GENOMICS 2.2: Structural and functional elements of the genome
1/67
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
68 Terms
What is the size range of prokaryote genomes?
Relatively small: 0,2-13 Mbp
What is the size range of eukaryote genomes?
Big: 3 Mbp - 700 Gbp
What is the genome organization and ploidy of prokaryotes?
Circular genomes with accompanying circular plasmids
Haploids (singular copy of the genome)
What is the genome organization and ploidy of eukaryotes?
Nuclear and oragnelles enclosed with membranes
Nuclear genomes: linear, organized in several chromosomes
Organelle genomes: circular, reminiscent of bacterial
Haploid (n) (gametos), diploid (2n), or polyploid (normally in plants)
What is the C Value?
The C value is the genome size of the haploid genome (in pg or Gb) → it allows us to compare genomes from different species
What is the C Value paradox?
The range of C values does not correlate with the complexity of organisms → we would expect that the more complex the genome the bigger it would be, but that is not always the case

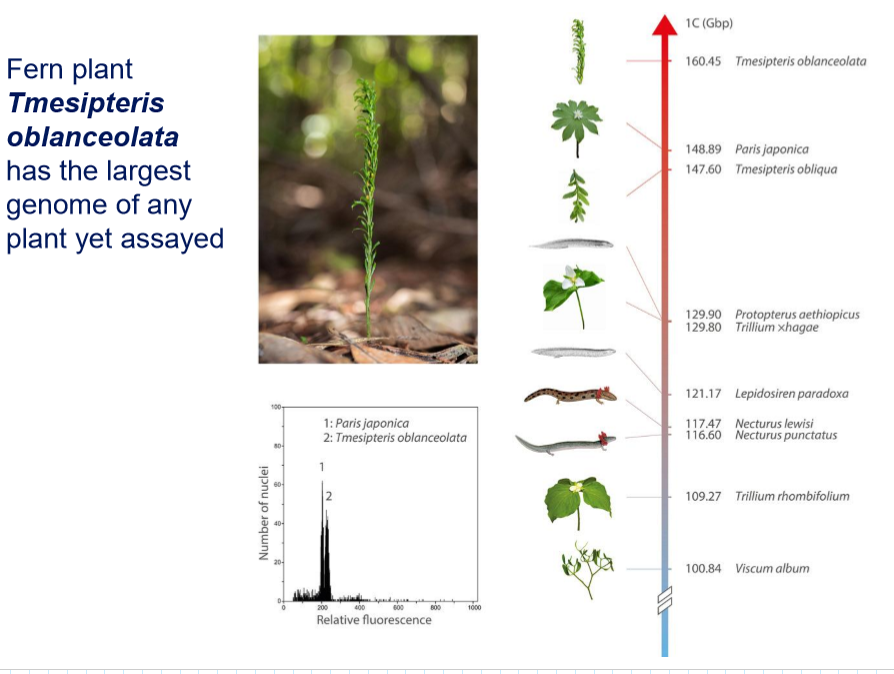
Which organism has the biggest genome? And the second biggest?
Tmesipteris oblanceolata → fork plant (160 Gbp)
Paris japonica → canopy plant
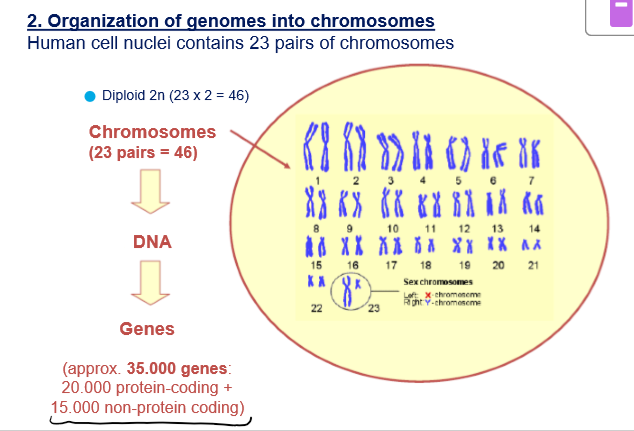
How are eukaryotic nuclear genomes organized?
Organized in chromosomes
Each chromosome is a linear DNA molecule
Number of chromosomes varies across species → similar to the C-value paradox, the number of chromosomes does not equal to complexity
Diploids: 2 pairs of homologous chromosomes → 2 chromosomes that have the same genes
In humans:
Haploid number of chromosomes
Diploid number of chromosomes
Haploid number of chromosomes: 23
Diploid number of chromosomes: 46
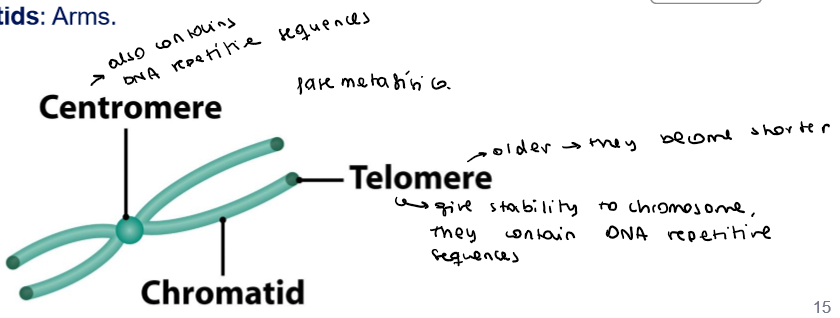
What are the parts of a metaphase chromosome?
