MAME Final
1/196
Earn XP
Description and Tags
Chapters 5-8
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
197 Terms
Taxonomy” and “Phylogeny” in prokaryotes based on the sequence of _____
the 16S rRNA gene (the rDNA sequence)
What are the steps to DNA analysis?
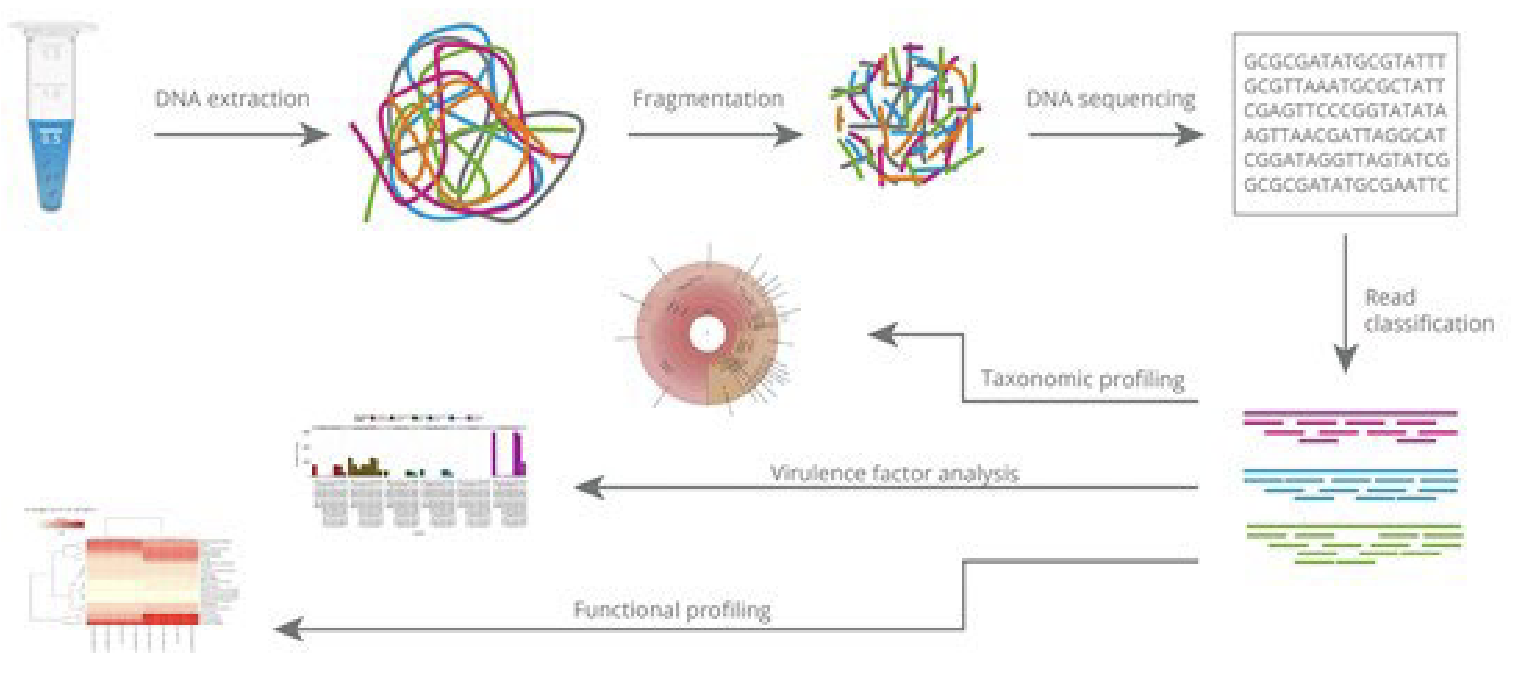
What happens in metagenomics?
Genetic materials (DNA, C) are extracted directly from samples taken from the environment (e.g. soil, sea water, human gut)
Filtering
Sequencing
Multiplication by cloning (shotgun sequencing)
These short sequences can then be put together again
using assembly methods to deduce the individual genomes or parts of genomes that constitute the original
environmental sample.This information can then be used
to study the species diversity and functional potential of
the microbial community of the environment
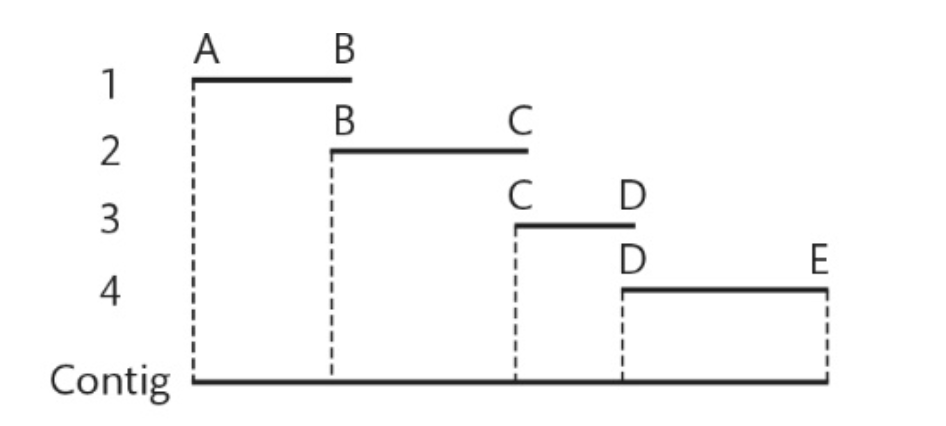
Assembling a contig from overlapping sequences.
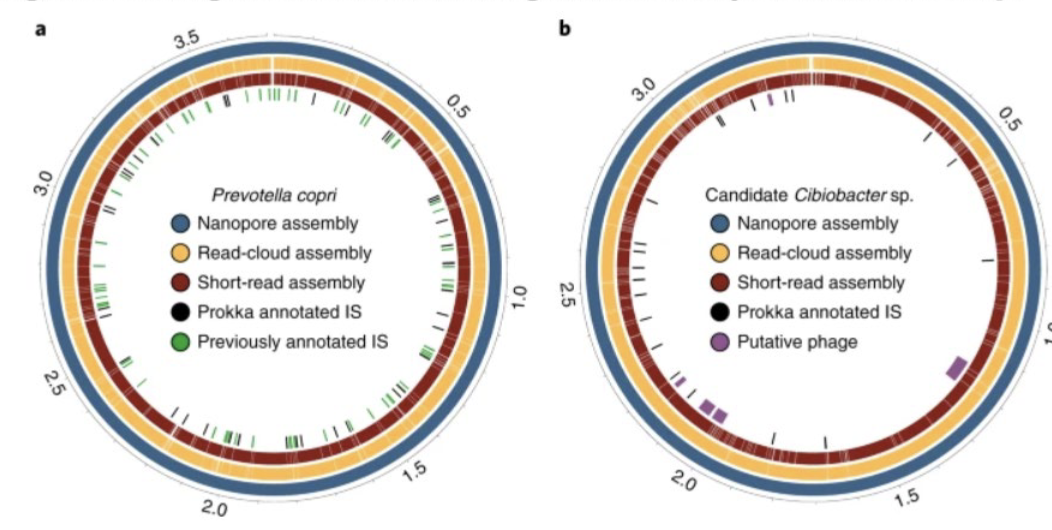
What do these rings represent?
In both plots, the outermost ring represents the complete, closed and circularized genome of the given organism. The middle and inner rings represent contigs from the corresponding read-cloud and short-read assemblies, respectively, that were mapped to the nanopore assembly. The inner track in each case displays annotated, predicted mobile genetic elements such as insertion sequences (ISs), transposases and prophage.
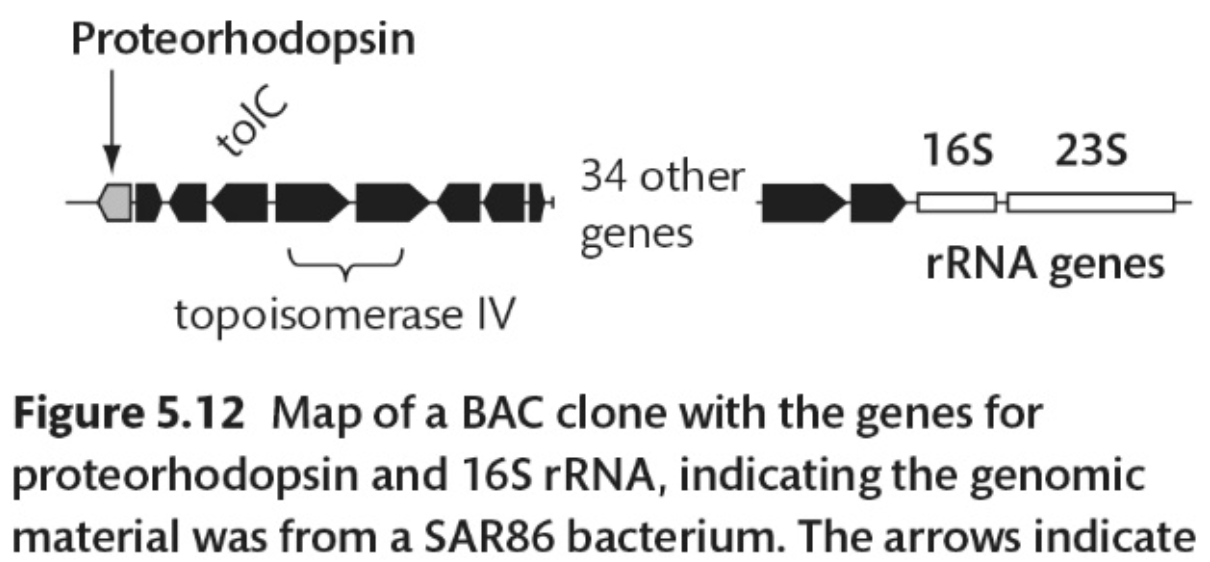
What do the arrows in this figure indicate?
The arrows indicate the direction of transcription for each gene.
