BSCI222 Exam 2 DNA Replication
1/55
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
56 Terms
Bacterial cell
Cell Division is part of the growth cycle of prokaryotes
How are cell growth and DNA replication related in bacteria?
Cell growth and DNA replication are coupled in bacteria: replication occurs alongside cell growth
Prokaryotic DNA replication
Occurs at a single origin of replication (oriC) in the prokaryotic circular chromosome
1 origin of replication, which creates 1 replication bubble that has 2 replication forks (1 fork on each side). Inside the bubble, DNA is copied and the bubble expands as replication occurs
origin of replication (oriC)
where DNA replication occurs in bacterial DNA
How does bacterial DNA replicate?
Begins at oriC
Proceeds bidirectionally around circular chromosome
replicon
A unit of DNA that is replicated; the stretch of DNA controlled by ONE origin of replication
When DNA replication starts, the helix opens up - this creates a “bubble” in the DNA. Inside the bubble, DNA is being copied and it expands as replication continues
How many replicons does prokaryotic DNA replication have?
Prokaryotic DNA replication has a single replication bubble (2 forks) = single replicon
replication fork = at the end of each bubble
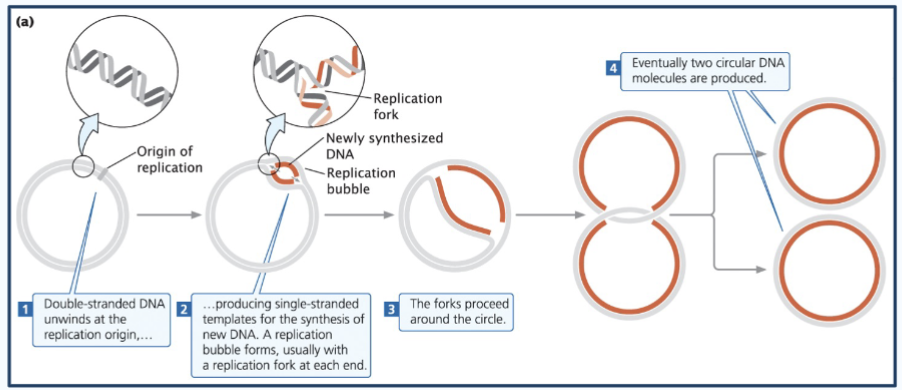
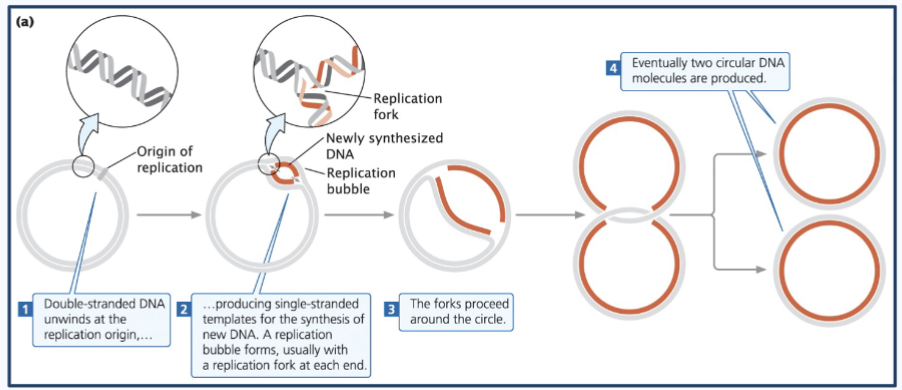
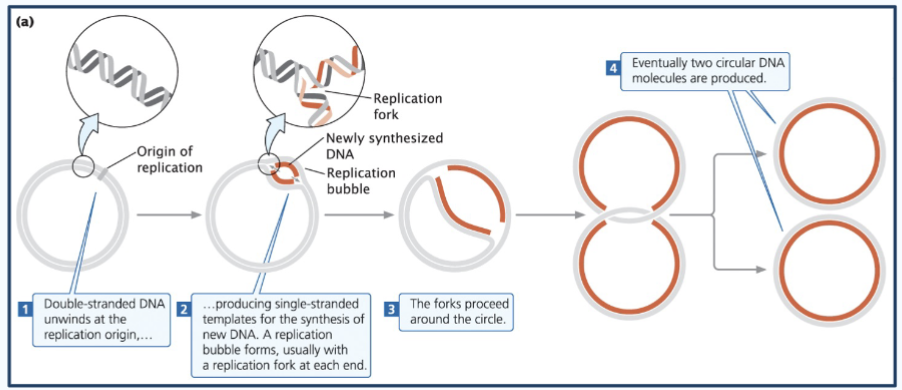
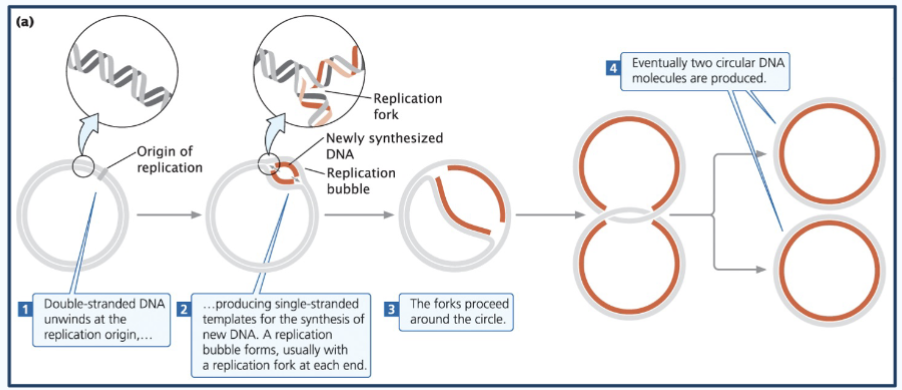
Steps of prokaryotic dna replication
1) The circular chromosome - Double-stranded DNA unwinds at the replication origin (origin of replication or oriC) - aka closed circle opens up at one point (the oriC)
2) This produces single-stranded templates for the synthesis of new DNA. A replication bubble forms, usually with a replication fork at each end.
