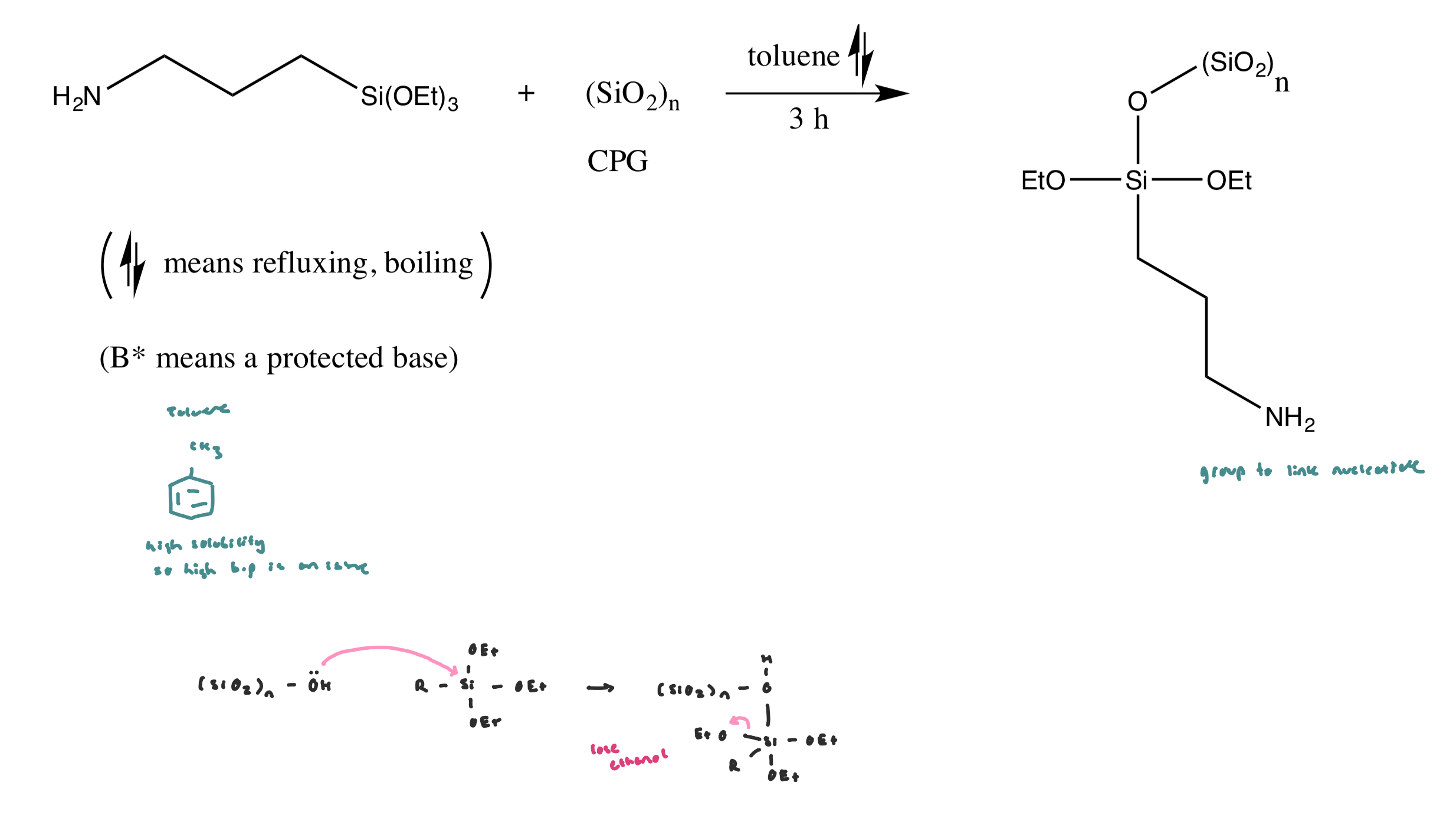
Functionalization of surface silanols (Si-OH)
Triethoxysilane reacts with surface OH groups, creating covalent Si-O-Si bonds, so the silica surface now has amine (-NH2) functional groups sticking out (to react later with other molecules)
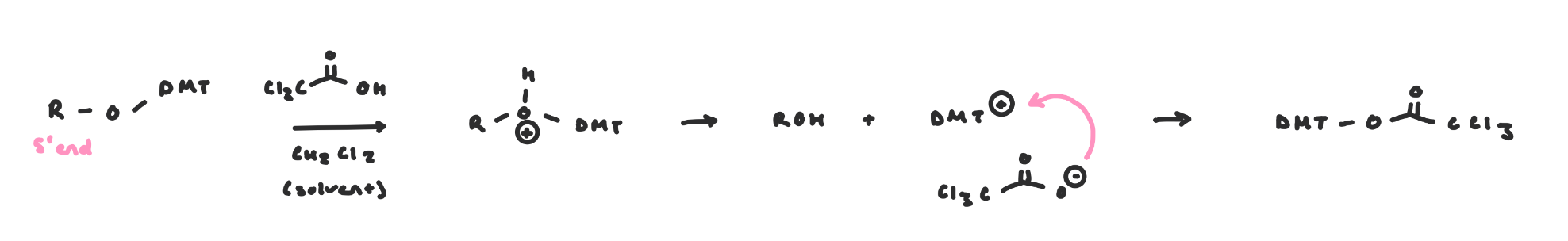
DMT Deprotection
Before coupling to the next nucleoside, the DMT group must be removed.
