MBS326R exam 3
1/138
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
139 Terms
Proteobacteria
a domain of bacteria that includes medically and scientifically important species such as salmonella; includes gram negative; facultatively or obligately anaerobic, chemolithoautotrophic, and heterotrophic
nonproteobacteria
a group of bacteria that contain photosynthetic bacteria (photosynthesizing); includes gram positive and gram negative; photoautotrophs and chemoheterotrophs
hyperthermophiles (NON PROTEO GRAM NEGATIVE BACTERIA): aquifex pyrophilus
growth optimum 85 c; rod shaped bacteria; grows best in environments with low oxygen levels
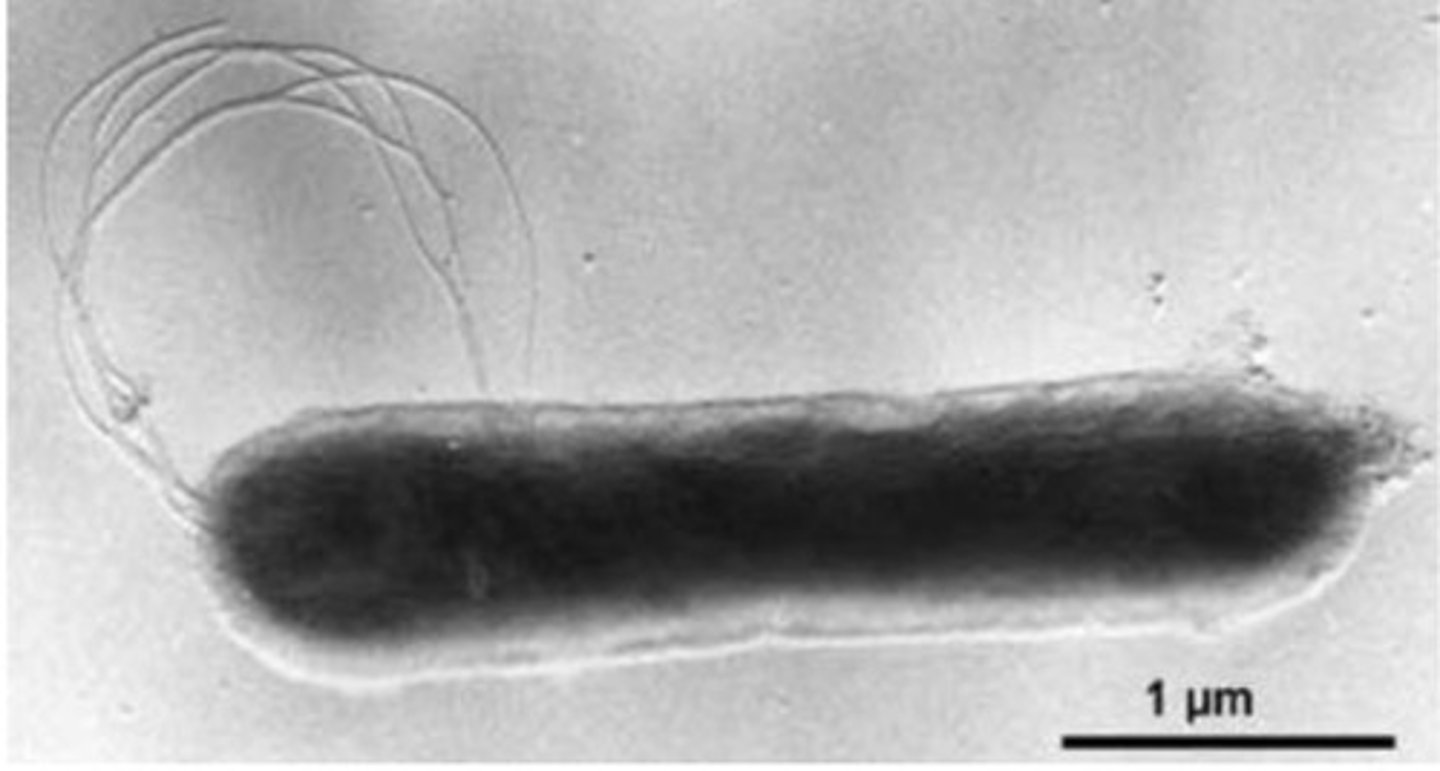
hyperthermophiles (NON PROTEO GRAM NEGATIVE BACTERIA): thermotoga
grow at high temps; rod shaped, wrapped with an outer envelope, toga like structure; can grow on methanol and acetate
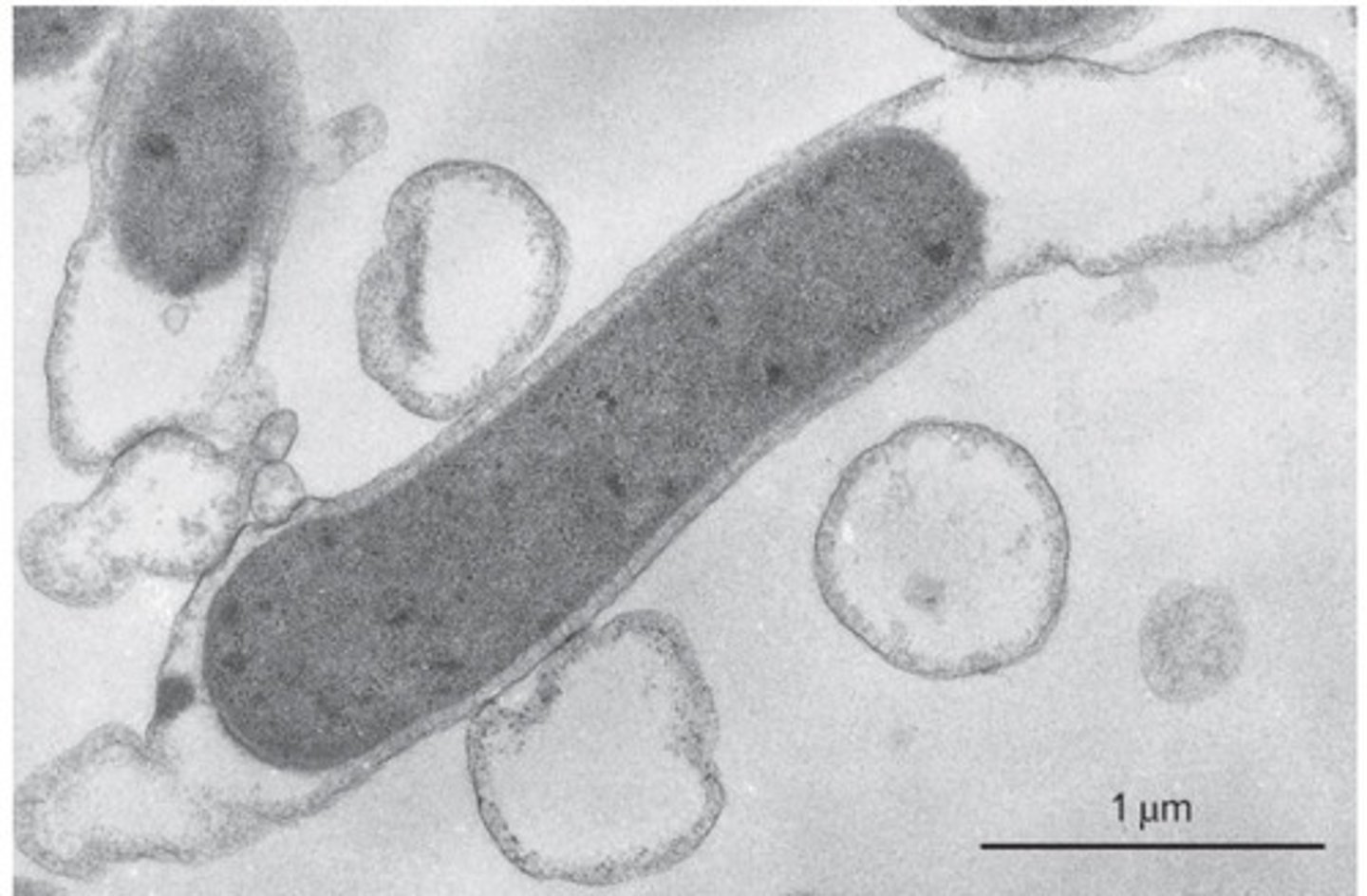
(NON PROTEO GRAM NEGATIVE BACTERIA): deinococcota
tough (resistant to drying out and radiation); can be spherical or rod shaped; stains as a gram positive (purple in color) but is gram negative due to second membrane
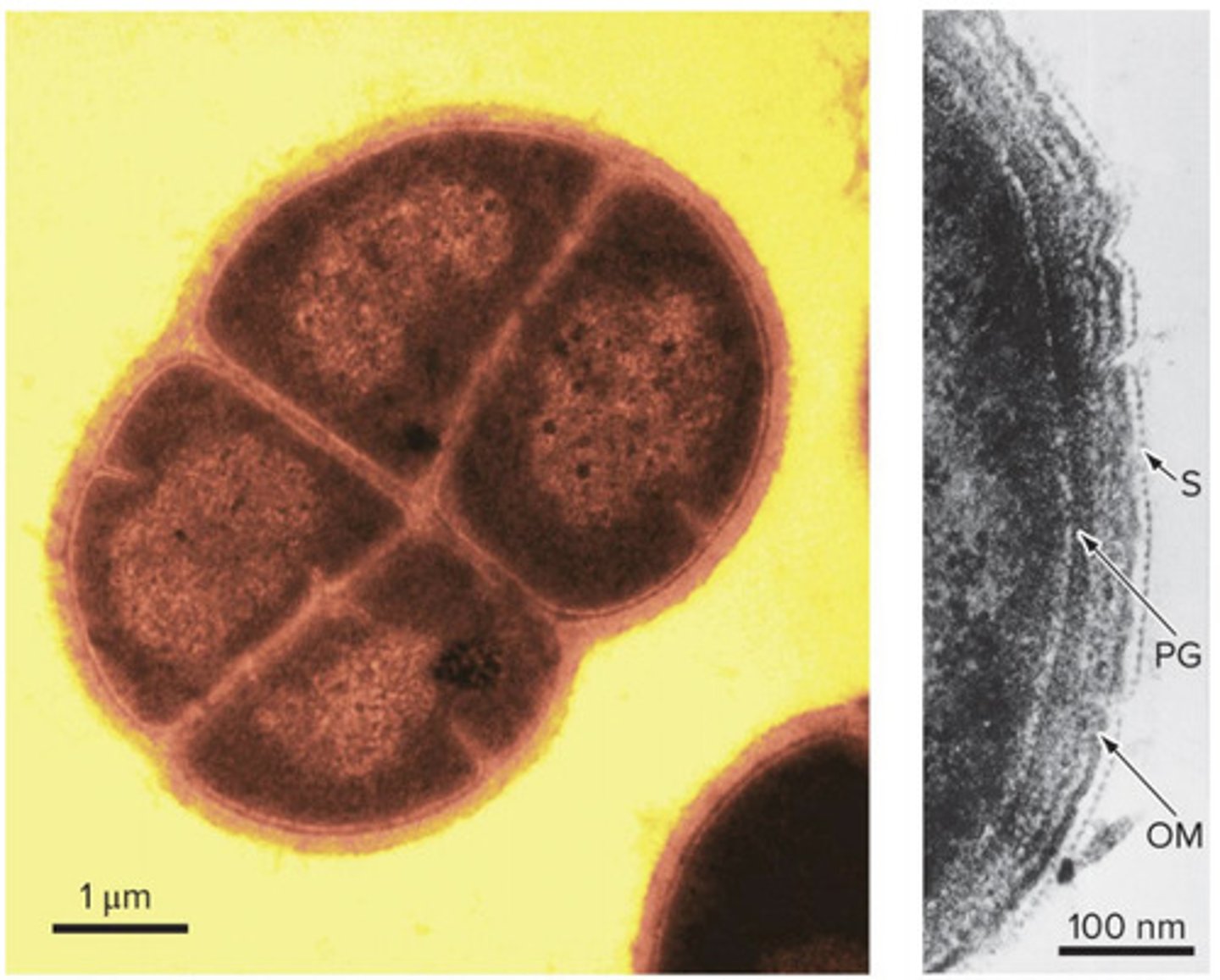
(NON PROTEO GRAM NEGATIVE BACTERIA): photosynthetic bacteria; cyano bacteria
carry out oxygenic photosynthesis; two photosystems; water as an electron donor; generates oxygen; largest and most diverse group of photosynthetic bacteria; light energy converted to chemical energy
(NON PROTEO GRAM NEGATIVE BACTERIA): photosynthetic bacteria; purple, green, and aerobic anoxygenic phototropic bacteria
carries out anoxygenic photosynthesis; one photosystem; alternate electron donor to water (H2 or H2S); live in environments where light is available and oxygen levels are low