3) The forks proceed around the bubble (The two replication forks move in opposite directions around the circular DNA)
4) Eventually, two circular DNA molecules are produced
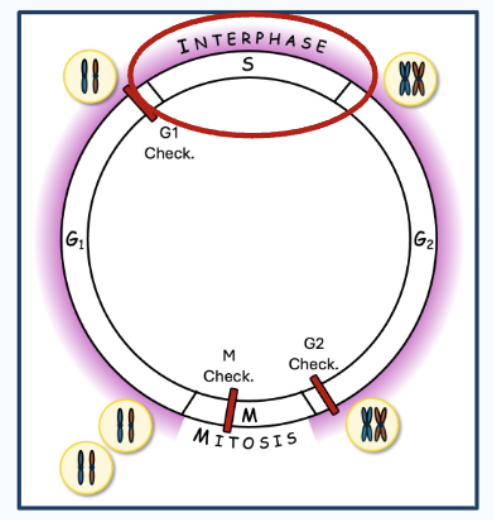
Eukaryotic cell cycle
Interphase = G1 (growth, cell makes necessary enzymes) + S (DNA replication; Each chromosome → becomes 2 sister chromatids) + G2
Then Mitosis (cell division) occurs at the end (Cell actually divides into two identical cells)
Products of meiosis exit the cell cycle (gametes do not re-enter the cell cycle; they wait to fuse in fertilization)
Interphase, including DNA replication during S phase, occurs prior to both mitosis and meiosis
Cell Division (Mitosis) is part of the life cycle of a cell
• Products of meiosis exit the cell cycle
• Entire genome replicated before cell divides (S-phase)
What must happen before the cell divides?
The entire genome must be replicated before the cell divides (S-phase)
Otherwise Daughter cells would get missing genes —> lethal / dysfunctional cells
Eukaryotic DNA replication
Multiple origins of replication per chromosome
Many large, linear chromosomes
Multiple origins per chromosome = multiple replicons
Ex. A mouse (mus musculus) 40 chromosomes with ~2.7 billion bp
- 25,000 replicons averaging 150,000 bp in length
What is true for both prokaryotic and eukaryotic DNA replication?
Daughter strand synthesis is semi-conservative - Each new DNA: 1 old strand + 1 new strand
DNA replication begins with unwinding double helix at origin
DNA is synthesized in 5->3’ direction (of daughter strand)
DNA strands are antiparallel (Two daughter strands synthesized in opposite directions)
Continuous synthesis on one strand (3’-end towards the fork)
Discontinuous synthesis on other strand (fragments)
Why is daughter strand synthesis semi-conservative?
