MCAT Biochemistry (Kaplan)
1/533
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
534 Terms
Amino acids have four groups attached to a central (α) carbon:
an amino group, a carboxylic acid group, a hydrogen atom and a R group
The R group determines
chemistry and function of that amino acid.
Twenty amino acids appear in the proteins of eukaryotic organisms
The stereochemistry of the α-carbon is`
1 for all chiral amino acids in Eurkaryotes
D-amino acids can exist in
prokaryotes
All chiral amino acids expect______ have an (S) configuration
Cysteine
All amino acids are chiral expect
glycine, which has a hydrogen atom as its R group
Side chains can be
polar or nonpolar, aromatic or nonaromatic, charged or uncharged
Nonpolar nonaromatic amino acids
glycine, alanine, valine, leucine, isoleucine, methionine, proline
Aromatic
Tryptophan, phenylalanine, tyrosine
Polar
Serine, threonine, asparagine, glutamine, cysteine
Negatively charged (acidic)
aspartate, glutamate
Positively charged (basic)
Lysine, arginine, histidine
Amino acids with _____alkyl chains are ______ and those with charges are _____; many others fall somewhere in between
long; hydrophobic; hydrophilic
Amino acids are
amphoteric; that is, they can accept or donate protons
The pKa of a group is the
pH at which half of the species are deprotonated; [HA] = [A-]
At low (acidic) pH, the amino acid is
fully protonated
At pH near the pI of the amino acid, the amino acide is
a neutral zwitterion
At high (alkaline) pH, the amino acid
is fully deprotonated
The titration curve is nearly _____ at the pKa values of the amino acid
flat
The titration curve is nearly_______ vertical at the pI of the amino acid
Vertical
Amino acids with charged side chains have an additional pKa value, and their pI is calculated
by averaging the two pKa values that correspond to protonation and deprotonation of the zwitterion
Amino acids without charged side chains have a pI around
6
Acidic amino acids have a pI
well below 6
Basic amino acids have a pI
well above 6
Dipeptides
have two amino acid residues
tripeptides
have three amino acid residues
Oligopeptides
have a “few” amino acid residues (<20)
Polypeptides
have “many” (>20)
Forming a peptide bond is a
condensation or dehydration reaction (releasing one molecule of water)
The nucleophilic amino group of one amino acid attacks the
electrophilic carbonyl group of another amino acid
Amide bonds are
rigid because of resonance
Breaking a peptide bone is a
Hydrolysis reaction
Primary structure
is the linear sequence of amino acids in a peptide and is stabilized by peptide bonds
Secondary structure
is the the local structure of neighboring amino acids, and is stabilized by hydrogen bonding between amino groups and nonadjacent carboxyl groups
α-helices
are clockwise coils around a central axis
β-pleated sheets
are rippled strands that can be parallel or antiparallel
Proline can interrupt
secondary structure because of its rigid cyclic structure
Tertiary structure
is the three-dimensional shape of a single polypeptide chain, and is stabilized by hydrophobic interactions, acid-baser interactions (salt-bridges), hydrogen bonding, and disulfide bonds
Hydrophobic interactions
push hydrophobic R groups to the interior of a protein, which increases entropy of the surrounding water molecules and creates a negative Gibbs free energy
Disulfide bonds
occur when two cysteine molecules are oxidized and create a covalent bond to form cysteine
Quaternary structure
is the interaction between peptides in proteins that contain multiple subunits
Proteins with covalently attached molecules are termed
conjugated proteins. The attached molecule is a prosthetic group and may be a metal ion, vitamin, lipid, carbohydrate, or nucleic acid
Denaturation
Both heat and increasing solute concentration can lead to loss of three-dimensional protein structure
Heat denatures proteins by increasing their average
kinetic energy, this disrupting hydrophobic interactions
Solutes denature proteins by
disrupting elements of secondary, tertiary, and quaternary structure
Isoelectric point of a neutral amino acid
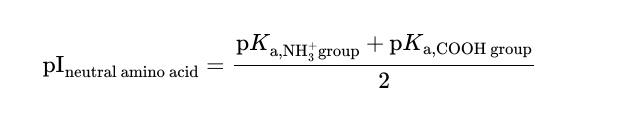
Isoelectric point of an acidic amino acid
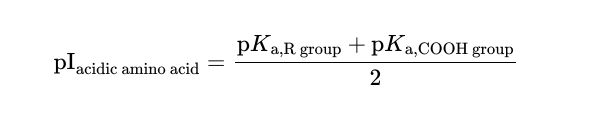
Isoelectric point of a basic amino acid
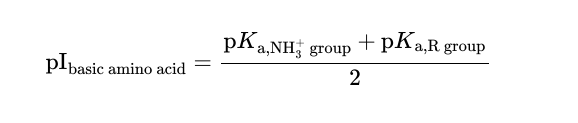
Enzymes
are biological catalysts that are unchanged by the reactions they catalyze and are reusable
Each enzyme catalyzes a
single reaction or type of reaction with high specificity
Oxidoreductases
catalyze oxidation-reduction reactions that involve the transfer of electrons
Transferases
move a functional group from one molecule to another molecule
Hydrolases
catalyze cleavage with the addition of water
Lyases
catalyze cleavage without the addition of water and without the transfer of electrons. The reverse reaction (synthesis) is often more important biologically
Isomerases
catalyze the interconversion of isomers, including both constitutional isomers and stereoisomers
Ligases
are responsible for joining two large biomolecules, often of the same type
Exergonic reactions
release energy; ΔG is negative
Enzymes lower the
activation energy necessary for biological reactions
Enzymes do not alter the
free energy (ΔG) or enthalpy (ΔH) change that accompanies the reaction nor the final equilibrium position; rather, they change the rate (kinetics) at which equilibrium is reached
Enzymes act by stabilizing
the transition state, providing a favorable microenvironment, or bonding with the substrate molecules
Enzymes have an
active site, which is the site of catalysis
Binding to the active site is explained by the
lock and key theory of the induced fit model
The lock and key theory hypothesizes that the
enzyme and substrate are exactly complementary
The induced fit model hypothesizes that the
enzyme and substrate undergo conformational changes to interact fully.