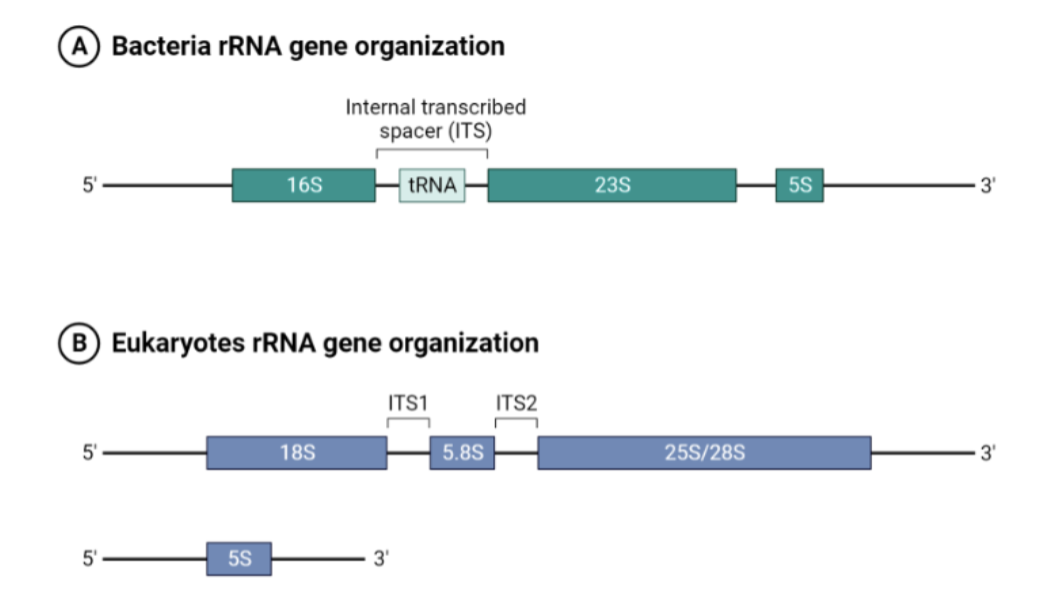
What does this figure show?
The structural organization of rRNA genes in bacteria and eukaryotes. In bacteria, the rRNA genes are present in a single operon containing 16S, 23S, and 5S rRNA genes. In contrast, eukaryotic rRNA genes are separated into multiple clusters, each containing 18S,
5.8S, and 28S rRNA genes, as well as a separate gene for 5S rRNA.
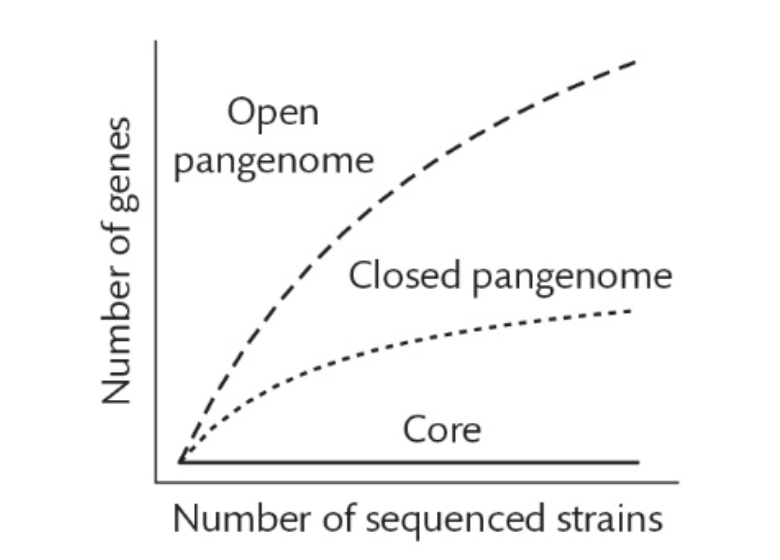
What assumption does this figure make?
It assumed the core genome for a bacteria with an open or closed pangenome is assumed to be the same size, but this is not necessarily the case.
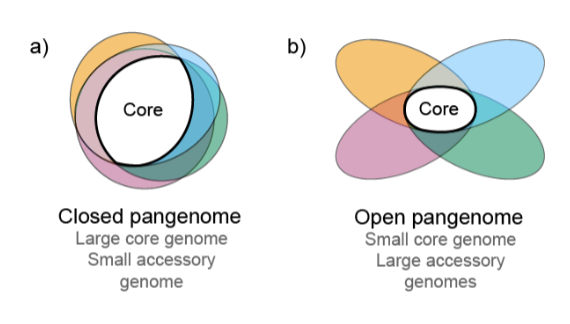
What is the difference between an open genome and closed genome?
A closed genome has a large core genome and small accessory genome, while an open is the opposite.
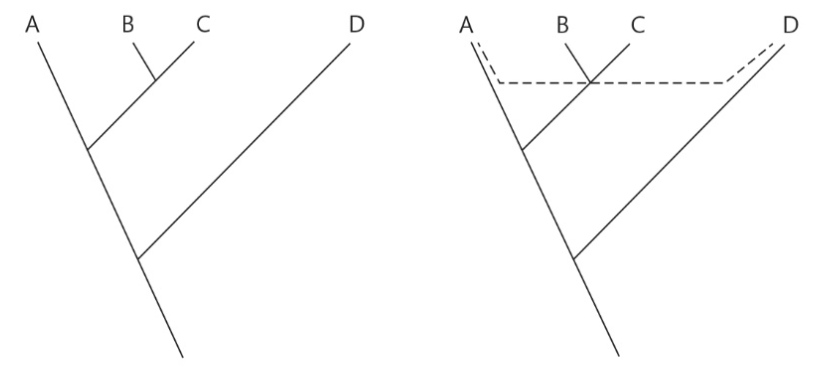
Which figure represents vertical gene transfer? Which represents horizontal gene transfer?
Left: Vertical gene transfer
Right: Horizontal gene transfer from Phylotype D to Phylotype A, independent of other genes
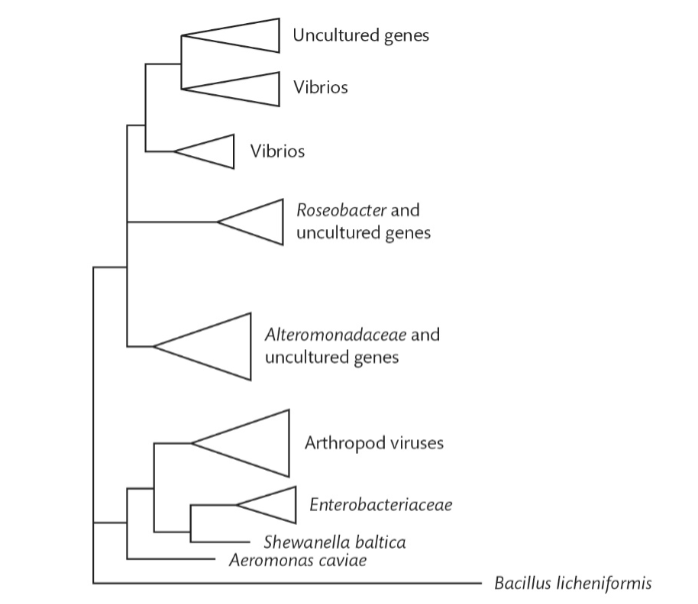
What is an example of horizontal gene transfer in this graph?
The presence of virus chitinases among the bacterial genes
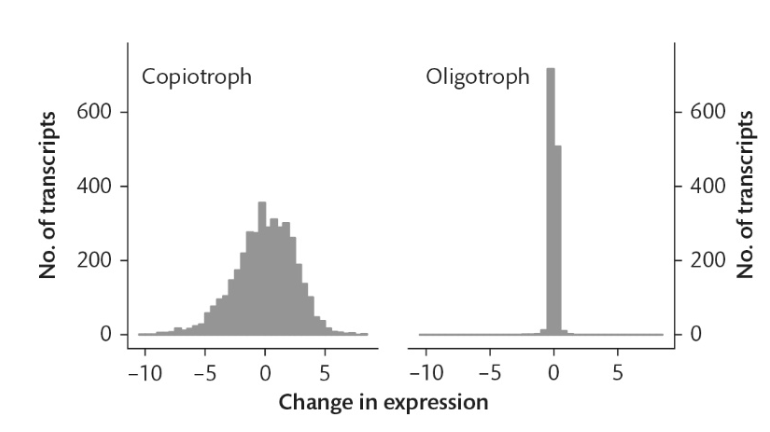
What does this graph demonstrate?
The change in growth rates of bacteria in oligotrophic and copiotrophic habitats
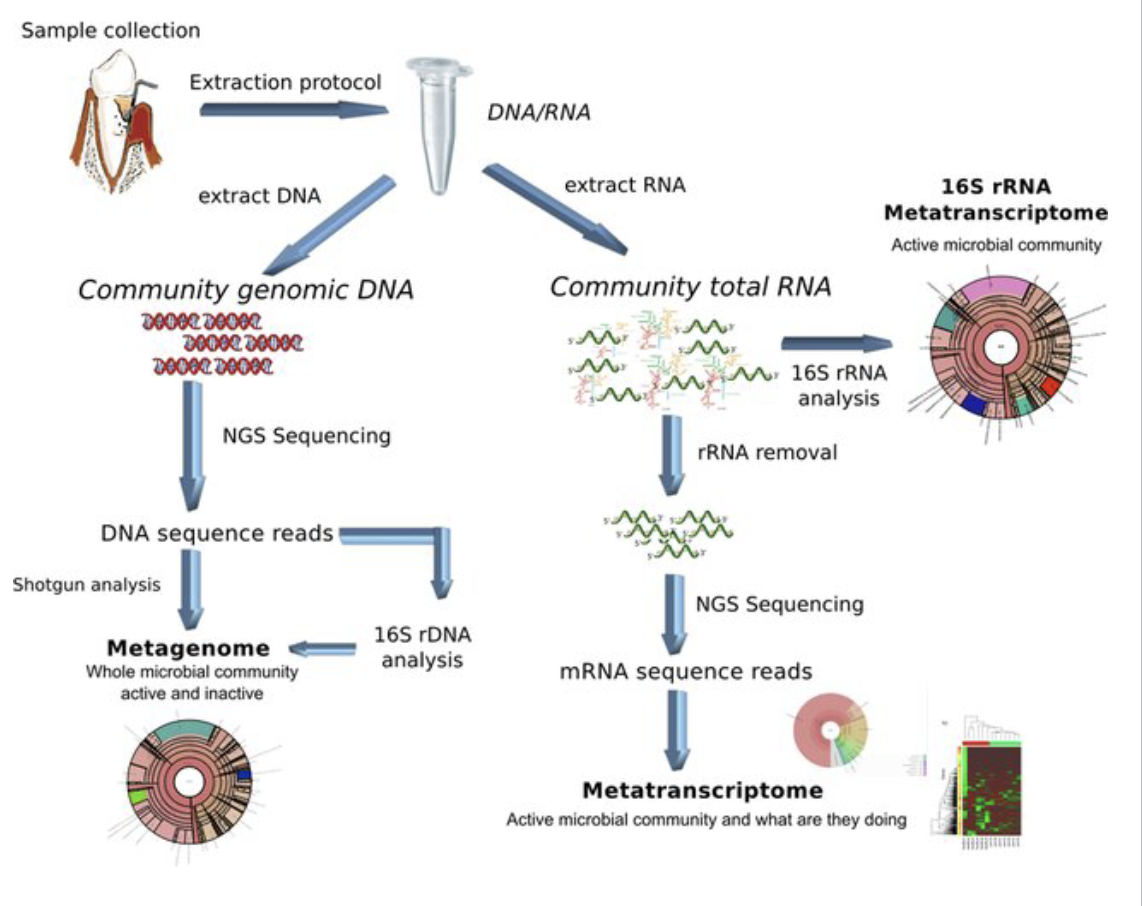
Between metagenomics and metatranscriptomics, which reads DNA sequences and which reads mRNA sequences?
