BILD 5 FINAL EXAM
1/37
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
38 Terms
P-Value vs Effect Size
P-values indicate whether a difference is statistically significant, but don’t describe how large that difference is
don’t capture whether difference is biologically meaningful
Effect size: quantity that captures magnitude of the effect
significant result could have tiny effect size
non-significant result could have large effect size
Type I vs Type II errors
Type I: false positive, erroneously conclude there is a difference when in reality, none exists
reject the null when null is actually true
Type II: false negative, fail to detect a difference when one does exist
fail to reject the null when null is false
Alpha and Type I error
The significance level (alpha) sets rejection fence:
when alpha = 0.05, we only reject the null in most extreme 5% of t-scores
when alpha = 0.01, we only reject null in most extreme 1% of t-scores
Lower alpha → stricter standard for rejecting the null
as we lower alpha, more cautious about rejecting null; reduces likelihood of type I error
Type I error rate = alpha!
alpha = probability of incorrectly rejecting the null
Trade-off between Type I and Type II error
Lowering alpha:
Good: more cautious about rejecting null → reduced type I error rate
Bad: might end up missing effect that DOES exist → increased type II error rate
With lower alpha, becomes harder to reject the null
less likely to reject null by mistake (reduce type I error)
makes it harder to reject null when you should (increase type II error)
Attempting to reduce one will increase the other, therefore error can never = 0; type I and type II errors are at odds with each other
Type II error and power + power analysis
Type II error (ß): if alternative is true, what’s the probability we miss the effect?
Power: if alternative is true, what’s the probability we detect the effect?
we want high power
study with low probability of type II error = high statistical power
power is opposite of type II error
Common target for statistical power is 80%:
if alternative is true (an effect DOES exist), we correctly detect it 80% of the time
Before conducting experiment, researchers perform power analysis:
before you start, you usually have some idea of expected effect size and SD
“what sample size do I need to achieve a desired power?”
Factors affecting statistical power
Sample size
higher the sample size, higher the power (directly proportional)
more likely we are to detect a difference if one exists
Significance level
higher the significance level, higher the power
if we increase alpha, don’t need as extreme of a t-score to reject the null
if alternative is true (truly is a difference), it is easier to discover it
Caveat: increasing alpha increases type I error
Effect size
higher the effect size, higher the power
easier to detect big difference compared to small difference
Standard deviation
higher the standard deviation, lower the power
large standard deviation: hard to detect an effect if it does exist
small standard deviation: easier to detect difference
The problem of multiple comparisons + Bonferroni correction
The more t-tests you do, the greater the chance of making at least one false positive
if significance level = 5%, each individual test has 5% chance of being a false positive (type I error)
Bonferroni Correction: divide alpha (ie 5%) by number of comparisons to determine a Bonferroni-corrected alpha
ex. if you perform 10 t-tests, new alpha threshold is 0.05/10 = 0.005
for any individual comparison to be significant, p-value must be < 0.005
P-hacking and Preregistration
P-hacking: repeatedly re-analyzing your data until you find a statistically significant result
undermines reproducibility of scientific research
perform more comparisons = significantly increase chance of at least one type I error (false positive)
Preregistration: specifying your study design, hypotheses, and analysis plans before collecting data
holds yourself accountable to original research objectives, reduces temptation to p-hack
avoids manipulating data to fit the narrative you want
Analysis of Variance (ANOVA)
Compare all the means simultaneously in one test: (compares 3+ groups)
Null: the mean is the SAME across all three groups
Alternative: the mean is NOT the same
at LEAST one group is different from the rest
To perform an ANOVA, R calculates:
an F statistic
a p-value, which tells us if F statistic large enough to be statistically significant
large F statistic = big deviation from null = low p-value
Variance and the F-statistic
Variance within each group: members of each group will differ due to random variation
Variance between groups: how different are sample means from each other?
