Double-Stranded DNA Break Repair Mechanisms
1/19
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
20 Terms
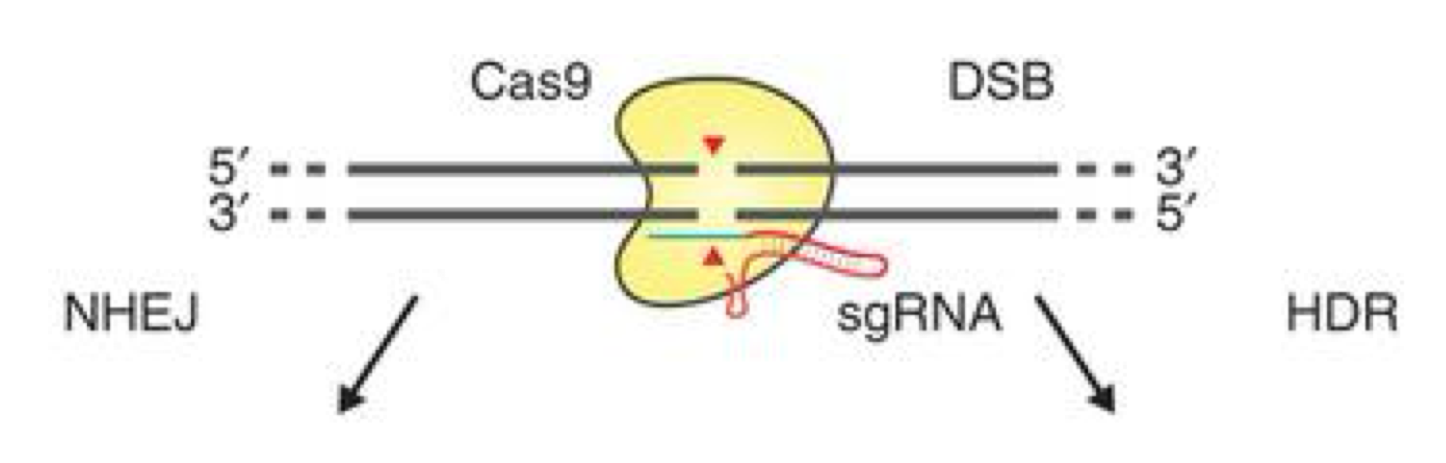
Nuclease-induced genome editing
1) use a guide RNA (sgRNA) to direct Cas9 to make a specific double-stranded DNA break within target sequence
2) when the cell detects the DSB, it will trigger DNA repair mechanism:
non-homologous end joining (NHEJ): sticks broken ends back together
homology-directed repair (HDR): using the homologous chromosome as template to repair the break and fill in any sequences that may be lost
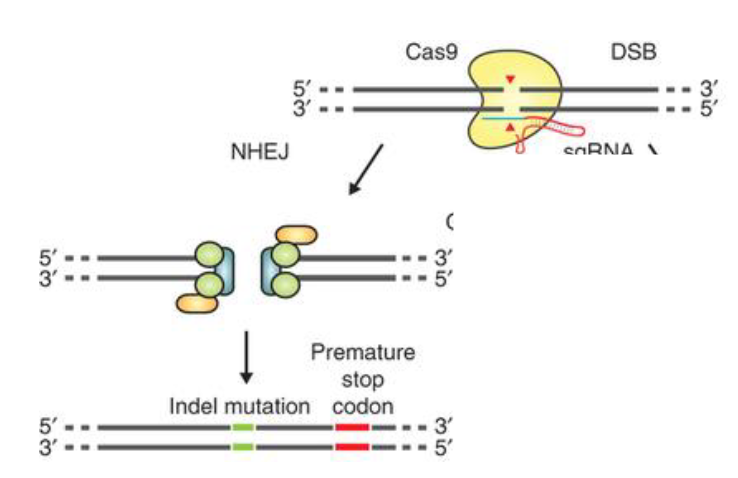
Non-homologous end joining (NHEJ)
results in small indels; can be used to create lof mutations (gene knockout)