(NON PROTEO GRAM NEGATIVE BACTERIA): bacteroidota
includes photolithoautotrophs and chemoheterotrophs; found in gut and oral cavity; up to 30% of human feces
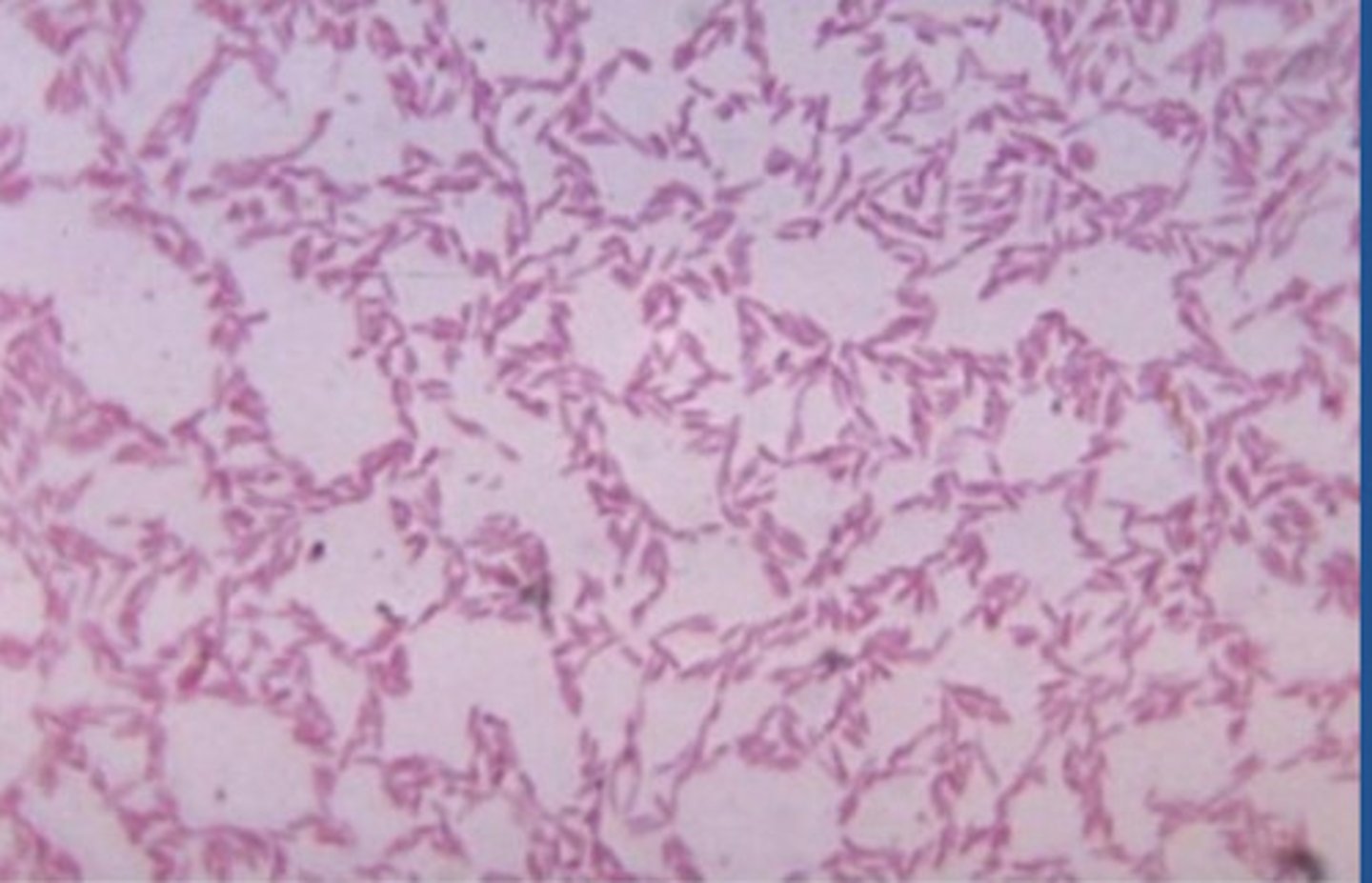
(NON PROTEO GRAM NEGATIVE BACTERIA): fusobacterioa
includes photolithoautotrophs and chemoheterotrophs; found in gut and oral cavity; spindle shape; associated with infections and diseases in humans
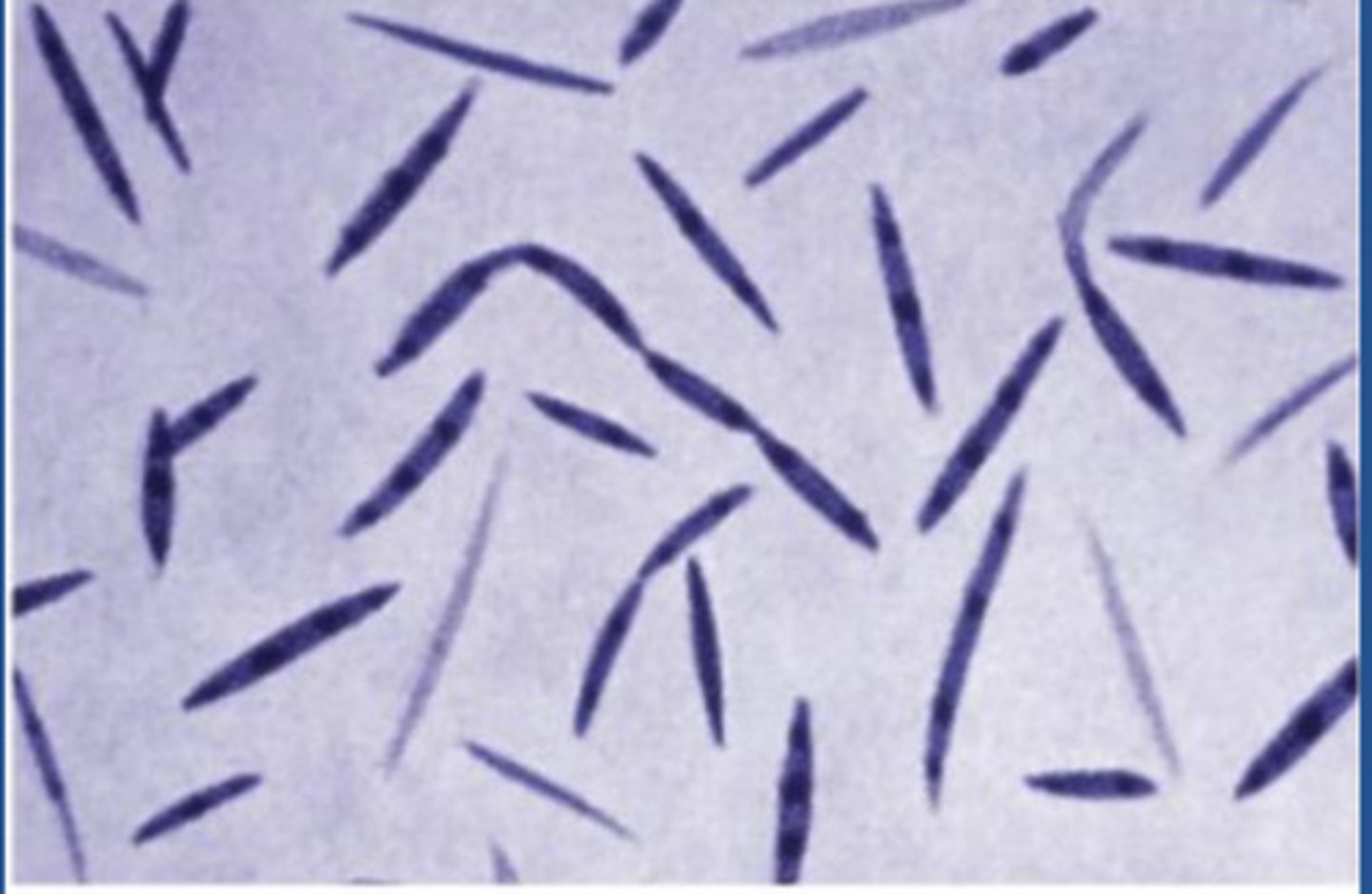
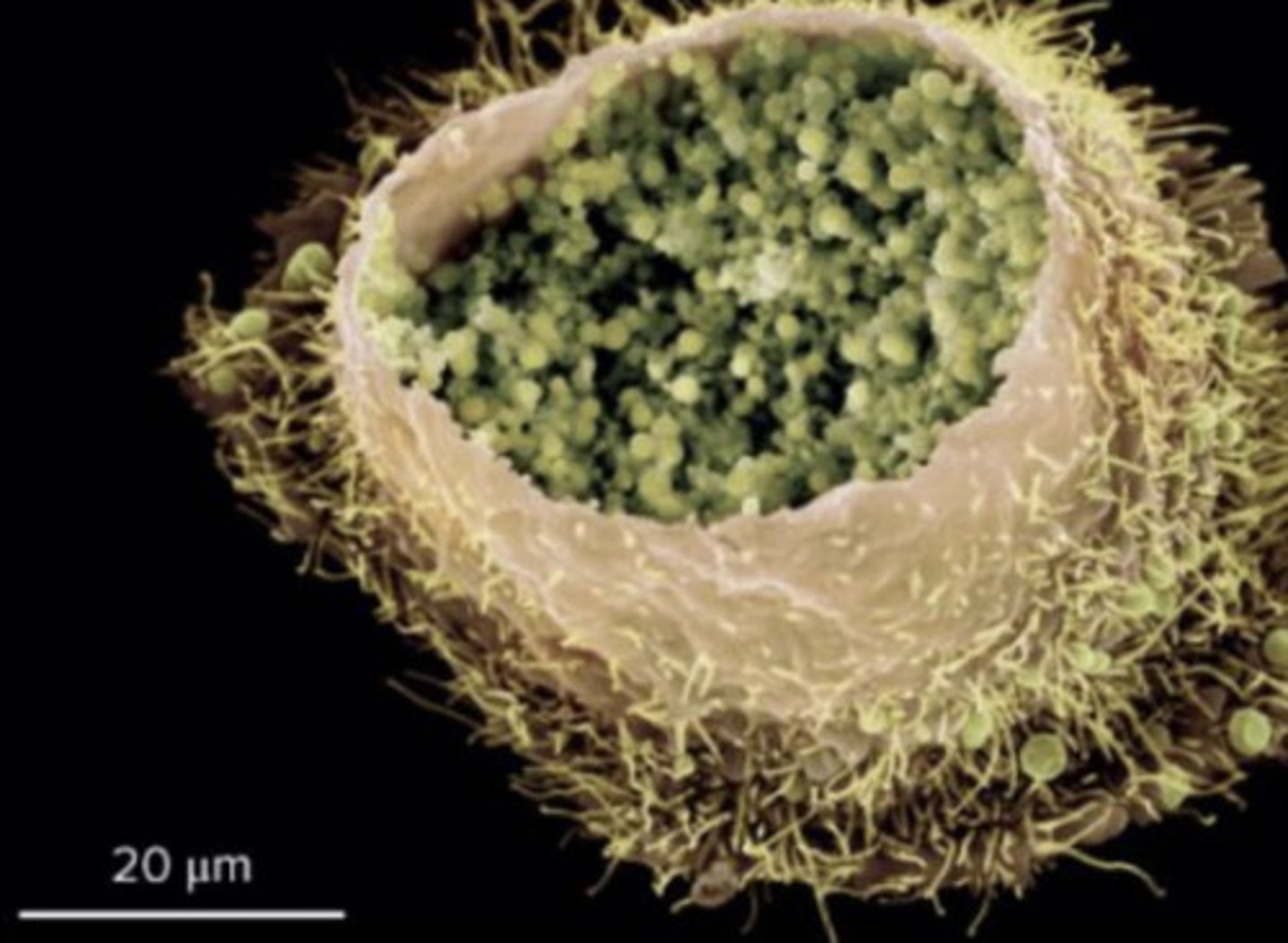
(NON PROTEO GRAM NEGATIVE BACTERIA): chlamydiae
coccoid shape (spherical); non motile (can't move); obligate parasites; small genome so can't metabolize carbs or synthesize ATP or NAD
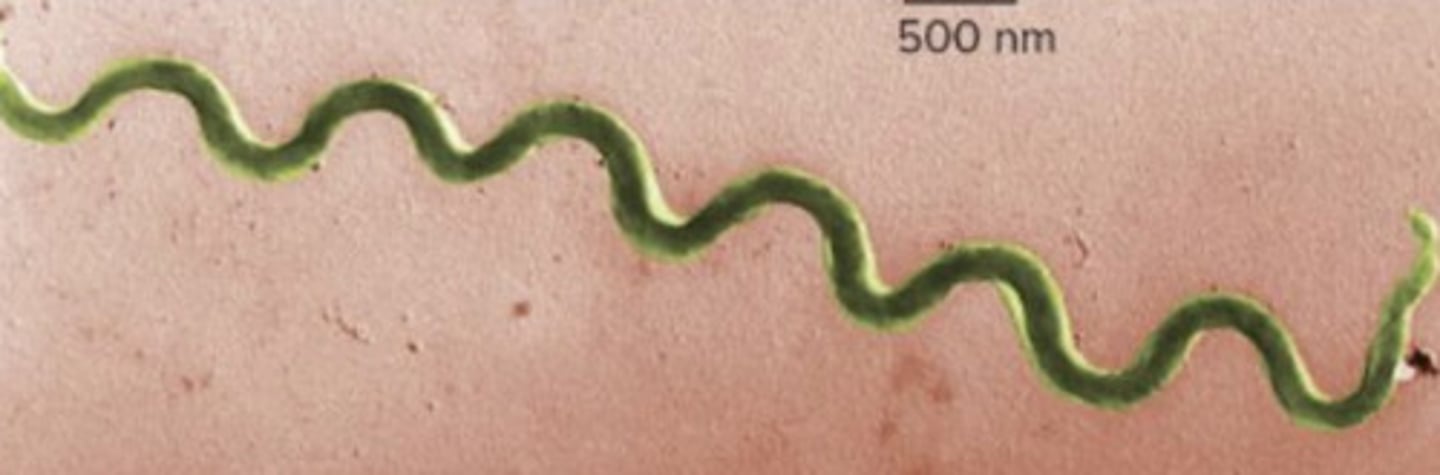
(NON PROTEO GRAM NEGATIVE BACTERIA): spirochaetota
chemoorganotropic bacteria (obtain energy by oxidizing organic compounds); periplasmic flagella; corkscrew like movement over surfaces
(PROTEOBACTERIA): alphaproteobacteria
oligotrophs (low nutrient environments); defined by relationships with nitrogen
example of alphaproteobacteria: rhodobacteria
uses light energy and has rs with nitrogen
(PROTEOBACTERIA): betaproteobacteria
oligotrophic soil bacteria; associated with ammonia oxidation; occupy diverse environments
1st example of betaproteobacteria: thiobacillus
soil, freshwater, and marine habitats; chemolithotrophs (oxidize sulfur compounds)
2nd example of betaproteobacteria: neisseria
non motile, aerobic, cocci; gonorrhea
3rd example of betaproteobacteria: bordetella
aerobic, motile, coccobacilli; chemoorganotrophs; whooping cough
(PROTEOBACTERIA): deltaproteobacteria
anaerobic: sulfur reduction, deep in soil
