L1: Subcellular turnover and microscopy
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
Degradation of cell components: Two mainpathways of protein-containing degradation
Ubiquitin-proteosome system
for single unfoldable proteins
Autophagy
less digestible proteins
organelles
macromolecule complexes
Ubituitin-proteasome system
For single unfolded polypeptides
Recognised as misfolded
tagged by covalent additios of poly-ubiquitin side chains
recognised by proteasome (barrel-shaped multiprotein complex)
fed through for degradation
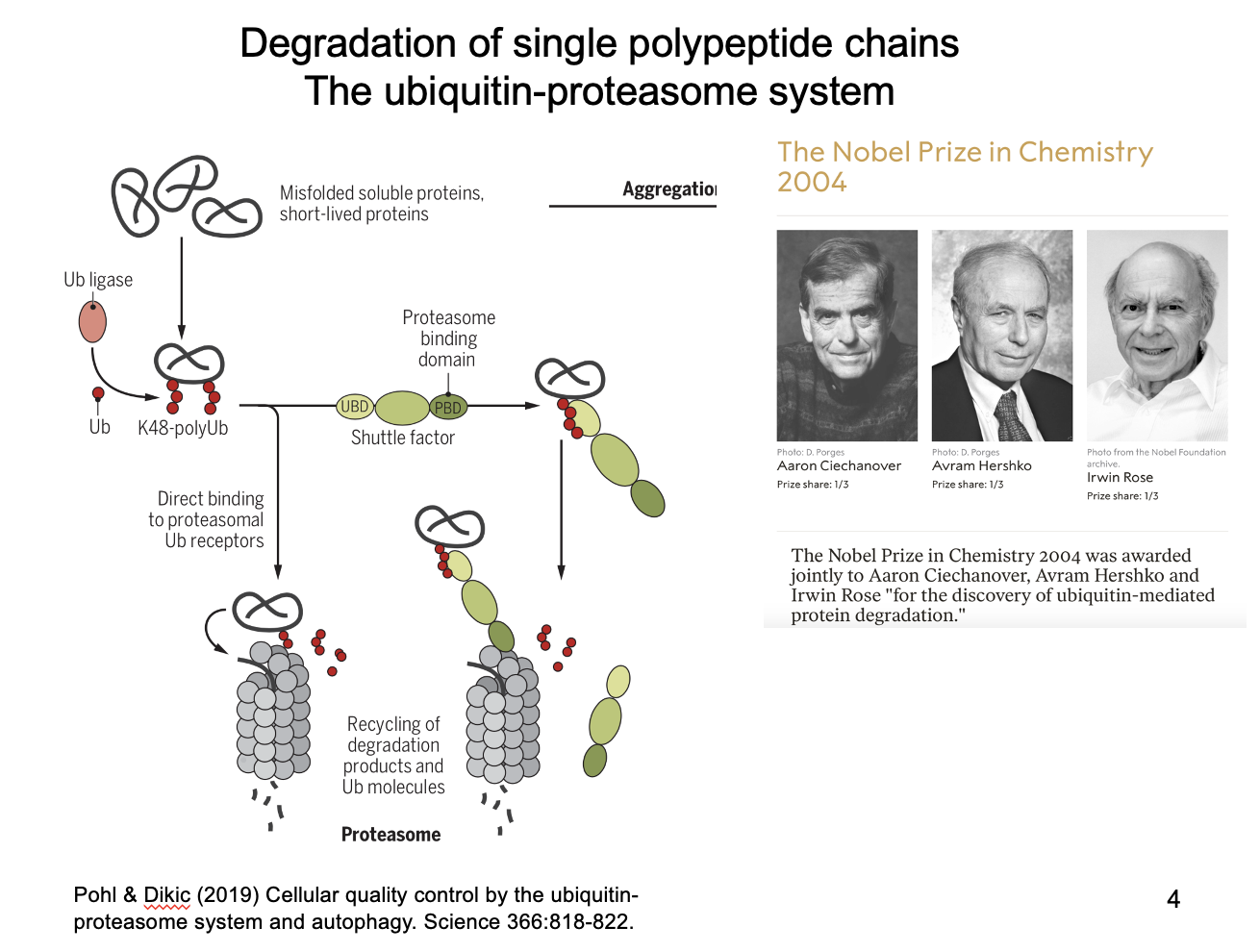
What is the protasome also used to degrade?
ER-associated degradation (ERAD)
ER lumen proteins are proteolysed into peptides
translocated retrogradely into cytosol
for proteasomal degradation
Autophagy and its role
‘self eating’
Lysosomes degrade cellular components→ form autophagosomes i the the lysosome
Are induced by amino-acid starvation
Its role:
suggesting a role is responsing to amino acid storages
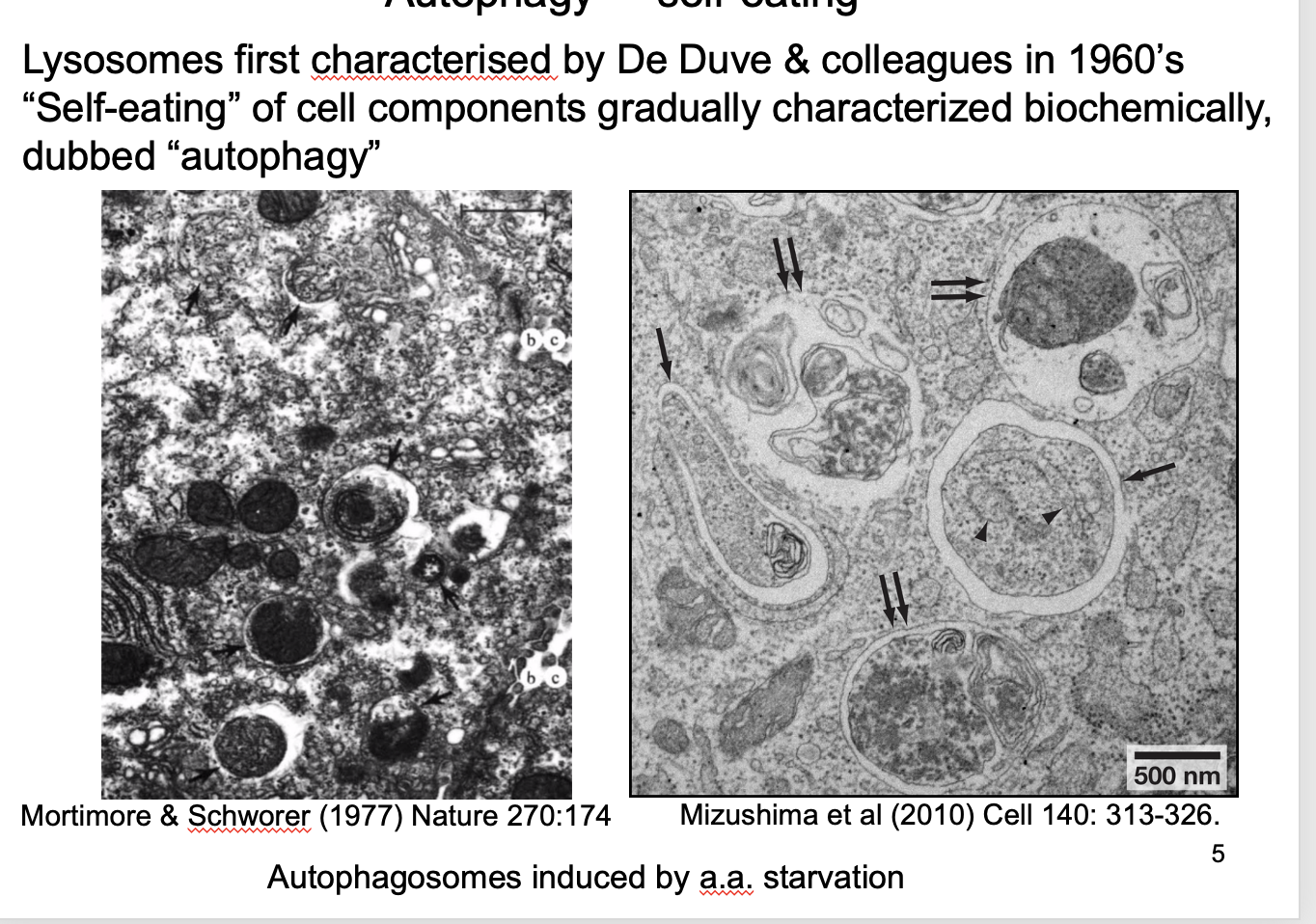
How discovered/ investigated
‘Autophagic' bodes’ accumulates in yeast vacuoles with imparied protease activity
Loss of ABs can be used to screen for mutants with defective autophagy
Recovery of ATG mutants (apg in yeast) or blockage using drugs identified
components of autophagy machinery
and defined steps depending of where autophagy was blocked
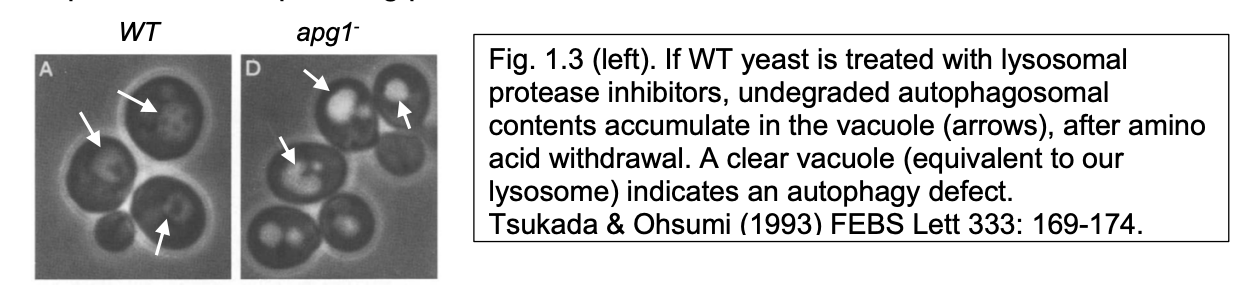

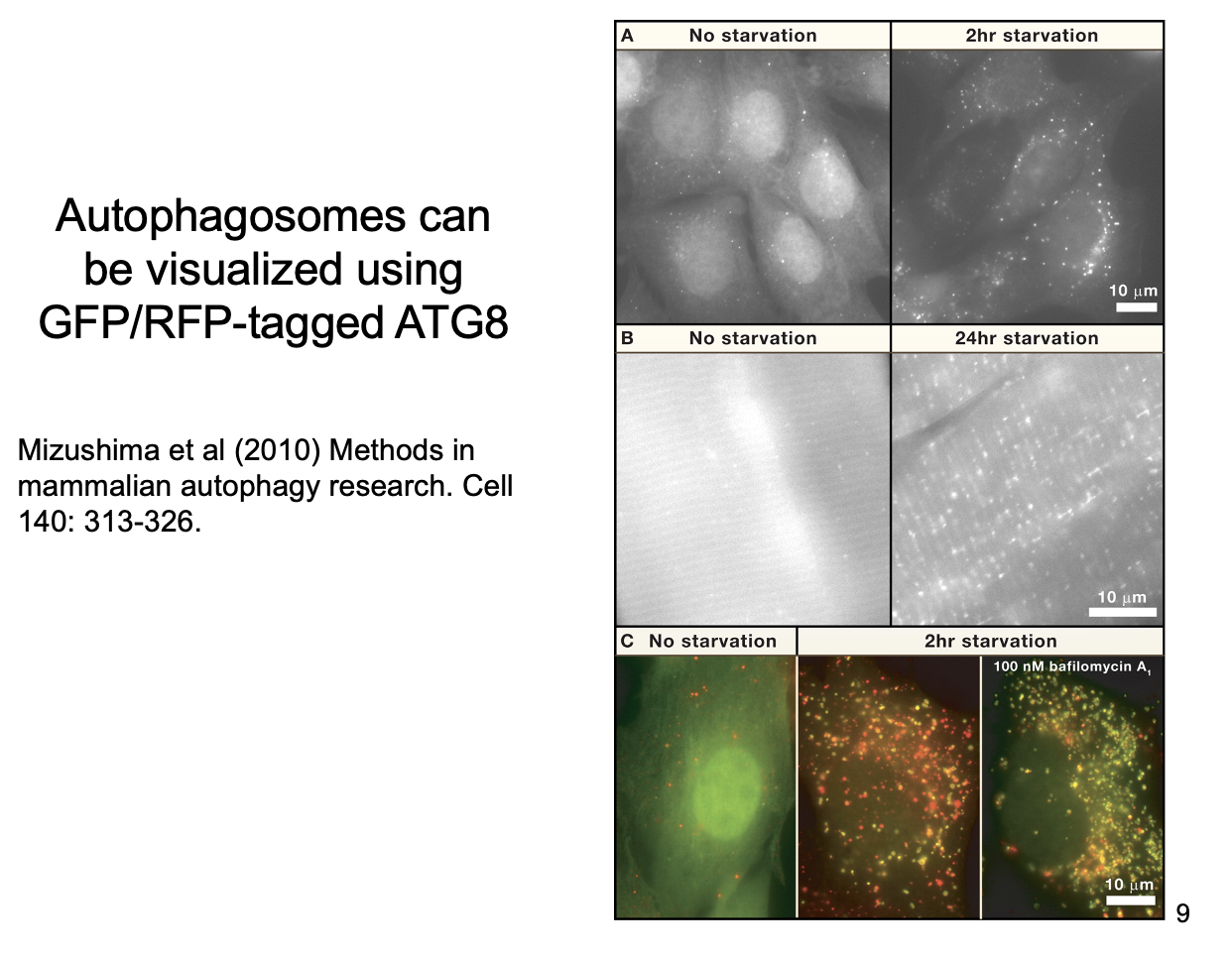
What are the steps in Autophagy
note: can use GFP to visualise the AB
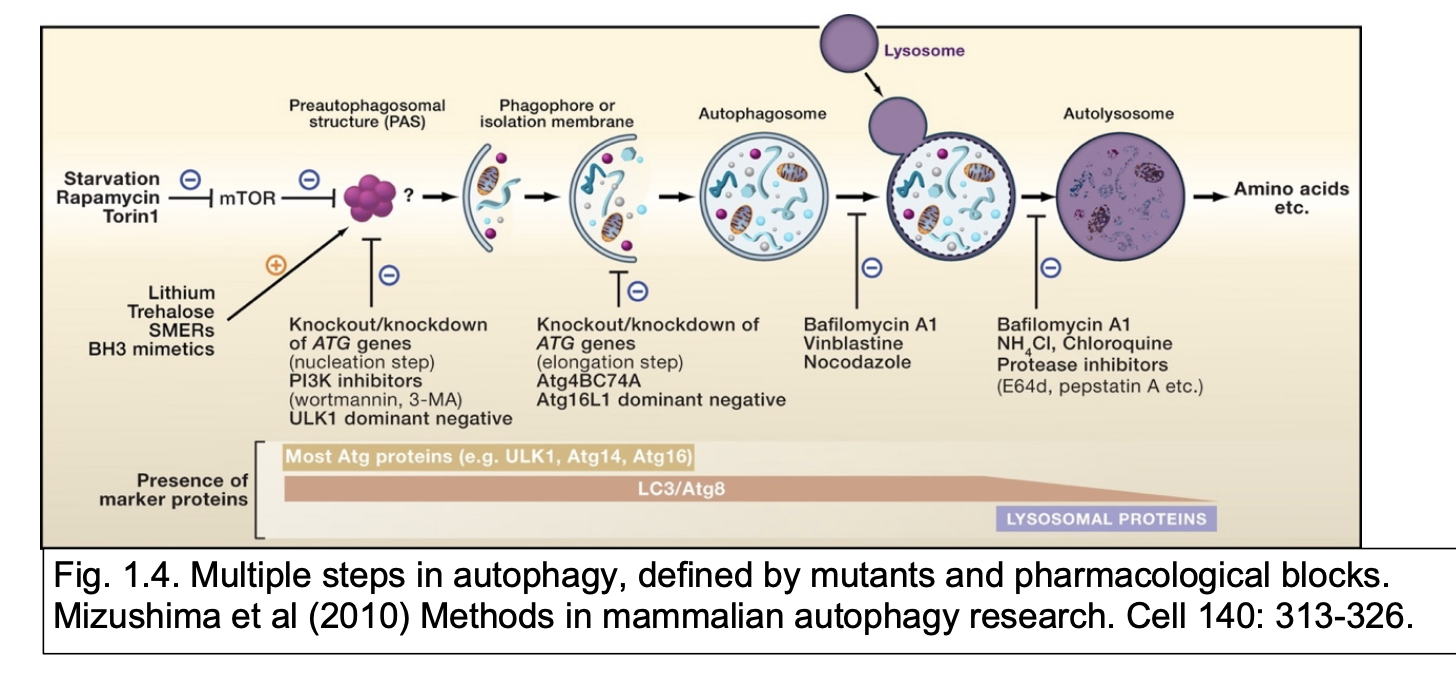
Note: Macroautophagy vs Microautophagy vs Chaperone-mediated
Macro→ normal ‘autophagy’
Micro→ invagination of autophagic substrates into autophagosomes
doubled up by lysosomes