proteins that recognize and bind to the broken ends
proteins that recruit and activate a nuclease, used to trim the broken ends
the DNA ligase that connects the broken ends together
NHEJ (flip)
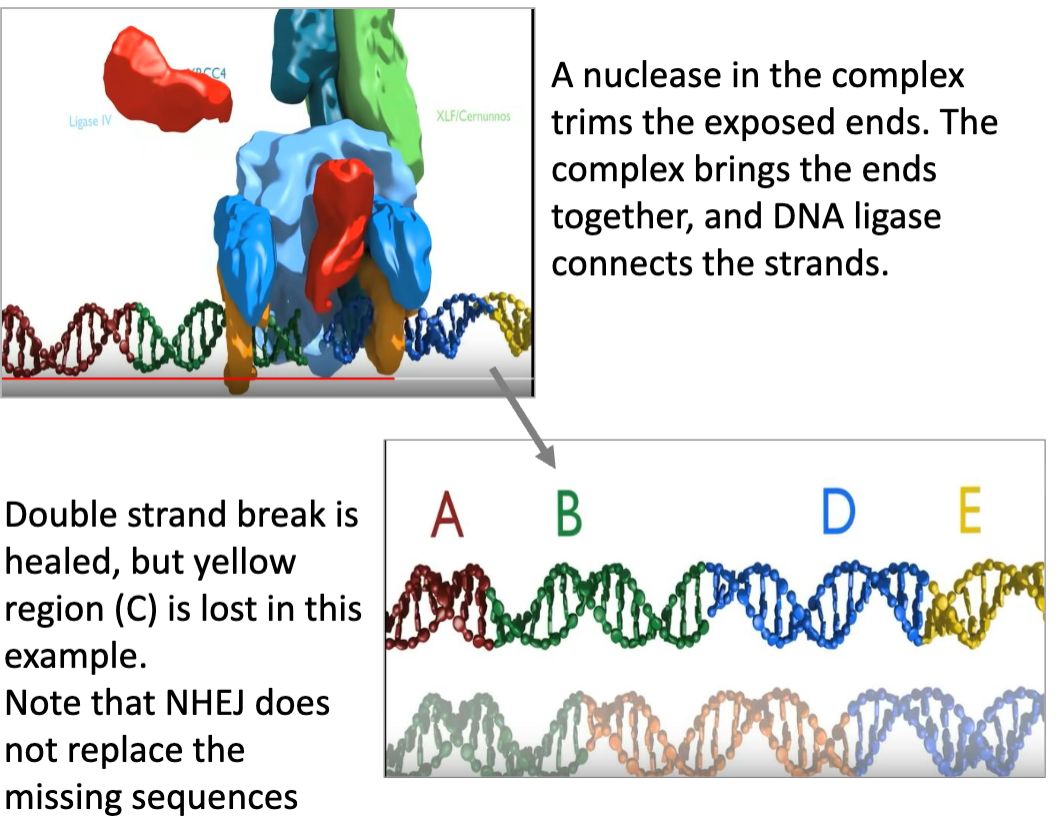
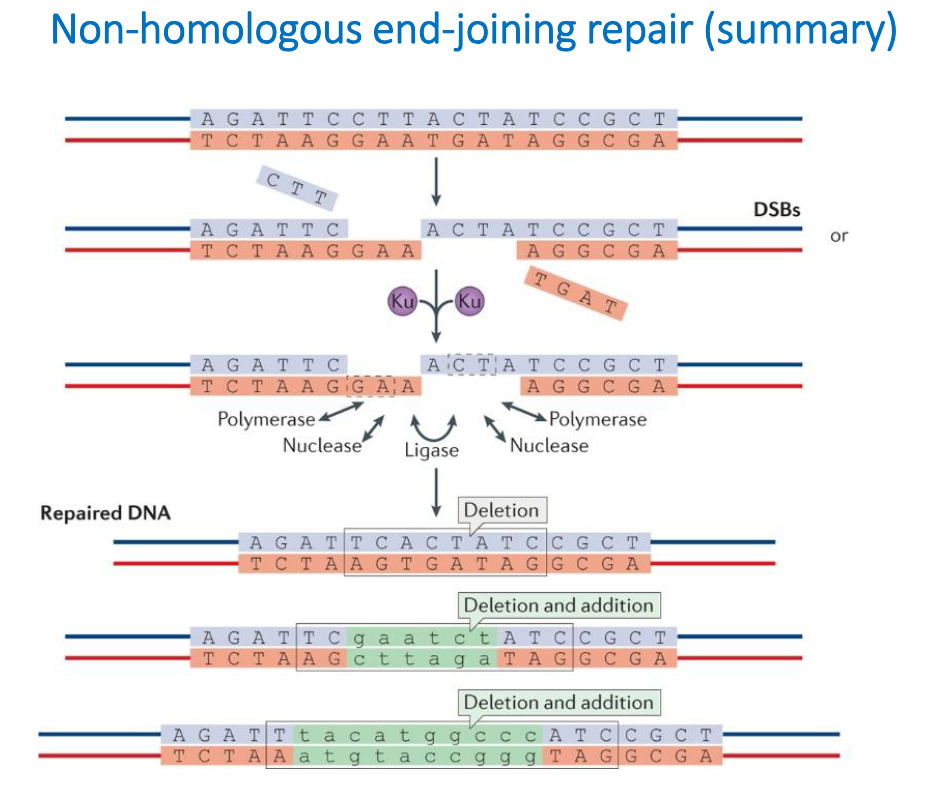
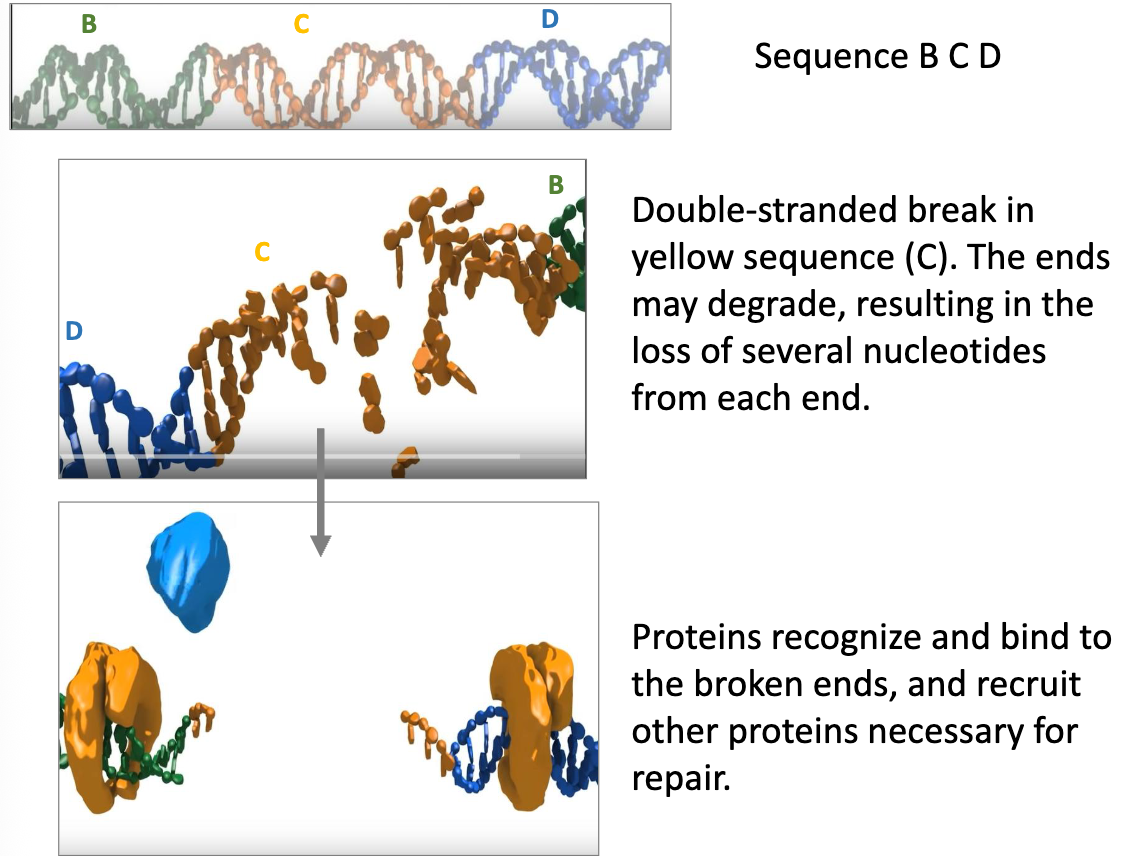
NHEJ Repair: Key Ideas
after a DNA double-strand break, there may be loss of nucleotides from either strand as the ends unwind and degrade
a complex of DNA binding protein kinases (the Ku proteins) recognize and bind to each broken end, recruiting a NHEJ polymerase, nuclease and ligases
this helps to connect the ends together and fill in any of the gaps