aerobic: not tied to sulfur; pathogens; related to soil
(PROTEOBACTERIA): gammaproteobacteria
largest bacterial class; based on metabolism and ecological niche; photoliths, enterics, pathogens
1st example of gammaproteobacteria: enterobacteriaceae
facultative anaerobes; chemoorganotrophs; found in soil environments where oxygen is limited
2nd example of gammaproteobacteria: pseudomonadales
motile; aerobic chemoorganotrophs; found anywhere; if you have infection it's most likely this
(PROTEOBACTERIA): epsilonproteobacteria
dealing with stomach/digestive tract
1st example of epsilonproteobacteria: campylobacter
can cause reproductive issues in cattle; can mimic host nerve cells to turn against host
2nd example of epsilonproteobacteria: helicobacter pylori
causes stomach ulcers and gastritis; microaerophile (can't grow under pH 4.5); produces enzymes that burn stomach lining
(FEATURE OF PROTEOBACTERIA): magnetotactic bacteria
means of navigation (Earth's magnetic field); occupy freshwater or marine sediments; highly motile
(GRAM POSITIVE BACTERIA): order actinomycetota
chemoorganotrops; facultative or strict anaerobes (require CO2); make hyphae
1st example of actinomycetota: bifidobacterium
human gut microflora; nonmotile, non sporing, and anaerobic
2nd example of actinomycetota: mycobacteriaceae
packed mycolic acids; hydrophobic and impenetrable to antibiotics bc of thick fat later
3rd example of actinomycetota: streptomycetales
hyphae help produce antibiotics and mineralization; largest bacterial genome
1st example of firmicutes: bacillus
endospore forming rods; produce antibiotics; motile with ton of flagella
2nd example of firmicutes: clostridiales
form heat resistant endospores; responsible for food spoilage; nerve related problems