Telomeres:
Tandem repetas of repetitive sequences
Provide stability
Centromere:
Appears as a constriction
Drive chromosomic movements during cell division
Chromatids:
Arms
How many genes do humans have? How many are protein-coding? How many are not?
Approx 35 000 genes
20 000 protein-coding
15 000 non-protein-coding
What is a karyogram?
It is a visualization of chromosomes by staining
In what phase do the chromosomes usually are in a karyogram?
In the metaphase
What is the banding pattern in this staining technique?
G-banding
Dark bands are AT rich
Pale bands are GC rich
The most common one
What is the banding pattern in this staining technique?
R-banding
Dark bands are GC rich
Pale bands are AT rich
What is the banding pattern in this staining technique?
Q-banding
Dark bands are AT rich
Pale bands are GC rich
(Different staining procedure than G-banding)
What is the banding pattern in this staining technique?
C-banding
Dark bands contain constitutive heterochromatin → to stain centromeres
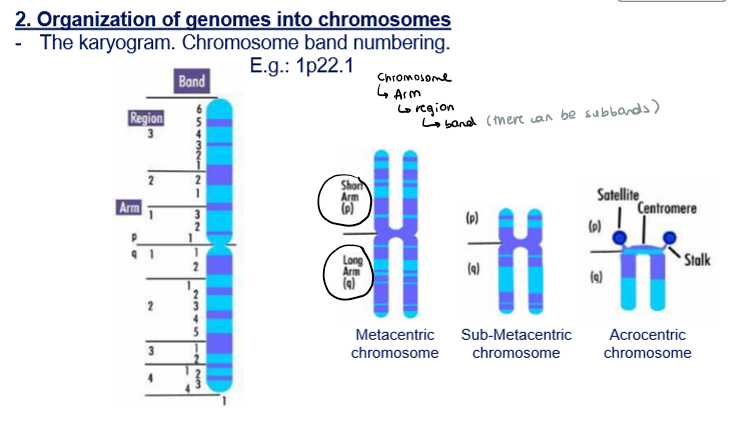
Which types of chromosomes are there?
Telocentric
Acrocentric
Sub-metacentric
Metacentric
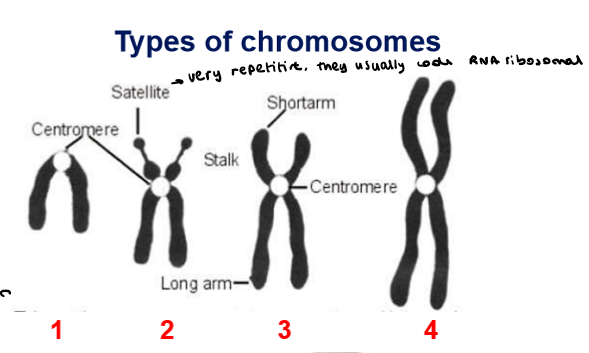
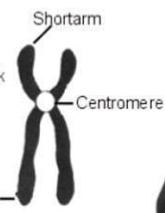
Which type of chromosome is it?
Sub-metacentric
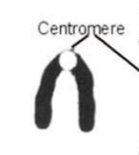
Which type of chromosome is it?
Telocentric
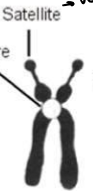
Which type of chromosome is it?
Acrocentric

Which type of chromosome is it?
Metacentric
True or false: the X and Y chromosome are the same size
False: they are not homologous
X: submetacentric medium chromosome
Y: small acrocentric chromosome
What are the characteristics of sex chromosomes?
Heteropyknotics: they stain differently than other chromosomes
Delay in duplication
Delay in ordering
Chromosome band numbering → what does each letter/number refer to?
1p22.1
1: chromosome
p: arm of the chromosome
Short arm (p)
Long arm (q)
2: region
2: band
1: subband
What is the karyotypes’ function?