the repaired DNA may have deletions or insertions as a result of the repair
this form of repair can be used to direct loss of function mutations in target genes
Homology Directed Repair
nucleases trim back the DNA to leave long 3’ tails
recombination proteins bring one of the 3’ tails over to the homologous sequences of the other chromosome, and unwinds the other DNA so that the tail hybridize
DNA polymerase can now extend the 3’ tail, using the other chromosome as a template
once the newly synthesized DNA overlaps with the other 3’ tail, the repair complex dissociates and the two tails can hybridize
DNA polymerase can fill in the gaps, and DNA ligase can connect the backbones
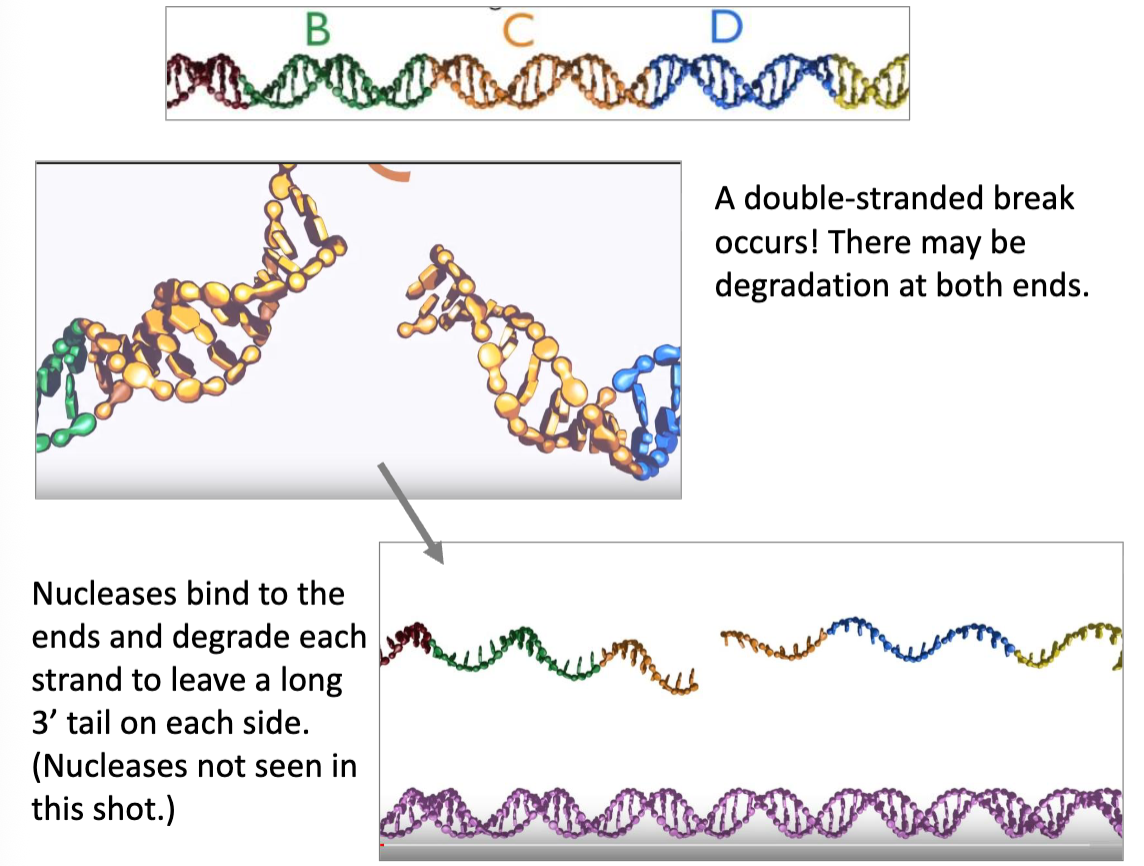
HDR (Flip)