F statistic = variance between groups / variance within groups
if F-statistic much greater than 1:
reject the null!
variance between groups are high and within groups are smaller; within each group, people are relatively similar
larger the F statistic, more likely we are to reject the null
smaller F statistic suggests our data is more consistent with the null
Analysis of Variance, R outputs
If p-value is 1.99e^-12:
probability of getting an F-statistic as large as ours, IF NULL IS TRUE
since p-value is < 0.05, we reject the null: at least one group is statistically different from the others
However, ANOVA by itself doesn’t tell you which group(s) is/are different
Tukey’s post-hoc Test and Interpretation
Post-hoc test: statistical test performed after initial test (ie, ANOVA) to find which pairs of groups are significantly different
Tukey’s post-hoc test: most common after ANOVA, if you reject the null
first, run ANOVA; if reject null:
run post-hoc test
TukeyHSD(result)
HSD stands for honestly significant difference
If p-value between groups are < 0.05, there is a significant difference!
draw significance bars to indicate statistical significance between groups
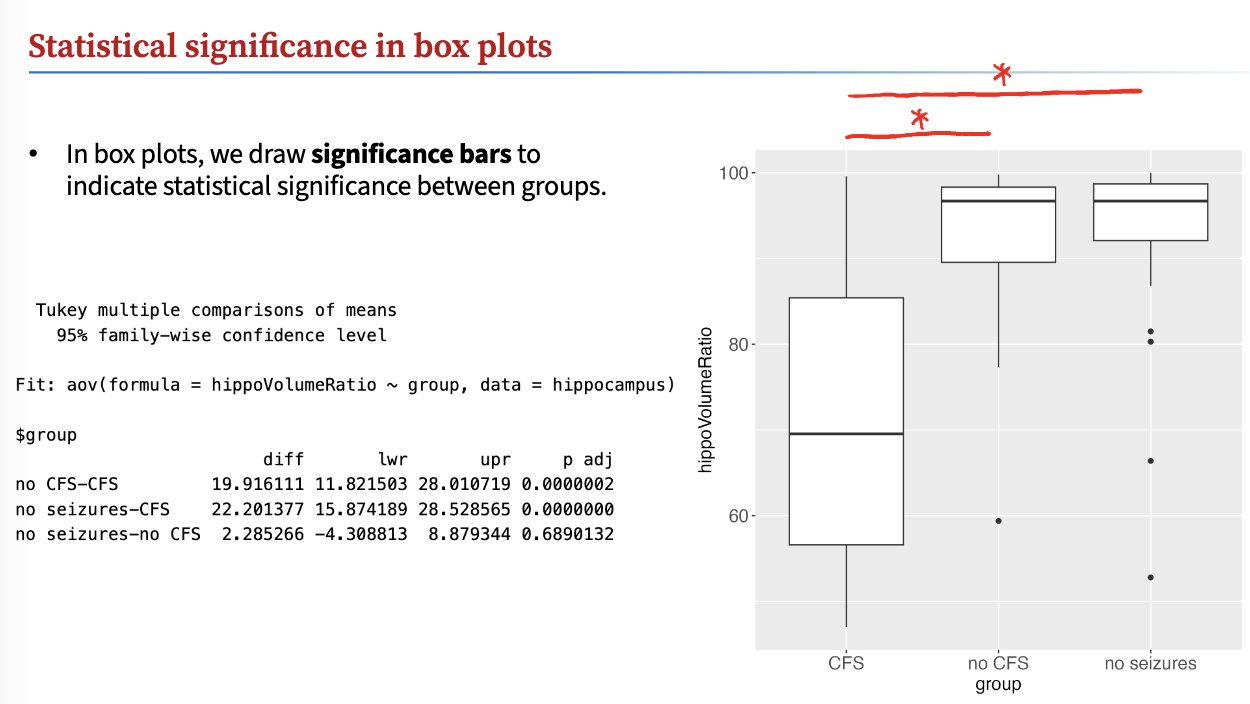

Verifying normality in ANOVA
For ANOVA, we try to verify the data meet the assumptions using AS FEW TESTS as needed
within each group, determine the residuals