1st example of cocci: staphylococcus
normal part of skin microflora; cause disease; bunch of circles together; facultative anaerobic, nonmotile
2nd example of cocci: enterococcaceae
normal part of gut microflora; can become opportunistic; round
3rd example of cocci: streptococcaceae
part of gut, respiratory, and mouth microflora; disease causing; grape like structures
baltimore classification system
A system used to classify viruses based upon their type of genome and replication strategy; 7 groups
DNA groups: dsDNA
+/- DNA transcribed to +mRNA, then translated to protein; +/- DNA replicates, then assembles into viral capsid
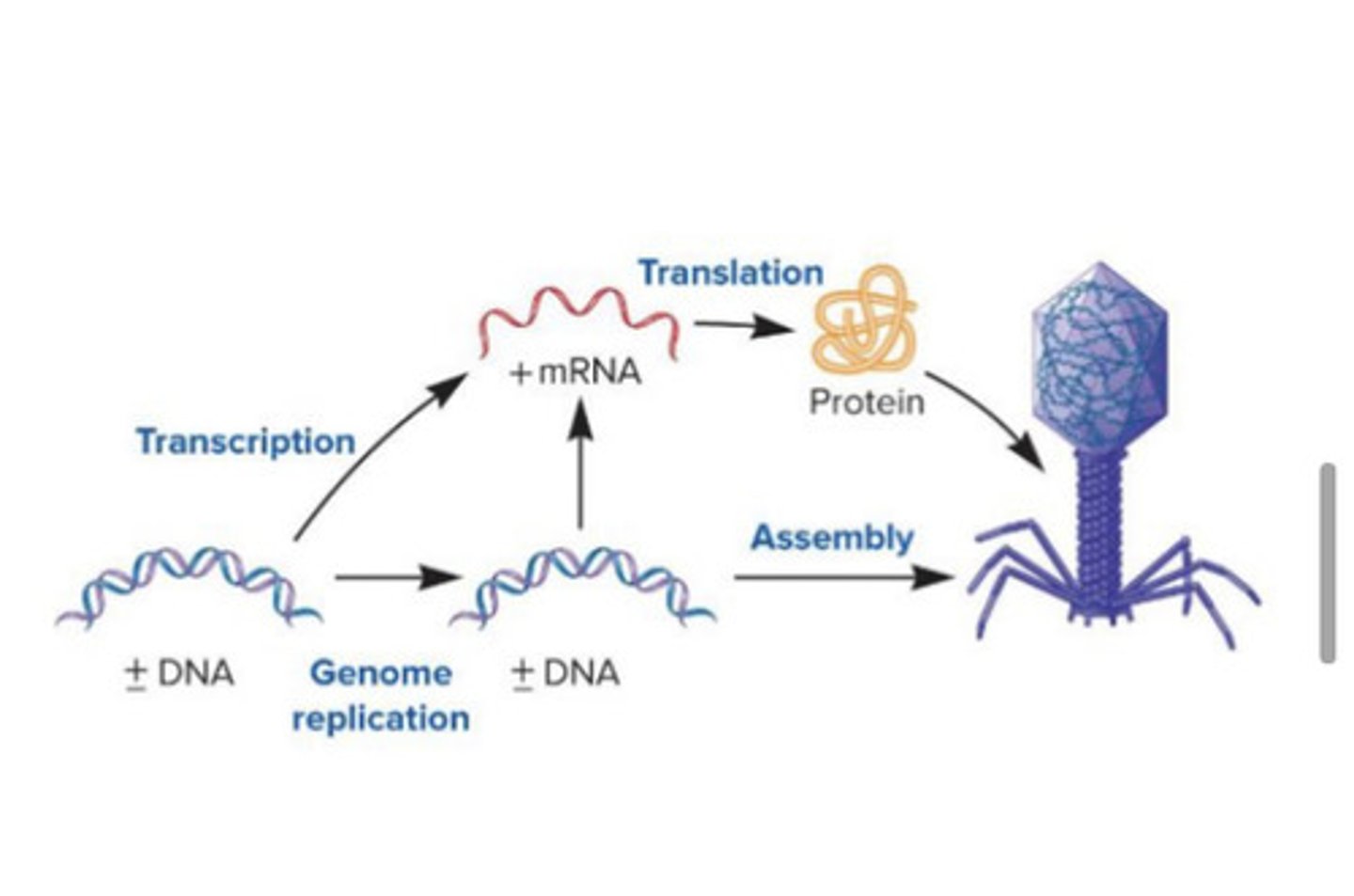
DNA groups: ssDNA
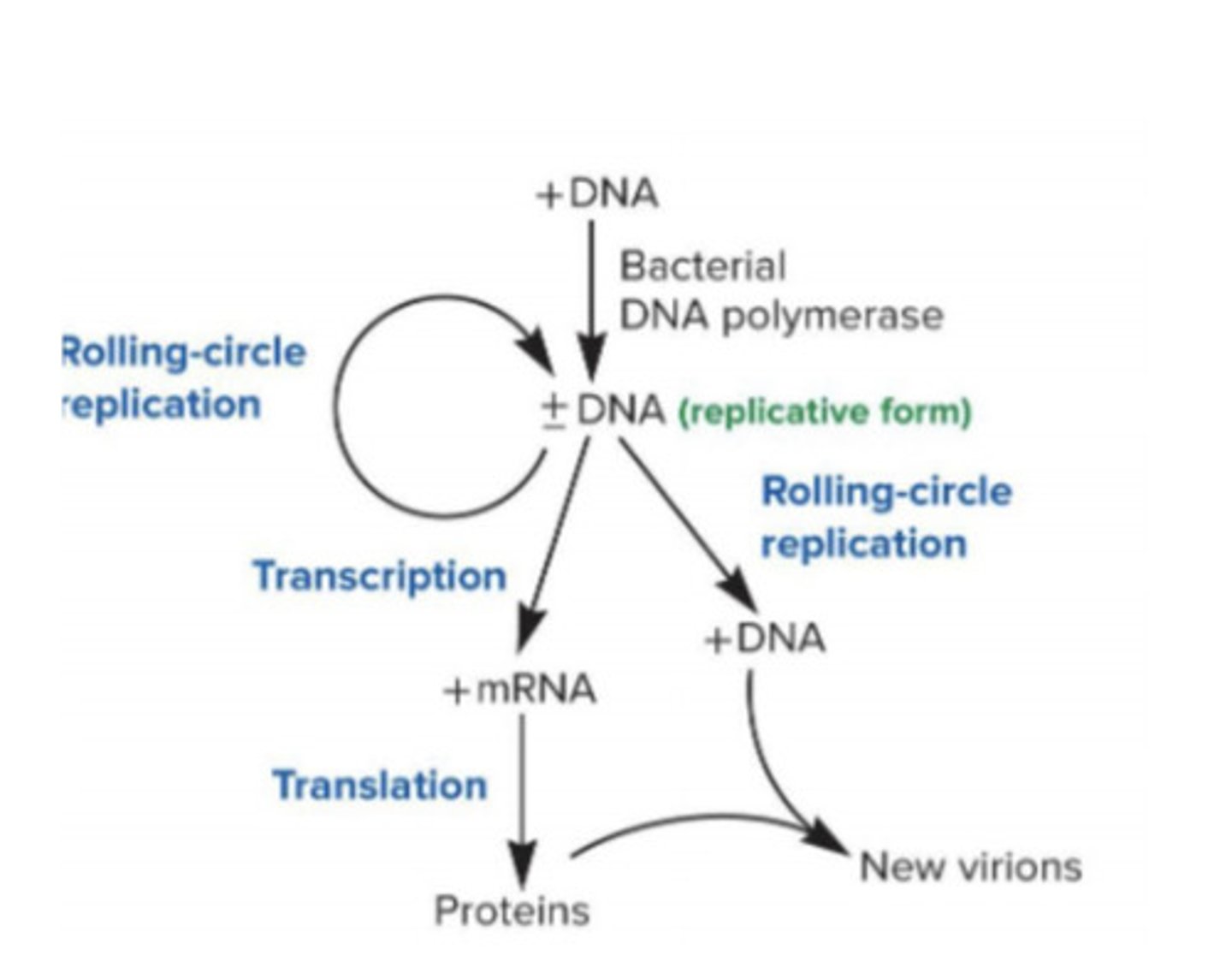
circular +DNA uses bacterial DNAP to synthesize +/- DNA via rolling circle replication; dsDNA then transcribed and translated to proteins to be assembled into viral structure; RCR turns dsDNA back into +DNA to be stored in capsid
RNA groups: dsRNA
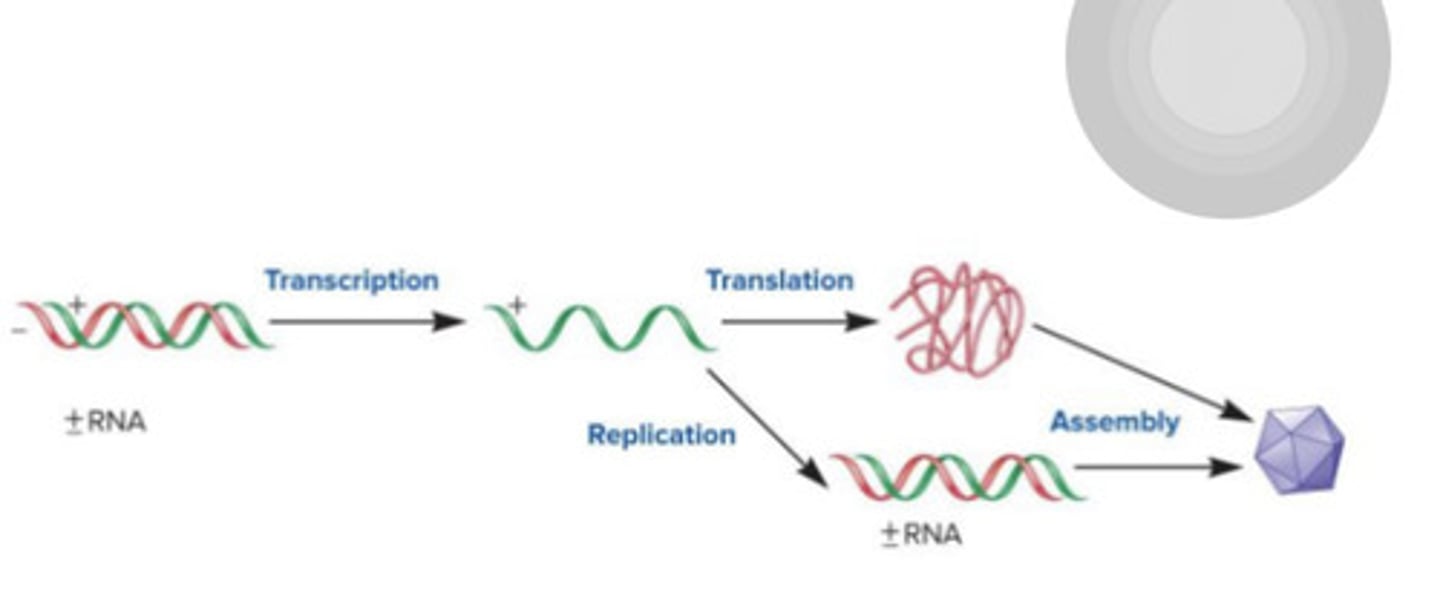
+/- RNA transcribed to +mRNA, to be translated to protein; +mRNA also replicated to create +/- DNA to be assembled into virus; host cells lack the right polymerase so viruses bring in a RNA-dep RNAP which allows for completion of cycle
RNA groups: +RNA
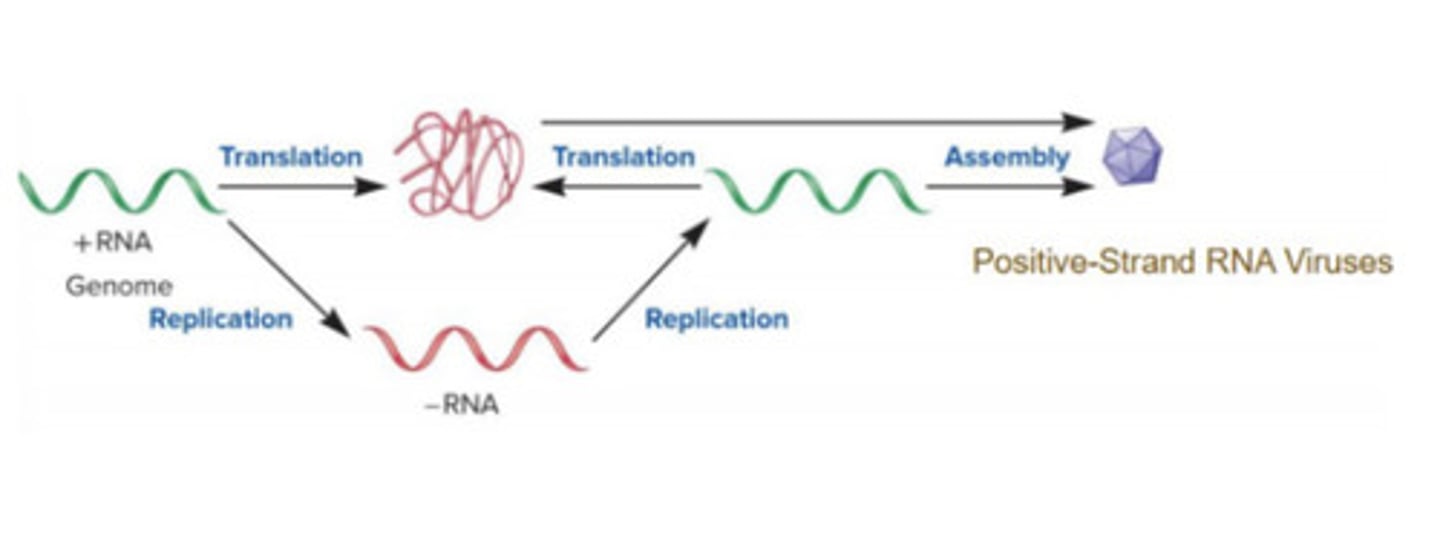
+RNA can be translated right away to create virus; +RNA then replicates to make -RNA, replicating back to +RNA (to produce more copies of +RNA); what is made first is the RNA-dep RNAP to create the -RNA strand
-RNA
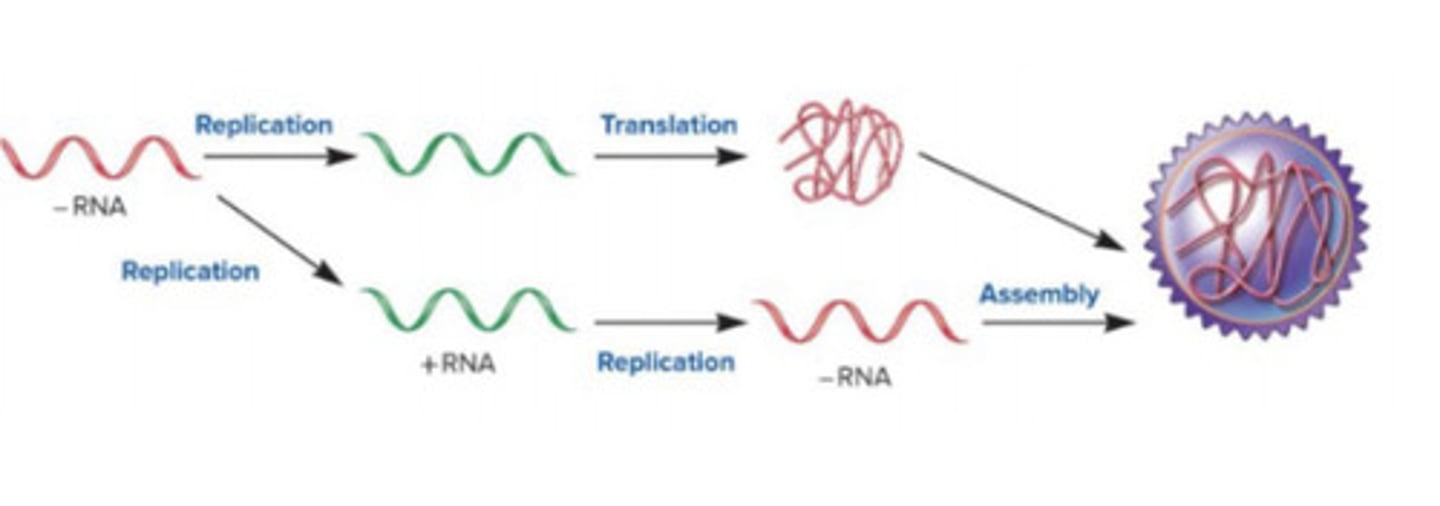