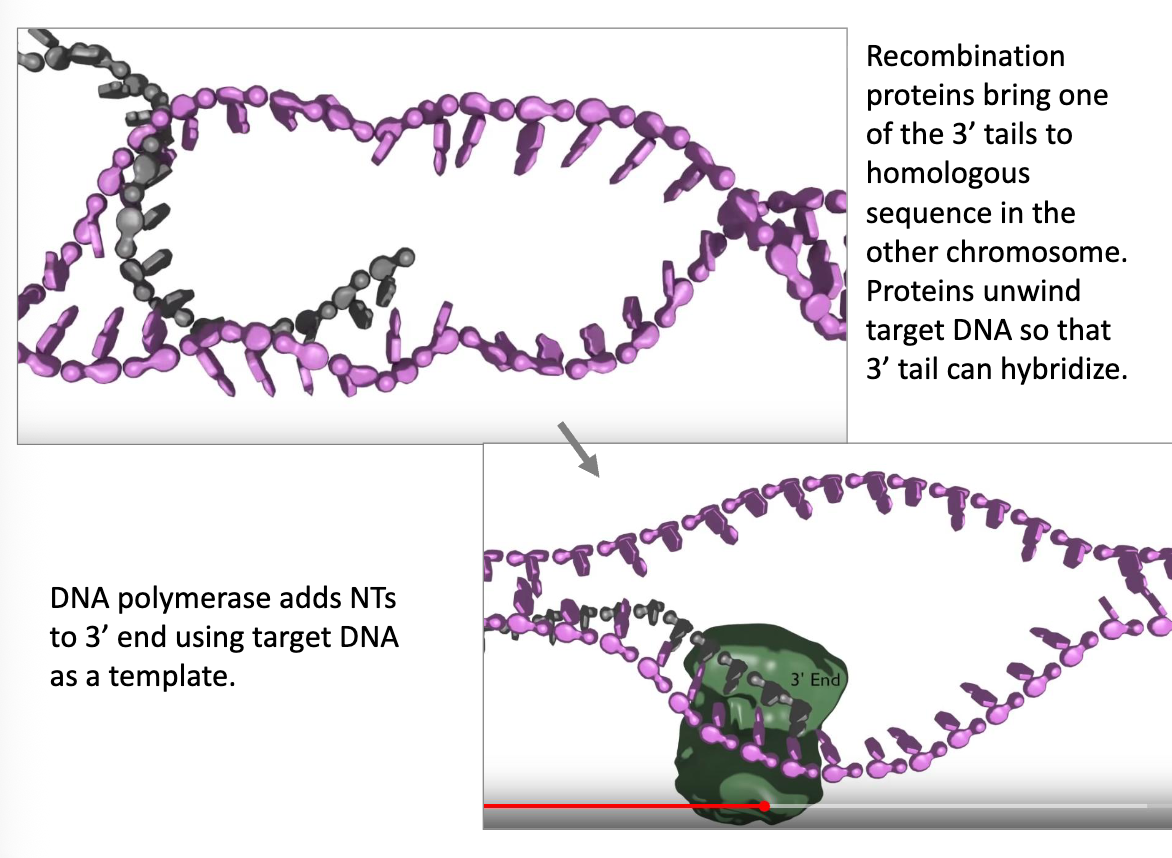
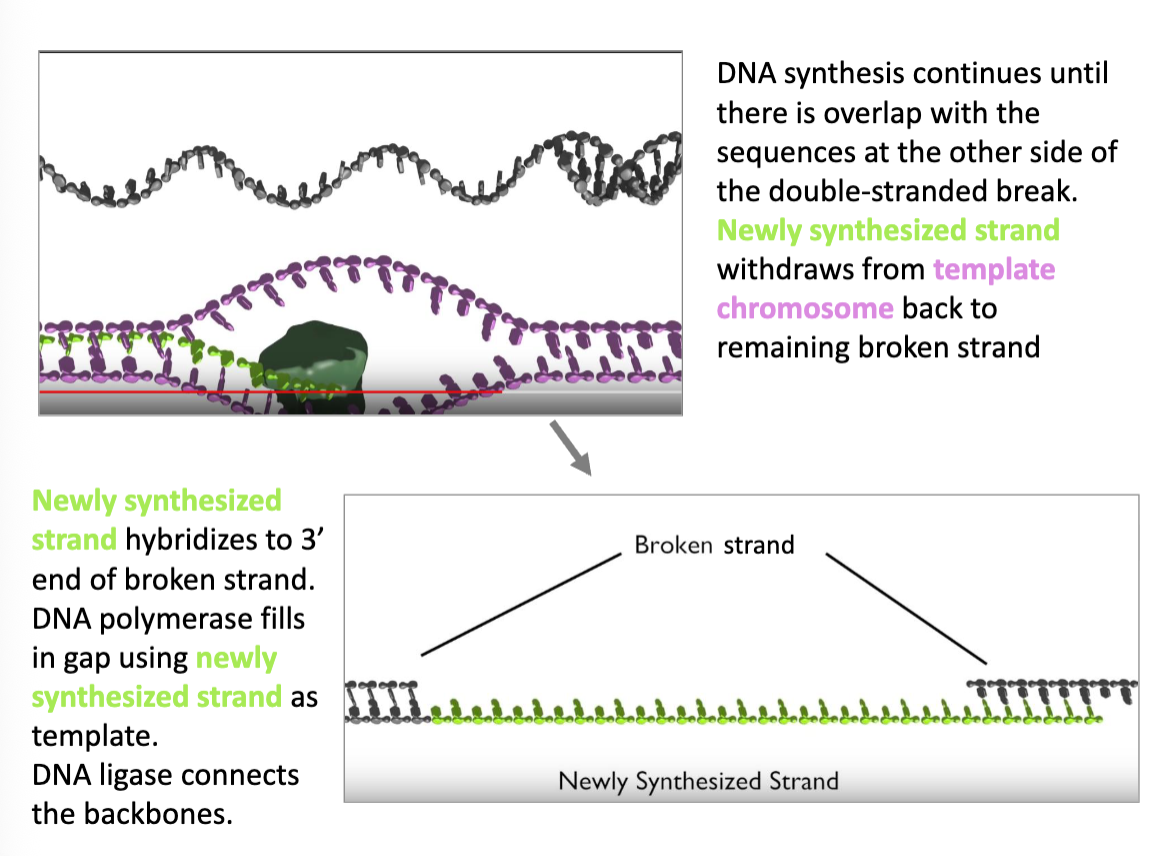
HDR Continued (flip)
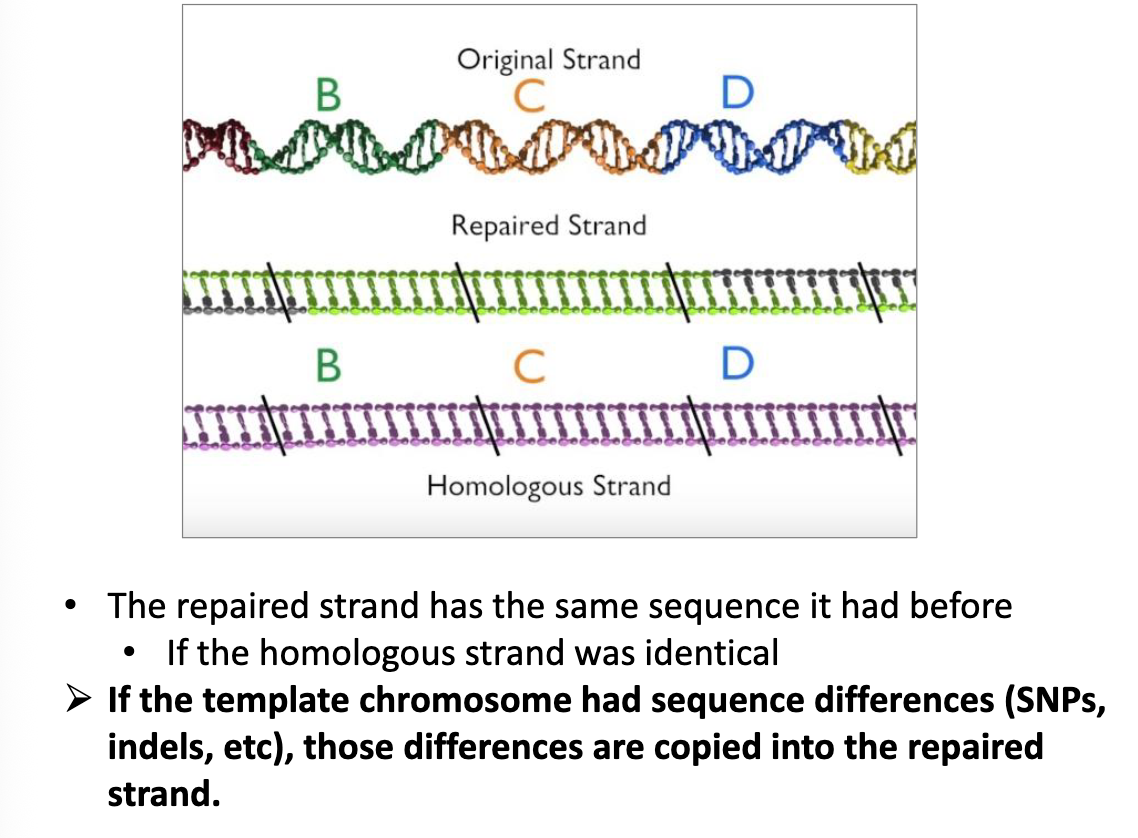
HDR: Key Ideas
after a DSB, nucleases bind to the ends and degrade one strand to leave a long 3’ tail
recombinase proteins allow the long 3’ ends to anneal to homologous sequences on the other chromosome. The homologous chromosome can then serve as the template as DNA polymerase further extends the 3’ tails
eventually the 3’ tails are long enough for the ends to have complementary sequences, and they can anneal to each other. DNA polymerase fills in the gaps, and DNA ligase connects the phosphate backbone
the repair would result in sequences that are copied from the template DNA (typically from the homologous chromosome)