They provide structural organization of individual genomes → you can detect deletions, insertions, duplications, translocations of large DNA fragments (you can detect big mutations)
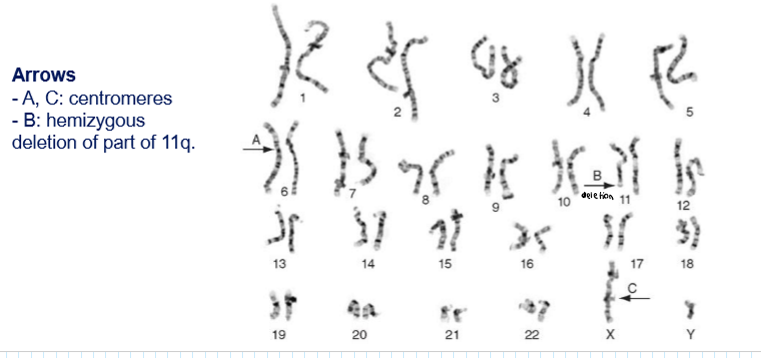
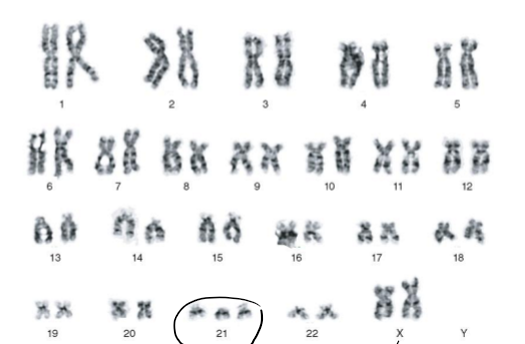
What can you say about the patient just seeing the karyotype?
Aneuploidy (abnormal number of chromosomes): trisomy of chromosome 21
2 X chromosomes: female
True or false: the GC content varies among species, across genomes and it is not uniform in a same chromosome
True
Eukaryotes have low range of variation: 35-45%
Prokaryotes have much wider range: 25-70%
What is an example of the % of GC content varying within a chromosome?
CpG islands: genomic region with high frequency of CG dinucleotides relative to the rest of the genome → (GCGCGCGCGC)n
This is very typical in the upstream promotor sequences → methilation occurs in C: the gene won’t be expressed
True or false: humans have low repetitive DNA
False, almost 50% is repetitive
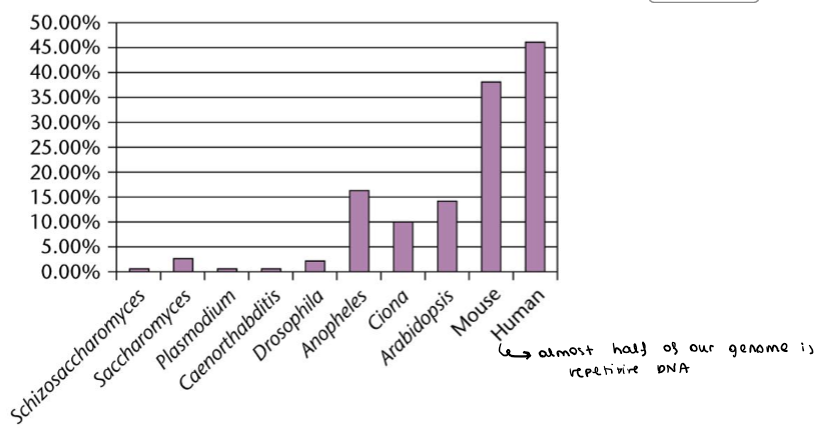
How can it be shown that a high % of genomic DNA is repetitive?
Through denaturing/renaturing experiments: repetitive sequences associate and dissociate much quicker, while non repetitive DNA takes longer
if the DNA is repetitive, when you do the hybridazion step, it will hybridaze much quicker (más concentración de esa secuencia, tendrá más probabilidad de encontrarse)
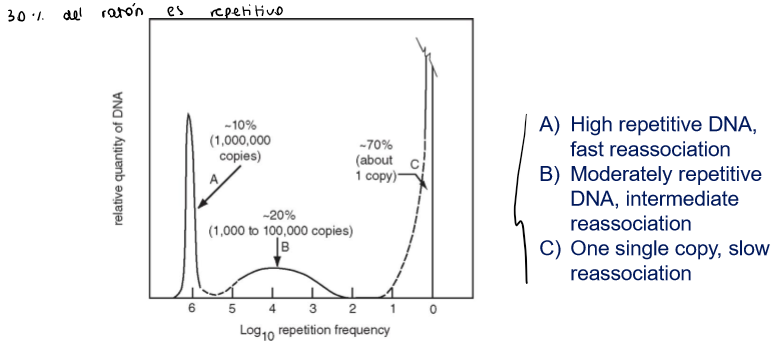
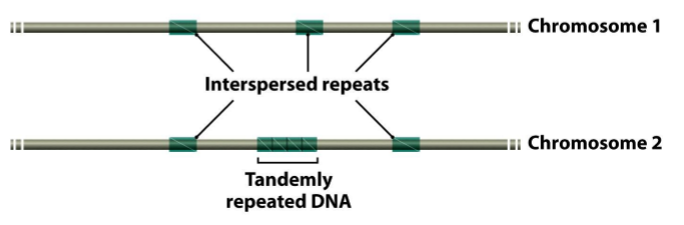
What are the two types of repetitive DNA?