Chaperone mediated→ translocation of chaperone-bound denatured substrates from the cytosol across the lysosome membrane into the lysosome
i.e the protein is translocated in directly
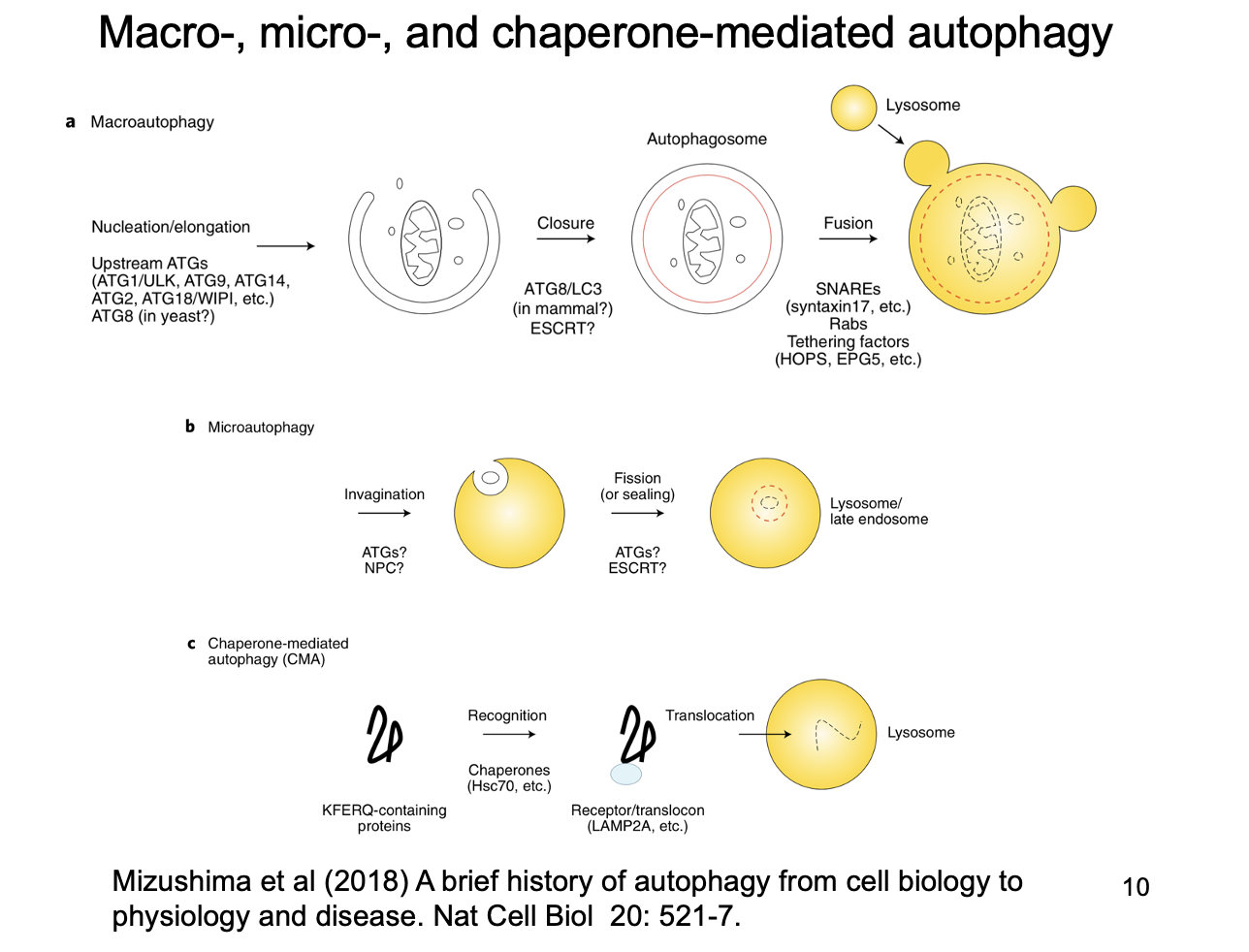
Autophagy can be divided into two types depending on what its substrates are
Constitutive
many cell components can be degraded
non specific
but, overall level is controlled by need (e.g need to recycle nutrients)
Substrate-specific
targeted
specific organelles are recognised by autophagy machinery
autophagy adaptors that recognise ‘eat-me’ signals on surface of substrate
when there is a physiolgical need to remove that specific organelle/substrate
e.g mitochondria autophagy→ mitophagy
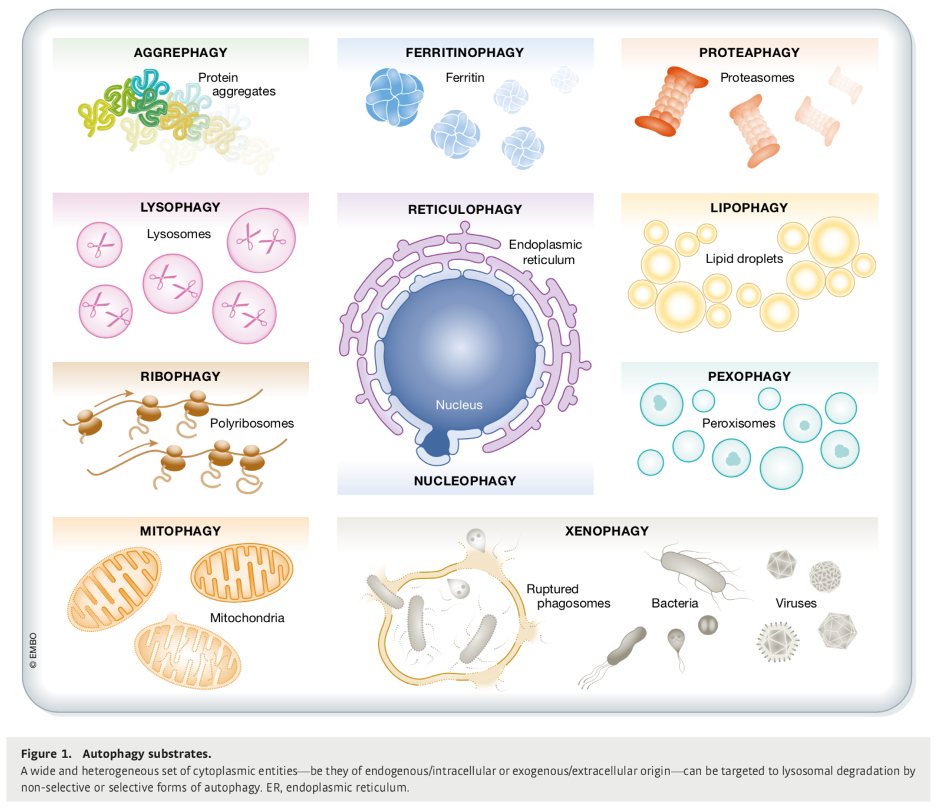
Types of ‘eat-me’ signals
Proteins covalently modified by covalent attachent of ubiquitin protein
decrepit organelles
toxic protein aggregates (found in many neurodegenertaive disease
intracellular pathogens
Roles of autophagy
Starvation
Aggregates (protein and RNA)
Organelles: mitophagy, ERphagy, lipophagy
What is autophagy controlled by
Must be appropriate substrates and conditions
Controlled by:
mTORC1
autophagy adaptors
How was the regulation of autophagy investigated and findings
Rapamycin:
Found to be inducer of autophagy
Investigated finding its cellular targets:
rapamysin-resistant mutants of yeast
Eventual findings:
Cellular target contained a large protein kinase, Tor
Tor→ ‘target of rapamysin’
mTor→ mammalian Tor
What is mTor and its role
Protein kinase
forms two alternative complexes:
mTORC1
mTORC2
These two have different effects but overall key in signalling
Sensing availability of energy and nutrients
Regulating cellular responses to nutrient availability
How does tapamycin work on this?