if you were to introduce a different template sequence, the repair could result in sequences copied from your engineered DNA template
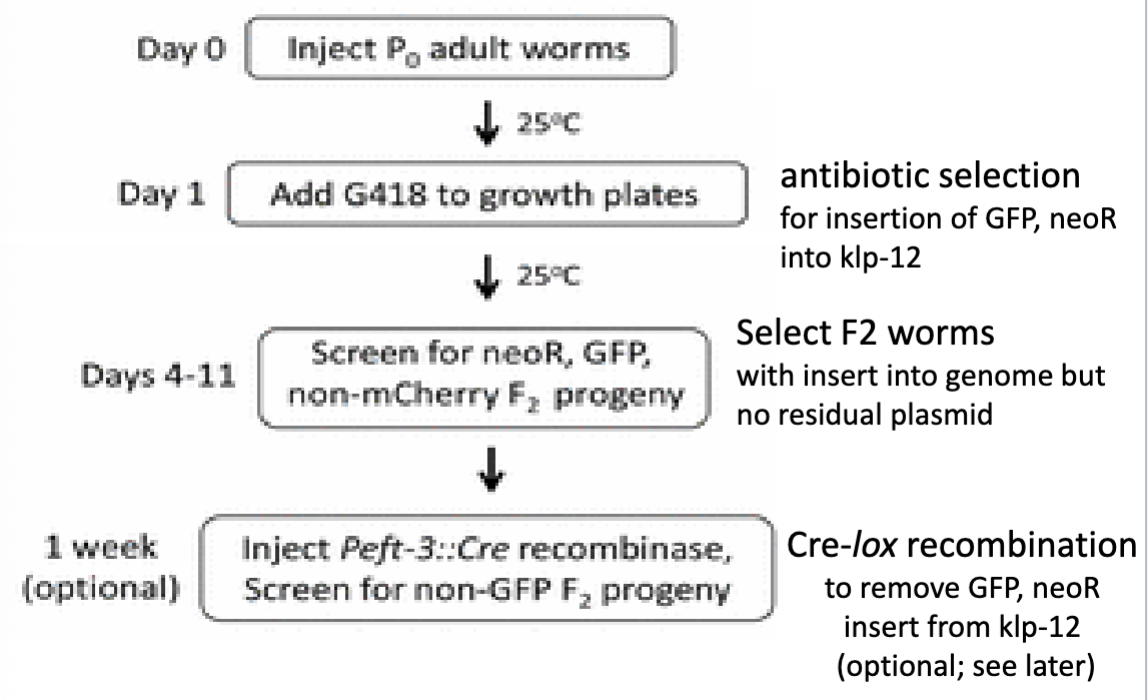
C. elegans Gene Knockout: Large-scale approach to knocking out C. elegans genes using CRISPR technology
Design stage: Identify good target sequences to use in the guide RNA
do the guide sequences have an optimal structure? (e.g. GC content, 3’ ends with GG, located next to PAM site within target”
are they unique? eliminate any sequences that have multiple targets
for KOs, generally best to target start of ORF
This became a database to help other researchers select guide RNAs for C. elegans research