Interspersed repeats: repeticiones intercaladas → most repetitive sequences are interpersed repeats
Tandemly repeated DNA: repeticiones seguidas, pequeños grupos donde no hay otras secuencias entre medio
How can the human genome be classified?
1/3 of the genome: genes and gene-related sequences (1200 Mb)
Exons (48 Mb): codificant region of the gene
Related sequences (1152 Mb):
Pseudogenes: nonfunctional genomic DNA sequences that closely resemble functional protein-coding genes but have lost their ability to encode proteins due to accumulated mutations
Gene fragments
Introns, UTRs
2/3 of the genome: Intergenic DNA (2000 Mb) → basically repetitive DNA
Interpersed repeats (1400 Mb):
LINE’s (640 Mb)
SINE’s (420 Mb)
LTR elements (250 Mb)
DNA transposons (90 Mb)
Tandemly repeats (600 Mb):
Microsatellites = simple sequence repeats (SSR) (90 Mb)
Various (510 Mb)
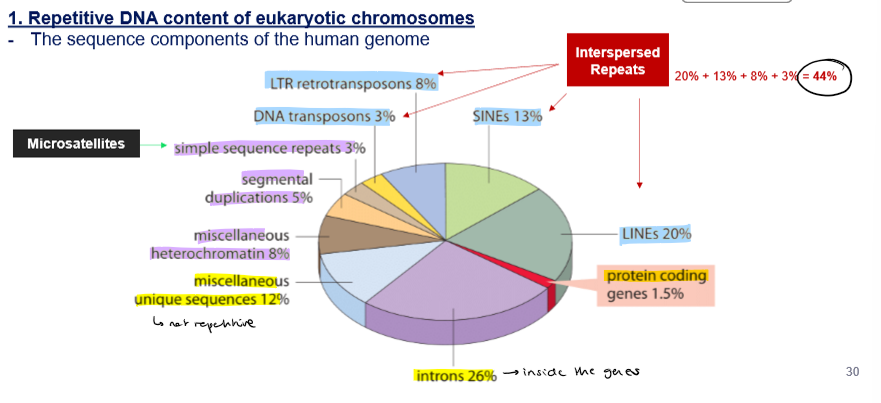
Which DNA content corresponds to these %?
8%
3%
13%
20%
3%
5%
8%
12%
26%
1,5%
Interpersed Repeats → 44%
8% → LTR retrotransposons
3% → DNA transposons
13% → SINE’s
20% → LINE’s
3% → SSR (microsatellites)
5% → segmental duplication
8% → miscellaneous (diverse) heterochromatine (condensed)
12% → miscellaneous unique (not repetitive) sequences
26% → introns (inside the gene)
1,5% → protein coding
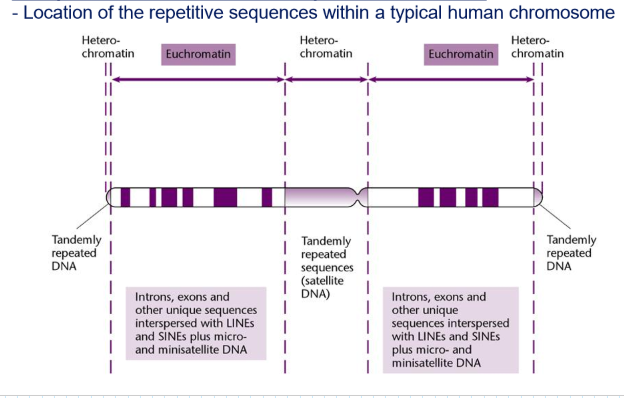
What is the importance of identying repetitive DNA?
Repetitive DNA influences the structure and function of the genome (chromosome rearrangement, transcriptional regulation)
Repetitive DNA: usually condensed → heterochromatin: gene silencing
Unique DNA: usually relaxed → euchromatin: gene expression
Importance in disease: recombination events resulting in duplications or deletions
What types of repetitive DNA are there?
Interspersed repeats (transposon-derived repeats)
Processed pseudogenes → An mRNA transcript from a functional gene is accidentally reverse-transcribed into DNA by an enzyme (usually from a LINE element) and then inserted back into the genome.
Simple sequence repeats
Segmental duplications
Blocks of tandemly repeats sequences
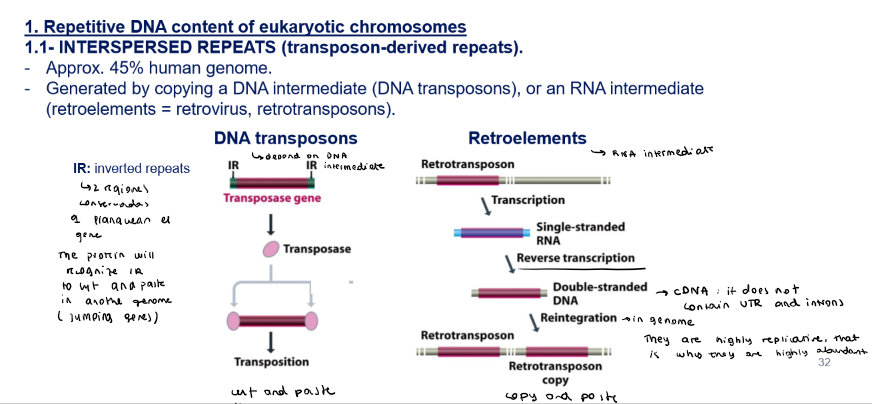
Repetitive DNA: how are interpersed repeats (trasposon-derived repeats) generated?