Rapamysin binds to FKPB12 (another subunit of mTORC1 complex)
makes mTor activity in mTORC1 sensitive to acute rapaysin exposure
Effects of mTORC1 vs mTORC2
mTORC1→ Promotes growth (blocks autophagy)
mTORC2→ enables autophagy?
Complementary roles
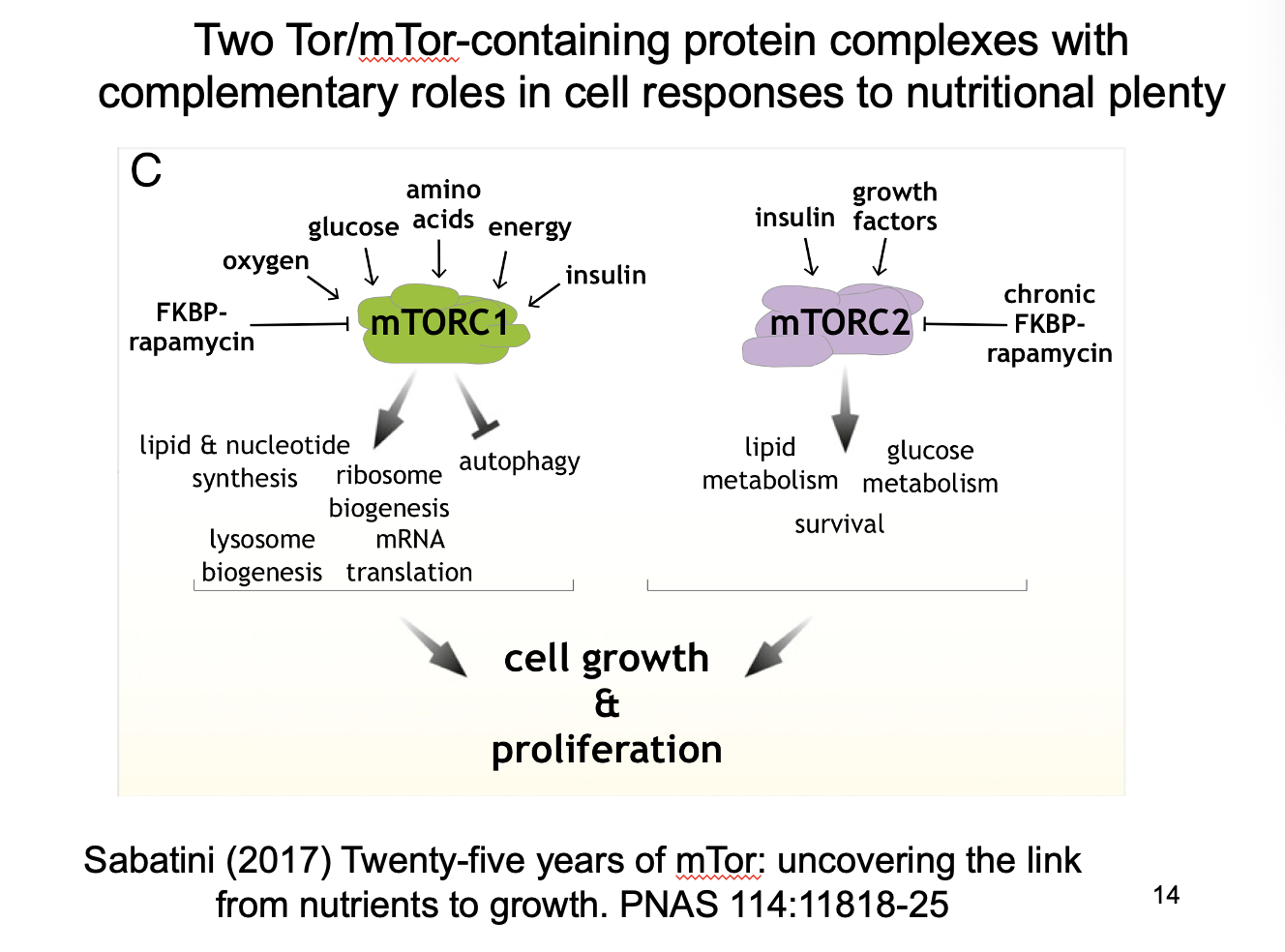
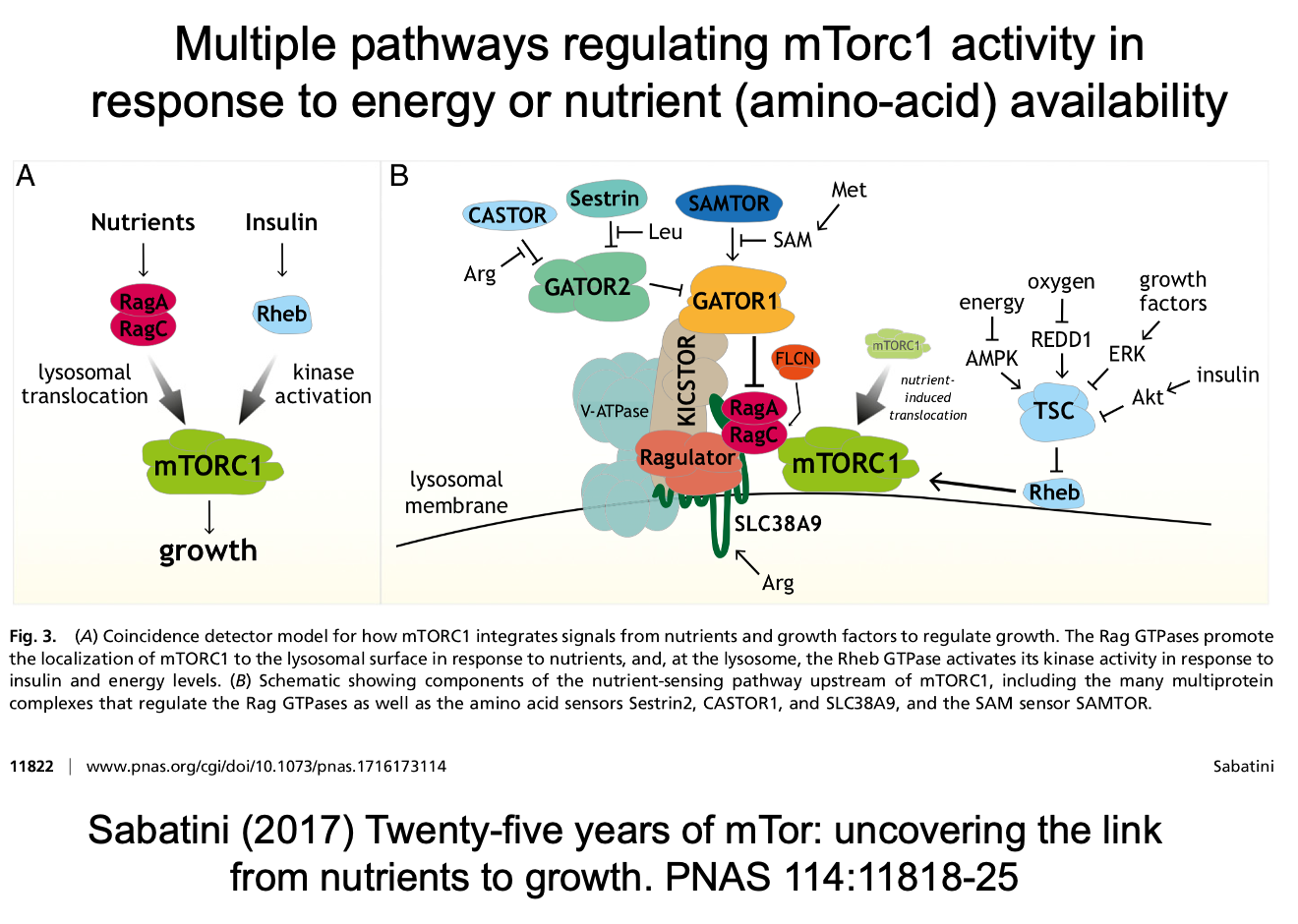
Nutrient regulation of mTORC1
There are multiple pathways in regulating activity in response to energy or aa availability:
Nutrients→ RagA/C (GTPases)→ lysosomal translocation→ mTORC1→ growth
Insulin→ Rheb→ kinase activation→ mTORC1→ growth
Amino acid detection
Converge on rag
different sensors for different aa
Ultimately sensing nutrients on the surface of the lysosome
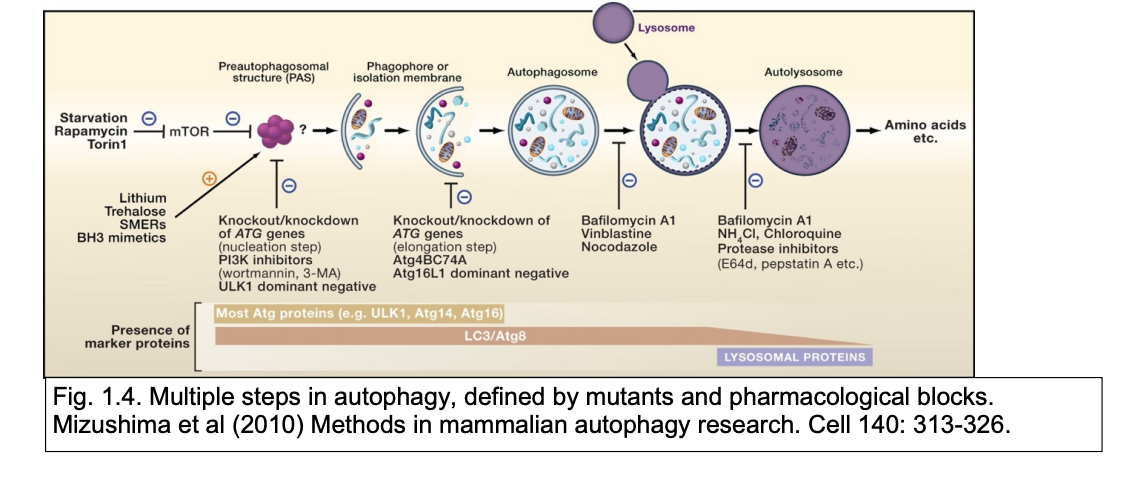
Regulation of autophagy by mTORC1
In general:
mTORC1 phsphylates some early players in the autophagy pathway
This is important in regulating constituitive autophagy
When high nutrients:
high mTORC1 actvity
Inhibits autophagy
promotes cell growth and high biosynthetic activity
In famine:
low mTORC1
disinhibits autophagy
breaking down cell components
provide new sources of aa, nucleotides, lipids for essential cell function in absence of external sources
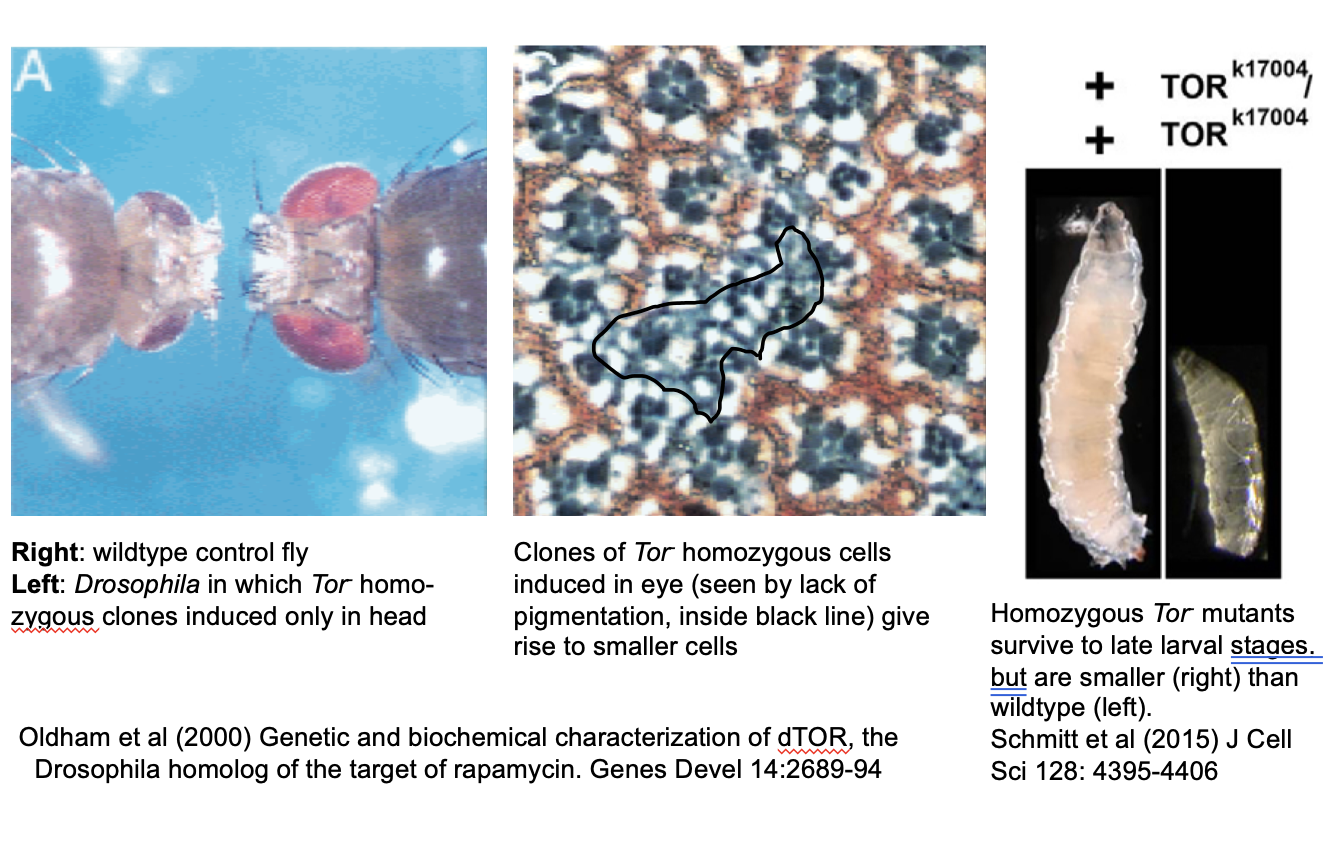
Evidence for mTORC1 role in autophagy: diseases caused by mutation to mTORC1
e.g Drosophila Tor mutants
Human TSC1/TSC2 mutations in tuberous sclerosis
number of growths/cysts in many organs
from excess mTORC1 activity
Neurodegneration (ALS, FTD) by various autophagy mutants
defective clearance of protein/RNA aggregates
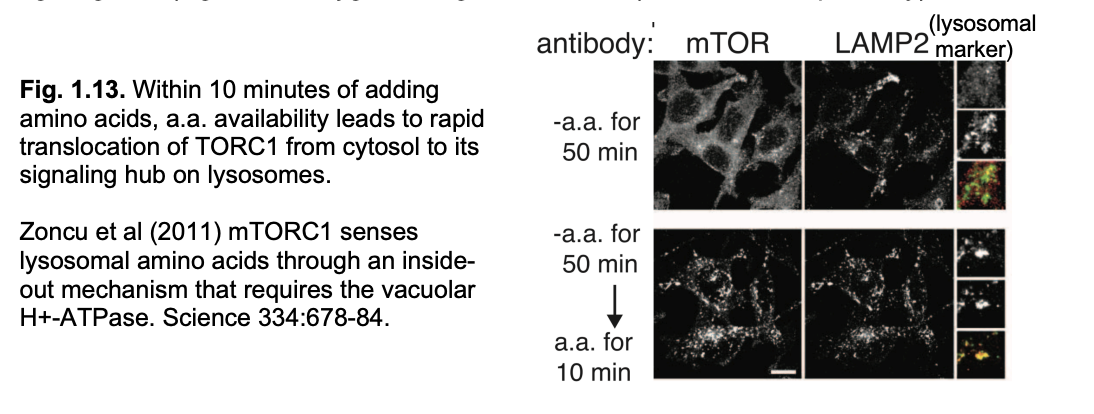
But how is mTorc1 itself regulated? Localisation
Localisation→ seen with immunofluorescent miscopy
In starvation→ mTORC1 is localised to the cytosol inactive
When aa available→ activate→goes to surface of lysosomes
Here is it regulated by all signallig machinery on the lysosome surface that respond to signals of
aa abundance (Via Rag/Regulator pathway)
glucose
oxygen
growth factors (via TSC/Rheb pathway)
Lysosomes themselves are dynamic
transported by MT based motors
aa availability can lead to relocalization from peri-nuclear to the cell periphery
Lysosomes as an amino acid store