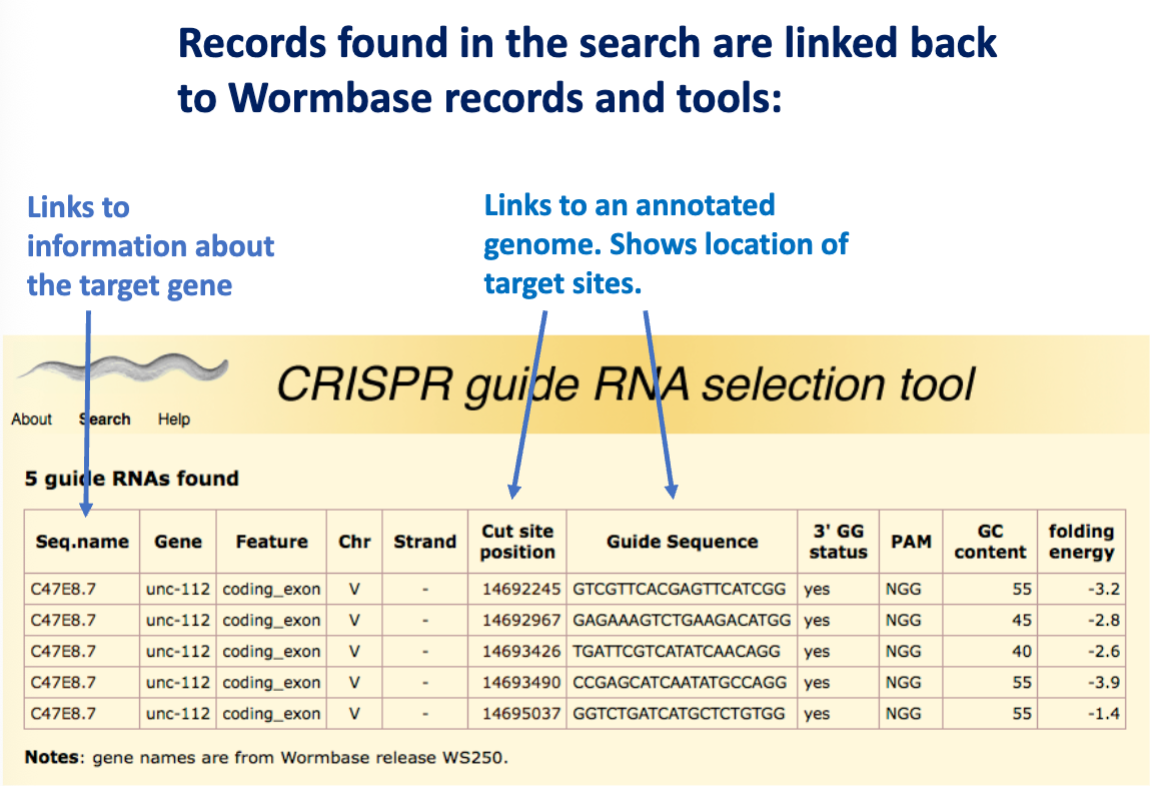
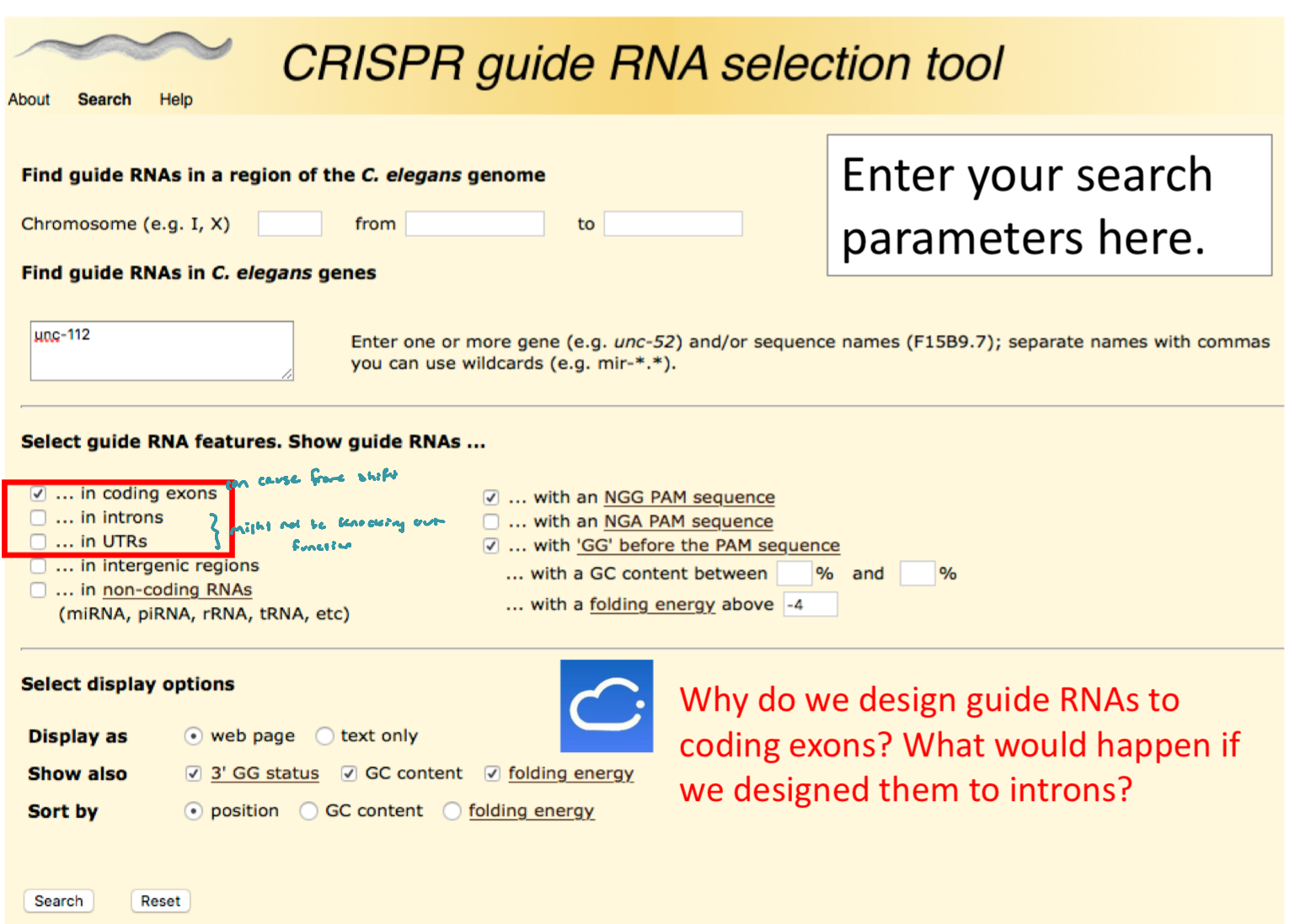

Chop-Chop
another tool to design sgRNAs
Why use a donor template to knock out a gene through homology-directed repair, when non-homologous end joining could also be used to create a loss-of-function mutation?
NHEJ results in random changes to the sequence, whereas HDR changes would be specific and known
ability to introduce selectable markers or reporter genes to help identify organisms with edited genomes
Design of the Donor Template
recall that it is being used to disrupt targeted genes and we need an easy way to identify C. elegans that have beeb succesfully edited