Use trichloroacetic acid, which proceeds via SN1 mechanism
These conditions don’t affect the other PGs, the phosphate triesters or the linkage to the CPG resin
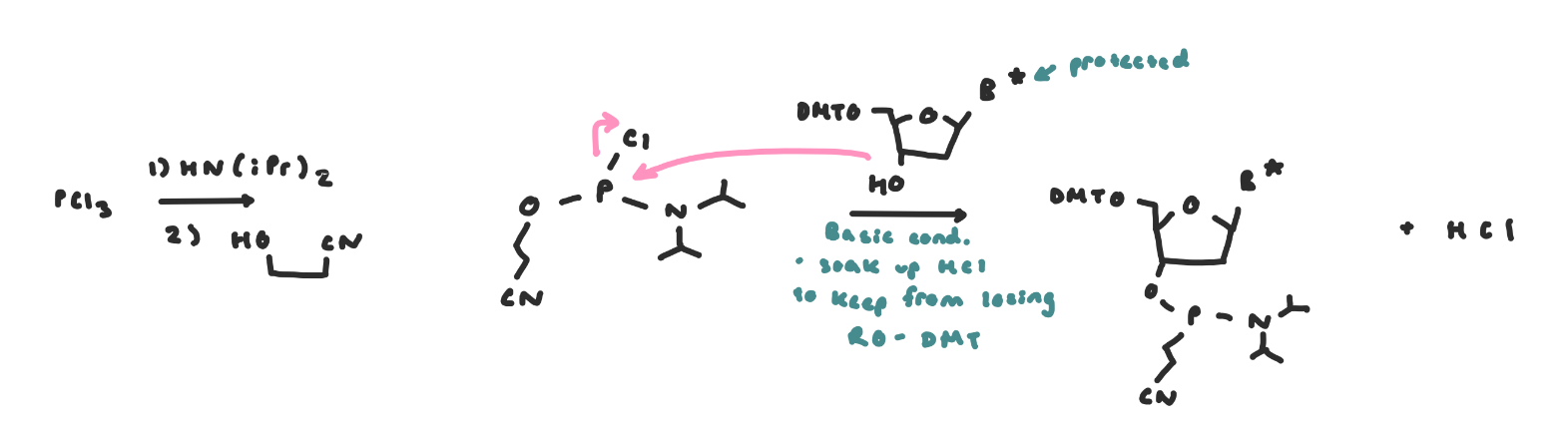
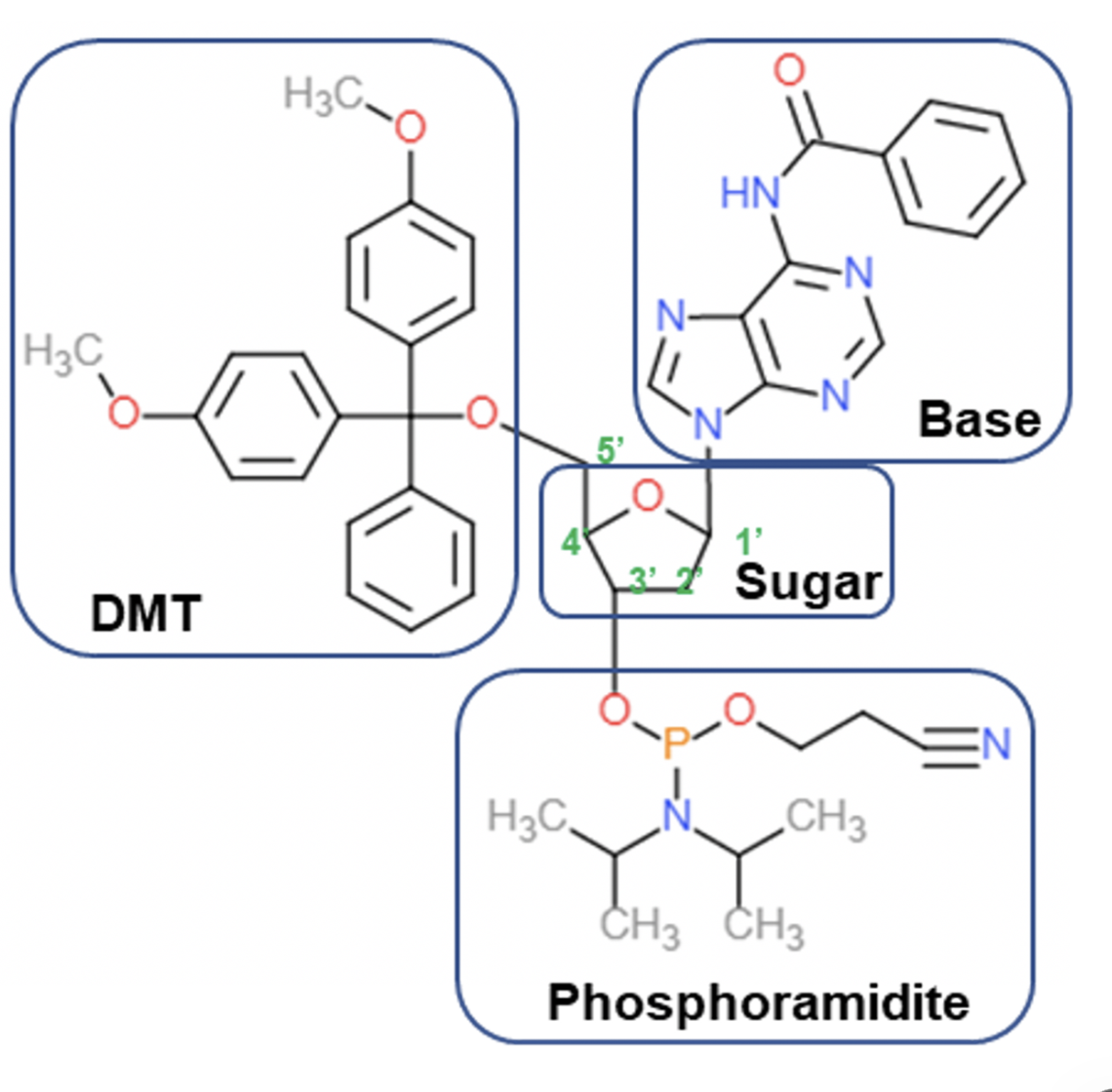
Phosphoarmidite
The incoming nucleoside, protected at its nucleobase and at its 5’OH is then activated at its 3’OH as a phosphoramidite using cyanoethylchloro-N,N-diisopropylphosphoramidite
on its own it’s not reactive enough to directly react with the 5’ OH on the growing DNA chain and has a poor leaving group