degerneative activity makes them rich stores of amino acids
must be tightly regulated by
nutritional state
local needs in the cell
Example of lysosome store regulation and evidence
Release aa to cytoplasm
activate mTORC1
EVIDENCE
mutating a channel that releases aa from lysosome to the cytosol
blocks mTORC1 activation
Conclusion:
one way TORC1 is regulatede if through lysosome aa efflux to signal nutrient availability
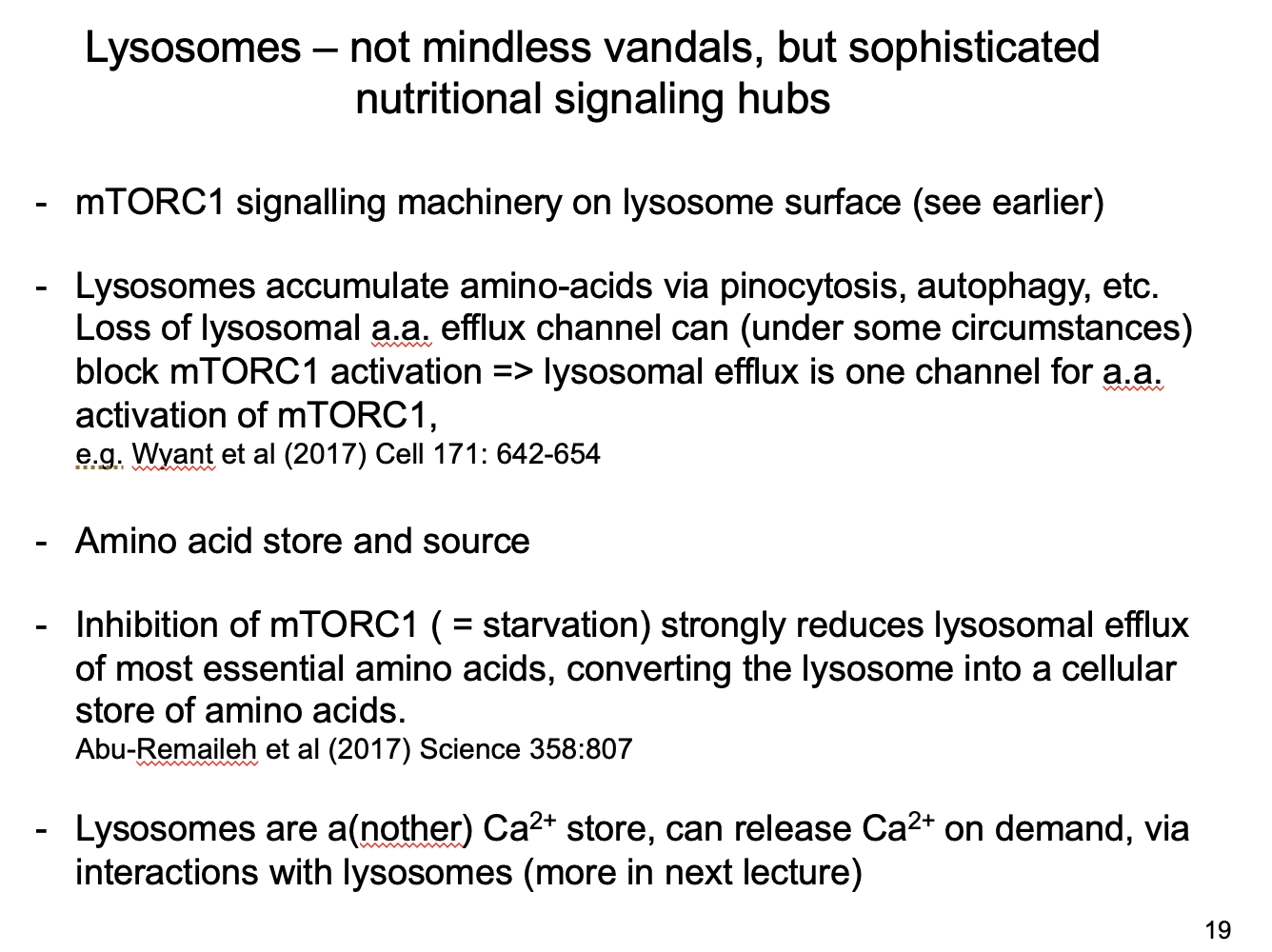
How can mTORC1 regulate the lysosome
In starvation
mTORC1 inhibition
reduces efflux of aa from lysosome
converts lysosome into a cellular store of essential aa for essential protein synthesis
Overall roles of lysosomes
Protein degradation (aminio acid source)
Amino acid store
linking store with cell nutritional status→ mTORC1 signalling
highly localised store for used for protein synthesis
A Ca2+ store
can release Ca2+ on demand
via interactions with lysosomes (see next lecture)

Other nutritional signalling pathways besides mTor
AMPK
ATP demand rise
ATP fall
AMP rises
activates AMPK (AMP-dependent protein kinase)
phosphorylates mutliple targets to restore mitochondrial function/biosynthesis
reduce ATP consumption
Insulin (physlogenetically ancient signallig pathway)
adapts metablsm and physiology
at cellular and organismal level
to nutritial states of high carb/lipid
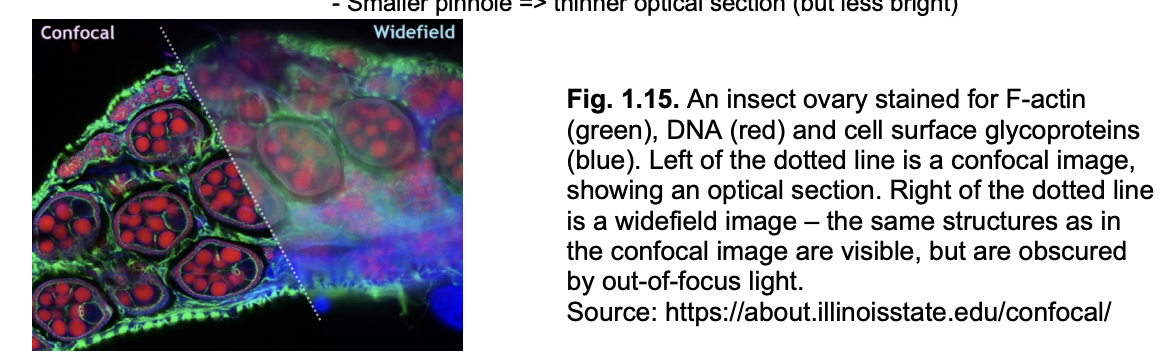
Seeing inside organelles: types of fluoresecent miscrocopy methods
Widefield
Confocal
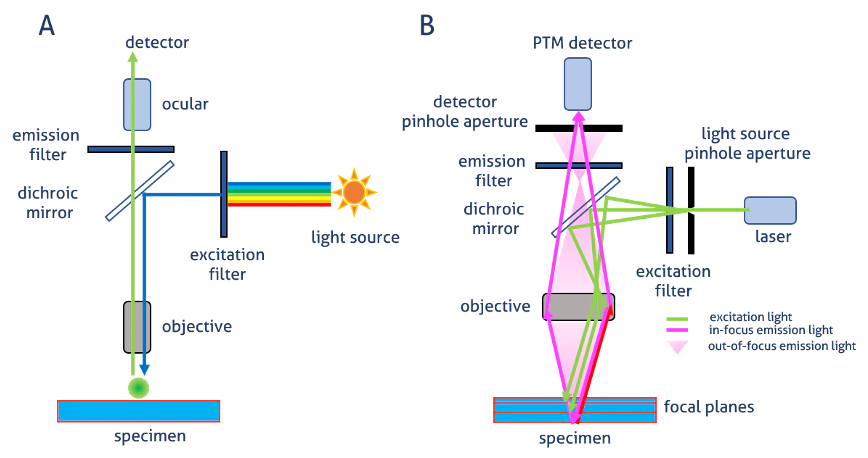
Widefield
whole smaple illuminated
out-of-focus light doesn’t form an image
but still reaches detector
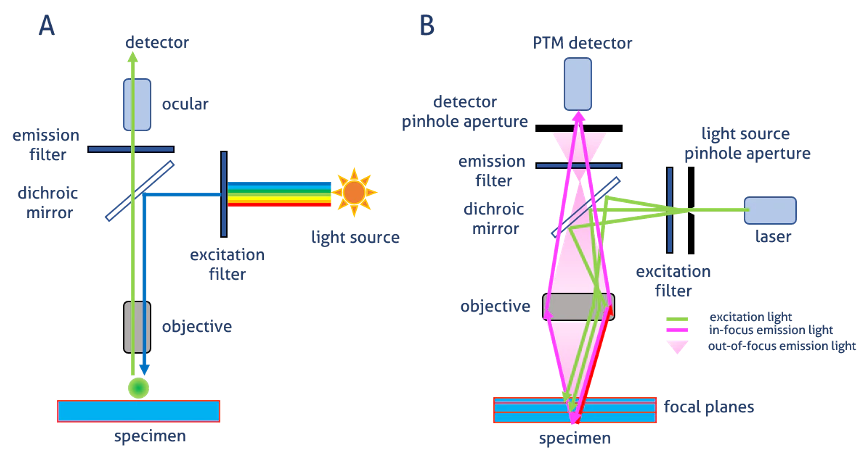
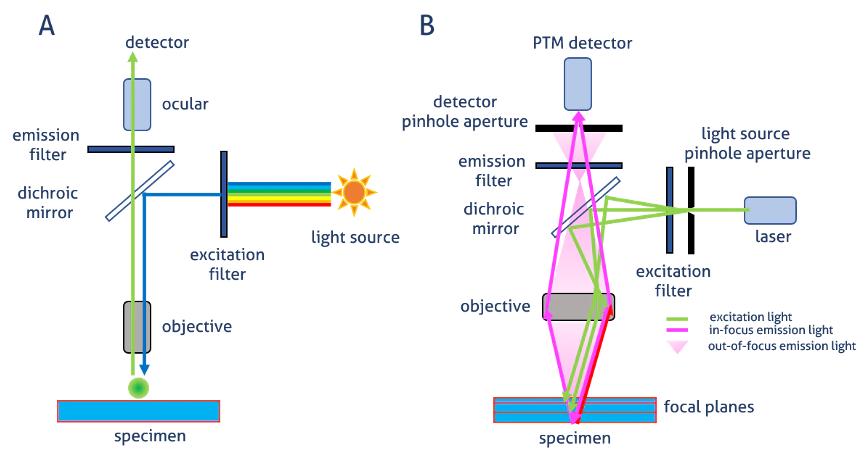
Laser scanning confocal microscopy
overall: light through a pinhole to cut out unfocused light
single point illuminated with bright laser
rapid scanning of laser across the sample builds up an image
Pinhole in the detector light path
allows all light from focus plane through
Out-of-focus light mostly excluded by pinhole→ image is an optical section
Smaller pinhole→ thinner optical section (but less bright)
In order to image sub-organelle resolution we need and why
Super-resolution microscopy:
most organelles at 1um to few um