note, in this study, 2 cuts were introduced in order to replace a longer stretch of sequences (requires 2 separate guide RNA constructs)
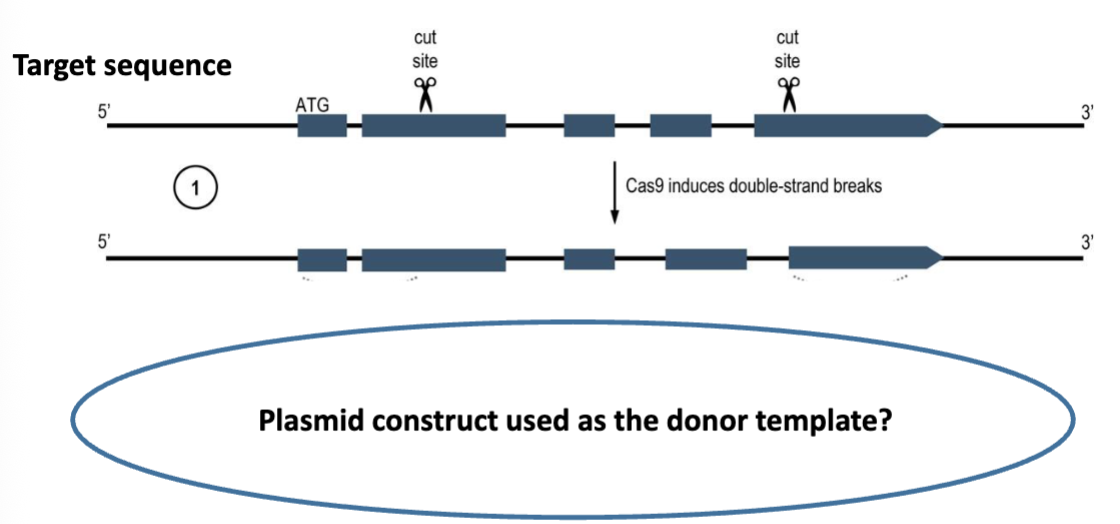
Design of the Donor Template 1
5’ homology arm and 3’ homology arm
in order for a template to be used in HDR, it must be flanked by sequences that are homologous to the targeted area
the homology arms should also cover the regions of the target sequence that will be cut
in this case, the template is being used to replace target gene sequences with selectable marker genes
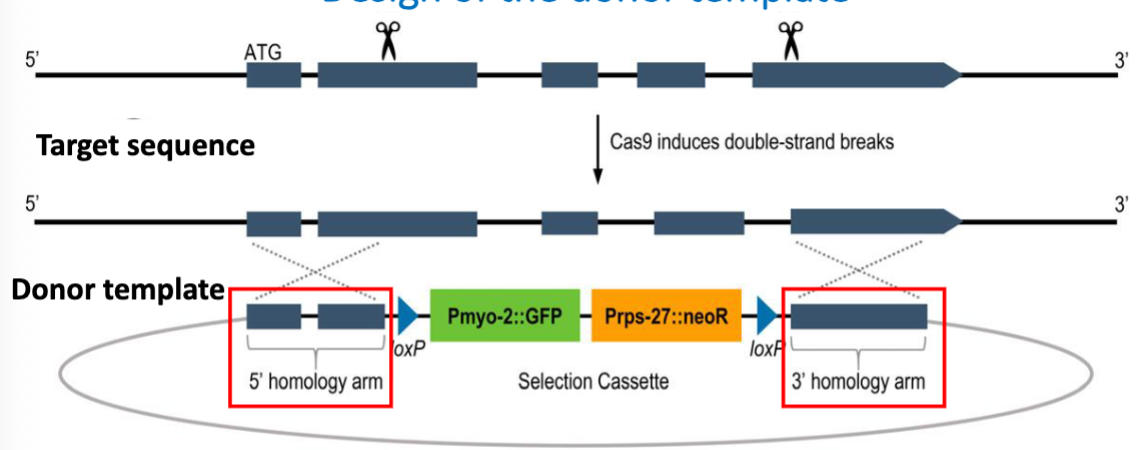
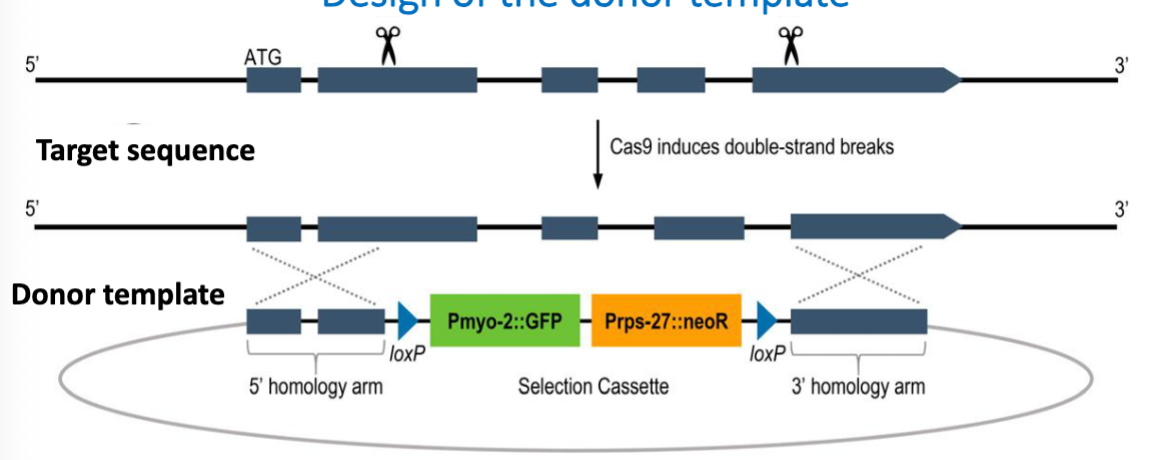
Design of the Donor Template 2
Selectable markers to detect transformed worms