The reaction at the phosphoryl chloride proceeds via an SN2-like mechanism on the P atom
figure: activation of incoming nucleotides
Coupling
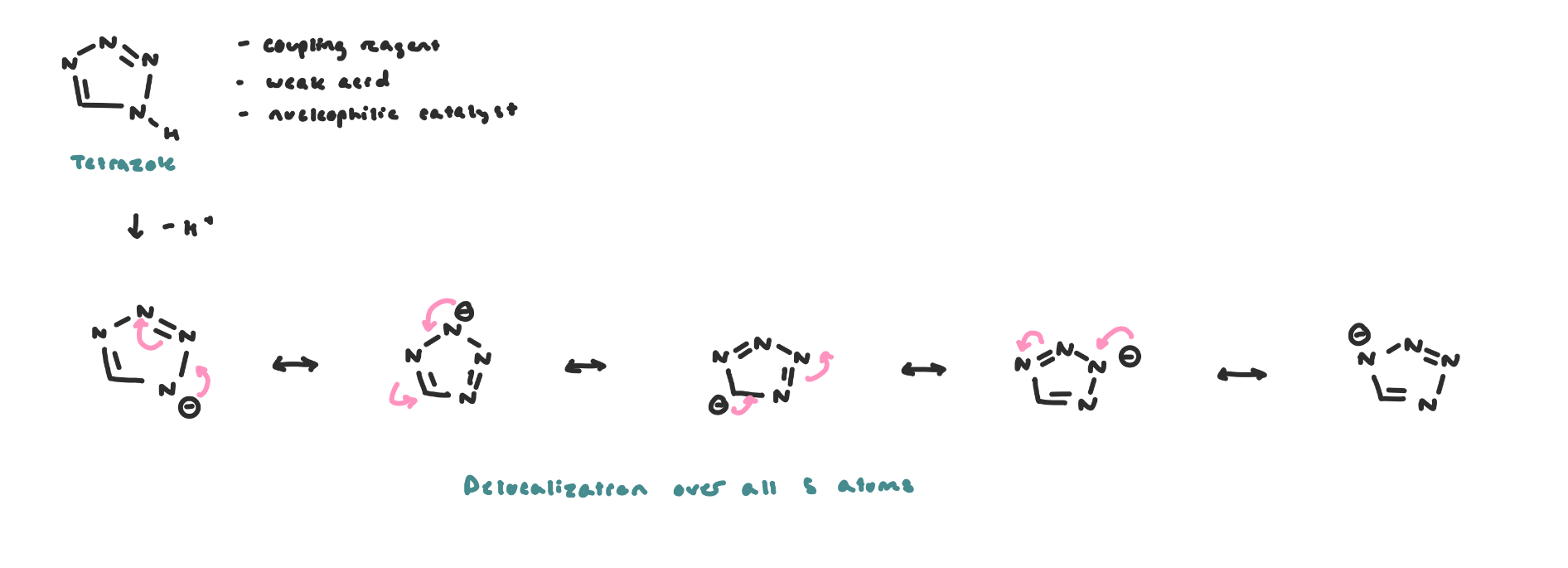
Coupling then occurs using tetrazole as the nucleophilic catalyst
Tetrazole has a pKa of 14.5 in CH3CN (solvent)
The charge is delocalized on all ring atoms
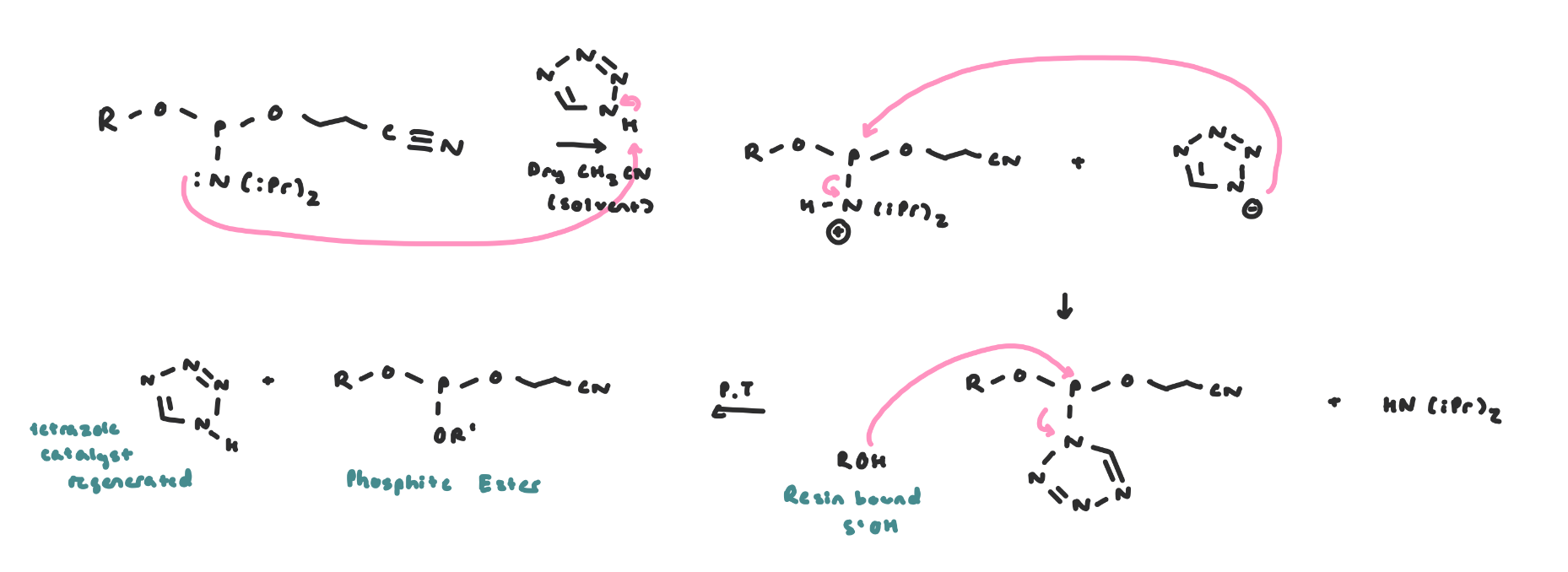
The coupling reaction likely proceeds via nucleophilic catalysis, where the tetrazole anion is the nucleophilic catalyst
Tetrazole protonates the amine on the phosphoramidite, turning the amine into a better leaving group