Each new DNA molecule keeps (conserves) ONE original strand and makes ONE new strand
Why? Because the two DNA strands separate, and each old strand acts as a template. This results in two daughter DNA molecules, each consisting of 1 old strand and 1 new strand
Semi-conservative: semi = “half”; conservative = original strand is kept
What direction is DNA synthesized in and why?
DNA is synthesized in the 5’ to 3’ direction of the daughter strand. This is because DNA polymerase can only add nucleotides to a free 3'-OH group
Why are DNA strands antiparallel?
Antiparallel = The two DNA strands run in opposite directions; one strand is 5’ —> 3’, the other is 3’ —> 5’
Two daughter strands are synthesized in opposite directions
What is 5’? What is 3’?
5’ phosphate
3’ OH group
Top strand: 5' ---------> 3'
Bottom strand: 3' <--------- 5'
What is the difference between the two daughter strands?
The two daughter strands are made differently because DNA polymerase can ONLY synthesize in the 5' → 3' direction but the template strands are antiparallel to each other
Continuous synthesis on one strand (3’-end towards the fork) = leading strand
DNA polymerase follows the replication fork
Builds DNA smoothly in one direction
Fork → → →
New strand: → → → (continuous)Discontinuous synthesis on other strand (fragments) = lagging strand
DNA polymerase must work away from the fork
So it builds DNA in small chunks = Okazaki fragments
Fork ← ← ←
Fragments: → → → (short pieces)Steps of prokaryotic DNA replication
Initiation
Bubble Expansion
Daughter Strand Synthesis
Termination
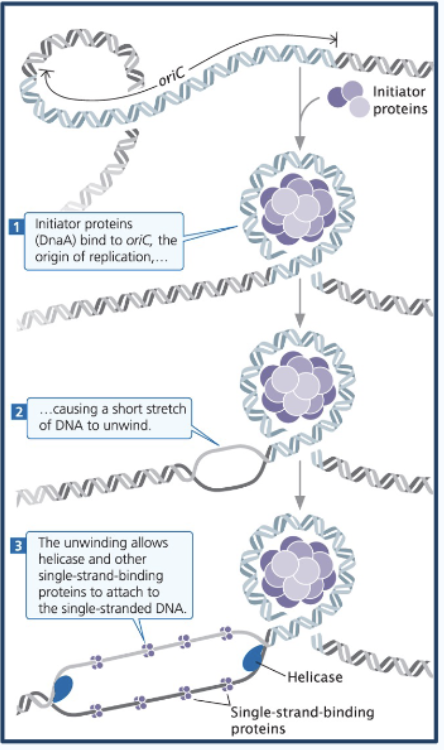
1) Prokaryotic DNA replication - Initiation
OriC sequences recruit initiator proteins (DnaA), which bind to the OriC at DnaA boxes (cooperative binding)
This opens DNA/separates strands at AT-rich DNA unwinding elements (DUEs), creating the replication bubble + 2 forks
OriC sequences recruit initiator proteins (DnaA), and they bind to DnaA boxes (cooperatively)
DNA strands unwind/separate at nearby DNA unwinding elements (DUEs) that are AT-rich regions
This unwinding allows helicase and other single-strand-binding proteins to attach to the single-stranded DNA
This forms the replication bubble
DNA unwinding elements (DUEs)
short AT-rich regions WITHIN the origin of replication that are the first to separate (melt) during DNA replication
Cooperative binding
binding of one molecule makes it easier for others to bind
This applies to DNA replication because
1. First DnaA binds a DnaA box
2. This makes it easier for MORE DnaA to bind nearby
3. Many DnaA proteins assemble together (cooperative binding)
2) Prokaryotic DNA replication - Bubble Expansion
After initiation opens the DNA, the replication bubble grows as DNA is unwound and copied
Helicase breaks hydrogen bonds between nucleotides (base pairs) and separates the two DNA strands; helicase moves in a 5’ —> 3’ direction of the parental strand
Single-strand binding proteins (SSBPs) bond to nucleotides (the single-stranded DNA) - covers 35-65 bases
Why? Because once DNA is opened, the strands want to snap back together; SSBPs prevent re-annealing (rejoining) or secondary structure
Gyrase (Type II topoismerase) - relieves strain double helix (tension ahead of the fork) by cutting both strands, lets DNA rotate, bonds them back together
As helicase unwinds DNA, twisting builds up ahead of the fork, which creates supercoiling (tension)
Name all of the proteins involved in bubble expansion
Helicase → opens DNA
SSBPs → keep it open
Gyrase → prevents twisting damage
Helicase
Enzyme that breaks hydrogen bonds between nucleotides
Moves on lagging in 5’->3’ direction (of parental strand)
Single-strand binding proteins (SSBPs)
Proteins that bind to nucleotides (the single-stranded DNA) - covers 35-65 bases
Why? Because once DNA is opened, the strands want to snap back together; SSBPs prevent re-annealing (rejoining) or secondary structure
Gyrase
Type II topoismerase
relieves strain double helix; cuts both strands, lets DNA rotate, bonds them back together
3) Prokaryotic DNA replication - Daughter Strand Synthesis
Reverse-complement of parental (template) strand
