CH9: Nucleic Acid Methods
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
what are restriction enzymes (endonucleases) and what do they do?
they are prokaryotic enzymes that recognize specific base sequences in double-helical DNA and cleave both strands at specific places
basically split DNA into specific fragments
the sequence they recognize is almost always palindromic (same fwd/bkwd)
describe the names of restriction nucleases
three-letter abbreviation for host organism
strain designation
roman numeral that distinguish multiple enzymes from same strands
how are the cuts different?
some cuts in a staggered way (top example), others cut straight across the DNA double strand (bottom example)
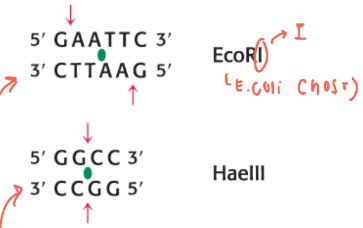
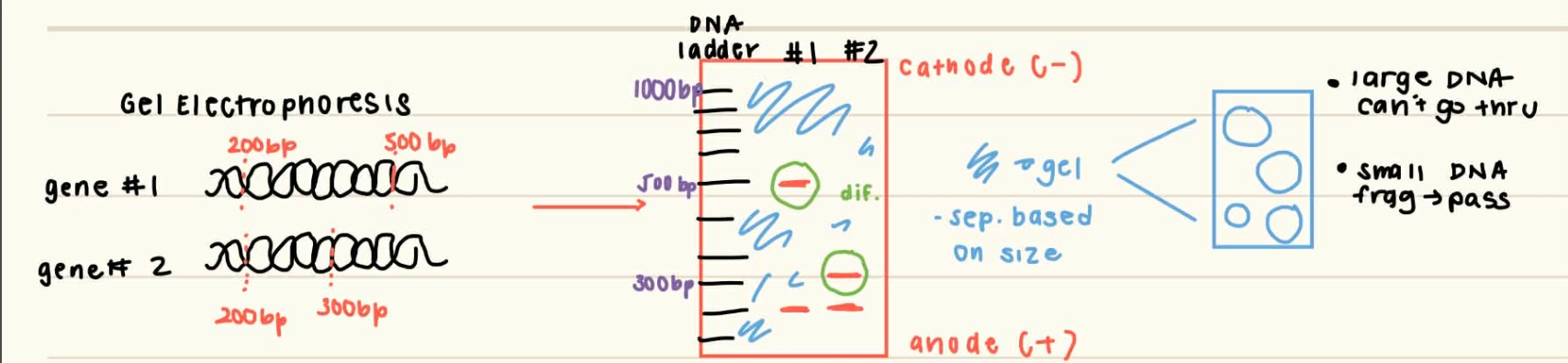
how can restriction fragments be separated and visualized? describe the system and what is used (draw it out)
separated through gel electrophoresis, which separates based on size
in agarose or polyacrylamide gels bc it forms solid porous gels, so only small frags can pass through
visualized by staining with DNA-binding dye like ethidium bromide
what is southern blotting and northern blotting? what is a DNA probe? describe the steps and why we use nitrocellulose membrane
southern blotting: technique where restriction fragment with specific sequence can be identified by hybridizing it with labeled complementary DNA strand
1. DNA cuts to frag, 2. run gel electrophoresis, 3. transfers to membrane (copying part), 4. add labeled DNA probe, 5. probe binds to matching DNA sequence, 6. detect where it’s bound w/ autoradiography or fluorescence imaging
finds specific genes
northern blotting: same procedure but for RNA
checks gene expression
probe is a labeled short stretch of single-stranded DNA with a known base sequence & binds to complementary sequence on fragment
32P labeled or fluorescently tagged
use nitrocellulose membrane bc it’s stable & easy to probe compared to fragile gel
describe the sanger method, what is it used for? what is ddNTPs what are the steps (5)?
controlled termination of replication = copying DNA but stopping the copy to figure out sequence
1. copy DNA,
2. random stops with (ddNTPs who lack 3’ OH so chain can’t continue)
3.separate fragments by size via capillary electrophoresis (shortest come out 1st)
4. detect colors (color order = DNA seq)
5. read sequence (smallest to largest frag, gives 5’-3’ of new strand)
what are the components of controlled termination of replication reactions? (sanger method)
DNA polymerase (makes new DNA strand based on template & adds complementary dNTPs to primer)
primer
template (DNA to be sequenced)
four deoxyribonucleoside triphosphates
small amount of 2’,3’-dideoxy analog (stops connectoin) of each nucleotide, each with different florescent label
what is the 2’,3’-dideoxy analog and what does it do?