Metagenomics is associated with DNA, and metatranscriptomics is associated with mRNA.
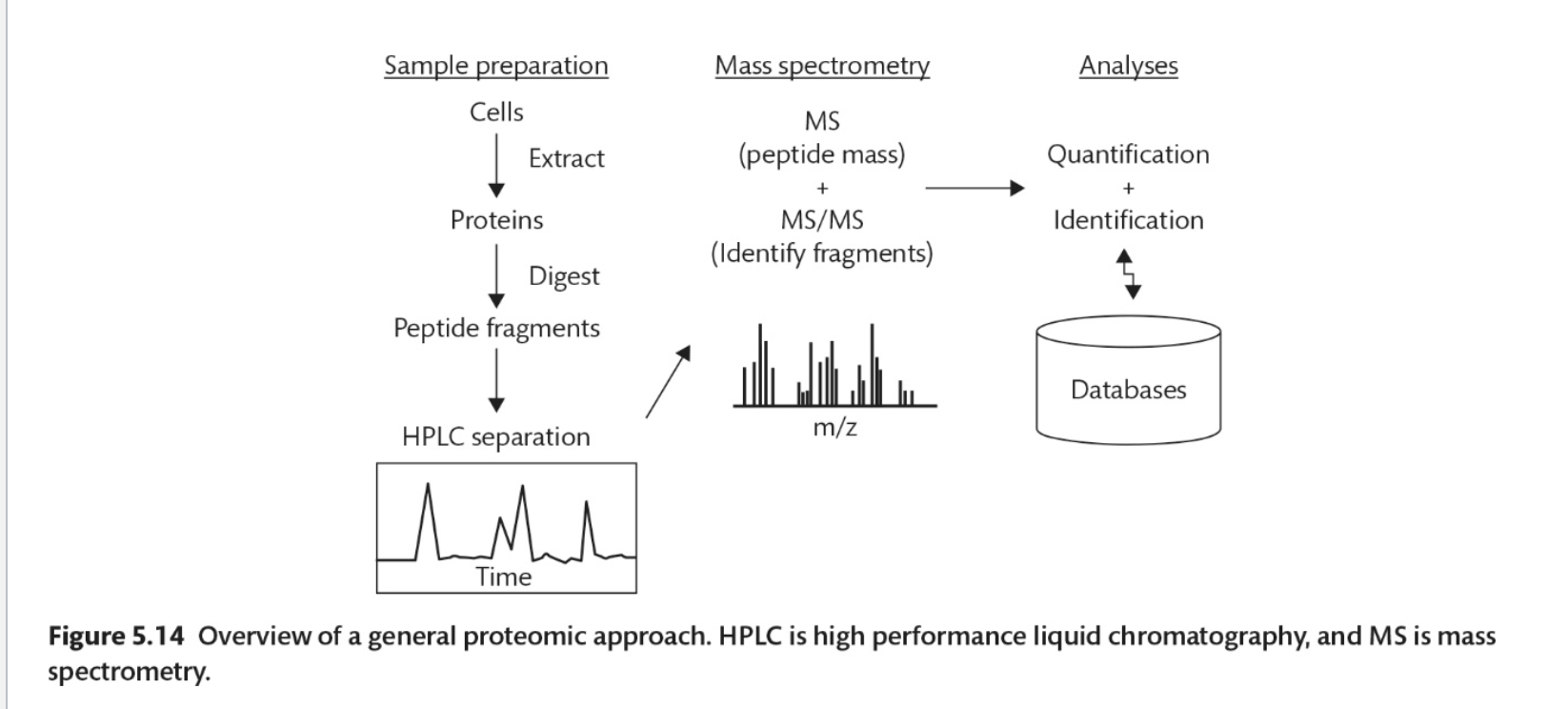
What are the steps to a general proteomic approach?
Sample preparation, mass spec, analyses
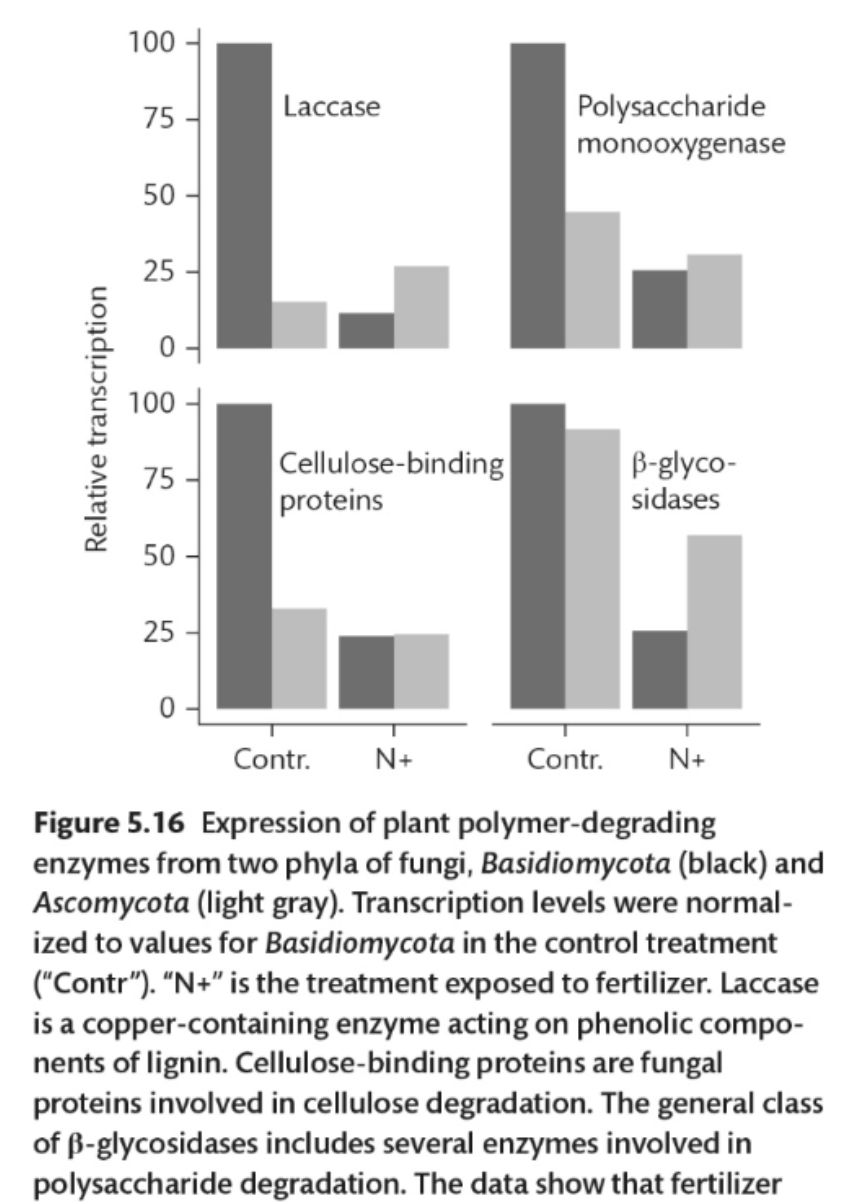
What do the data in this figure show?
The data show that fertilizer caused a decrease in expression of the four proteins, although response varied with the fungus and the protein.
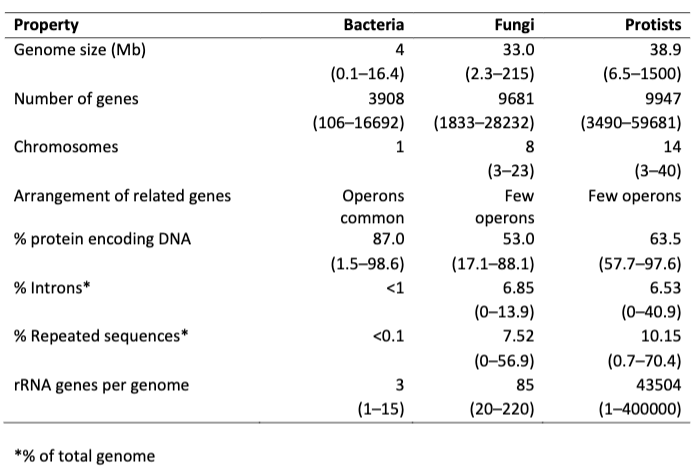
What does this table (5.1) show?
The genome structure of bacteria and eukaryotic microbes. The values are the mean and the range in parentheses.
