Biol 213 Chapter 7
1/111
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
112 Terms
In addition to its role as the genetic material that is inherited each generation, DNA also encodes instructions for
cellular function: DNA makes RNA makes Protein
One big difference between cells is the
type and amount of proteins they contain
This can be regulated at many different steps in gene expression
The chemical structure of RNA differs slightly from that of DNA:
1) Ribose, not deoxyribose. The extra OH makes a big difference!
2) Uracil substitutes for thymine because oxidative deamination of C produces U, and it would not be recognized as a mutation.
3) RNA mostly occurs as a single nucleotide chain, although parts of it can be folded into secondary structures.
RNA molecules can fold into specific structures that are held together by
hydrogen bonds between different base pairs
mRNA
codes for proteins
rRNA
form the core of the ribosome and catalyze protein synthesis
miRNA
regulate gene expression
tRNA
serve as adaptors between mRNA and amino acids during protein synthesis
other small RNAs
used in RNA splicing, telomere maintenance, and many other processes
transcription is key to
• understanding why different cells in a plant or animal have different properties
• understanding how levels of specific proteins are regulated
• understanding how many signals and drugs affect cells
• increasing yields of a protein in biotechnology
• decreasing levels of an enzyme to for metabolic engineering
• thinking of ways to turn off a ‘bad’ gene in medicine
transcription provides ____ of genetic information
amplification
genes can be transcribed with very different
efficiencies
levels of gene expression are tissue-specific
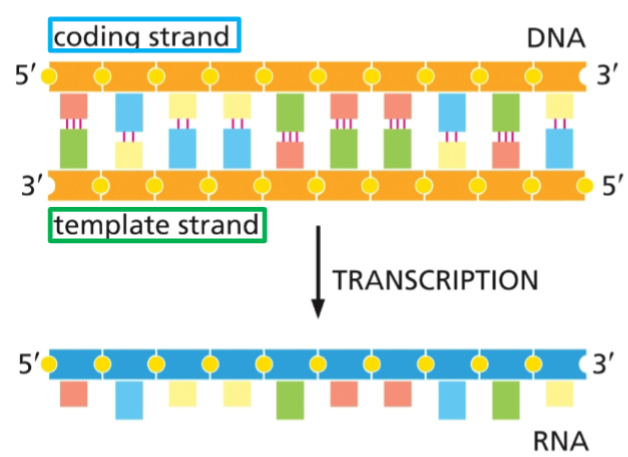
sense, or coding strand
By convention, DNA is drawn with the 5’ end on the left side of the top strand. This places the strand with the same sense as the RNA on top.
The bottom strand (running from 3’ to 5’) is what RNA polymerase actually copies into RNA
start in direction where promoter sequence is
On an individual chromosome, either DNA strand can be
used as a template(but not at the same time)
in transcription, RNA is synthesized by
RNA polymerase
Nucleotides added to 3’ end of RNA strand (5’ to 3’)
RNA sequence is dependent on complementary base pairing (A-U, G-C)
no primer, helicase, or topoisomerase needed
RNA polymerase error rate is
in 104 Compare to DNA pol
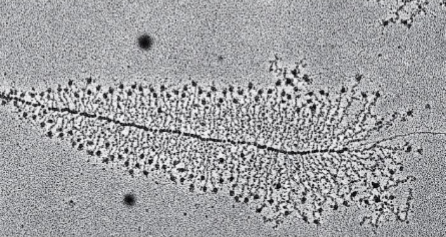
Genes can be simultaneously transcribed by many
RNA polymerase molecules
why is this a bacterial cell
because of the dark little dots connected to mRNA and many ribosomes present
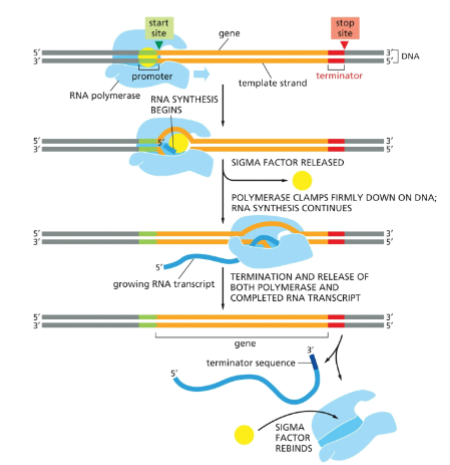
What controls where RNA polymerase initiates and terminates transcription?
promoters and terminators
promoter
DNA sequence that is recognized by RNA polymerase as a start point.