slightly larger than the theoretical limit of resolution of light microscopy
(half the wavelength of visible light (diffraction limit))
So organelles can just about be seen with light but
sub-organelle is hard to distinguish
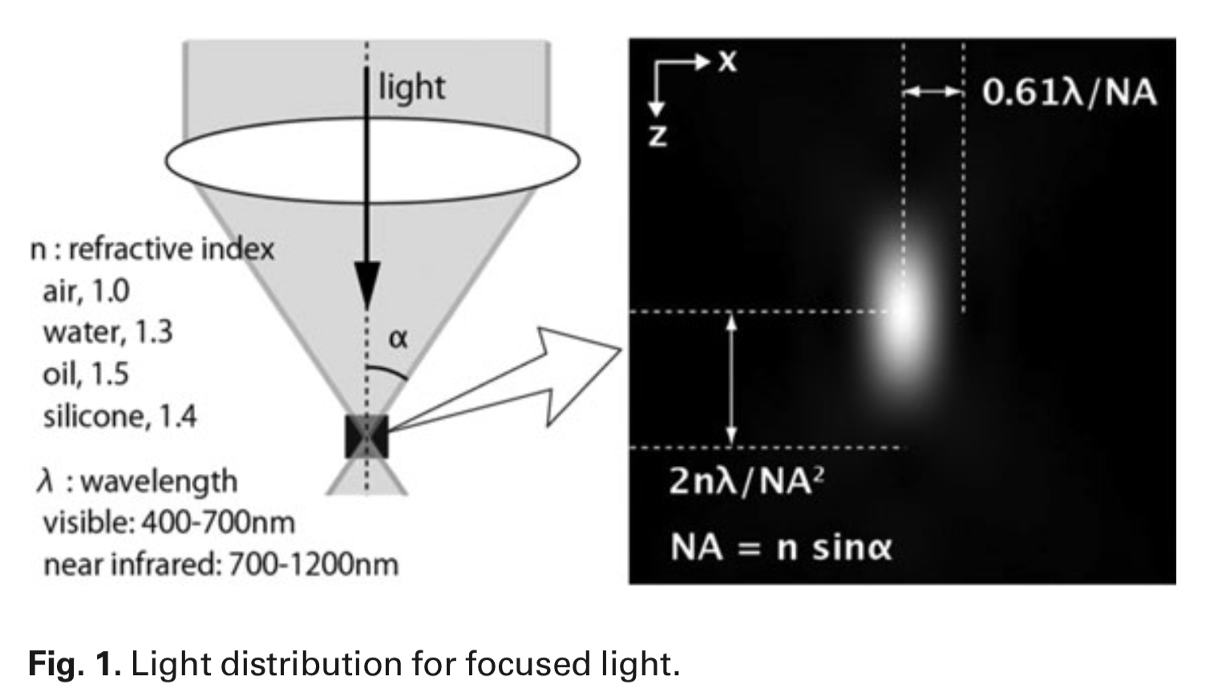
Point spread function and what is it dependent on
limit of resolution
forms a spread-out image
with the point source being the centre of the PSF
What dependent on:
refractive index
wavelength
numerical aperture of objective
optical properties
Two types of Super-resolution microscopy
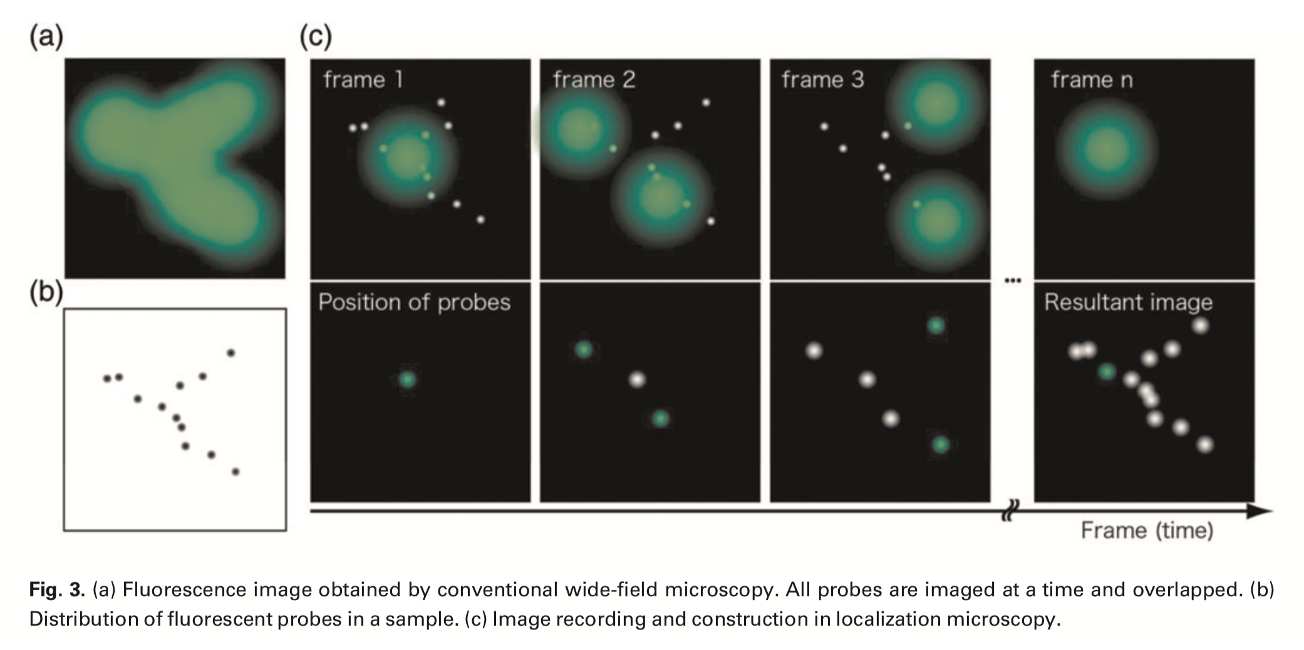
Single-molecule localisation microscopy (SMLM)
Stimulated depletion Emission Microscopy (STED)
What is the basis of SMLM
Activate one molecule at a time
With photo-activatable or photo-switchable dyes/proteins
ativated randomly at low efficiency by short pulse of a specific wavelength
produces a single molecule pixel
Record where PSF of this single molecule is
Photoswitch it off by a different wavelength
Process repeated (and a different pixel will be activated)
The middle of the pixel must be where the actual image is
repeat many times to get idea of all of the pixels i nthe image
resolution of this method
resolve a few 10s of nm
Limitations of this method
number of rounds of photoswitching
hard to apply repeats to live imaging
takes time
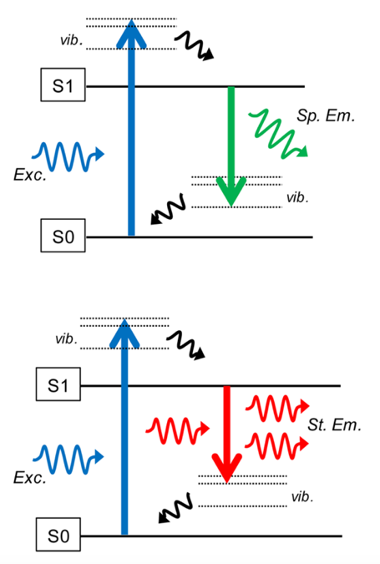
STED: stimulated depletion Emission Microscopy: concept
Use a doughnut-shaped ring of high-intsensity far-red light
this suppresses fluorescene of the outer parts
increases the resolution
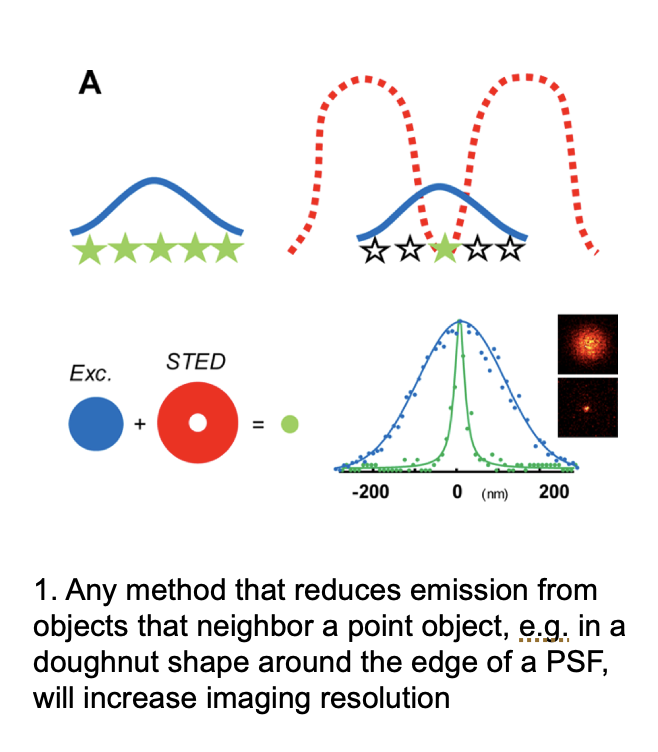
How does this red ring work to for this?
Normal fluoresecence:
return of an electron to S0 ground state gives normal fluorescence
With red ring
smaller energy loss for falling electrons
longer-waventlgeth is outputted
which is not detected (not visible light)
Pros and cons
Pros:
does not need repeated round
resolution→ 50nm
increasing power of doughnut can decrease this further
Cons/limits and solution
limit to how far power can increase without bleaching fluorphores
solution: use chemically modified proteins whose fluorescence survives better than GFP
Uses a lot of energy and heat??