They can be generated by:
Copying a DNA intermediate → DNA transposons
A transposable gene is flanked by two inverted repeats (IR)
A transposase will recognize the IR and cut and paste the gene in another part of the genome
Copying a RNA intermediate → retroelements (RNA intermediate) = retrovirus, retrotransposon
A retrotransposon will go through transcription and generate a single-stranded RNA
It will go through reverse transcription and generate double-stranded DNA → cDNA: it doesn’t contain UTR and introns
It will then reintegrate in the genome randomly (copy and paste) → they are highly replicative, that is why they are highly abundant in the genome
Repetitive DNA: how much of the human genome interpersed repeats do represent?
45% approx
What are long terminal repeats (LTR)?
DNA sequences found flanking retrotransposons and retrovirus (common to viral retroelements)
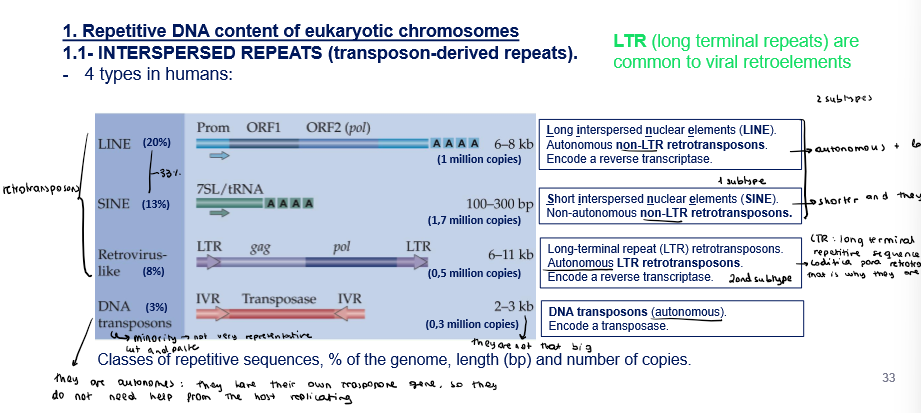
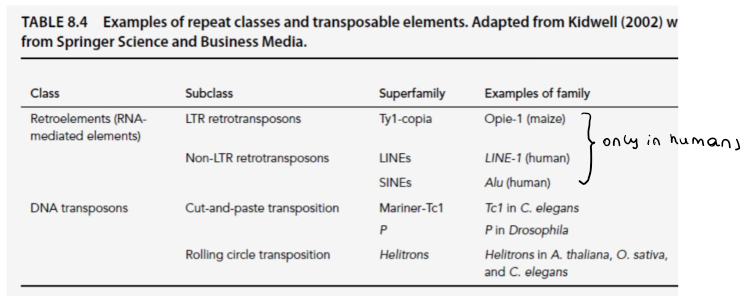
Repetitive DNA: what are the 4 types of interpersed repeats in humans?
Retrotransposons:
Long interpersed nuclear elements (LINE): 20%
Autonomous non-LTR retrotransposons
Encode a reverse transcriptase
Short interpersed nuclear elements (SINE): 13%
Non-autonomous non-LTR retrotransposons
Long-terminal repeat (LTR) retrotransposons: 8%
Autonomous LTR retrotransposons
Encode a reverse transcriptase
Transposons:
DNA transposones: 3%
Autonomous
Encode a transposase
Repetitive DNA: Are interpersed repeats present in eukaryotic genomes?
Yes
Repetitive DNA: what are pseudogenes?
A nonfunctional gene copy
Repetitive DNA: what are conventional pseudogenes?
An initial mutation inactivates the function
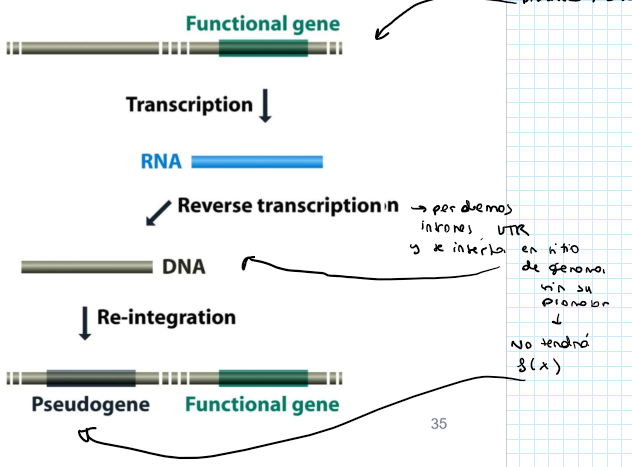
Repetitive DNA: what are processed pseudogenes?