antibiotic resitance gene (neoR), to select for worms that have been edited sucessfully
GFP used to track this mutation in crosses
a marker for having at least one deleted copy of the gene
fluorescence intensity reflects whether the worm is heterozygous or homozygous for the deletion (2x GFP)
Pmyo-2 and Prps-27 are promoters
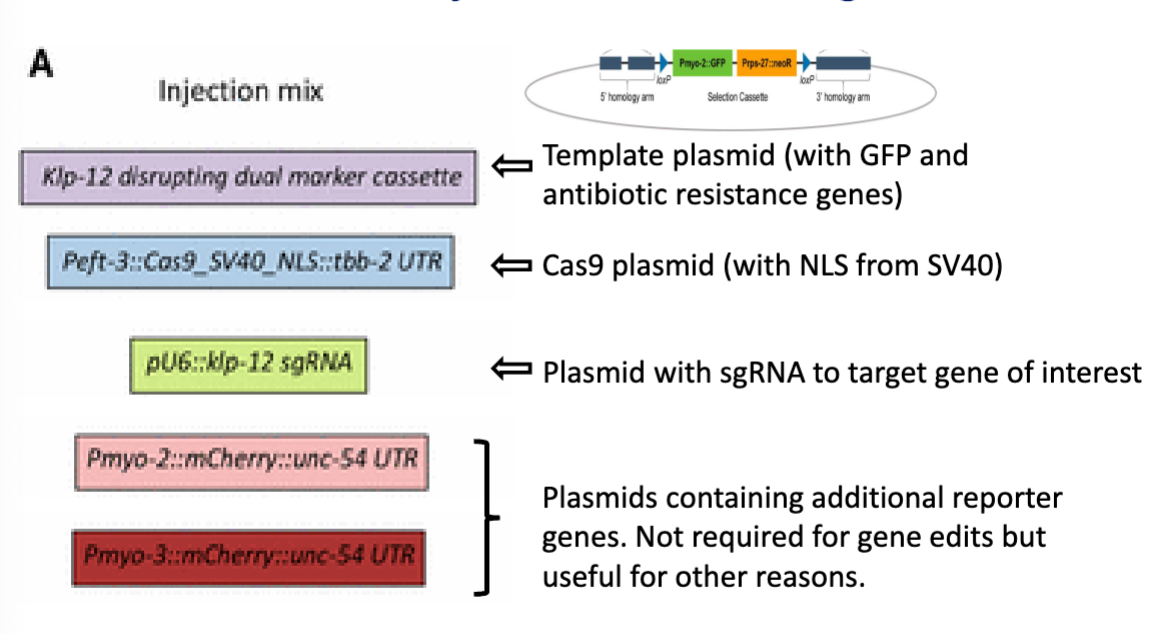
Plasmids injected into C. elegans
slide is showing the components you inject into C. elegans to carry out a CRISPR gene knockout (each plasmid has a specific role)
need Cas9 plasmid with NLS signal to cut DNA
need guide RNA to tell Cas9 where to cut
need a repair template to disrupt the gene (used during DNA repair after Cas9 cuts)
Screening Process for Successfully Edited Worms