Chain elongation occurs until RNA polymerase reaches a _____, at which RNA is released and the RNA polymerase dissociates from the DNA
terminator site
almost always transcribed, allows termination
similarities for transcription in bacteria and eukaryotes
inititation
elongation
termination
what do bacteria have in transcription that eukaryotes dont
sigma factor (eukaryotes have transcription factors instead)
sigma factor
subunit of RNA polymerase Responsible for the recognition of the promoter sequence and the tight binding of the RNA polymerase to the DNA
Producing mRNA molecules is more complex in
eukaryotes
Multiple stages of gene expression in eukaryotes. Each stage offers opportunities to regulate expression.
Which stage of gene regulation occurs before transcription? epigenetics regulation
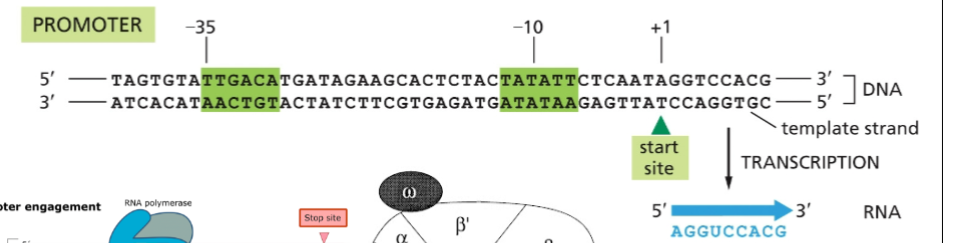
Bacterial Transcription Initiation
The sigma subunit of bacterial RNA polymerase recognizes the -10 and -35 sequences and directs the rest of the RNA polymerase to bind there. Sigma factor is released after transcription begins.
ex. -10 = 10 bases before we start
negative because zero is first base at start
Eukaryotic promoters contain sequences that promote the binding of the general transcription factors. The only one we will worry about is the
TATA box at -30, relative to the transcription start site
Eukaryotic Transcription Initiation
RNA Pol II requires general transcription factors
TATA-binding protein (TBP) is a subunit of TFIID; involved in the recognition of the promoter
Assembly of transcription initiation complex; enables the recruitment of RNA Pol II
Phosphorylation of RNA Pol II by TFIIH, releases RNA Pol II from the transcription initiation complex and allows transcription proceeds
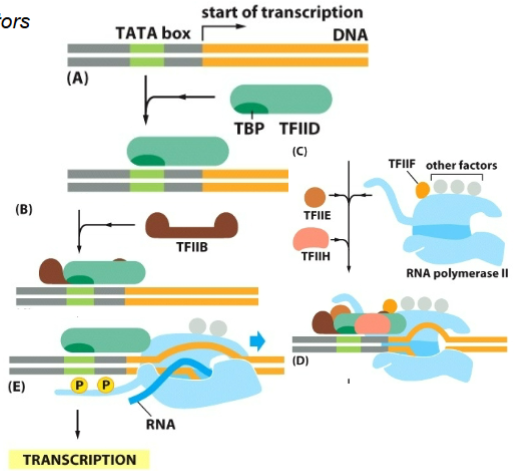
transcription is not initiated without
TFIIH (transcription factor 2)
transcription factor 2 order
D first then that recruits B then H
Transcriptional Elongation
RNA Polymerase Unwinds DNA to access the template strand
• Only exposes ~10-20 DNA nucleotides at a time.