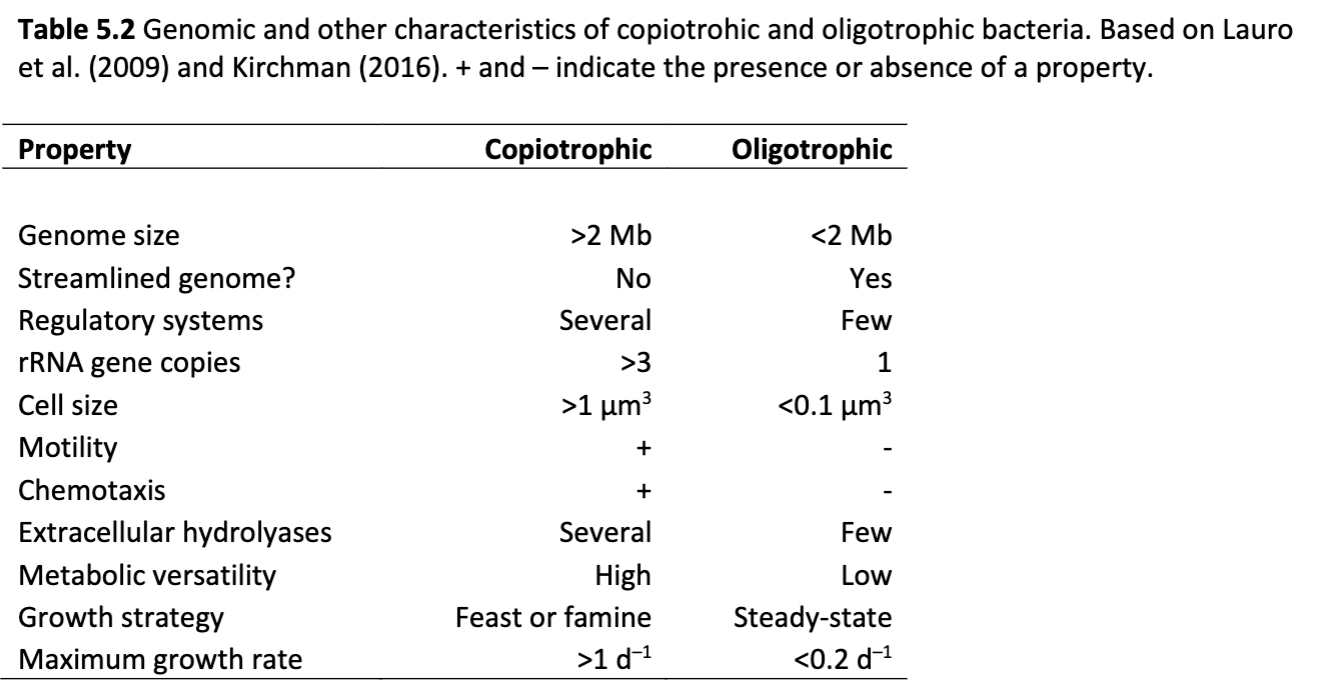

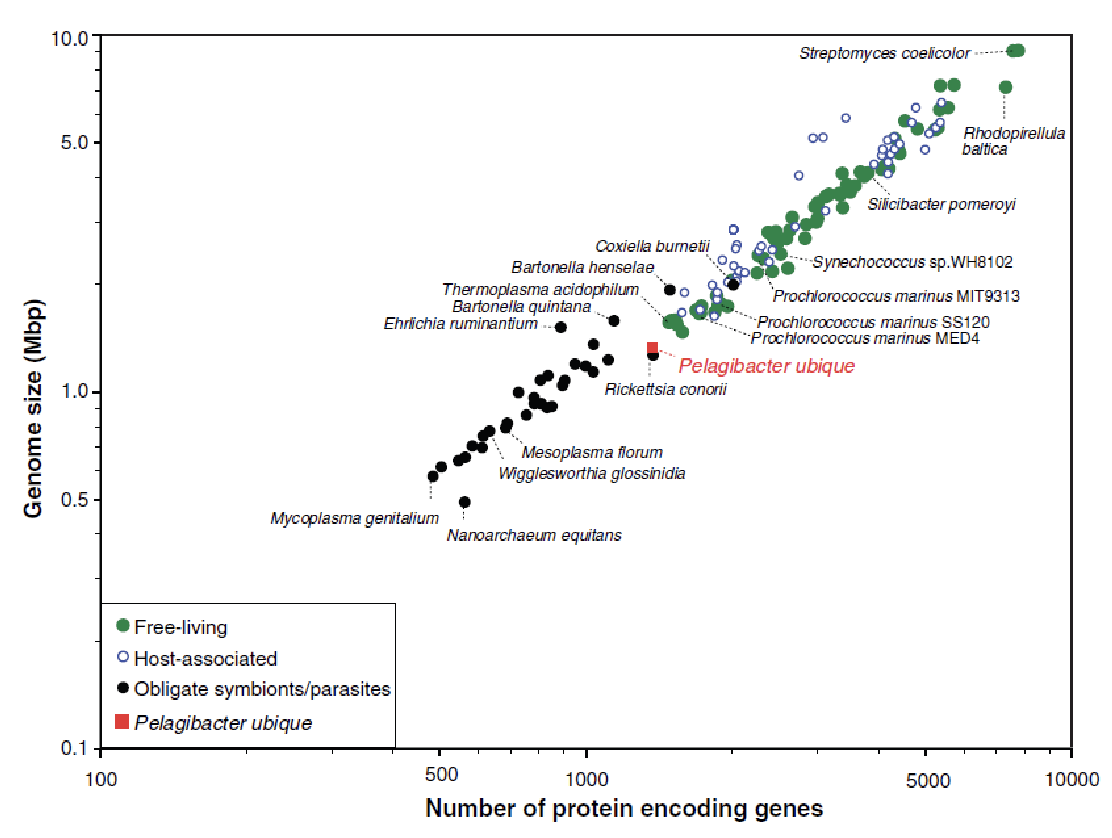
Using information from this graph, do oligotrophic bacteria have a streamlined genome?
Yes, but copiotrophic bacteria do not.
What are gene products of mRNA translation called?
polypeptides
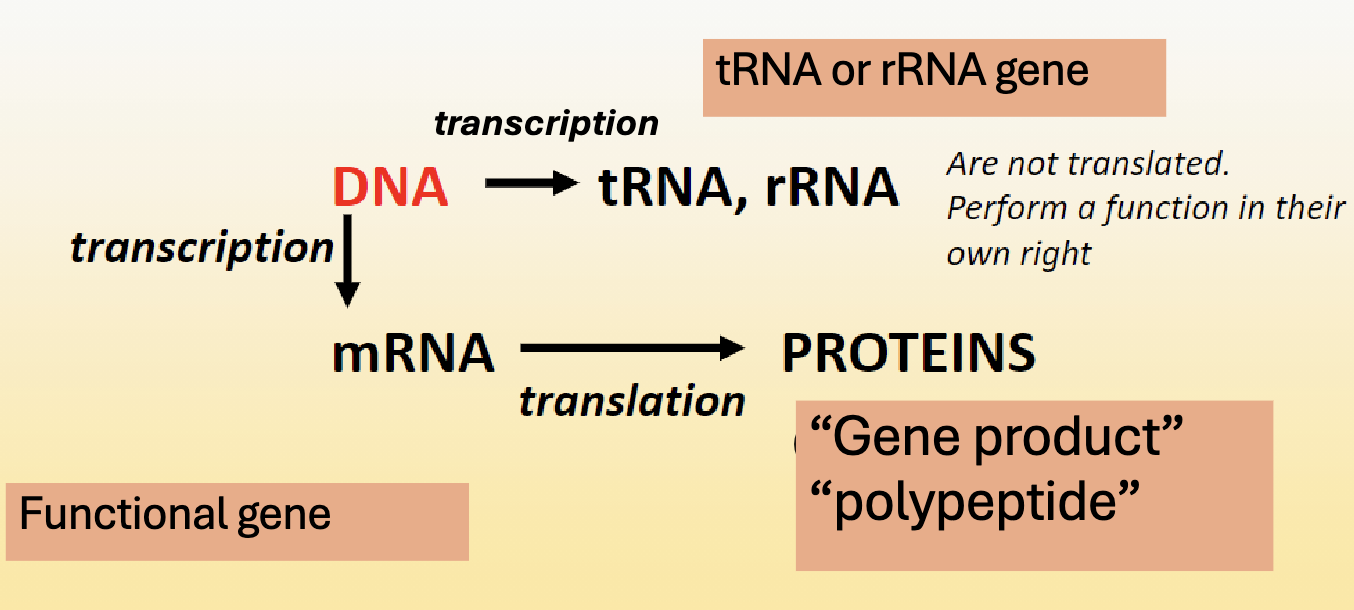
What is another term for barcoding genes?
phylogeny markers, and everyone has a phyologenetic marker gene.
What is the barcoding gene for E.coli? How many Mbp was it? How many bp is the entire 16S rRNA gene? What is the length of the E.coli genome?
What is the barcoding gene for E.coli? 16S rRNA gene.
How many bp was it? ~1,500 bp
What is the length of the E.coli genome? ~5 Mbp
What is bacterial DNA in?
Chromosomes and plasmids. In most natural bacteria, there is only 1 chromosome.
True or False: Archaea can have plasmids, but bacteria cannot.
False. Both can have plasmids.
What traits are used to separate microbes into phylotypes?
Morphology, physiology, average nucleotide identity, rRNA genes
Describe the difference between phenetic and genotypic classifications:
Phenetic classifies into phenoypes based on what it can do and what it looks like.
Genotypic categorizes based on % nucleotide similarity to others.
What is the typical length of an amplicon used for taxonomy and phylogeny?
~200-300 bp
What is phylogenetic classification generally based on?
It’s usually based on the direct comparison of gene sequences and/or sequences of gene products
Ribosomal proteins and ribosomal RNAs form _____.
Ribosomes. Ribosomes direct translation to convert information in mRNA to proteins
What is a conserved region?
Slow-evolving regions enable the design of universal PCR primers that can amplify genes from various different taxa. They are necessary for structure.
What is a variable region?
Sequences in the fast-evolving regions that reflect differences between species and are
useful for taxonomic classification
What is barcoding/metabarcoding
“Barcoding” or “Metabarcoding” = sequencing a gene that is relatively conserved among all species in that group of organisms, for determining taxonomy and/or phylogeny
_________ is an essential gene (like 16S) that has regions of conserved sequence and variable regions. Variable regions contain sequence subjected to evolution, just like V regions in 16S; can be used for phylogeny studies.
mtCOI
What are the steps for barcoding?
1. Isolate total genomic DNA from one (barcoding) or for metabarcoding, all organisms present in the ecosystem
2. Amplify a portion of the conserved gene (16S, 18S-ITS-23S, CO1)
3. Sequence that amplicon
a. To get full-length amplicon sequence, subclone and use vector-specific primer for sequencing
b. To sequence the PCR product directly, remove dNTPs and PCR primers. Then add one primer matching one end of the amplicon and subject to sequencing. (“tag sequencing” “NGS seq”)
What are the main parts of the SAR11 readings?
Of the >90 phyla found in the biosphere, only a few are abundant in any particular habitat. Only a few bacteria are widespread, AKA ubiquitous within oceanic habitats.
Relative abundances as determined by culturing ≠ relative abundances
determined by sequencing (i.e., “relative abundance” means % of total that is alpha-proteobacteria, % of total that is y-proteobacteria)SAR11: the most abundant bacteria in global oceans AND is ubiquitous
SAR11 clade belongs to the alpha-proteobacteria class, one cultured
representative is Pelagibacter ubique
Giovanonni et al. (2005) – really big deal, first time cultured a representative from the SAR11 clade
Culture- independent/dependent is way more reliable in giving good idea of community diversity, community structure.
independent
What are the advantages of the SAR11 clade’s small organism size?
Less likely to be preyed upon →Better at taking up organic matter from the env. b/c of…
Large Surface to Volume ratio (small cell, larger S/V ratio)
Other advantages, e.g., with respect to Reynold’s number
What is the genome size of SAR 11
1,308,759 base pairs. This is the smallest of any cell known to replicate independently in nature.
What methods were used to determine genetic diversity in Sargasso Sea bacterioplankton?
The picoplankton samples were collected using tangential flow filtration on Durapore 0.1 um flurocarbon membranes from a depth of a 1-2 m deep hydrostation. RNA genes were then amplified, and 12 cloned 16s rRNA genes.
What did phylogenetic analysis show from the 1990 paper?
It showed that one cluster of sequences (the SAR 7 cluster) branched within the radiation of oxygenic phototrophs.
The most abundant class of bacterial ribosomal RNA genes detected in seawater DNA by gene cloning belong to _____ —an α-proteobacterial clade
SAR11
On average, the SAR11 clade accounts for ______ of the cells present in surface waters and nearly _____ of the cells present in the mesopelagic zone. In some regions, members of the SAR11 clade represent as much as 50% of the total surface microbial community and 25% of the subeuphotic microbial community.
a third, a fifth
True or False: P.ubique has the smallest number of genes (1354 open reading frames) for any free-living organism
True
How were Proteorhodopsin genes first discovered?