Then, tetrazole anion attacks P, and we have a reactive intermediate where tetrazole is a better leaving group than NR2 and phosphorus becomes more electrophilic
Coupling: now the 5’OH of the growing chain attacks, tetrazole leaves and the new P-O bond forms
Coupling Mechanism
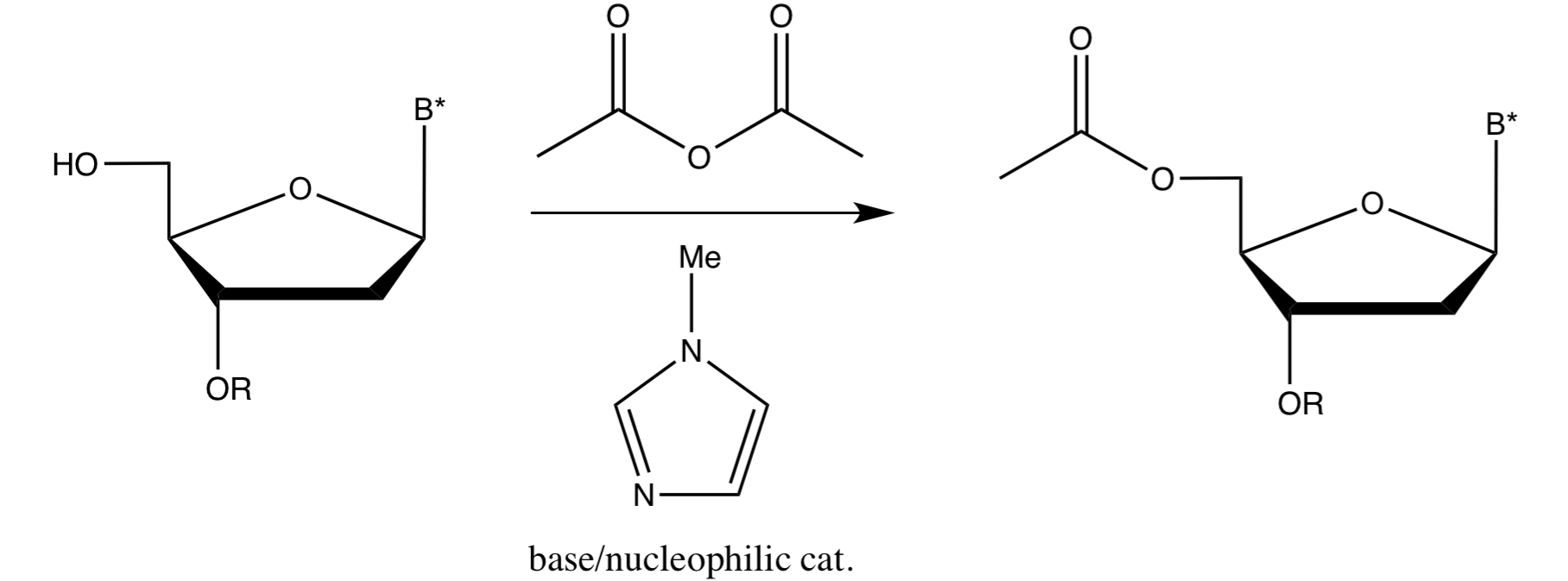
Capping
Following each coupling, capping is often done using acetic anhydride and 1-methylimidazole (nucleophilic catalyst)
This makes purification easier, as most of the impurities are kept small and easy to separate from the large product