cannot serve as mRNA to make proteins since negative so must bring in a RNA-dep RNAP to make +RNA; -RNA is replicated into +RNA; then translated into protein and replicated back to -RNA to assemble into virus
retroviruses: +ssDNA-RT
+ssRNA is reverse transcribed into -DNA; reberse transcriptase converts ssDNA into dsDNA
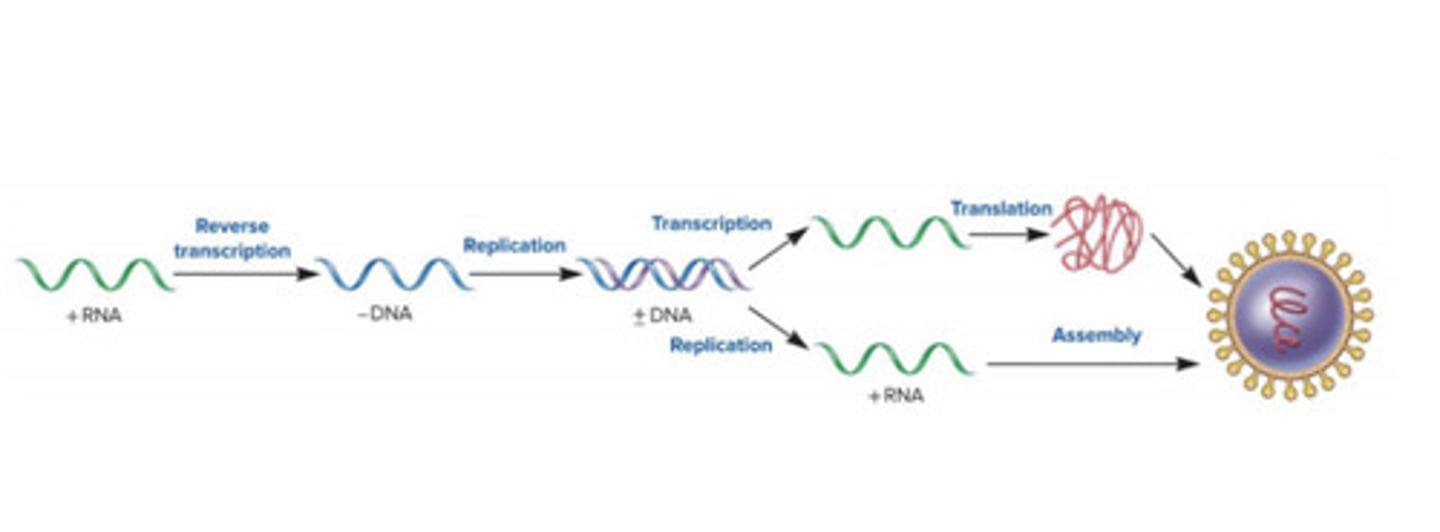
retroviruses: dsDNA-RT
dsDNA is transcribed to ssRNA, then replicated to dsRNA; RT then converts dsRNA back into dsDNA and is then transcribed to mRNA
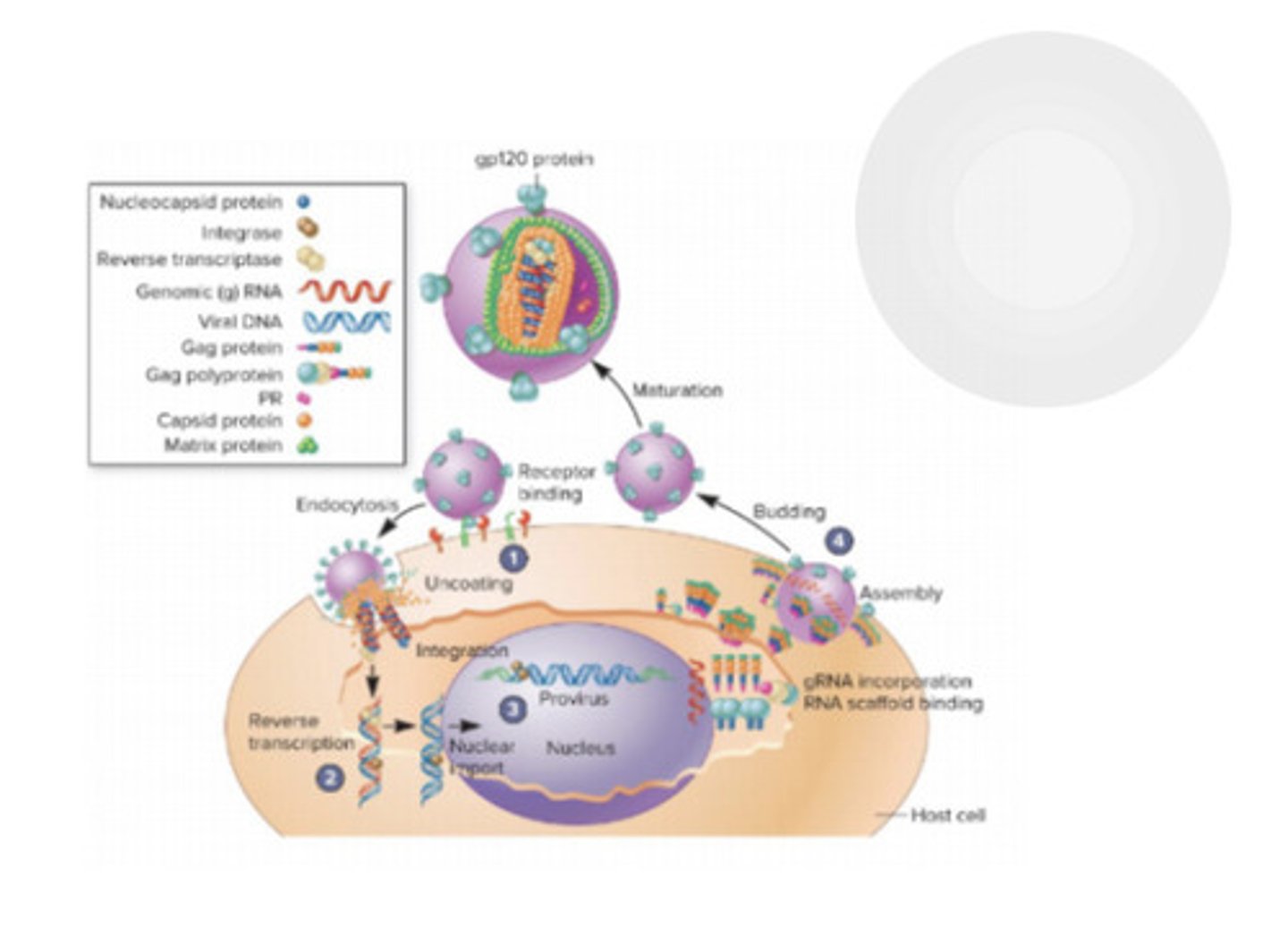
reverse transcriptase
lots of roles; is error prone with no proofreading abilities
T4 = virulent/lytic dsDNA phage
generalized transduction; produce virally encoded DNA-dep DNAP; attachment = phage long tail fiber contacts E coli's outer membrane
Lamba bacteriophage = temperate bacteriophage
specialized transduction; after infecting host can be either lytic or lysogenic cycle; linear dsDNA at the start then circularizes after injecting into host cytoplasm
Latent infection = lysogenic cycle
will occur if high levels of cll/clll; won't show symptoms right away
lytic infection
will occur if no high levels of cll/clll; could show symptoms right away
viruses
most are eukaryotic; can be enveloped or not
bacteriophage
infect bacteria
archaea
extremophiles; microbial dark matter (never been grown in lab)
methanogenesis
last step in anaerobic degradation of organic compounds; if no oxygen available churn out methane; consequence is too much methane
methanotrophs
solves problem of too much methane by oxidizing methane and turning to CO2
examples of archaea
thermoacidophiles, methanogens, haloarchaea
archaea structural adaptations
monolayer; ether stability
archaea metabolic adaptations
sulfur and methane
protista features