Some enzymes require
metal cation cofactors or small organic coenzymes to be active
Enzymes experience saturation kinetics
as substrate concentration increases, the reaction rate does as well until a maximum value is reached
Michaelis-Menten and Lineweaver-Burk plots represent
this relationship as a hyperbola and line respectively
Enzymes can be compared on a basis of their
Km and vmax values
Cooperative enzymes display a
sigmoidal curve because of the change in activity with substrate binding
Temperature and pH affect an enzyme’s activity
in vivo
changes in temperature and pH can result in
denaturing of the enzyme and loss of activity due to loss of secondary, tertiary, or, if present, quaternary structure
In vitro
salinity can impact the action of enzymes
Enzyme pathways are highly
regulated and subject to inhibition and activation
Feedback inhibition
is a regulatory mechanism where by the catalytic activity of an enzyme is inhibited by the presence of high levels of a product later in the same pathway
Reversible inhibition
is characterizes by the ability to replace the inhibitor with a compound of greater affinity or to remove it using mild laboratory treatment
Competitive inhibition
results when the inhibitor is similar to the substrate and binds at the active site.
Competitive inhibition can be overcome by adding more substrate. Vmax is unchanged, Km increases.
Noncompetitive inhibition
results when the inhibitor binds with equal affinity to the enzyme and the enzyme–substrate complex. vmax is decreased, Km is unchanged.
Uncompetitive inhibition
results when the inhibitor binds only with the enzyme-substrate complex. Km and Vmax both decreases
Mixed inhibition
results when the inhibitor binds with unequal affinity to the enzyme and the enzyme-substrate complex. Vmax is decreased, Km is increased or decreased depending on if the inhibitor has higher affinity for the enzyme or enzyme-substrate complex
Irreversible inhibition
alters the enzyme in such a way that the active site is unavailable for a prolonged duration or permanently; new enzyme molecules must by synthesized for the reaction to occur again
Regulatory enzymes can experience
activation as well as inhibition
Allosteric sites
can occupied by activators, which increase either affinity or enzymatic turnover
Phosphorylation (covalent modification with phosphate) or glycosylation (covalent modification with carbohydrate)
can alter the activity or selectivity of enzymes
Zymogens
are secreted in an inactive form and are activated by cleavage
Michaelis-Menten rates
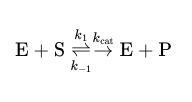
Michaelis-Menten equation
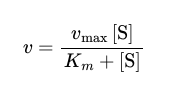
Turnover number (Kcat)
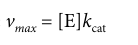
Structural proteins
compose the cytoskeleton, anchoring proteins, and much of the extracellular matrix
The most common structural proteins are
collagen, elastin, keratin, actin, and tubulin
They are generally fibrous in natur
Motor proteins
have one or more heads capable of force generation through a conformational change.
They have catalytic activity, acting as ATPases to power movement.
Muscle contraction, vesicle movement within cells, and cell motility are the
most common applications of motor proteins.
Common examples include myosin, kinesin, and dynein
Binding proteins
binds a specific substrate, either to sequester it in the body of hold its concentration at steady state
Cell adhesion molecules (CAM)
allow cells to bind to other cells or surfaces
Cadherins
are calcium-dependent glycoproteins that hold similar cells together
Integrins
have two membrane-spanning chains and permit cells to adhere to proteins in the extracellular matrix. Some also have signaling capabilities
Selectins
allow cells to adhere to carbohydrates on the surfaces of other cells and are commonly used in the immune system
Antibodies (or immunoglobulins,Ig)
are used by the immune system to target a specific antigen, which may be a protein on the surface of a pathogen (invading organism) or a toxin
Immunoglobulin contain
a constant region and a variable region; the variable region is responsible for antigen binding
Two identical heavy chains and two identical light chains form a
single antibody; they are held together by disulfide linkages and noncovalent interactions
Ion channels
can be used for regulating ion flow into or out of a cell. There are three main types of ion channels