Capping is done before oxidation
Only free 5’OH, those that have not reacted with the previous nucleoside, will be capped; the rest are left unchanged
Phosphite Oxidation
Following each coupling, the ensuing phosphite is oxidized to the phosphate, because the phosphite is not sufficiently stable to survive repeated cycles
Phosphite ester → oxidation → phosphate ester
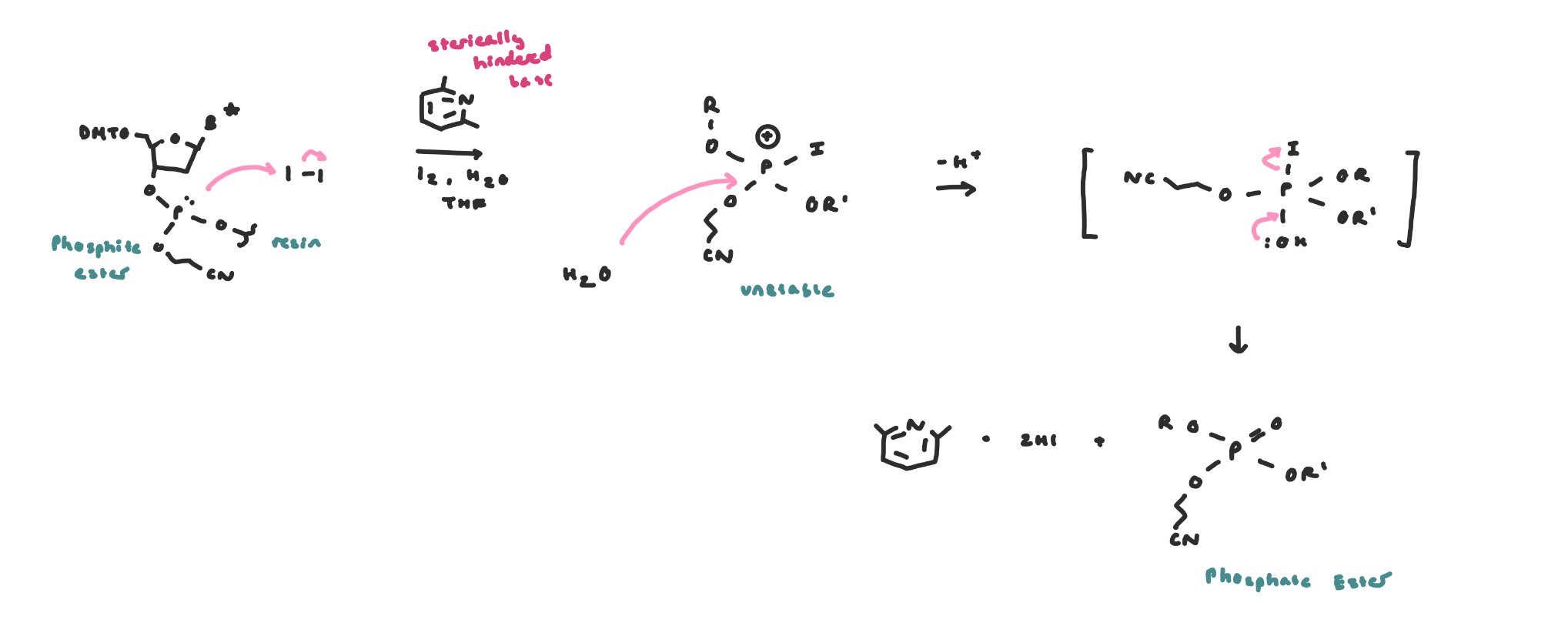
After all the couplings…
Once all couplings are complete, the last DMT is removed (using trichloroacetic acid).