plasmalemma (cell membrane); vacuoles; cilia/flagella; free living and in freshwater; grow in environments where oxygen is limited
prions
made from spontaneous generation of mutant protein form; cause neurodegenerative disease; cause proteins to misfold
eukaryotic microorganisms
need certain conditions to survive; include protists and fungi
what differs between eukaryotic cells
cell wall.
plant cells have cellulose and fungal cells have chitin
eukaryotic cells common features
larger than bacterial and archaeal cells; diverse to adapt to environments; cell wall composition is diverse
encystment
formation of dormant cysts in harsh conditions
excystment
escape from cyst in good conditions
trophozoites
actively growing and replicating protists
the protists: plant like protists
possess chlorophyll, perform photosynthesis, and contribute to carbon fixation
examples: diatoms, euglenozoa, chloroplastida
the protists: fungi like protists
decompose organic material, reproduce with spore formation and form multicellular structures
examples: myxogastria (acellular slime molds) and dictyostelia (cellular slime molds)
the protists: animal like protists
differ in terms of motility (pseudopods, cilia, flagellum)
examples: amoebozoa (important in terms of ecological relationships)
fungal groups
decompose organic matter
fermentation, antibiotics
fungal groups: zoosporic fungi
produce motile spores; must infect host cells to complete life cycle; polar tube for host invasion
fungal groups: zygomycetous group
alternative mechanisms of spore dispersal; sexual reproduction occurs when environmental conditions are not favorable; free living and parasitic
fungal groups: dikarya group