Primase binds to single-strand DNA and makes RNA primer, which gives DNA polymerase a primer (starting point for DNA synthesis) - primer is a free 3’-OH end
DNA polymerase III adds DNA to 3’-end of primer. It also does proofreading, removing mistakes using 3’ → 5’ exonuclease (an enzyme that removes nucleotides one at a time from the END of a DNA or RNA strand); exonuclease works backward from 3’ to 5’ (if wrong base, it removes it)
DNA polymerase I removes the RNA primer using a 5’ —> 3’ exonuclease and replaces it with DNA (adds DNA nucleotides to adjacent DNA strand aka the DNA strand right next to where the primer was removed
(the growing daughter strand))Ligase makes a phosphodiester bond, joining fragments into one continuous strand
Does not add a nucleotide
Reverse-complement of parental (template) strand
New DNA is made based on the template
Uses base pairing rules:
A ↔ T
G ↔ C
“Reverse” = opposite direction
“Complement” = matching bases
Example
Template: 3' - A T G C - 5'
New strand: 5' - T A C G - 3'
4) Prokaryotic DNA replication - Termination
termination = when DNA replication stops; the replication forks finish copying the DNA and the chromosome is fully duplicated
Some prokaryotes have terminator sequence (Ter). The terminator protein (Tus) binds to the Ter sequence and blocks helicase, preventing unwinding
One fork hits the Ter + Tus complex and the fork stops (stalls at the terminator). The second fork continues until it meets the first and that is when replication is complete
Fidelity
accuracy of DNA replication
Fidelity and error repair of DNA replication in prokaryotes
Prokaryotic DNA replication has a very low error rate
DNA polymerase III does proofreading
- Polymerase has high processivity (very fast)
- High fidelity enzyme (1 error in 100,000 bp)
- Distortion in double helix during synthesis triggers exonuclease
- Enzyme removes several bases and tries again
• Proofreading reduces error rate to 1 in 10,000,000 bp
• Post-replication - Mismatch repair enzymes scan and repair
- Fixation of errors (mutations) very rare
What enzyme does proofreading for prokaryotic DNA replication?
DNA polymerase III does proofreading; while building DNA, Pol III is constantly checking: “Did I add the correct base?”
Has high processivity (very fast)
High fidelity enzyme (1 error in 100,000 bp) aka very accurate in DNA replication
If a mistake is made, then exonuclease is triggered to remove several bases and try again
DNA polymerase = enzyme with multiple activities
It has both Polymerase activity (build DNA) and Exonuclease activity (remove DNA)
DNA polymerase is an enzyme that synthesizes DNA, while exonuclease refers to a function (often within the same enzyme) that removes nucleotides for proofreading or repair.
Distortion in double helix during synthesis triggers exonuclease
When DNA polymerase adds the wrong base, the DNA shape becomes slightly distorted
1) DNA polymerase adds wrong nucleotide
2) DNA helix becomes distorted
3) Polymerase detects the distortion
4) Switches to exonuclease mode; 3’ → 5’ exonuclease removes bases
5) Removes the incorrect base (sometimes a few bases)
6) Tries again with the correct nucleotide
Proofreading reduces error rate to…
1 in 10,000,000 bp
What happens after DNA replication is finished?
Mismatch repair enzymes scan and repair (find mistakes that proofreading missed) - ex. A paired with C instead of T
Fixation of errors (mutations) very rare aka very rare for DNA polymerase + mismatch repair enzymes both miss a mistake, which causes the mistake to be expressed as a mutation
If not repaired —> mistake passed on; if repaired —> no mutation
Fixation
When a DNA error (mutation) becomes permanent in the genome because it is not repaired and is copied during subsequent DNA replication
Error is copied in the next round of replication → becomes permanent, so a mistake becomes a permanent mutation
1) Eukaryotic DNA replication - Initiation
Origin of replications (origins) recruit origin recognition complex (ORC) proteins. ORC bind to origins during the G1 phase of the cell cycle (before replication starts).