RO-DMT → acid → ROH
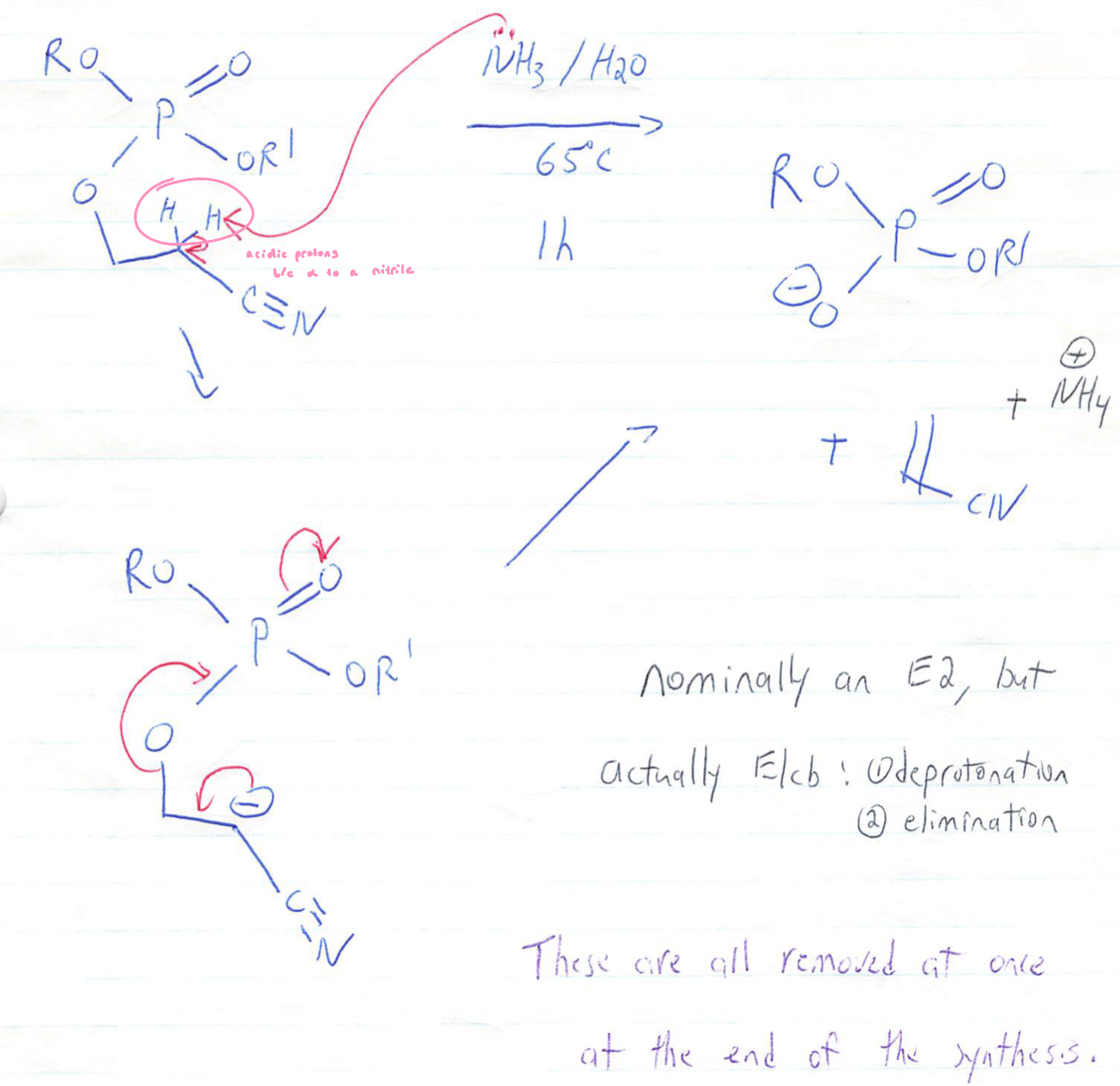
Concentrated ammonium hydroxide (NH4OH) at 65ºC for one hour:
cyanoethyl removal is via an E1 cb (elimination via conjugate base) reaction
All cyanoethyls are removed together at the end of the synthesis, not after each cycle
Base removes a proton → forms a carbanion → carbanion eliminates → kicks out the phosphate oxygen
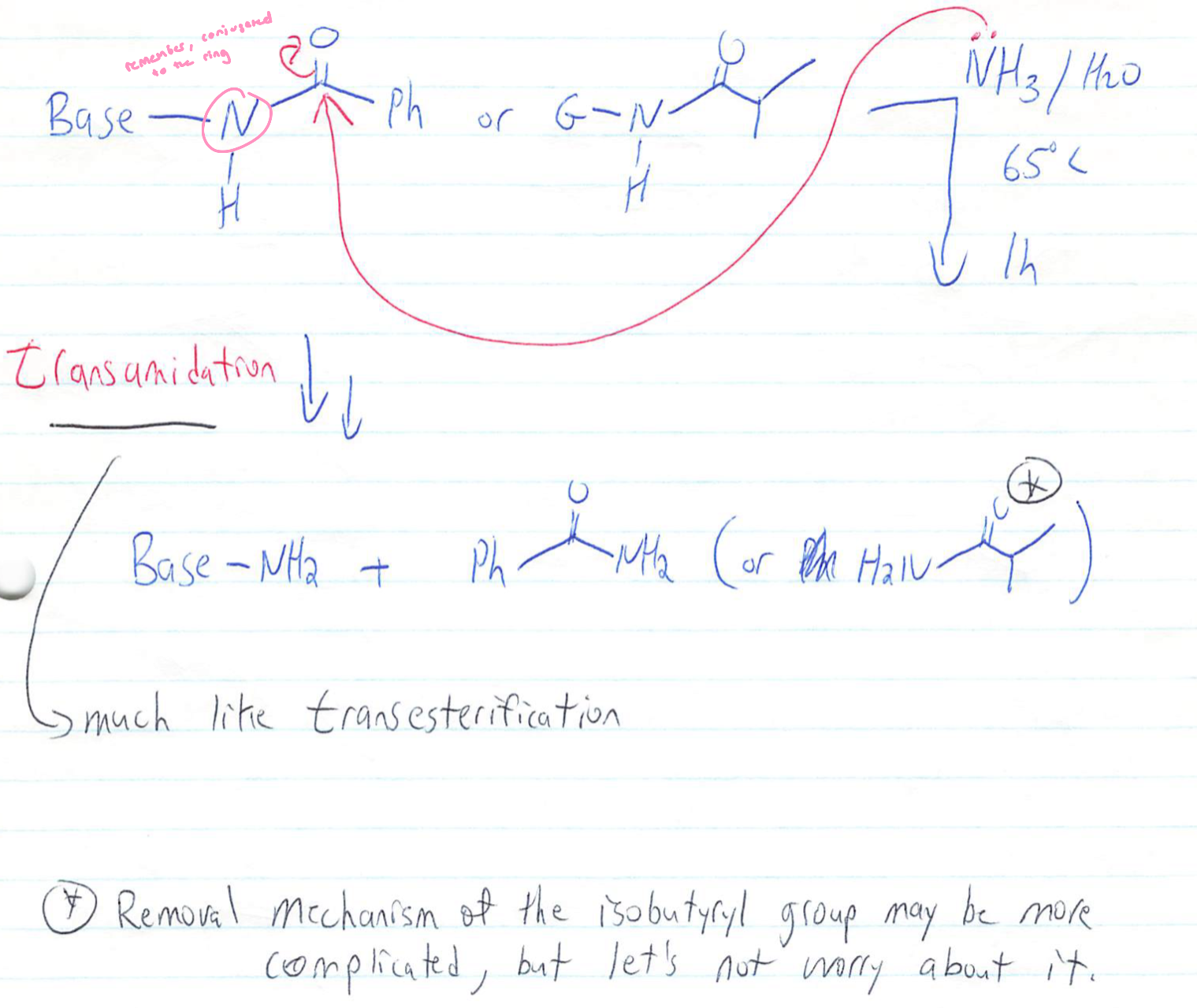