Residuals: difference between each data point and its group mean
analyzing residuals can show non-normality
Steps:
1) use aov function and save output to object
2) extract residuals as vector
3) make plot of residuals
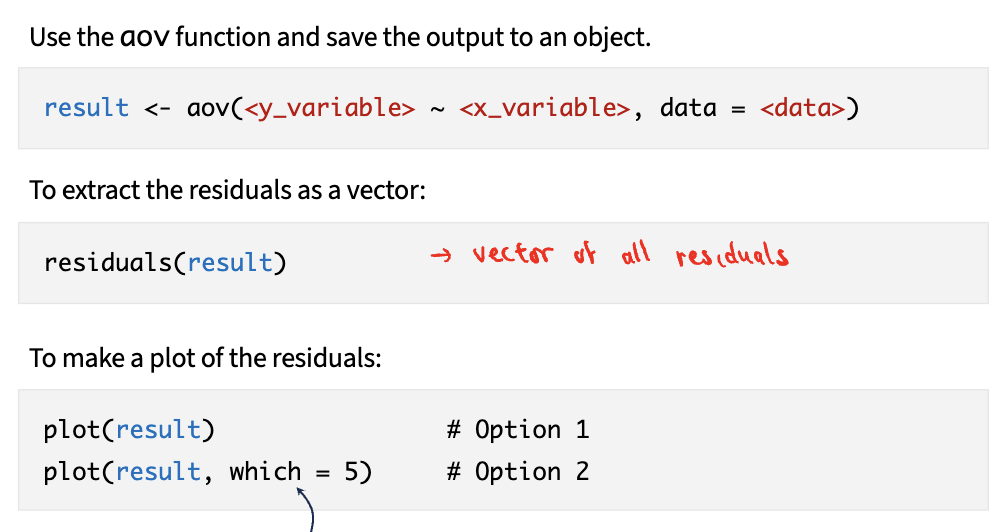

Non-parametric tests
Sometimes, your data may never meet the assumptions of a t-test/ANOVA (parametric test) regardless of transforming
can use non-parametric tests, which do not assume normality
tests other attributes of data, ie median, rank order
generally have lower power
Paired t-test → Wilcoxon signed rank test
2-sample t-test → Mann-Whitney U test
ANOVA with Tukey’s post-hoc test → Kruskal-Wallis test with Dunn’s post-hoc test
Covariance
After finding (x - mean) (y - mean) for every data point, we calculate average
more points in positive quadrants, positive covariance = positive association
more points in negative quadrants, negative covariance = negative association
equally many points in both positive and negative quadrants = zero association
Only sign of covariance is meaningful:
same data, different units → different covariance
covariance changes even though underlying relationship between variables did not change
Pearson correlation coefficient
Normalize covariance by using r, called Pearson correlation coefficient
is not sensitive to units
r tells us if two variables rise and/or fall together
Does NOT imply a causal relationship between two variables!