Proteorhodopsin genes were first discovered through the cloning and sequencing of large genomic DNA fragments from seawater
True or False: Proteorhodopsin genes are found in easily cultured bacteria.
False. Proteorhodopsin genes have not been
found in cultured bacteria, and on the basis of
environmental sequence data, it has not yet been
possible to reconstruct the genomes of uncultured bacterial strains that have proteorhodopsin genes. Although the metabolic effect of proteorhodopsins is uncertain, they are thought to function in cells for which the primary mode of metabolism is the heterotrophic assimilation of dissolved organic carbon.
What is MALDI? Why is it used to analyze proteorhodopsin?
matrix-assisted laser desorption/ionization (MALDI) mass spectrometry. This technique is effective with the presence of mostly arginine-containing peptides in the tryptic digest of proteorhodopsin, which are favoured by the MALDI ionization process
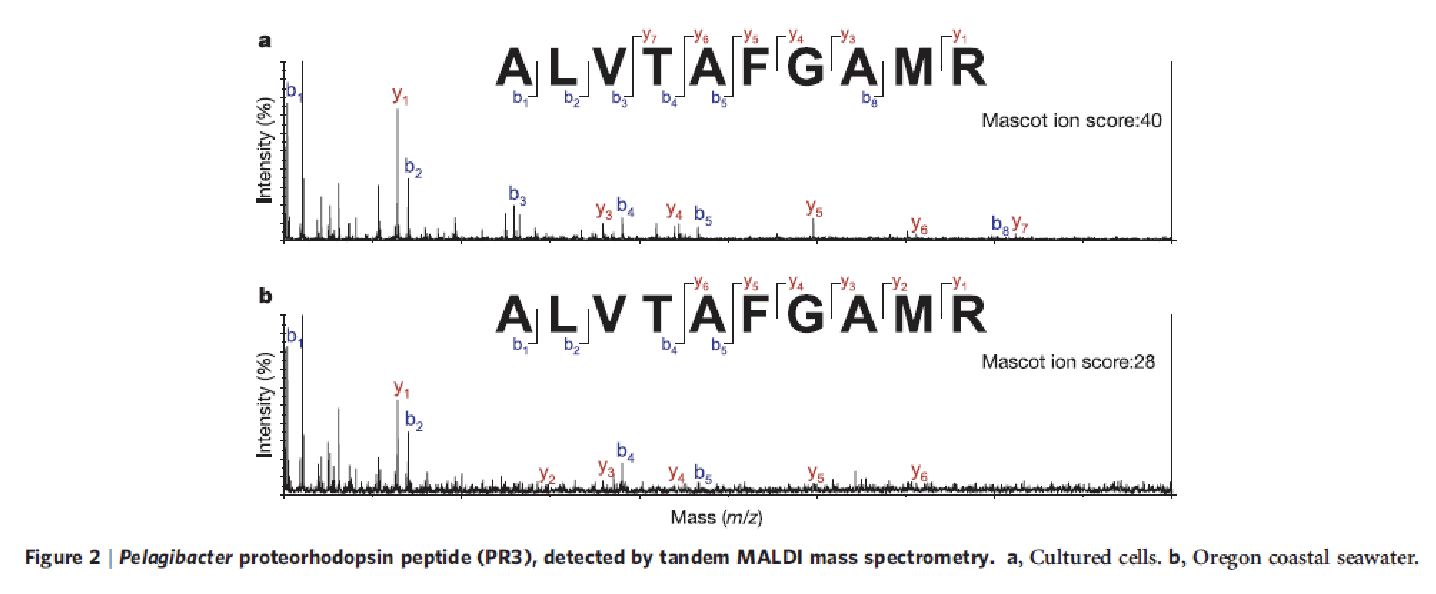
What was the primary objective of the Wilhelm 2007 paper?
Reconstruct info about specific
uncultured organisms from fragmentary environmental DNA sequences.Used the genome of an isolate of the marine alphaproteobacterium SAR11 ('Candidatus Pelagibacter ubique'; strain HTCC1062), from cold water
Compared variation in SAR11 metagenome sequence data from the Sargasso Sea, a warm, oligotrophic ocean gyre.
Define “core genome” and “pan genome”
a "core-genome" consists of genes that are always present, and a "pan-genome" of genes that are variably
present.
What does ecological data suggest about the SAR11 clade?
ecological data suggests that the SAR11 clade consists of a few ecotypes, which can be differentiated either phylogenetically, or by their appearance in the environment at different depths and seasons
A scientific study has this goal:
“We isolated and sequenced genomes from diverse SAR11 cultures that represent three major lineages and encompass the full breadth of the clade.”
What would the data from this expand our knowledge of?
The new data would expand observations about genome evolution and gene content that
previously had been restricted to the SAR11 Ia subclade, providing a much broader perspective on the clade’s origins, evolution, and
ecology.
At what depths were samples collected for the Gill 2022 sargasso sea paper?
From the surface and at depth just below the chlorophyll max.
Marine bacteria are amongst the most abundant organisms in the global ocean and are
responsible for ____ of primary productivity and nutrient fluxes in marine
environments
Greater than half
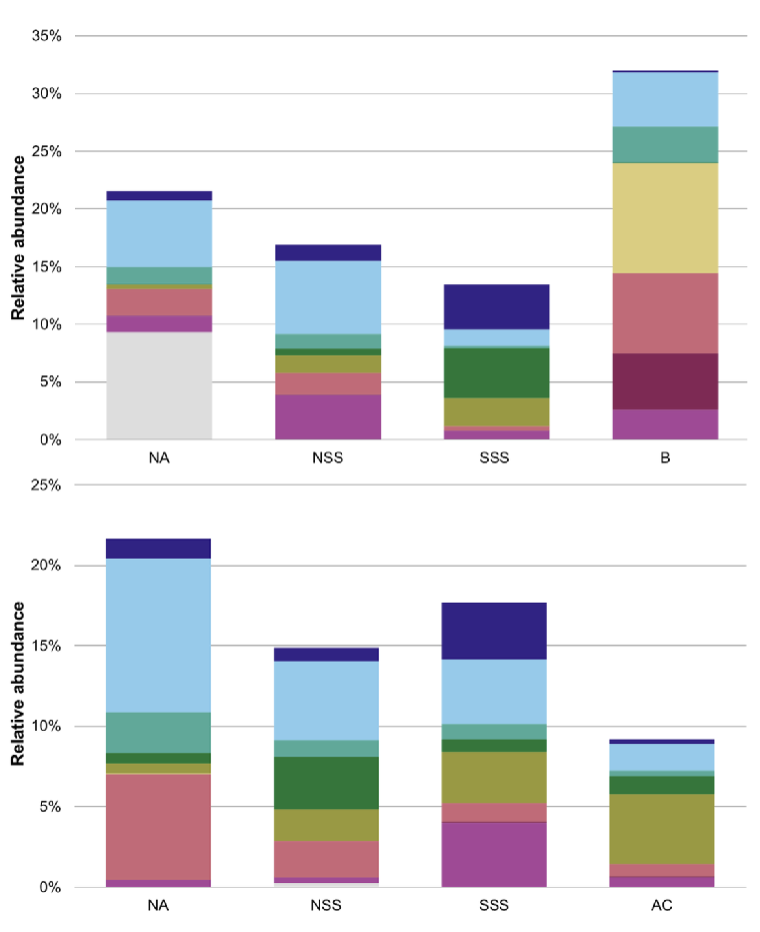
What were the two most abundant microbial groups found in the Gill 2022 study?
Procholorococcus and synechococus. (definitely spelled wrong but we just have to know the gist)
What indices are used to determine species diversity?
Simpson Index*
Shannon Index
Chao1 Index
*Simpson Index:
the greater the value, the greater the sample diversity.
As we sequence more and more 16S genes, we get a better idea of diversity in an ecosystem. Give an example where this is the case.
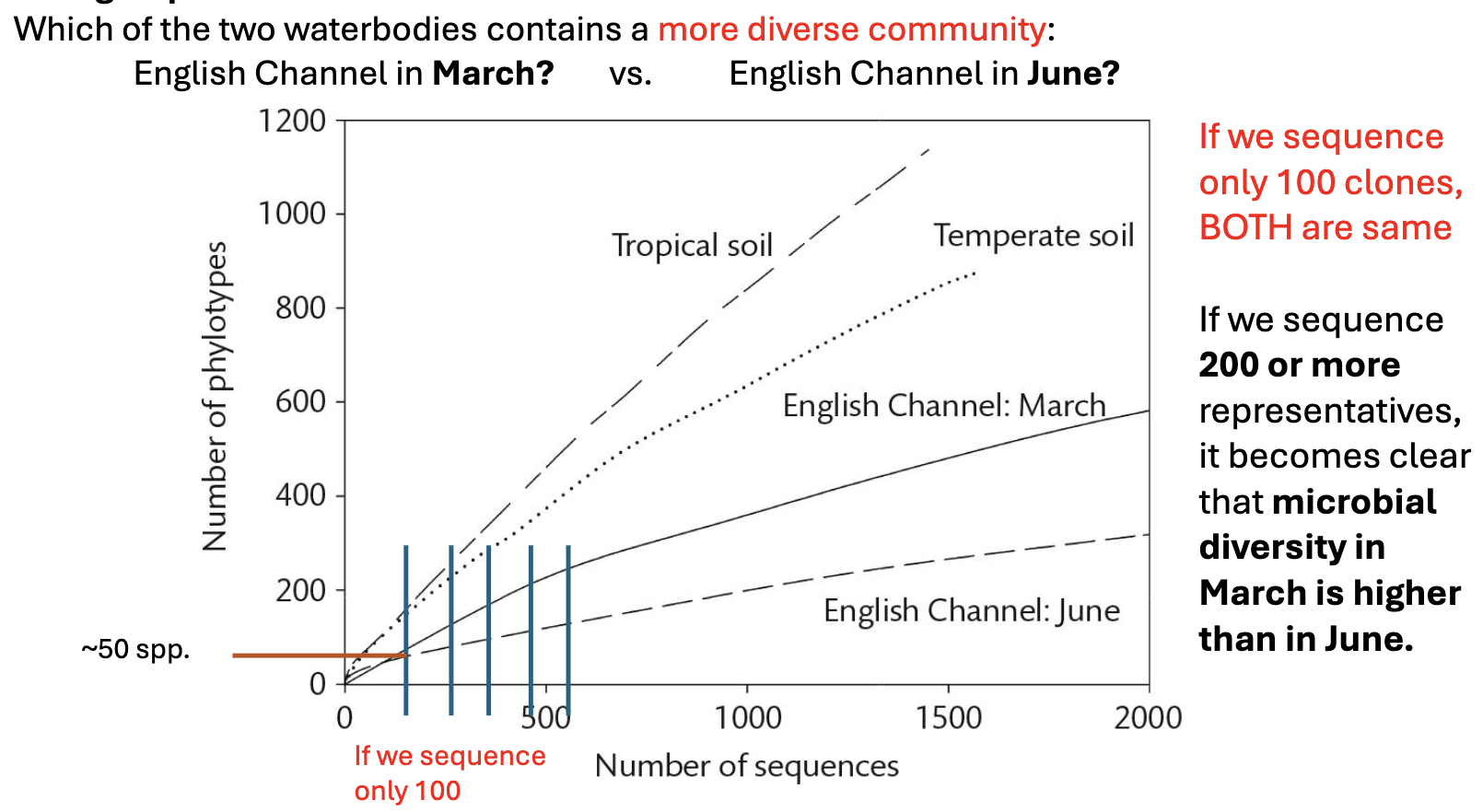
On earth, including in the water, there are ______ species of bacteria.