Other types of super-resolution approaches
Structured Illumintion Microscopy
gentler on live preparations than STED
faster than SMLM
comutationally complex
depth limited to few um
How is works:
contrast raw images (below the diffraction limit) from Moire patterns *above the diffraction limit)
Take single image in different oritentations to get more and more info of what the original pattern was with computational analysis
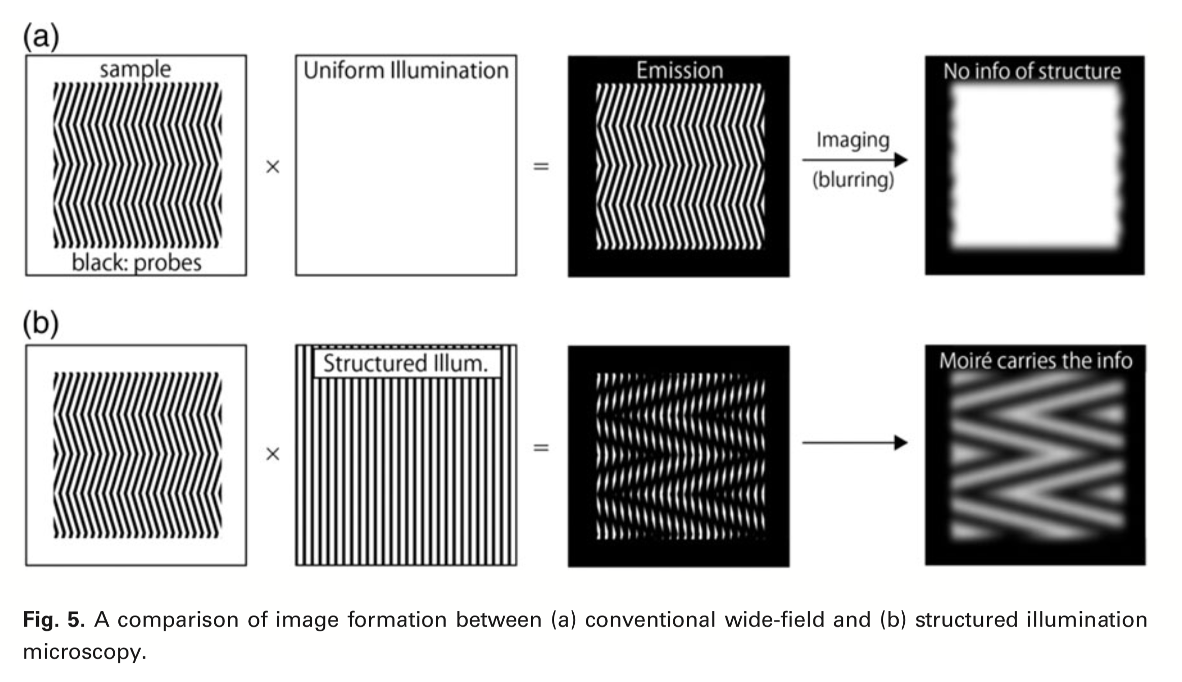
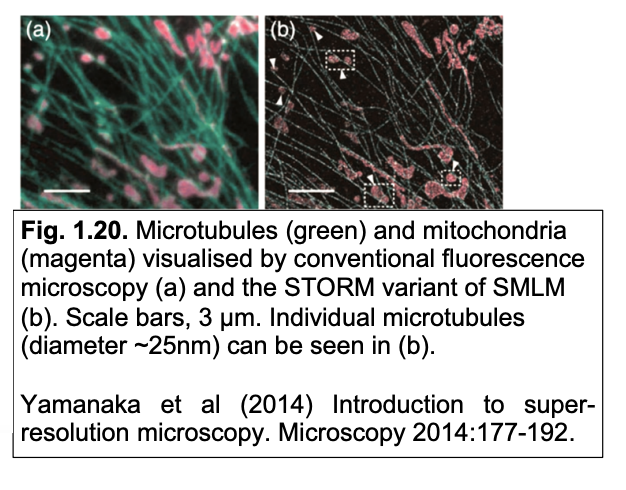
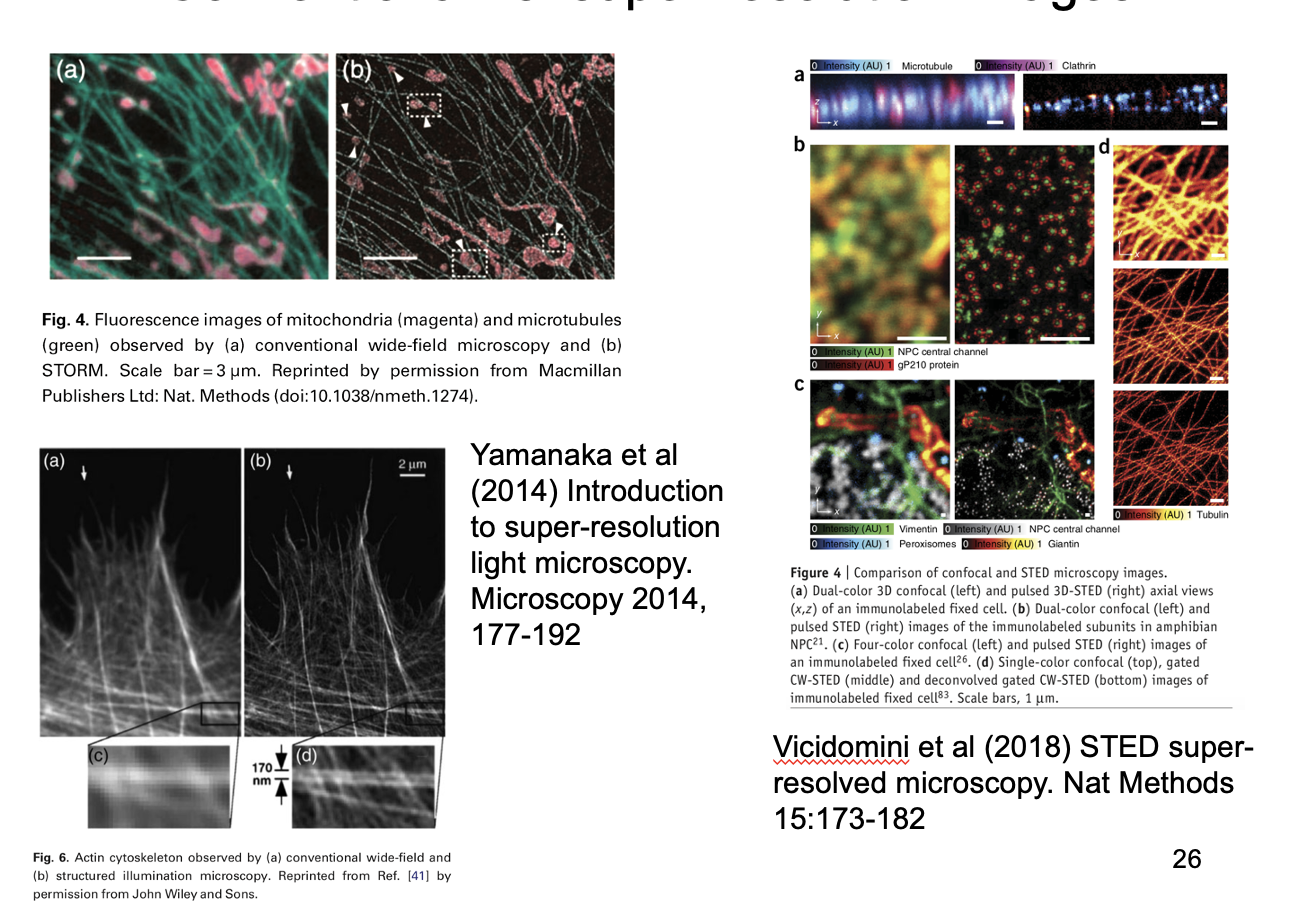
Results: Conventional vs super-resolution images
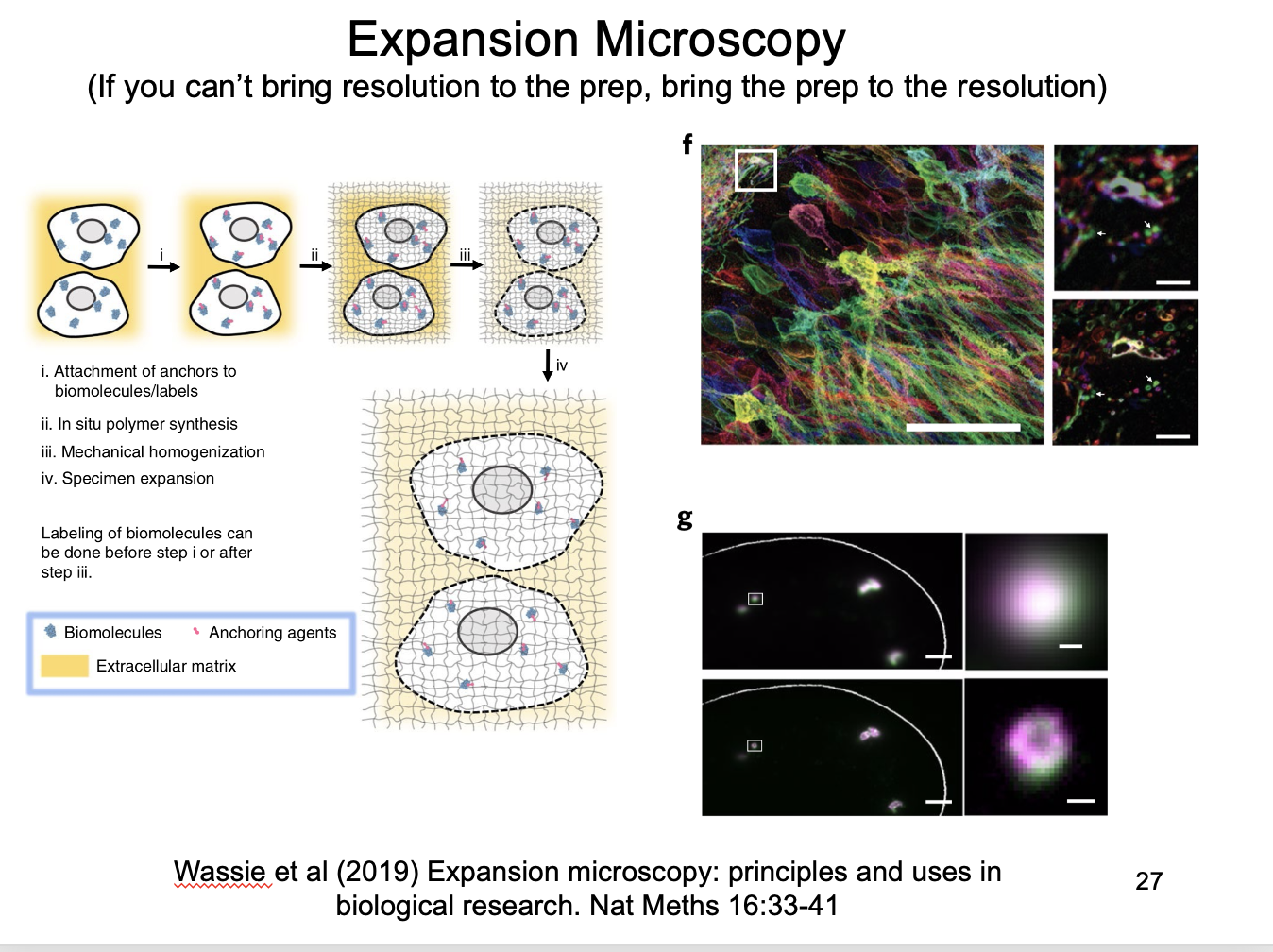
Instead of super-resoluton: Expansion microscopy
If you cannot bring resolution to prep, bring the prep to the resolution
impregnate preparation with a gel that expands on hydration
with light protease treatment
to allow uniform expansion of cell components
More microscoptic methods for viewing organelles
Light sheet
Lattice light-sheet
why are these methods used instead of confocal sometimes?