r only tells us whether two variables rise and/or fall together
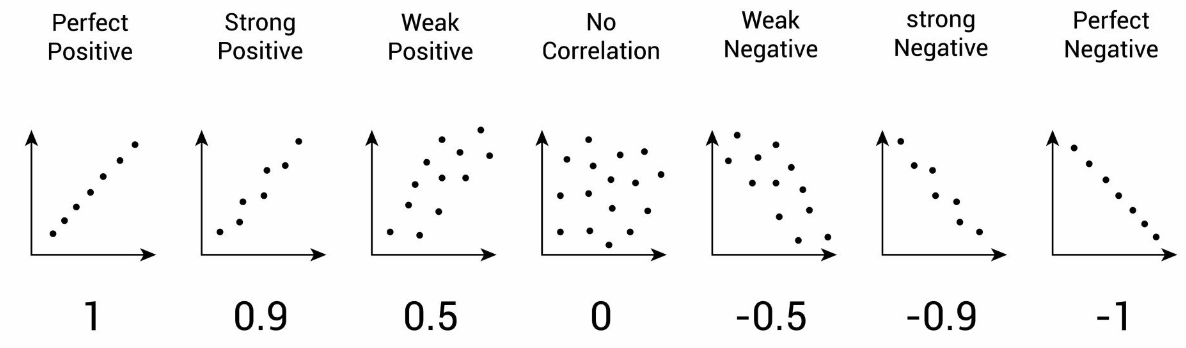
Correlation vs Regression
Correlation: goal is to quantify the association between two numerical variables
NOT trying to fit a line to the data
JUST seeing if two variables rise/fall together
Regression: goal is to predict the value of one variable from another
clearly defined independent and dependent variable
we ARE trying to fit a straight line to the data
Linear model/regression model
Goal: predict the dependent variable from the independent variable
draws straight line through the data, creates equation y = mx + b
Sample statistics (b1 and b0): estimate the population parameters (ß1 and ß0)
class 20
Least Squares Approach
Residual: difference between each y-value and model’s prediction
across all data points, calculate sum of squared residuals
better the model fits our data, the smaller the sum of squared residuals
y-value should be closer to model’s prediction
Best fit line (aka regression line): one that minimizes sum of squared residuals
say this line was obtained using the “least squares approach”
Slope of the regression line
Slope is the biology!
ex. plant mass = 0.92 x nitrogen levels + 10.7
for every 1 ppm increase in nitrogen, plant mass increases by 0.92g
Y-intercept has no biological significance
you should avoid extrapolating too far beyond the data
The better our model, the lower SS(model) is compared to SS(mean)
SS measures error of the model
Lower SS(model) → our model’s predictions are closer to the actual data
class 20
R² and its meaning
R²: proportion of the total variance of y that is explained by x
The sum of square went down from 16.05 → 8.58, 47% decrease!
there is a 47% decrease in error when we take nitrogen levels into account
47% of the total variance in plant mass is explained by the soil nitrogen content
SS(mean) is often called SS(total)
represents the total variability in the y variable
SS(model) is often called SS(residuals)
represents the residuals of the final model

Null distribution of b1 values + comparing t-score with the null
theoretical distribution of all possible b1 values IF THE NULL IS TRUE
due to random sampling, even when the null is true, the slope of our sample (b1) won’t be exactly zero
the larger the magnitude of the t-score, the more likely we are to reject the null
Comparing our t-score with the null
if null is true, we expect the t-score to be close to zero
p-value: probability of getting OUR slope / OUR t-score or a more extreme value of the null is true
if p < alpha, REJECT the null
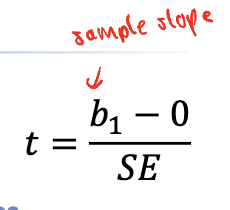
Interpreting linear regression in R
Only care about the slope!
Since p-value of bite slope (11.677) is 0.0393:
We reject the null, there is a relationship that exists between the variables
IF THE NULL WERE TRUE, we have a 0.039 chance of getting a t-score (or a slope) as extreme as ours
p < alpha
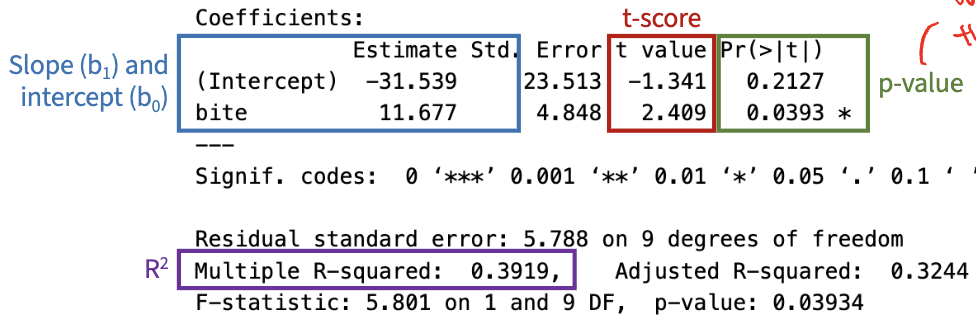
Overfitting
when model tries TOO hard to match the data
even if the model perfectly describes the data we have, it will perform poorly on new, unseen data
we never expect R² to be 1
if you draw a squiggly line, unlikely that it is the true relationship
Comparing Residuals
Residuals: difference between each measurement and model
when relationship is truly linear, some points are above and some are below the regression line
there is no pattern to the residuals; are normally distributed
when relationship is non-linear, there is a pattern to the residuals
residuals are NOT normal
Transforming residuals for regression
1) Plot the residuals and run a KS test on the residuals
if normal, linear regression is valid
if non-normal:
2) make a histogram of the x and y variables
is one variable clearly skewed?