• Connects RNA nucleotides using DNA as a template.
• Produces the RNA transcript in a 5’ to 3’ direction
• Typically producing the RNA transcript at ~40 nucleotides per second
Bacterial transcriptional terminators often have extensive secondary structure just before the
stop side
the mechanisms of termination are different in
prokaryotes and eukaryotes
prokaryotic termination
• Rho- dependent or independent termination.
• Rho – factor or G-C hairpin
• RNA polymerase reads through a “termination sequence”
• This cause the RNA polymerase to dissociate from the DNA.
• The RNA is immediately ready for translation….boom
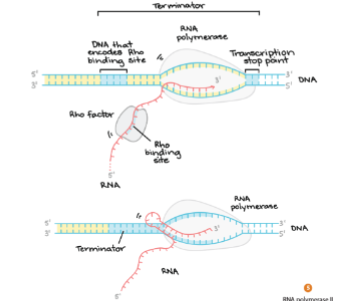
Eukaryotic termination
RNA polymerase reads through a special termination sequence known as a Polyadenylation sequence (AAUAAA)
The end of RNA transcript is then bound by proteins causing the RNA polymerase to dissociate from the DNA
after termination, the RNA needs additional processing
RNA polymerase during transcription for prokaryotes
have a single type but differing sigma factors for specificity
RNA polymerase during transcription for eukaryotes
have three types
RNA pol I: most rRNA genes
RNA pol II: protein-encoding genes (makes mRNA)
RNA pol III: tRNA, 5S rRNA, small structural RNA genes
initiation during transcription for prokaryotes
RNA pol has the sigma factor protein subunit
initiation during transcription for eukaryotes
RNA pols require general transcription factors
Transcript processing during transcription for prokaryotes
transcripts are generally NOT processed
Transcript processing during transcription for eukaryotes
mRNAs are processed
is Packing of DNA into nucleosomes in eukaryotes or prokaryotes
eukaryotes
Transcription takes place in the ___ in eukaryotic cells
nucleus
translation (protein synthesis) takes place in the _____ in eukaryotic cells
cytosol
Eukaryotic RNA must be transported from nucleus to cytosol before
leaving the nucleus, mRNA must be processed
RNA capping in eukaryotic cells
modification of 5’ end, 7-methylG
Polyadenylation in eukaryotic cells
modification of 3’ end, polyA tail
longer tail= shorter time
splicing in eukaryotic cells
removal of introls
Prokaryotic RNA is generally not
processed
Phosphorylation of RNA Pol II by TFIIH also allows
RNA processing proteins to assemble on its tail
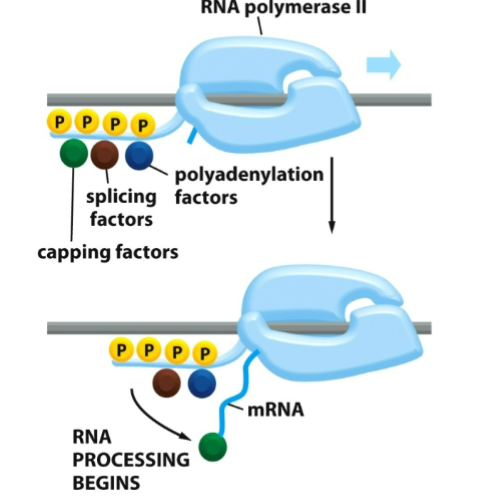
Capping and polyadenylation increases
stability of eukaryotic mRNA molecules
Gene organization differs between eukaryotes and prokaryotes
For most bacterial genes, the DNA sequence that encodes them is co-linear with the RNA that is produced. In contrast, most eukaryotic genes have non-coding, intervening sequences, or introns, that must be removed before a functional mRNA is made.