ORC anchors Helicase on double helix (helicase placed on DNA) - still G1 phase
Helicase activity triggered at start of S-phase (DNA synthesis)
Licensing factors determine which origins will be active; only certain replication origins are allowed (“licensed”) to start replication, and this is tightly controlled.
2) Eukaryotic DNA replication - Replication bubble expansion
MCM2-7 proteins are activated during the S phase to become helicase
Helicases move in opposite directions (two helicases in opposite directions), expanding the replication bubble
Class I Topoisomerase cuts single strand to relieve strain (While helicase unwinds the DNA, tension aka supercoiling builds ahead of the fork. Class I Topoisomerase fixes this by cutting one strand to let the DNA relax, and then rejoins it)
MCM2-7
The helicase in eukaryotes. MCM2-7 proteins load onto DNA at the origin during G1. They form a helicase complex that is initially inactive. But in the S phase, during actual DNA replication, MCM2–7 becomes active and starts unwinding DNA
Class I Topoisomerase
Cuts a single DNA strand to relieve strain (While helicase unwinds the DNA, tension aka supercoiling builds ahead of the fork. Class I Topoisomerase fixes this by cutting one strand to let the DNA relax, and then rejoins it)
3) Eukaryotic DNA replication - Daughter Strand Synthesis
Eukaryotes have many different DNA polymerases that grow the daughter strand
DNA polymerase alpha has primase activity, starts synthesis; makes short RNA polymer and adds DNA to 3’-end
DNA polymerase delta joins complex for lagging strand synthesis
DNA polymerase epsilon carries out leading strand synthesis
RNA primers are removed by FEN1, a 5’->3’ exonuclease
DNA polymerase alpha
Acts like (RNA) primase in prokaryotic DNA replication; Makes a short RNA primer and then adds a short stretch of DNA
DNA polymerase delta
Joins complex for lagging strand synthesis; it extends DNA okazaki fragments and repeatedly starts after each primer
DNA polymerase epsilon
carries out leading strand synthesis
FEN1
A 5’ —> 3’ exonuclease enzyme that removes RNA primers
Nucleosome dynamics
In eukaryotes, DNA is wrapped around histone proteins → forms nucleosomes (DNA is packaged on nucleosomes, which helps compact it)
However, DNA unwinding (helicase) disrupts nucleosomes, so histones temporarily come off of DNA
Histones re-associate on newly synthesized DNA (old histones + new histones bind back to DNA)
Nucleosomes reform on both double helices (Both daughter DNA molecules get nucleosomes)
Why do chromosomes shorten during replication?
Unlike bacteria, eukaryotes have linear DNA/chromosomes: Ends of DNA = difficult to fully replicate
Removing primer at the end of a strand leaves unreplicated DNA (no 3’-OH group)
The last RNA primer on the lagging strand cannot be replaced.
Telomere
repeated sequences that protect ends of linear chromosomes; has a G-rich 3' overhang that is bound by a telomerase enzyme
Chromosome shortens by…
70-100bp each time
How is this problem fixed?
At the end of a chromosome, one strand sticks out: G-rich 3’ overhang = loose tail of DNA
The telomerase enzyme binds to the G-rich 3’ overhang
Telomerase extends the end to allow second strand synthesis (so replication can be completed)
Telomerase
Enzyme that adds more repeats to make the 3' overhang longer, so that it can act as a template for synthesis of the second strand
It includes a reverse complementary RNA sequence (carries its own RNA template. This RNA is complementary to the telomere sequence and guides the addition of new DNA repeats).
How does telomerase work?
Telomerase binds 3’ overhang
↓
Uses its RNA template
↓
Adds DNA repeats to the end
↓
Allows completion of replication
Telomeres protect chromosome ends, and telomerase extends them using an internal RNA template to prevent DNA loss during replication.
What happens without telomerase?
DNA polymerase can’t copy the very end, and DNA gets shorter every time
Where is telomerase active?
Active in single-cell eukaryotes and in germ cells (renewal)
Termination of DNA replication
Prokaryotes have a specific Termination signal (Ter/Tus sequence). Tus binds to Ter and the protein blocks one of the forks. But eukaryotes have no specific sequence; in Eukaryotes, DNA replication stops when the forks run into each other ❌ No specific sequence