~ 1012
What can the Redfield ratio be used to explore?
How primary production and respiration affect the concentrations of major nutrients in oxic ecosystems
Give examples of primary producers:
Cyanobacteria, Diatoms, dinoflagellates,
Kelps, Land plants
Give examples of secondary producers:
Proteobacteria, Humans, dogs, cats, geese
__________ is the process by which organisms trap light energy (photons) and store it as chemical energy in the form of ATP and/or reducing power in NADPH.
Phototrophy
There are two major types of phototrophy:
chlorophyll-based chlorophototrophy and rhodopsin-based retinalophototrophy.
Microbes account for ~50% of global primary production. What group is responsible for ~50% of aquatic primary production?
Cyanobacteria
Rates of primary production are governed by what?
Light intensity and quality, as well as nutrients like ammonium, nitrate, phosphate, and iron.
Give examples of eukaryotic microbes that can dominate the algal community.
Diatoms, dinoflagellates, coccolithophorids, and phaeocystis. Some are toxic while others are photoheterotrophic.
True or False: Primary production is the most important process in the biosphere.
True (ch. 6 intro). This is especially true for oxygenic photosynthesis.
True or False: Photoheterotrophy and mixotrophy are very different types of metabolism.
False. They are essentially the same, with photoheterotrophy referring to bacteria, while mixotrophy is applied to protists.
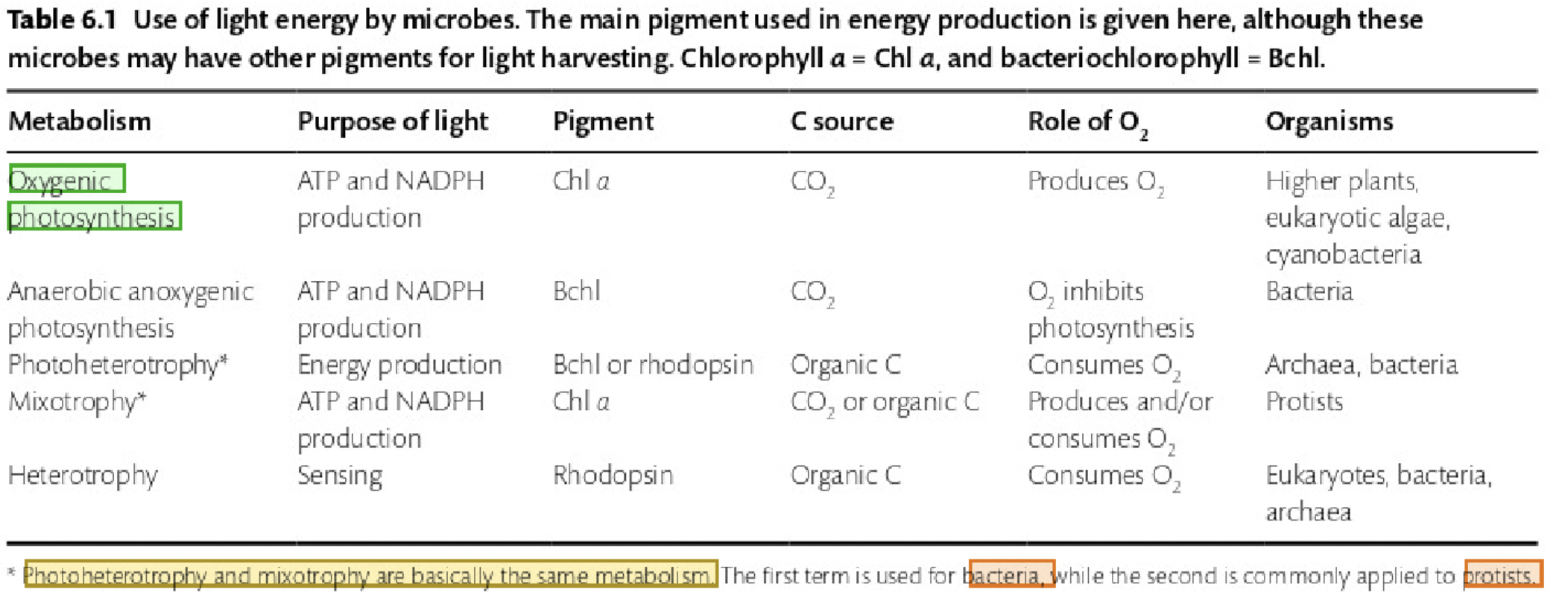
True or False: There are no known photoautotrophic archaea.
True, though some halophilic archaea use light energy using rhodopsin.
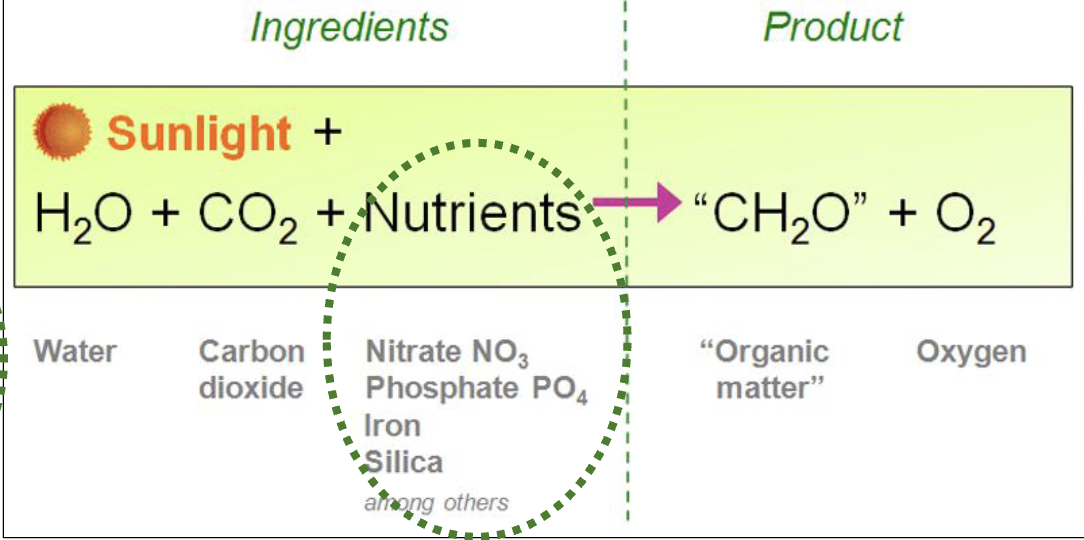
Light-driven primary production is based on photosynthesis and can be summarized with what equation?
CO2 + H2O + nutrients + light ——> biomass + O2

Summarize the process of oxygenic photosynthesis:
Light = energy source
CO2 = source of C
H2O = source of electrons
Light harvested by accessory pigments is transferred to chlorophyll a where the water is “split” to synthesize ATP and NADPH, a reaction that produces oxygen. ATP and NADPH are then used to fix CO2 to produce organic material.
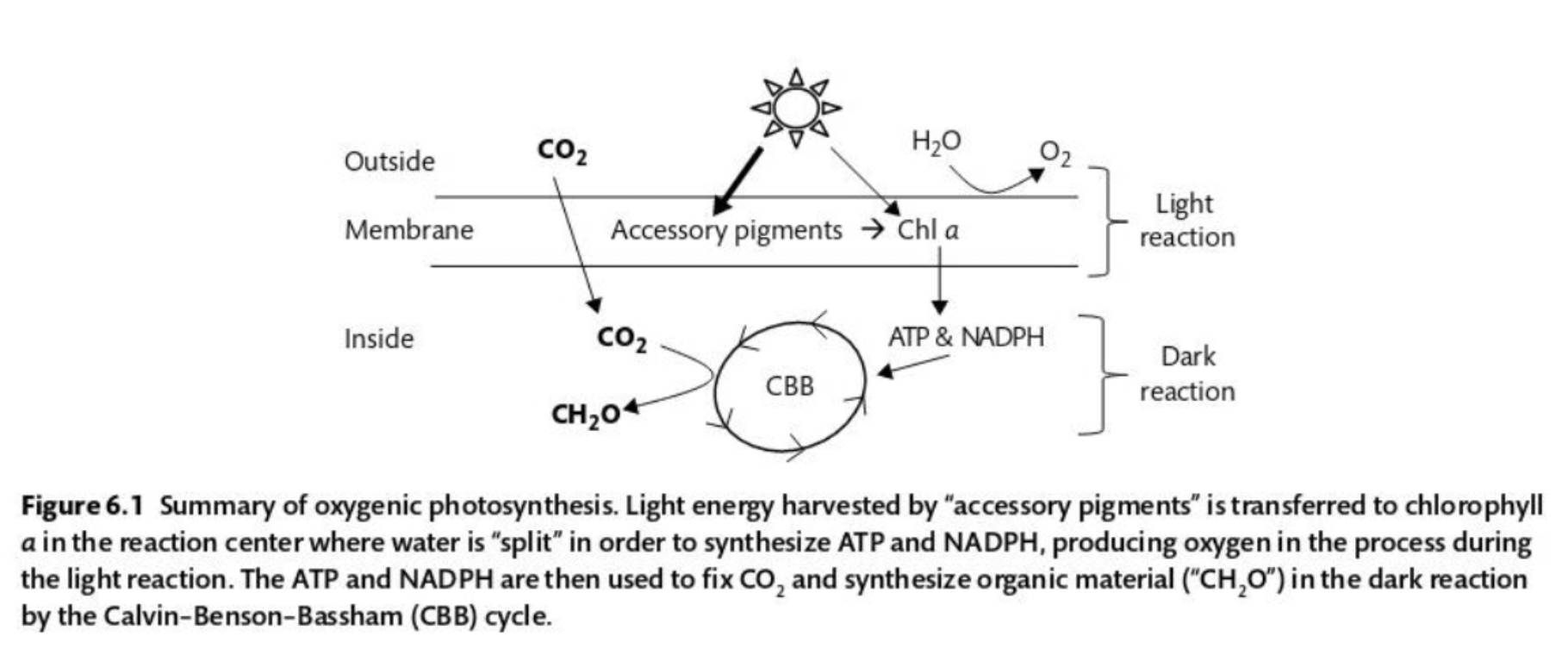
What percent of cholorophyll a molecules in phytoplankton are used for light harvesting?