decreases the photodamage from confocal
by using a single plane of light
in contrast→ confocal illuminates the whole plane
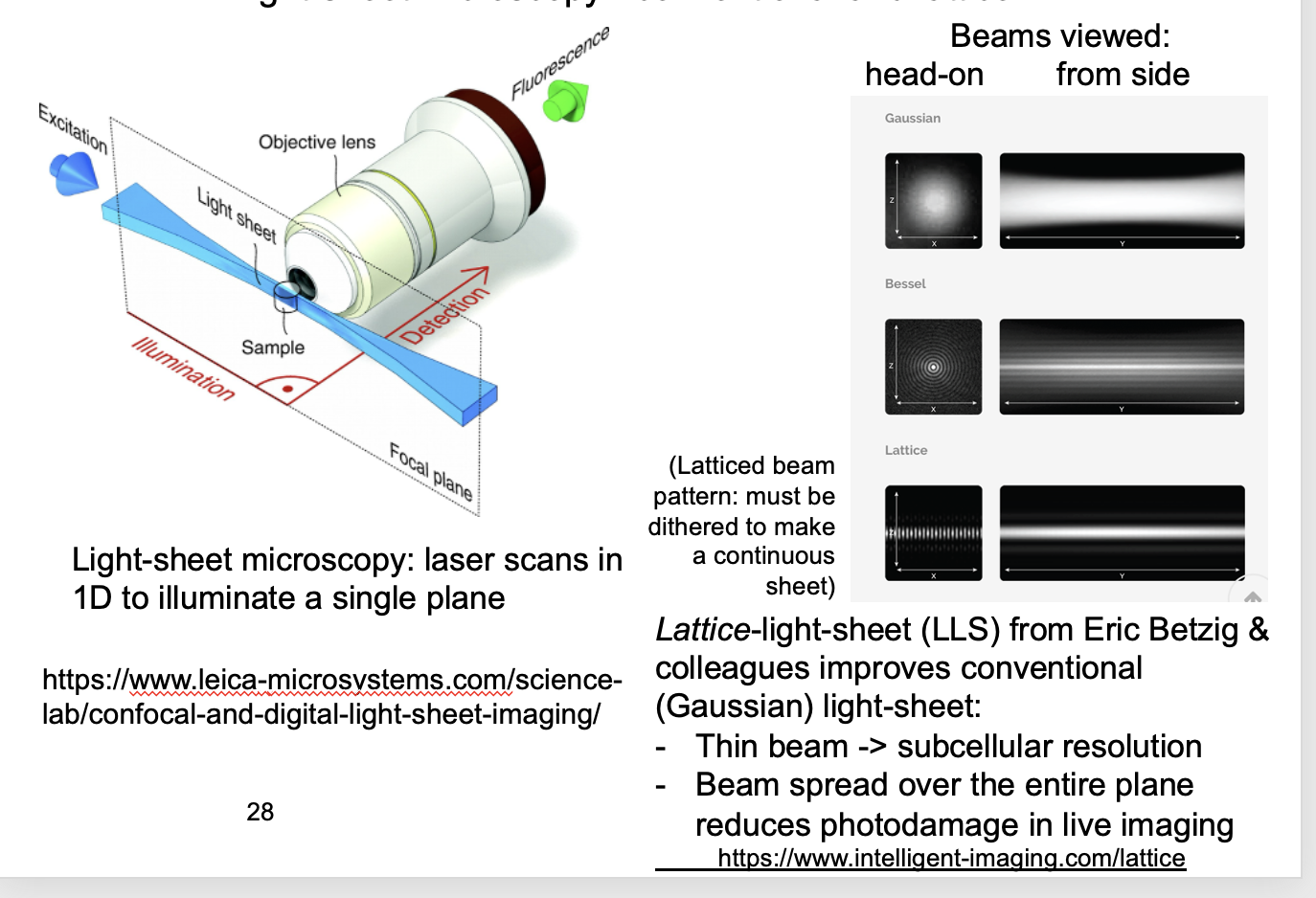
Light sheet
sample if illuminated by sheet of light perpendicular to the imaging objective
(rather than via)
good alternative to confocal microscopy for imaging single planes
SPIM→ Single-plane illumination microscopy
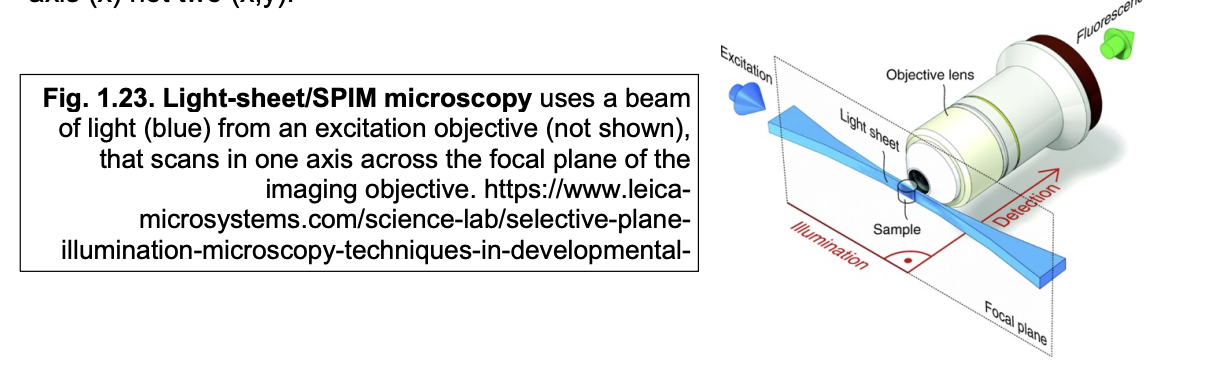
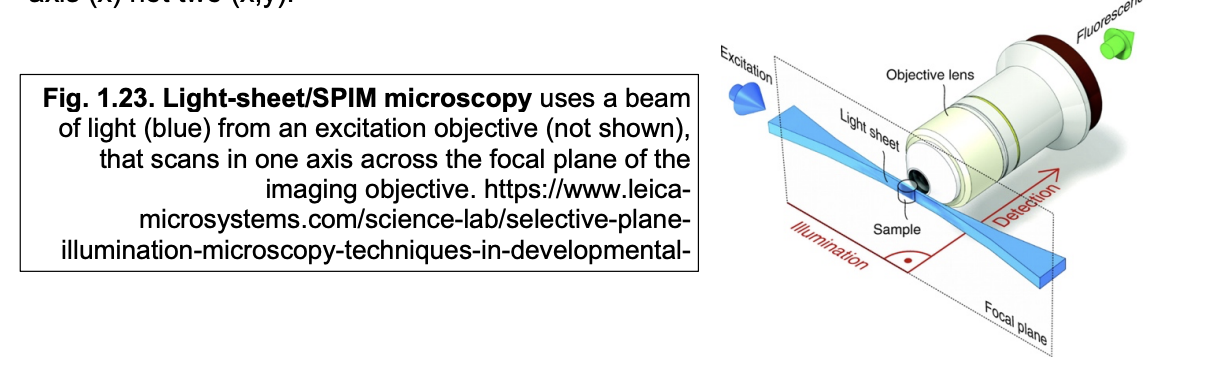
Advantages of SPIM
reduces photodamage by confining illumination to the focal place
much faster than confocal
since laser only has to scan in one axis
x (not x and y)
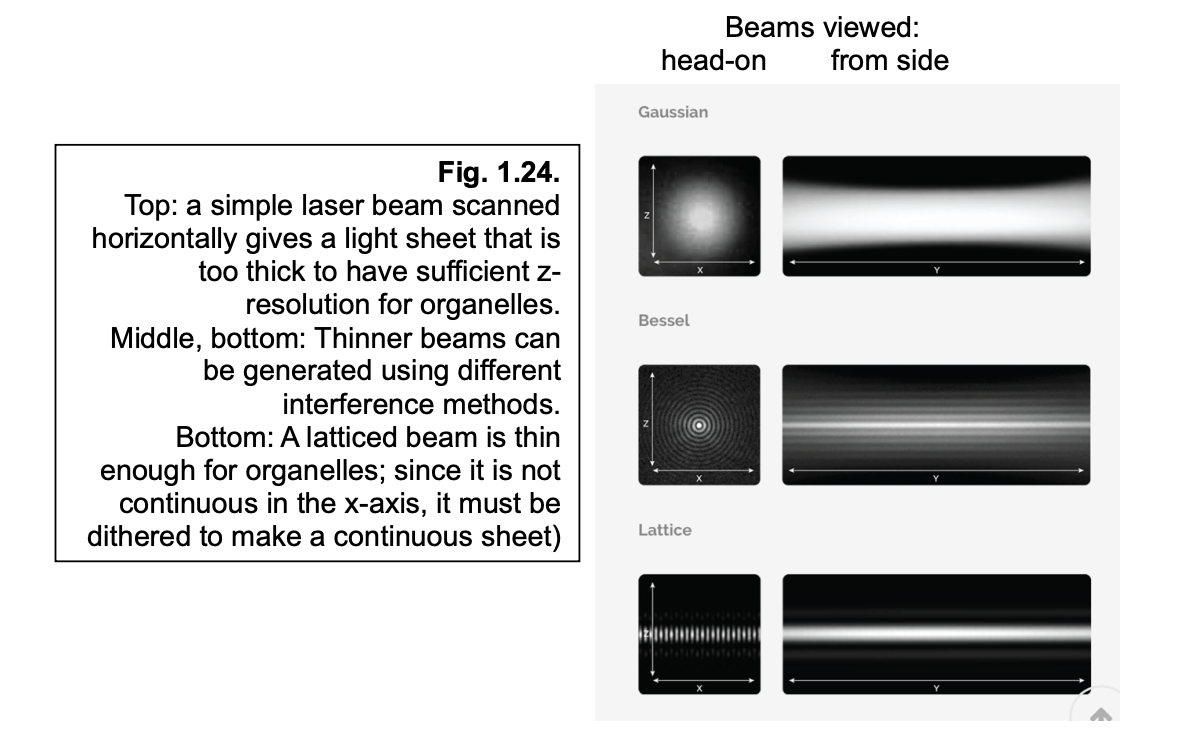
Lattice light sheet
uses multiple beams (lattice)
to spread a thin light-sheet out across the entire imaging place
since it is not continuous in the x-axis→ it must be dithered to make a continuous sheet
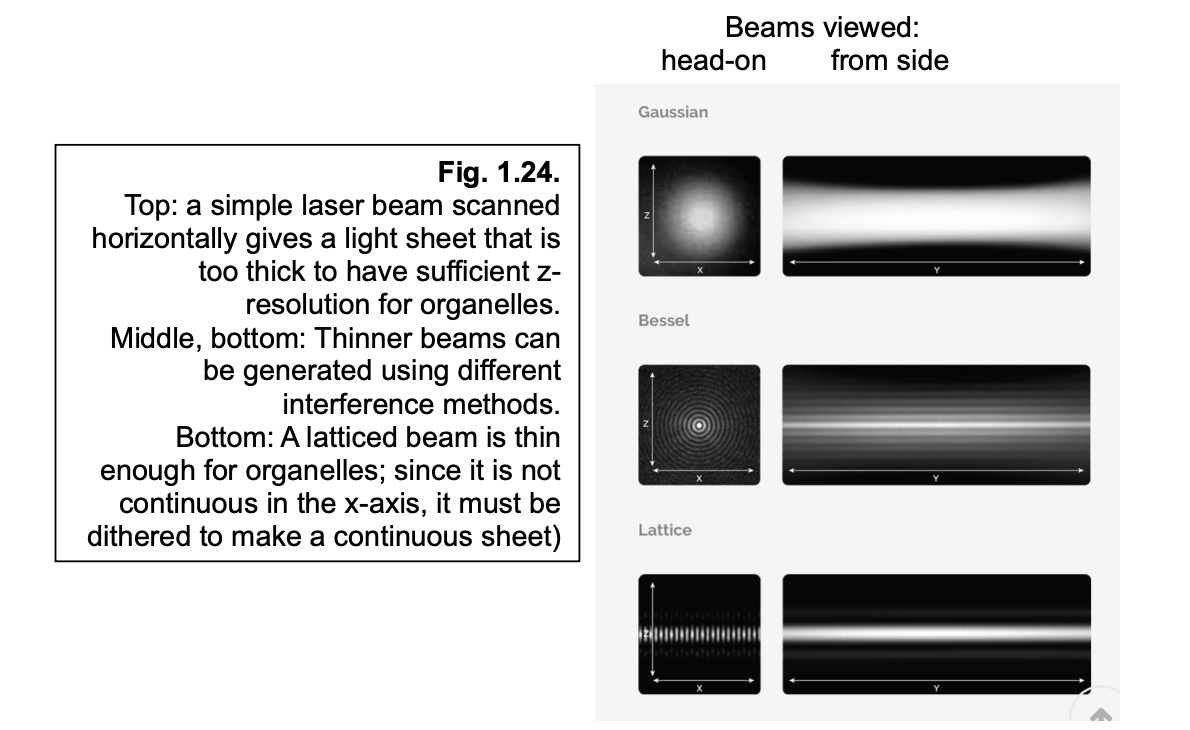

Pros
further gains in speed and phototoxicity

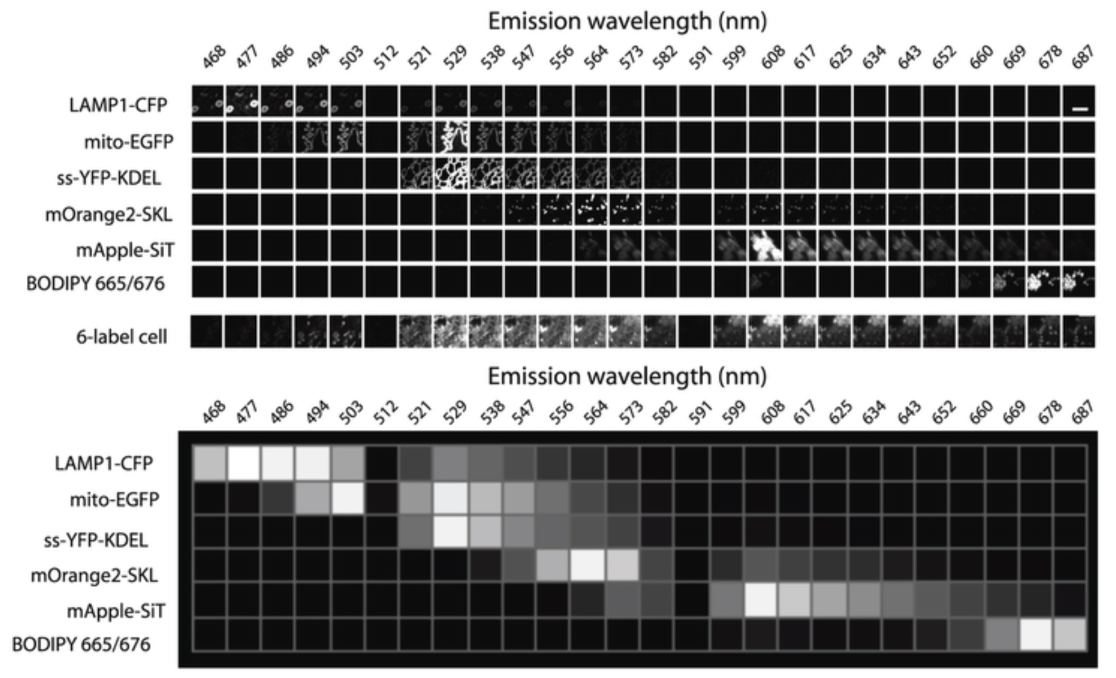
How can multiple organelles be live imaged: why can’t just use 6 different markers for 6 organelles?