They arise by abnormal retrotransposition event on a functional gene → chat: An mRNA transcript from a functional gene is accidentally reverse-transcribed into DNA by an enzyme (usually from a LINE element) and then inserted back into the genome.
They lack introns and is inactive (no upstream regulatory sequences, no expression)
Repetitive DNA: what are sinple sequence repeats (SSR)?
They are tandemly repeated DNA
Minisatellites: n repeats lager than 13 bp (up to 500 bp) (clusters of up to 20 kb)
Microsatellites: n repeats of up to 13 bp (clusters of up to 150 bp) → more abundant, that is why when we say SSR we usually refer to microsatellites
What is a polymorphism?
It is the presence of genetic variation wiithin a population, upon which natural selection can operate
Repetitive DNA: why microsatellites have a high mutation rate?
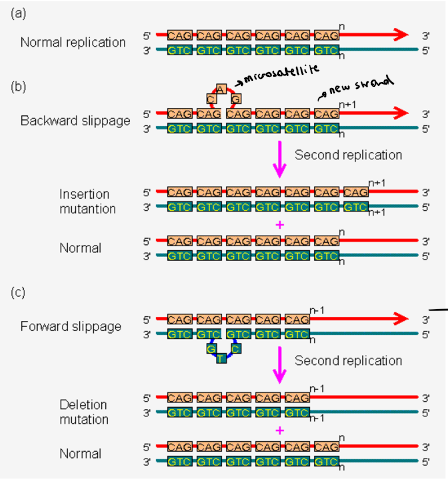
It is because of replication slippage: occurs at the repetitive sequences when the new strand miss pairs with the template strand → it causes microsatellite polymorphisms
During replication, DNA strands are opening and closing, because they are so similar, they can bind incorrectly, making a mismatch/loop
It can be a:
Backward slippage: the new strand has the loop (addiotional nt) → we add a repetition
Forward slippage: the template has a loop (the new strand will have fewer nt) → we lose a repetition
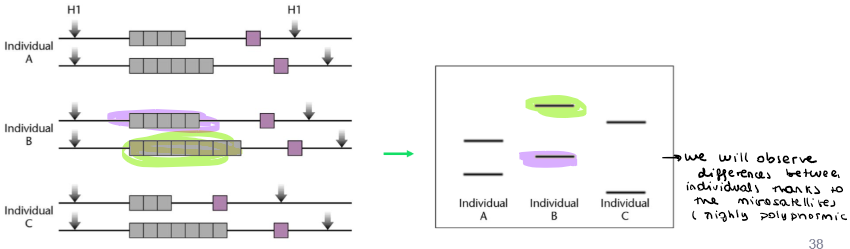
Repetitive DNA: why can microsatellites be used in genetic profiling of individuals?
Because they are highly polymorphic elements → high variation and high heterozygosity
For example, each microsatellite averages 10 allelic variants
Repetitive DNA: how microsatellites are used to identify individuals? Simple Sequence Lenght Polymorphisms (SSLPs analysis)
The FBI have identified 13 different SSRs → at least everyone should have one (in 2017 they incremented it to 20)
They have designed a set of 13 different primers corresponding to the SSRs → depending on the different combination of microsatellites each individual has, different bands will appear in the gel
It will allow us to differentiate genomes
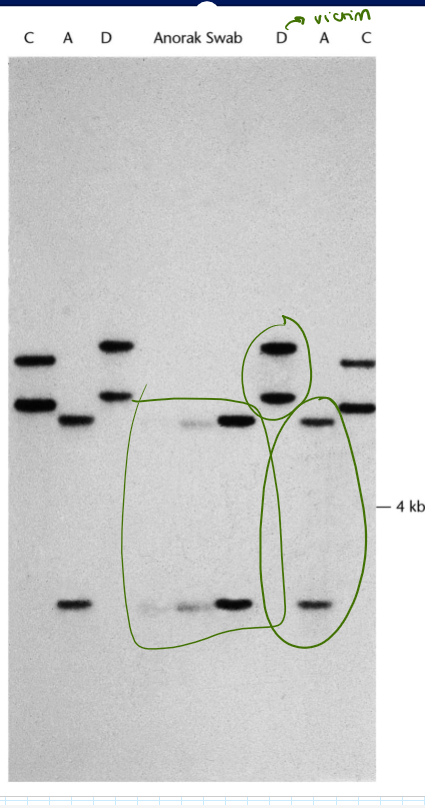
D: victim
A: suspect 1
C: suspect 2
Who is the killer?
A, because the anorak swab matches
Repetitive DNA: what are segmental duplications (low copy repeats)?
They are two genomic regions sharing >90% nt identity over a span of 1kb
Common in plant and animal genomes
5% of genomic DNA are segmental duplications
They make difficult the sequence assembly and the genetic dissection
Unknown function other than redundancy
Compared with other mammals, the genomes of — and other — show an enrichment of large, interpersed — —.