More than 99%. It is in the reaction centers of chlorophyll a that light energy is converted to chemical energy.
True or False: Higher land plants have greater diversity of accessory pigments than phototrophic microbes.
False. These accessory pigments enable phototrophs to harvest the wavelengths of light found in lakes and oceans, especially green light (450-550 nm).
What pigments absorb the green part of the light spectrum?
Fucoxanthin (a carotenoid found in diatoms and some other eukaryotic algae)
Peridinin (another carotenoid, in dinoflagellates)
Phycoerythrin (made by cyanobacteria, red algae, and cryptomonads)
These pigments are why phototrophic microbes can be yellow, red, brown, or shades of green not seen in higher plants
What is an example of primary production without photosynthesis?
Chemolithoautotrophy aka chemosynthesis
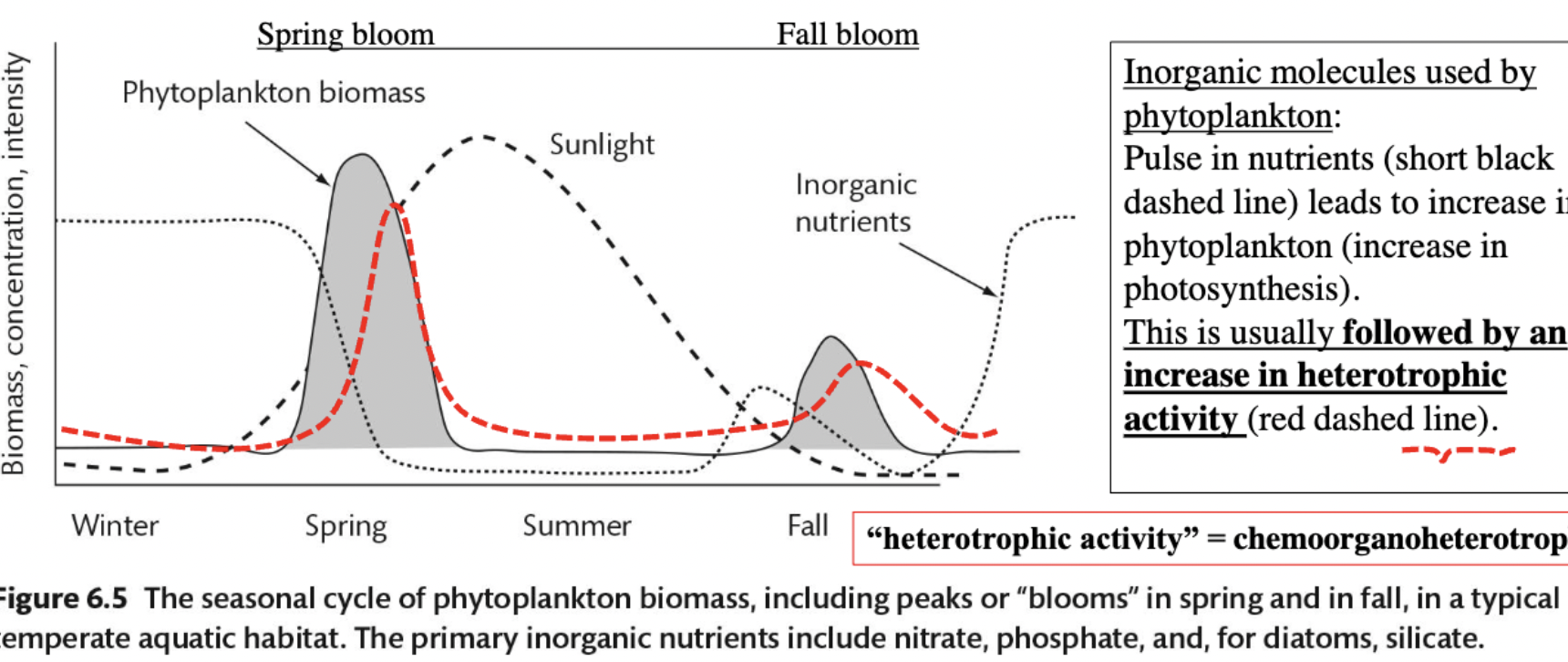
At what time of year does plankton biomass, concentration, and intensity become the greatest?
This peak occurs in spring, where the increase in the nutrients used by phytoplankton results in a phytoplankton bloom, subsequently increasing heterotrophic activity.
What processes fall under the blanket term “phototrophy”
Phototrophic Primary production (1oP)
Photoautotrophy: Oxygenic photosynthesis
Photoheterotrophy: Rhodopsin-based photoheterotrophy
or Anoxygenic phototrophy (Aerobic OR Anaerobic Anoxygenic phototrophs)
-- Only certain types of Anaerobic Anoxygenic phototrophs can fix CO2Mixotrophy: e.g. Karenia brevis & Pfiesteria: dinoflagellates that can consume prey
Overall, this applies to Fixing Carbon: CO2 + H2O + light → CH2O + O2
Primary Production alone promotes an increase in D.O. HOWEVER, heterotrophic activity, e.g. respiration performed by the heterotrophic prokaryotes and eukaryotes, often follows closely behind the peak in 1oP. What factors affect O2 and CO2 budgets, and are they negative or positive effects?

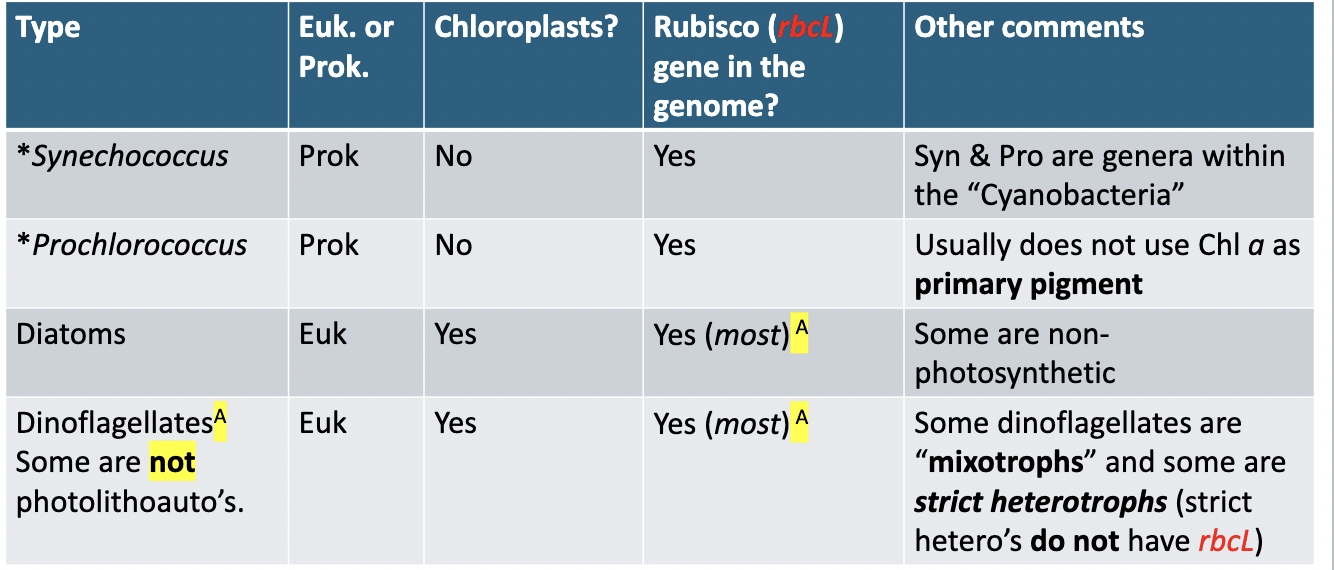
Does procholorococcus use chlorophyll a as its primary pigment?
Not usually
True or False: Some dinoflagellates are mixotrophs and some are strict heterotrophs.
True
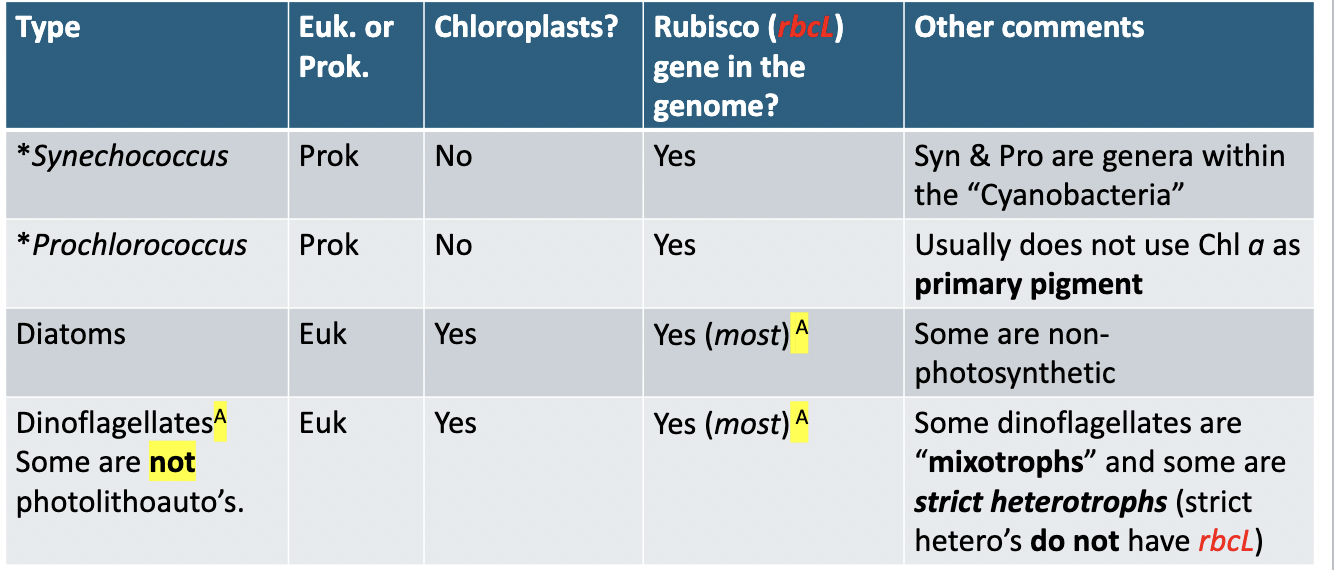
The famous genus Pfiesteria contains mixotrophic species. How do they get their food?
P. piscidia preys on other microbes and on the red blood cells of fish, and makes toxins that are harmful to fish & humans.Using peduncle, sucks out contents of prey cell.
What genus of dinoflagellates are symbionts of corals?