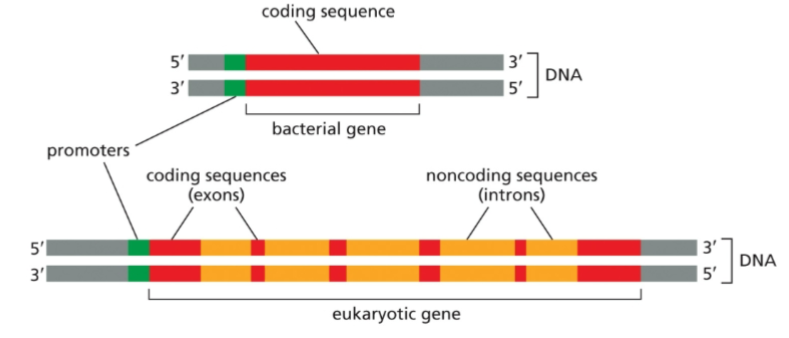
Introns are removed by
RNA splicing
An entire gene is transcribed (exons plus introns) but removal of the introns begins immediately after
capping occurs
Capping and splicing both occur while RNA polymerase continues to transcribe DNA
RNA splicing is performed largely by
catalytic RNA molecules (small nuclear RNAs, snRNAs) with help from a few proteins (snRNPs). Spliceosome
The splicing RNA molecules recognize intron- exon boundaries or junctions (sequence-specific recognition – only a small amount of bases) and cut out introns as a lariat structure
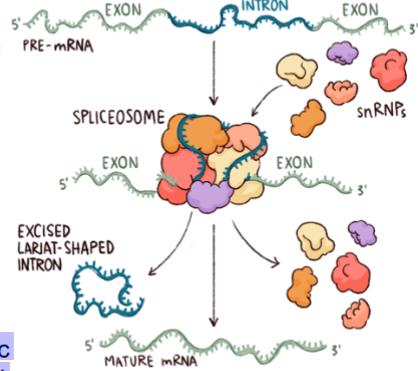
spliceosome
ribonucleoprotein complex
The sequence of only a few small parts of an intron are critical for
splicing
An intron in a pre-mRNA molecule forms a branched structure during RNA splicing, and is removed by a complex structure called a
spliceosome
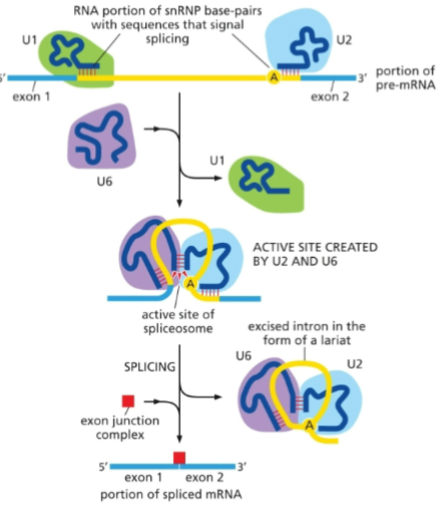
Small nuclear ribonuclearproteins (SnRNPs) carry out
different stages of intron splicing
Exons are ligated back together precisely and marked with proteins (red box) to indicate
splicing has been completed
Some pre-mRNAs undergo alternative RNA splicing to produce
different mRNAs and proteins from the same gene.
ex. striated muscle mRNA, smooth muscle mRNA, brain mRNA, etc.