Humans
Primates
Segmental duplications
Repetitive DNA: what are blocks of tandemly repeated sequences?
Occurs in telomeres and centromeres in the form of heterochromatin
Telomeres:
Human telomeric repeats: (TTAGGG)n
Telomere lenght:
11 kbp at birth
< 4 kbp in old age
Centromeres:
a 1-4 Mb region with alpha-satellite repeats (171 bp repeats)
They have high density beacuse they are very repeated → they are satellite bands in density gradient centrifugation of genomic DNA
True or false: gene density does not vary
False: it depends on the sequence
Gene content: what is the gene definition?
The one gene - one functional RNA hypothesis: the gene is a portion of DNA required for the expression of a functional gene product (an RNA or a protein)
Gene content: the — is the basic atomic unit of inheratance
Transcript
What is an open reading frame (ORF)?
The part of DNA or RNA that ha sthe potential to be translated into a protein
From start codon to stop codon
Gene content: what are the types of genes?
Protein-coding genes:
Major category
Minimum ORF 90 bp (30 aa)
Non-protein-coding genes:
tRNA → protein synthesis
Function: The "adaptor" molecule. It physically brings the correct amino acid to the ribosome during translation (protein synthesis). It reads the codon on the mRNA and drops off the amino acid.
rRNA → protein synthesis
Function: The "factory floor." It is the main structural and catalytic component of the ribosome (the machine that builds proteins). The ribosome is actually made of rRNA and proteins.
snoRNA: small nucleolar RNA → rRNA processing
Function: "rRNA processing." These work in the nucleolus (a part of the nucleus) to chemically modify and cut up rRNA molecules to help them mature into functional ribosomes.
snRNA: small nuclear RNA → splicing
Function: "splicing." These are the main components of the spliceosome. The spliceosome is the complex that cuts out introns from pre-mRNA and joins the exons together to make the mature mRNA.
miRNA: microRNA → regulation
Function: "regulation." These are short RNAs (about 22 nucleotides) that bind to specific messenger RNAs (mRNAs) and block them from being translated or mark them for destruction. They are a key part of gene regulation (turning genes off).
lnc-RNA: long non-coding RNA
Function: A catch-all category for long RNA molecules (longer than 200 nucleotides) that don't code for protein. They do a huge variety of jobs, including silencing entire chromosomes (like X-chromosome inactivation) and controlling transcription.
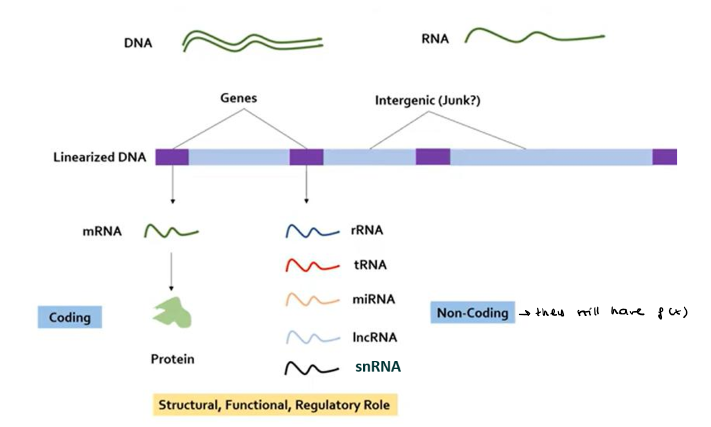
What are ESTs?
Expressed sequence tags
Short sequence of DNA that is generated by sequencing one or both ends of a cDNA clone
They are a subtype of cDNA library but not complete
Gene content: how can we identify exons?