has no 3’ OH so when added, it blocks further polymerization & terminates the chain
present in low enough concentration that chain termination takes place only occasionally
creates fragments of different lengths
what is oligonucleotide synthesis? (technique name? and how it works)
occurs by sequential addition of activated monomers to growing chain linked to a resin (solid)
(solid-phase method)
DNA is built one nucleotide at a time, growing chain is attached to solid support (like glued to table)
Scientists add one base —> wash —> add next base —> repeat
what are activated monomers and what are they for?
they are protected deoxyribonucleoside 3’-phosphoradmidites
protected means parts of blocked temporarily so only correct bond forms
they are DNA building blocks used in synthesis
how can newly synthesized DNA be seen? (2 ways)
32P (radioactive phosphorus)
fluorescent markers
What can we use oligonucleotide synthesis/solid phase methods for? (3)
as a probe (short DNA that finds matching sequence)
as a primer (short DNA strand that starts DNA replication)
synthesizing new tailor-made genes ( design/build custom dna seq)
what are the protecting groups for deoxyribonucleoside 3’-phosphoramidite, where are they located? what do they block? (2)
5’ -OH is blocked by dimethoxytrityl (DMT) at the TOP end of nucleotide
prevents nucleotide from connecting at wrong place
removed so nucleotide can attach to growing chain
3’-Phosphoryl oxygen atom is blocked by b-cyanoethyl (bCE) at the BOTTOM end of nucleotide
prevents unwanted reactions at 3’ end
later is removed for chain elongation
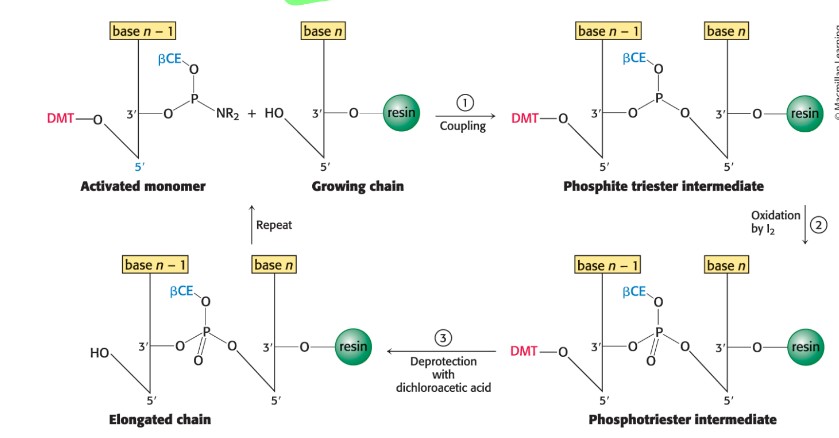
what are the steps in oligonucleotide synthesis via phosphite triester method?
deprotection/detritylation: 5’ DMT protecting group removed from previous base to let next base to connect
coupling: a new, activated phosphoramidite nucleotide is added to chain
oxidation by I2: phospite triester linkage is converted into a stable linkage (P gains =O)
capping: unreacted strands are capped with acetylation to prevent truncated sequences in future cycles
what is PCR? what do these reactions require? (3)
polymerase chain reaction (PCR) = method for amplifying specific DNA sequences
PCR rxns require
pair of primers that hybridize w/ flanking sequence (regions on either side) of the target
all four deoxyribonucleoside triphosphates (dNTPs used to add next nucleotides)
a heat stable DNA polymerase (bc sample exposed to heat)
what are the 3 steps of a PCR cycle?
denaturation: 2 strands of parent DNA separated by heating solution to 95C for 15s (heat stable dna pol Taq can survive this)
annealing: solution is cooled to 54C to let primers anneal (stick/zipper) to DNA ( new bonds formed)
extension: solution is heated to 72C to let Taq polymerase elongate both primers in direction of target sequence
how much is desired sequence amplified by in PCR?
after n cycles, desired sequence is amplified 2^n-fold
what are four applications of PCR in the real world?
diagnose infections
detect certain cancers early & monitor chemotherapy
solve forensic cases
study molecular evolution
polymerase chain reaction….