most diverse fungal group; may be filamentous or unicellular; always without flagella
fungal groups: dikarya: ascomycota
create tiny spores inside sacs called asci; found in freshwater, marine and terrestrial habitats
white nose syndrome: white hyphae will grow around bat muzzles
fungal groups: dikarya: basidiomycota
club fungi; saprophytes decay plant matter
innate immunity
first line of defense; resistance to any foreign microbe; general mechanisms such as skin, mucus
acquired immunity
if innate system cannot find foreign microbe, this comes into play; tailored to specific foreign microbe; has memory (effectiveness increases on repeated exposure)
antigens
any substance recognized by the immune system; enables immune system to rid the host of foreign invadors
barriers in innate resistance: skin
mechanical barrier to microbial invasion
barriers in innate resistance: mucosal membranes
protective covering that resists penetration and traps microbial invaders
complement system
barrier breached through damage (e.g. cut yourself) turn to complement system. triggers inflammation and recruits WBC; lysing microbial cell membranes; opsonization (tagging microbe for destruction)
"knockout blow"
bone marrow - B cells
originate and develop in bone marrow; await introduction to the antigen to which they will make targeted antibody
thymus - T cells
mature T cells enter bloodstream where they await activation by innate immune cells
mast cells
specialized tissue cells that trigger local inflammatory reactions and are responsible for many allergic symptoms
dendritic cells
in the tissues; responsible for processing foreign matter and presenting it to lymphocytes
natural killer cells and innate lymphocytes