By alignments of cDNA and EST to the genome
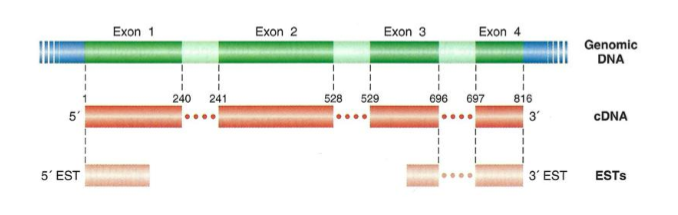
Gene content: we use extrinsic (BLAST) and intrinsic (DNA patterns such as start and stop codon) algorithims to define:
Intron/exon boundaries (GT/AT) → GT al inicio del intrón y AT al final (splicing sites)
Exons (noncoding 5’ and 3’ UTR, ATG, Stop)
Regulatory elements (basal-TATA-box, proximal, distal)
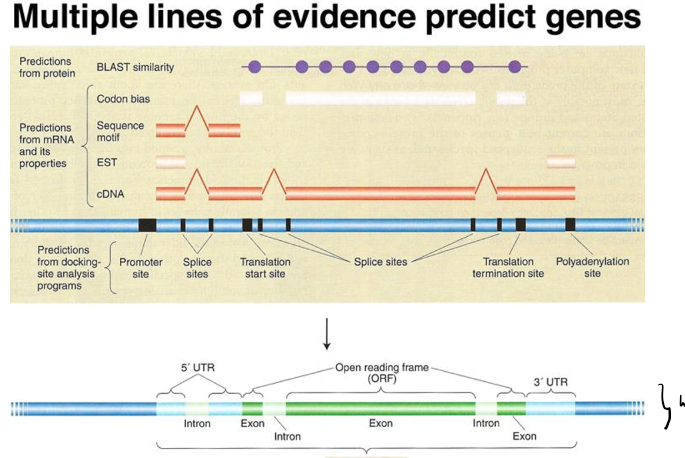
Gene content: more complex → — exons
Give an example of a complex gene:
more
Titin (connectin, muscle elasticity): has 364 exons
Gene density: what does it mean to have a compact genome? Give an example of an organism that has a compact genome and an organism that has a non-compact genome:
To have a high number of protein-coding-genes and a low number of repetitive DNA
Compact genome: yeast
Non-compact genome: maize
Gene content: what does the Gene Ontology classify?
It classifies the molecular function, the biological process and the cellular component of a gene
Gene content: why non-protein-coding genes (ncRNA) are more difficult to identify in a genome than exons?
They are more difficult to predict
Not represented in cDNA libraries (no polyA)
Conserved the secondary structure (not codons): no sequence similarity searches
La función depende de cómo se pliega el ARN, no de qué nucleótidos exactos tiene. La secuencia puede cambiar mucho siempre que los pares de bases que forman la estructura se mantengan. La secuencia cambió, pero la estructura sigue igual → la función se conserva → la evolución lo permite. BLAST busca similitud de secuencia, no de estructura.
Less known function and distribution
Regulatory regions: what are cis-regulatory modules (CRMs)?
They are promotors, enhancers and silencers → no tienen una secuencia fija, por eso necesitamos métodos experimentales para identificarlas
Regulatory regions: what is an experimental tool that can be used for identification of cis-regulatory modules (CRMs)?
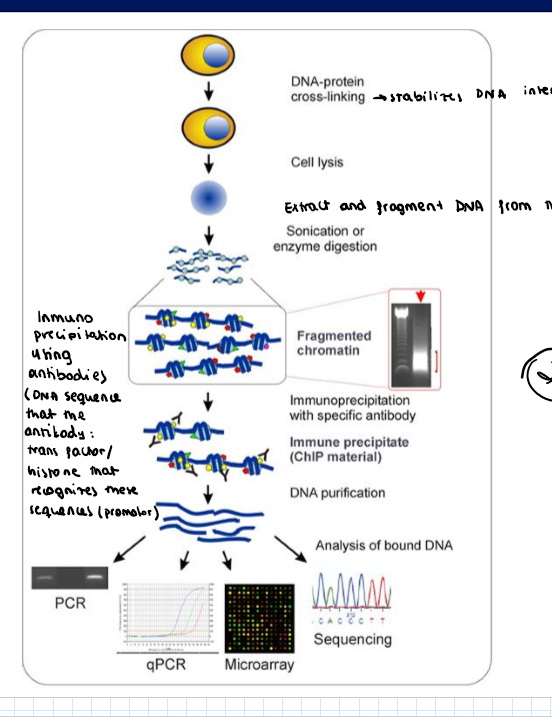
Chromatin Immunoprecipitation: ChIP-seq or ChIP-chip → where are transcription factors or histones bound in the genome?
DNA-protein cross linking: stabilizes DNA interacction
Cell lysis: free DNA and proteins
DNA fragmentation by sonication or enzyme digestion
Inmunoprecipitation:
add an specific antybodu against a transcription factor or a histone modification
DNA purification: you eliminate the protein and only save the DNA that was bound to the antibodies
You sequence these segments
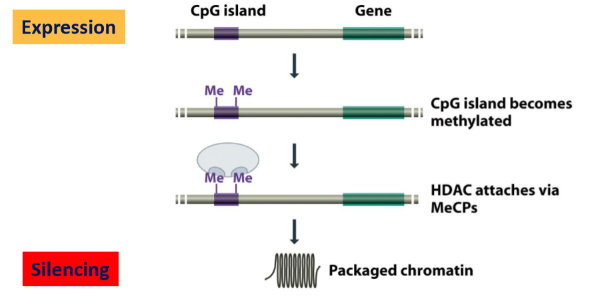
Regulatory regions: how can CpG islands determine gene expression or silencing?
CpG islands: high proportion of C-G
When not methylated: active promotor → gene expression
When methylated:
C-methylation
Recruitment of Methyl-CpG-binding proteins (MeCP)
Recruitment of Histone-deacetylase (HDAC): they remove acetyl group of hystones, now they are + charged and bind more to DNA (-)
Chromatin will be more compact → gene silencing