is one variable mostly normal but has a clear outlier?
transform skewed variable or remove the outlier
3) run a KS test on the new model residuals
Chi-squared goodness-of-fit test
Used to perform hypothesis tests on categorical data:
Step 1: calculate a chi-squared (x²) test statistic
Step 2: predict the null distribution of x²
Step 3: compare our value of x² to the null distribution
Basically, compare observed frequency of category to expected frequency of category
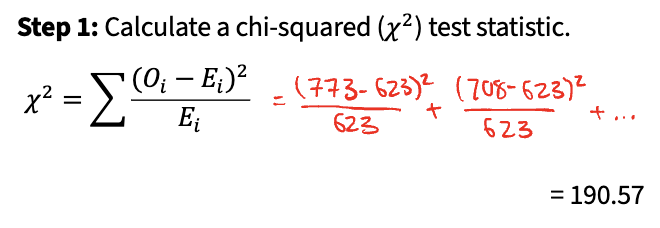
Statistical Tests Summary
One sample t-test: 1 numerical
Paired t-test: two categories; 1 numerical, 1 categorical
Two-sample t-test: two categories; 1 numerical, 1 categorical
ANOVA: 3+ categories; 1 numerical, 1 categorical
Linear regression: 2 numerical
Chi-squared goodness of fit: 1 categorical
Correlation is also appropriate for quantifying association – it does NOT attempt to predict Y from X
Manipulative vs Natural Experiments
Manipulative: experimenter actively controls the independent variable
more directly establish causality between independent and dependent variabels
however, might not be practically and/or ethically feasible
Natural/Observational: experimenter relies on pre-existing differences in independent variable
challenging to implement control group, randomly assigning subjects to treatment vs control groups, and blinding
Random vs Systematic Error
Random: fluctuations in our data that occur just by chance
ie natural variation in blood pressure between individuals
Systematic: error that consistently skews our data in one direction
ie meditation group starts to eat more healthily and exercise more
includes confounding variables, non-random assignment, experimental bias
Minimizing systematic error
Control groups: isolate the effect of the treatment (IV) on DV
to minimize confounding variables, keep control group maximally similar to treatment group
Random assignment: each experimental subject should have equal chance of being assigned to treatment or control group
Blinding: if participants or researcher are aware of treatment, may consciously or subconsciously influence their behavior
Single-blind: participants unaware if they have been assigned to treatment or control
Double-blind: BOTH participants AND researchers are unaware who is assigned to treatment or control
Sample Size and Random Error
If we only give each cream to one person, exposes us to LOTS of random error
any observed difference could be significantly affected by natural variability between the two people
When you increase sample size, you give the skin creams to more people
increasing sample size reduces the influence of random error
averages out random variation across more people
any one individual now exerts a weaker pull on the overall mean
When increasing sample size, standard error of the mean decreases!
smaller n → larger SE
larger n → smaller SE
standard error is a measure of the random error!