Requires 6 different fluorescent protein tags
however:
with so many tags→ emission spectra cannot be separated using silter sets alone
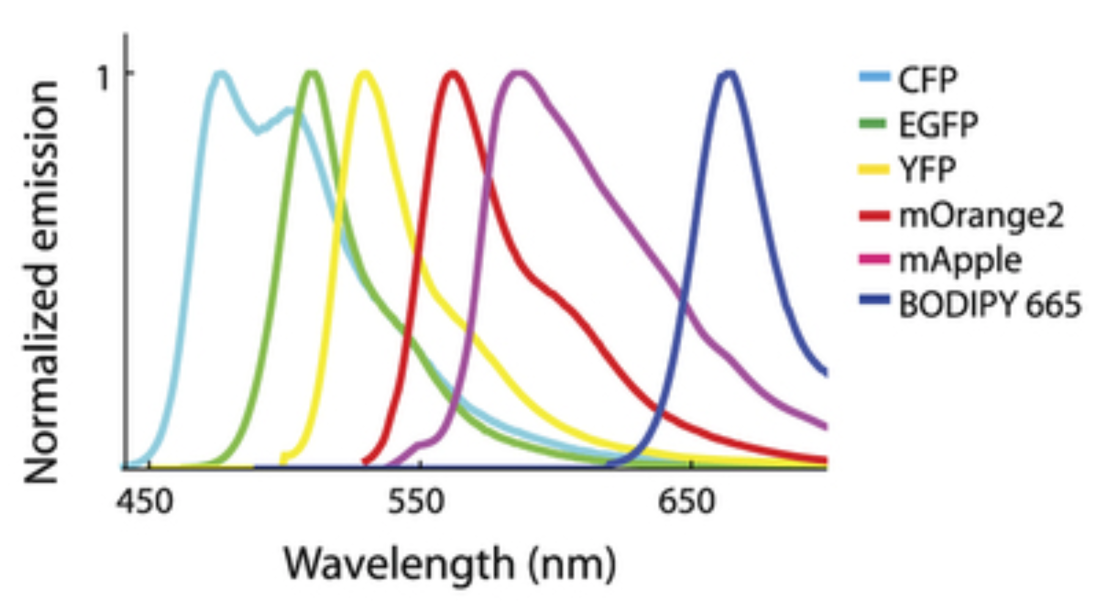
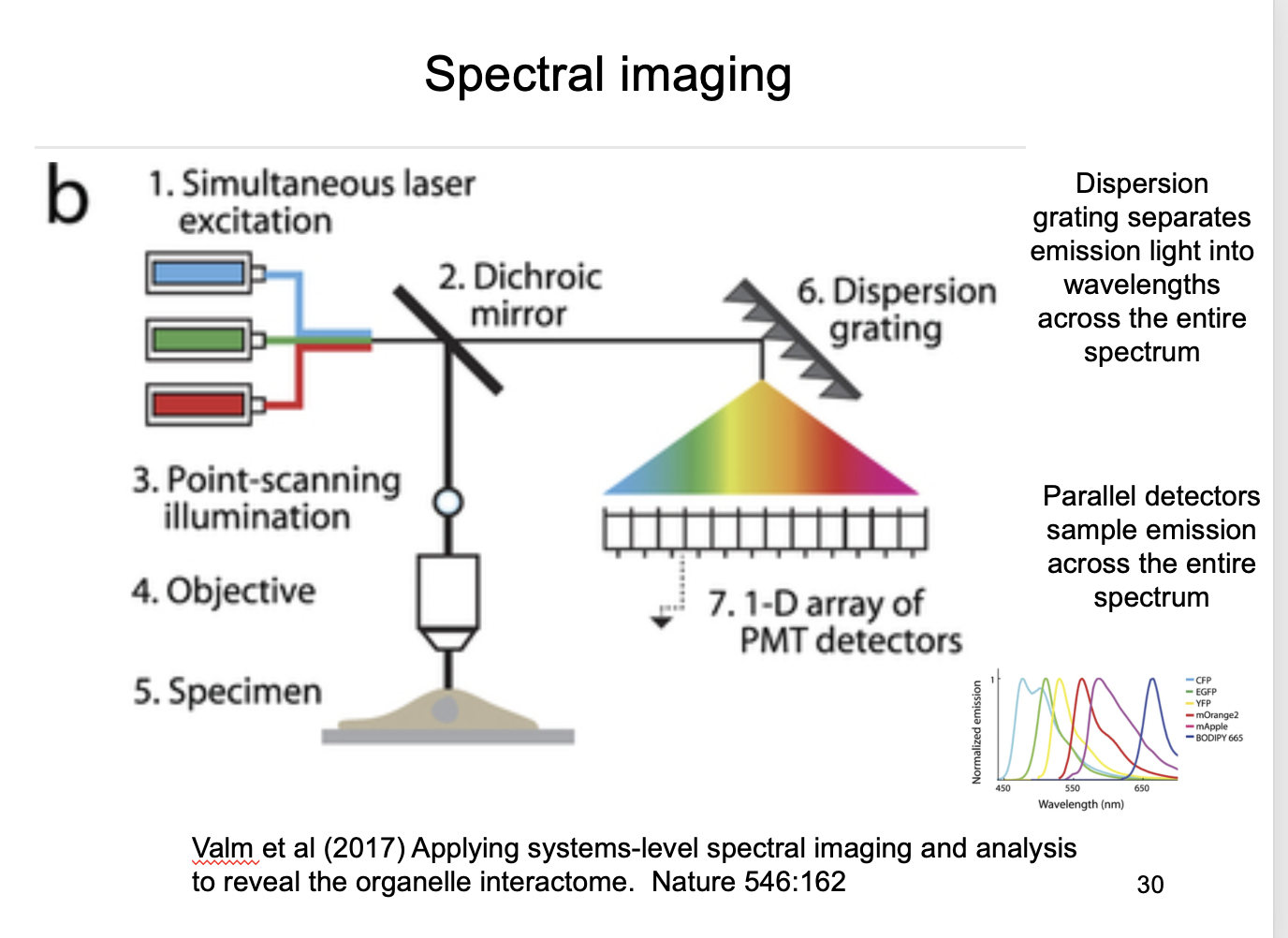
Solution to this: Spectral imaging
Dispersion grating separates emission light into wavelentghs across the entire spectrum
parallel detectors ample emission across the entire spectrum
Code of colour sorresponds to different organelles from 6 fulorophores
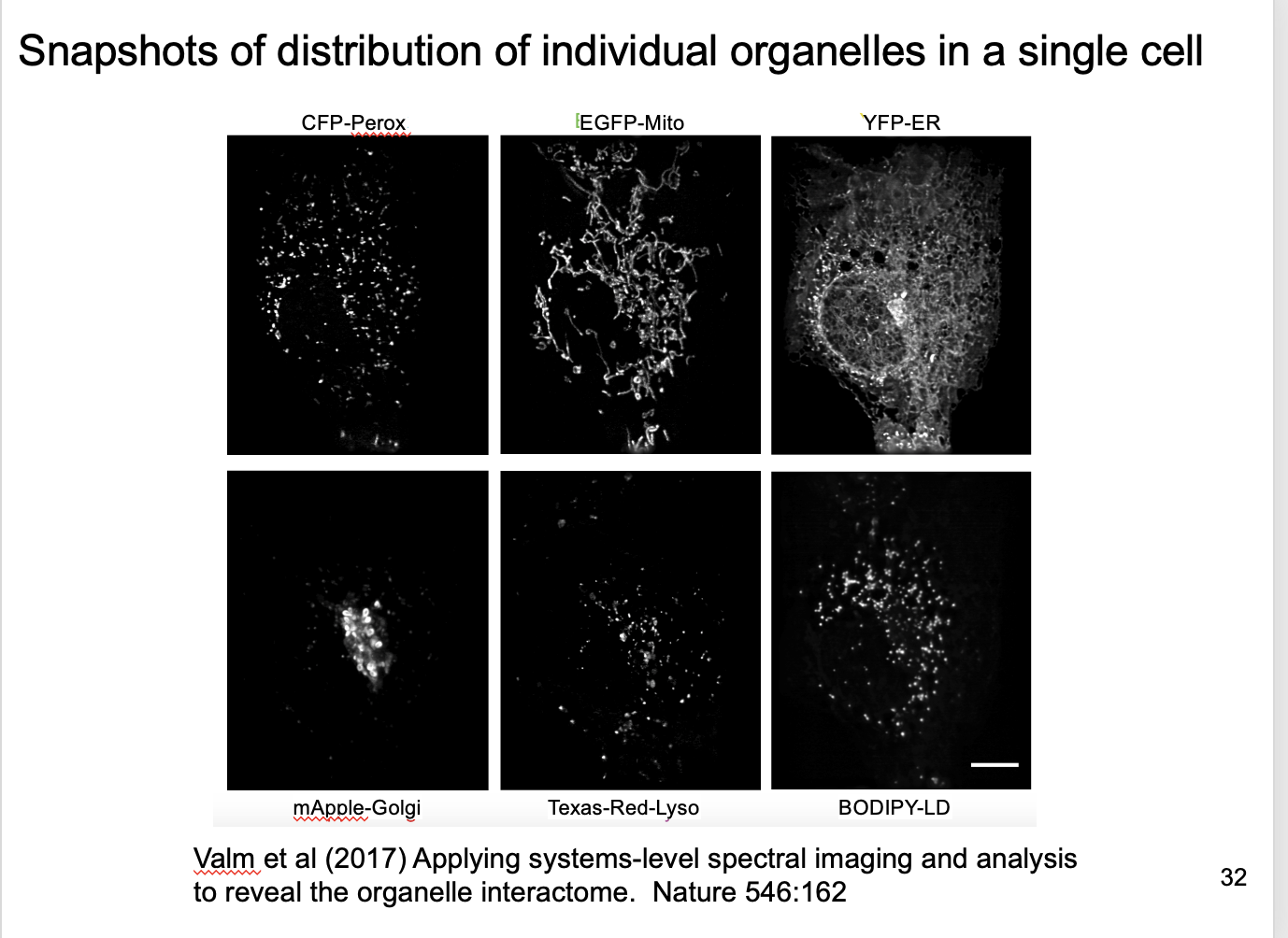
How to decode this code for the organelles (Spectral unmixing)
Each channel detects mix of signals from 6 fluorophores
here 26 simulatenous equations to find the 6 unknowns