rRNA is transcribed and processed in the
nucleolus
Ribosomes and mRNAs travel from the nucleus to the cytoplasm through
nuclear pores
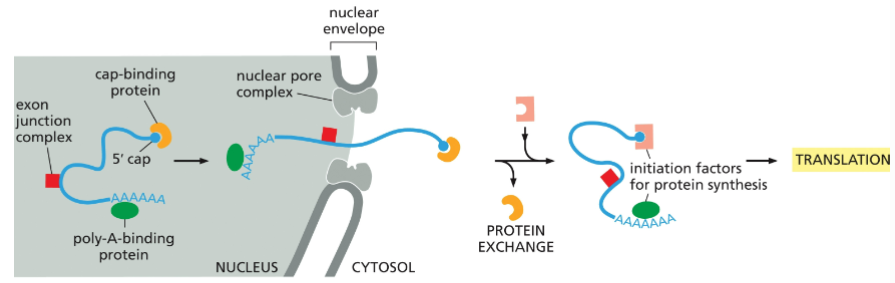
Mature mRNAs are selectively exported from the nucleus
Only mature mRNA is exported - no introns, broken strands or incorrectly spliced variants. These are distinguished by the binding of SSspecific proteins associated with each step of RNA processing:
• polyA-binding proteins
• Cap-binding complex
• Nuclear transport receptor
In bacteria, transcription and translation are closely coupled because ribosomes can
access mRNAs are they are being transcribed
translation is key to
• understanding, and predicting the effects of, many mutations
• genetic engineering - making a precise modification to a protein
• understanding the mechanism of action of many antibiotics
template for translation
mRNA
template for transcription
DNA
There are no tRNA for stop codons, thus
61 tRNA
The trinucleotide code has
redundancy Third position “wobble”
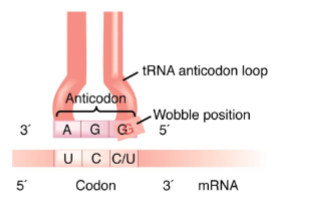
all amino acids are encoded by multiple codons except for
Methionine and Tryptophan
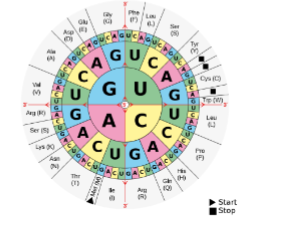
how do you read this
read inside to out
stop codons
3 total
UAA
UAG
UGA
Transfer RNAs (tRNAs)
adapter molecules that link codons with amino acids
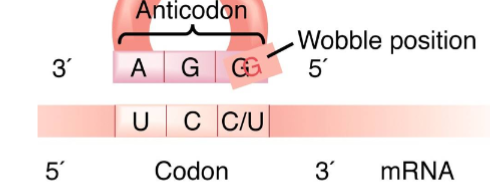
hydrogen holds the
anticodon to codon bonds
The wobble position of a codon is its third base, but this corresponds to the
first base of the anticodon.
remember the antiparallel annealing of oligonucleotides
transcription
is only the addition of nucleotides
tRNA adds anticodons from
3’ to 5’ while mRNA is in 5’ to 3’
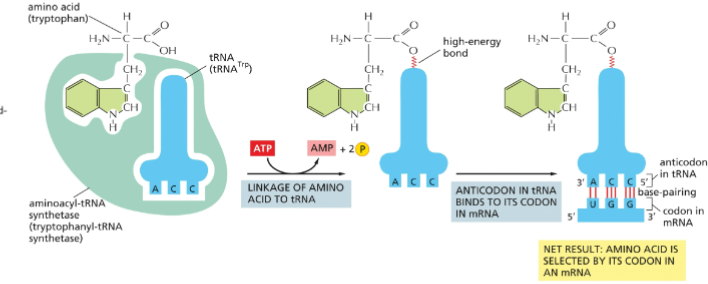
Aminoacyl tRNA synthetases
bind to tRNAs and charge them with the appropriate amino acid.