do not express antigen specific receptors; innate response, no memory
endocytosis/phagocytosis
process by which phagocytic cells recognize and kill cellular microbes
recognition of microbe by phagocyte: opsonin dependent
protein coated
recognition of microbe by phagocyte: opsonin independent
microbe associated molecular patterns
inflammation
defense reaction to tissue energy
acute inflammation is immediate response of body to injury
cardinal signs of inflammation
redness, warmth, pain, swelling
granuloma
when phagocytic cells can't destroy pathogen and mass of cells formed in attempt (basically if infection isn't dealt with)
characteristics of adaptive immunity
1. discrimination: between non self and self
2. specificity - activated T and B lymphocytes respond to specific non self antigens
types of adaptive immunity: antibody mediated (humoral)
B cells act and circulate antibodies that bind toxins to destroy them
types of adaptive immunity: cell mediated (cellular)
T cells attack target cells infected with intracellular pathogens
can be CTL or CD8+
recognition of foreignness: endogenous antigen protein
class 1 binds to peptides by sampling proteins in cytoplasm
recognition of foreignness: exogenous antigen processing
class 2 binds fragments that come from antigens outside cell and present to T helper cells
CTLS: effector cells
go to work
CTLS: memory cells
sleep
B cell biology
become plasma cell once activated; made out of membrane bound antibodies
antibodies
don't kill a pathogen but can mess with it's ability (to replicate, etc)
four polypeptide chains: constant C regions
sequence does not vary
four polypeptide chains: constant V regions
form antigen binding sites