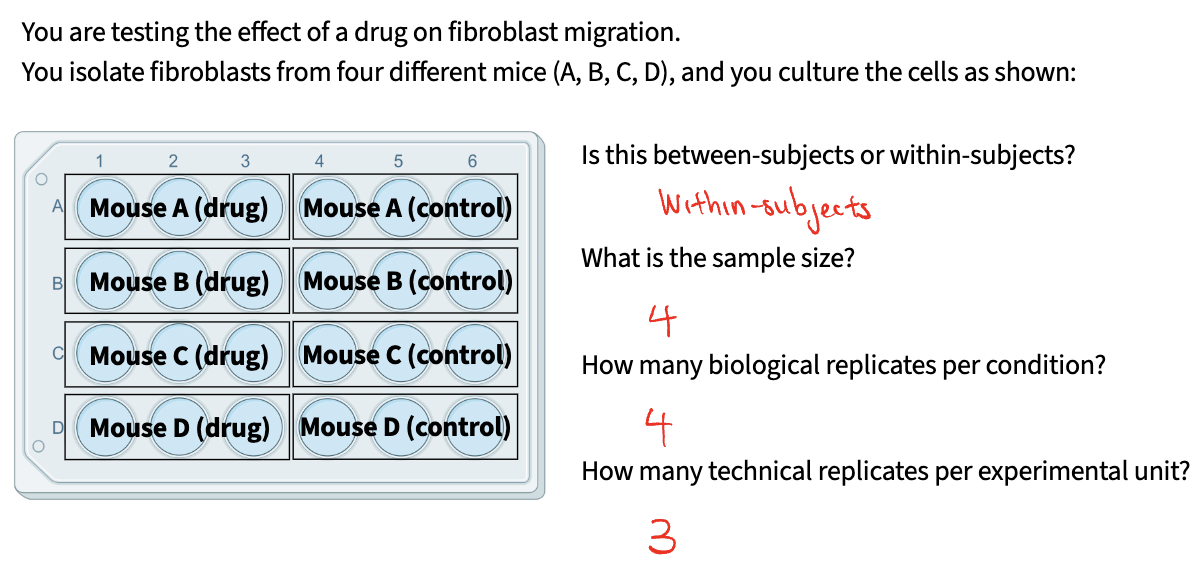
Experimental Unit + Between-subjects vs. within-subjects
An individual or group that is assigned a treatment independently of every other unit
Between-subjects / between-groups: each experimental unit experiences EITHER one condition OR the other
unpaired
independent groups design
Within-subjects / within-groups: each experimental unit experiences BOTH conditions
paired
repeated measures design
Sample size and replicates (biological vs technical)
Sample Size: total number of experimental units across entire experiment
Biological replicates: number of experimental units that experience each condition
captures biological variability
Technical replicates: number of measurements you take per experimental unit
increases precision of measurement
does NOT capture any additional biological variation
Pseudoreplication and how to avoid it
Pseudoreplication: when you erroneously report the sample size as being higher than it actually is
ex. if you take 4 measurements per plant, you think sample size = 6 plants x 4 measurements = 24
not correct! 4 measurements are technical replicates
To avoid pseudoreplication:
Ask yourself: how do I increase sample size?
add more plants!
adding more experimental units / biological replicates
DO NOT take more measurements from each plant!
adding more technical replicants!
increasing sample size = capturing more biological variability
Measurement Validity
Does my experimental system actually address the research question that I’m asking?
To validate, you need additional control groups:
Negative control: checks if experimental system can detect a lack of change when we expect it to
ie mice receive an injection of DMSO solvent WITHOUT a supplement
establish baseline comparison WITHOUT any treatment
Positive control: checks if experimental system can detect a change when we expect it to
ie mice receive an injection of a supplement KNOWN to raise blood insulin levels
verifies our system CAN detect a postive result
For both positive and negative controls, we KNOW what to expect
check experimental system is behaving as it should
Comparing positive and negative controls
If there is no difference between the positive and negative controls, this undermines the validity of our data.
injection was faulty?
technique for measuring insulin is broken or not sensitive enough?
incorrect dosage – all supplements are delivered at too low a concentration?