nucleobases are deprotected via transamidation (remember the amine groups were protected)
Ammonia replaces the protecting group
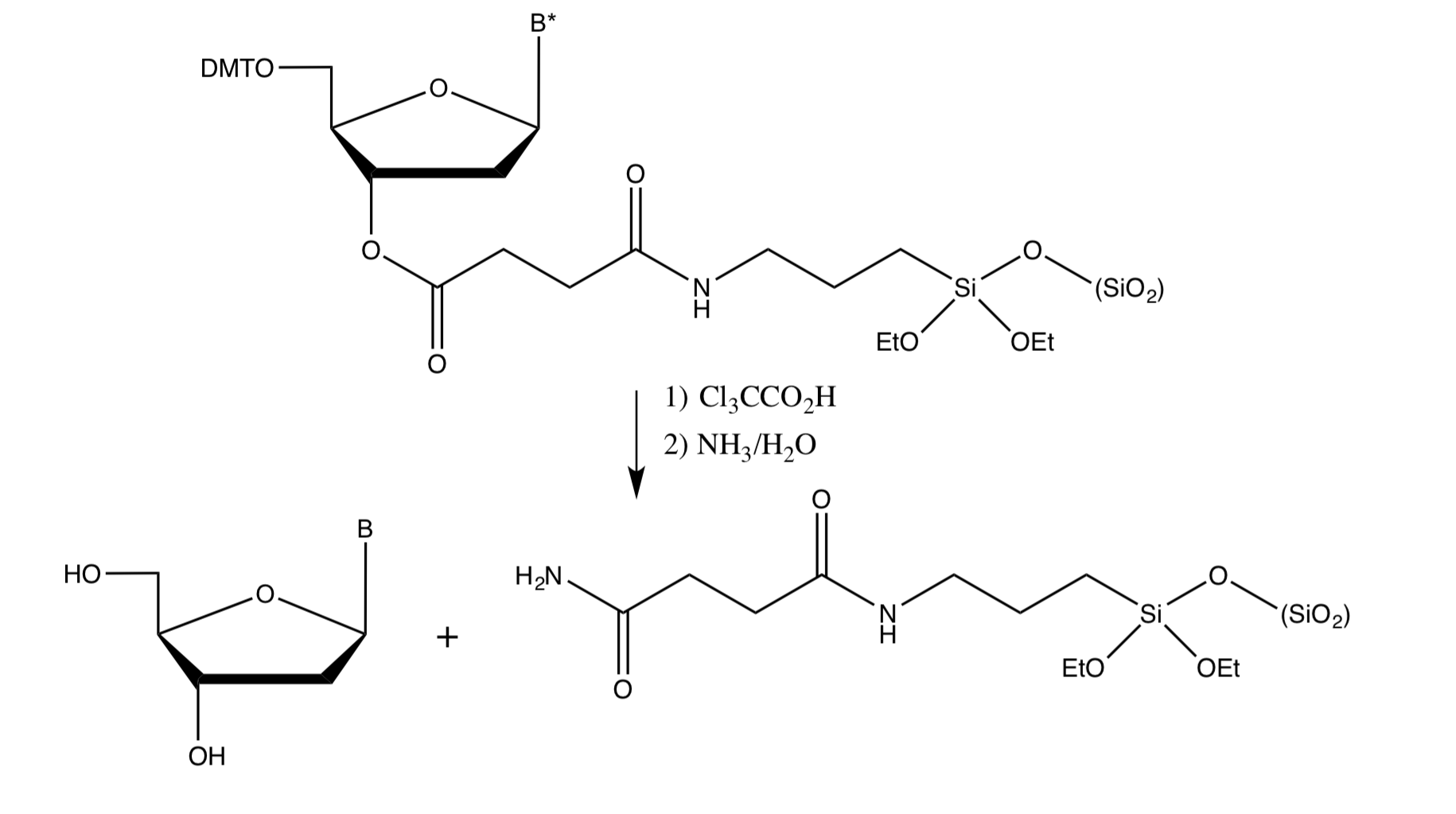
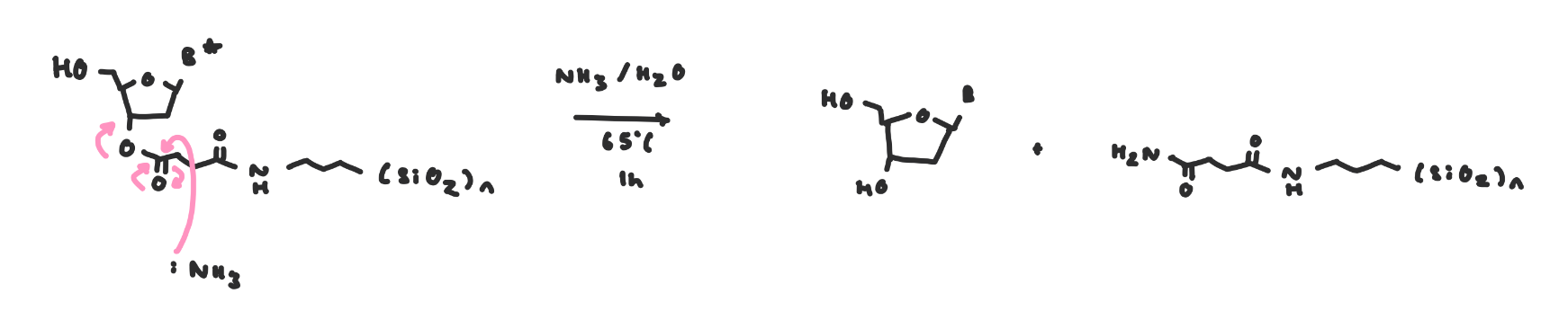
The cleavage from the resin is ester hydrolysis
Deprotection of Phosphotriester (Cyanoethyl Removal)
Deprotection of nucleobases
Cleavage
Summary of solid phase DNA synthesis:
Link nucleotide to CPG protected at the nucleobase (purine/pyrimidine) and the 5’OH
Protect the incoming nucleotide at its nucleobase and its 5’OH
Incorporate the phosphoramidite into incoming nucleotide
Do the phosphoramidite coupling
Cap unreacted 5’OH’s
Oxidize the phosphite to the phosphate
Deprotect the new 5’OH
Repeat steps 2-7 for each nucleotide
Remove final DMT; then cleave oligo from the CPG and remove protecting groups from the nucleobases and the cyanoethyls from phosphotriesters