tRNA synthetase
recognizes nucleotides at the anticodon and the 3’ amino acid-accepting arm to provide specificity. Need a different enzyme for each amino acid
The ribosome is a
ribonucleoprotein complex
eukaryotic ribosome
80S
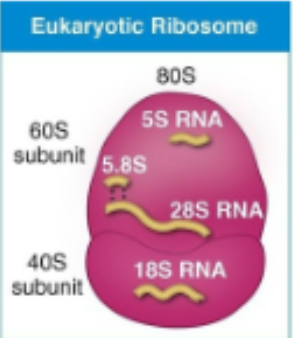
prokaryotic ribosome
70S
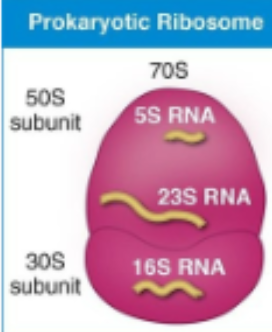
Protein synthesis takes place at the
ribosome
The two ribosomal subunits come together on an ___ during protein synthesis
mRNA molecule near its 5’ end (small and large subunits)
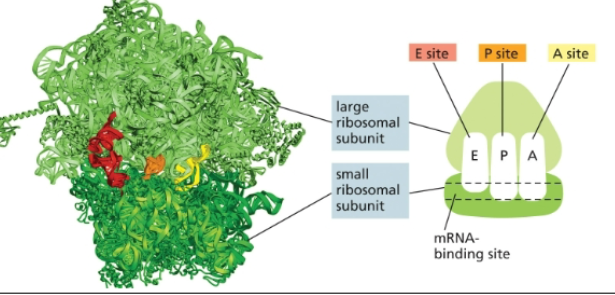
Small ribosomal subunit
matches tRNAs with codons
large ribosomal subunit
catalyzes formation of peptide bonds
Three binding sites of protein synthesis
A site
P site
E site
A site (aminoacyl-tRNA)
charged tRNA binds to its codon
P site (peptidyl-tRNA)
condensation of amino acids
E site (exit)
where the “uncharged” tRNA is ejected
RNA can be translated in three possible reading frames depending on
where the decoding process begins
this means DNA can provide 6 possible reading frames
the site of translation inititation is crucial!
the first peptide bond is formed in
translation elongation
Translation takes place in a
four-step cycle, which is repeated over and over during the synthesis of a protein
ribozyme
an RNA molecule that possesses catalytic activity
Ribosomes are ribozymes, because the catalytic activity that forms peptide bonds is the rRNA molecule
Proteins are synthesized on
polyribosomes
It is not an accident that these look like circular mRNAs, as proteins at the 5’ cap and the 3’ polyA tail interact with initiation complexes to ensure ribosomes translate only intact mRNAs.
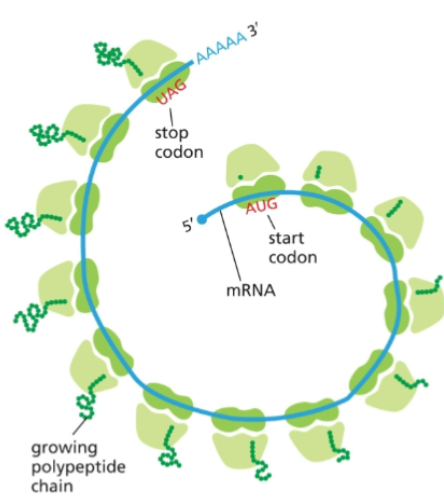
Release factors bind stop codons triggering the release of
the newly synthesized protein
the ribosome then dissociates from the mRNA
translation initiation study guide
• Translation begins with the codon AUG, which codes for methionine.
• An initiator tRNA molecule charged with methionine (distinct from the other methionine-carrying tRNA) binds to the P site of a small ribosomal subunit. It is the only tRNA that can bind directly to the P site, and this binding is assisted by proteins called translation initiation factors
• The loaded ribosomal subunit binds to the 5’ end of an mRNA, recognizing the cap, and moves 5’ to 3’ along the mRNA until it encounters an AUG codon (Kozak sequence).
• The initiation factors dissociate making room for the large subunit to bind forming a complete ribosome
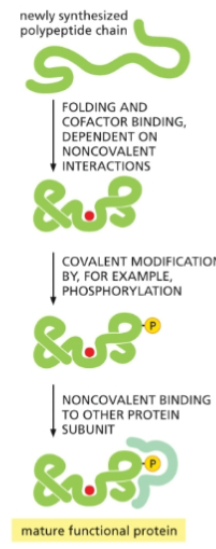
Many proteins require post-translational modifications to become
fully functional