Symbiodinium
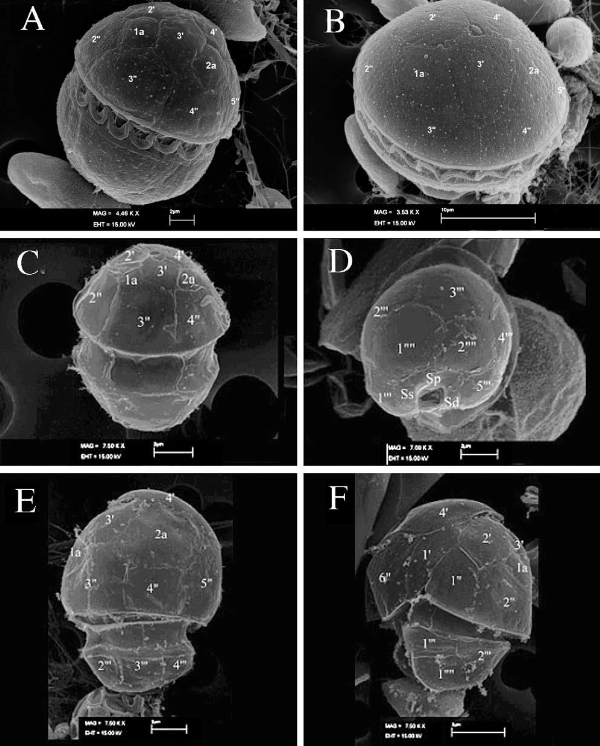
What is the significance of this group of dinoflagellates?
They are genetically and morphologically similar to Pfiesteria, the “fish killer” mixotroph.
What are the phylogenetic marker genes for each group of organisms?
16s rRNA- prokaryotes, in the mitochondria and chloroplasts of eukaryotes
18s rRNA- eukaryotes
mtCO1 - prokaryotes and eukaryotes
Describe the major cyanobacteria genera Prochlorococcus:
Prochlorococcus is a genus of very small (0.6 μm) marine cyanobacteria with an unusual pigmentation
Prochlorococcus microbes are among the major primary
producers in the ocean, responsible for a large
percentage of the photosynthetic production of oxygenAverage genome size is about 2,000 genes. In
contrast, eukaryotic algae have over 10,000 genes
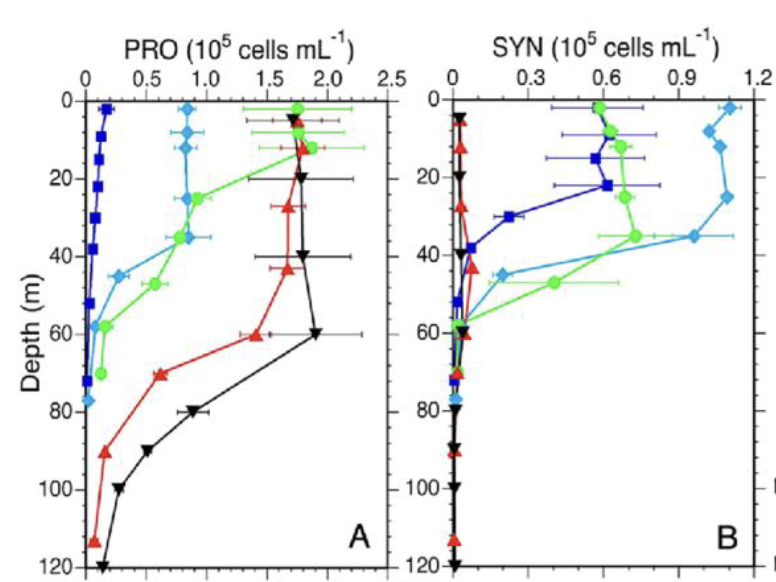
What does this figure show?
Mean depth profiles of cell abundances for picophytoplankton populations during Cycles 1-5. PRO = Prochlorococcus; SYN = Synechococcus.
What is the smallest known photosynthetic organism?
Prochlorococcus. It is 0.5 to 0.7 μm in diameter. Prochlorococcus is also presumably the most abundant photosynthetic organism on earth, found from surface depths to 200 m.
How often do Prochlorococcus divide?
Once a day in the subsurface layer of oligotrophic areas,
where it dominates the photosynthetic biomass.
What amount of global carbon fixation do Prochlorococcus and Synechococcus account for?
About 1/3, even though they comprise less than 1% of ocean mass.
What systems enable oxygenic photosynthesis in cyanobacteria?
• have photosystems I and II
• Some use chlorophyll a as main pigment
• prochlorophytes have chlorophyll a and chlorophyll b
• most appear “blue-green” due to presence of phycocyanin
• But the presence of phycoerythrin in many marine isolates
gives them a red or brown coloration
Although similar to eukaryotic photosynthetic organisms,
cyanobacteria are different because they do NOT have
chloroplasts (these are true bacteria without organelles).
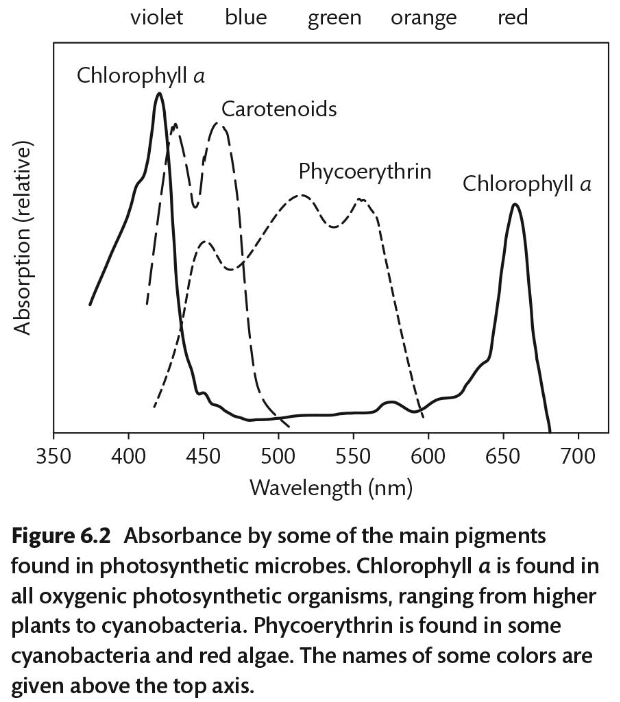
What wavelength of light does Phocerythrin absorb? What do carotenoids absorb?
Phycoerythrin: green and orange
Carotenoids: blue
Is CH2O particulate?
Yes. It is integrated into biomass, now part of the PARTICULATE pool of organic matter, as part of the bodies of the algae/cyanobacteria and the grazers that eat them.
How do scientists measure primary production rates?
By measuring the rate by which CO2 is fixed, the rate of biomass production, or the abundances of primary producers by biomass values or cell numbers.
Scientists can also measure light-driven primary production photosynthesis rates by measuring rates of O2 production in the light
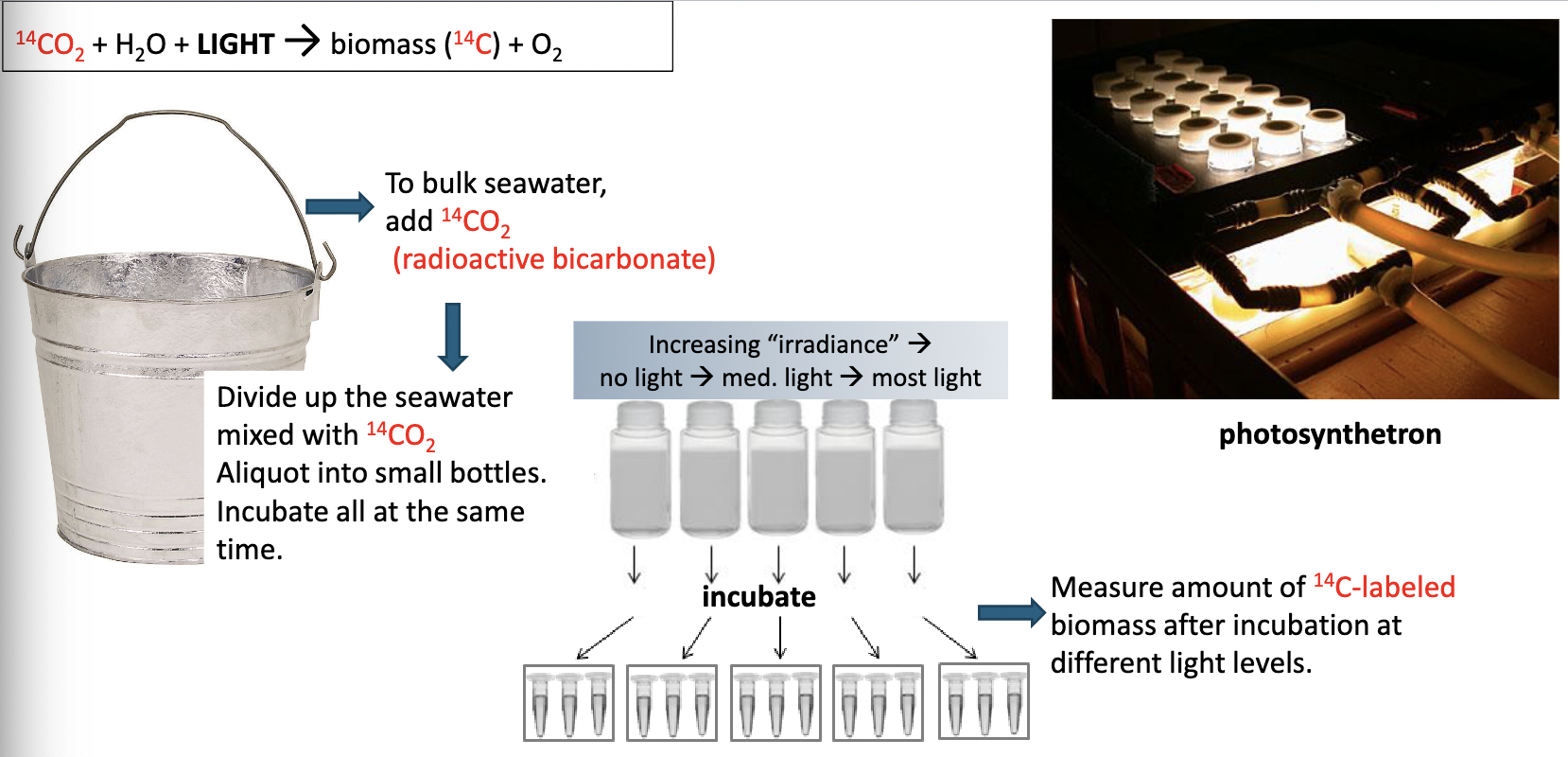
What is carbon fixation?
When inorganic carbon is “fixed” into non-gaseous
form (C gets incorporated into an organic molecule)
*Converts ADP to ATP → energy storage
*Pi (inorganic phosphate) is needed
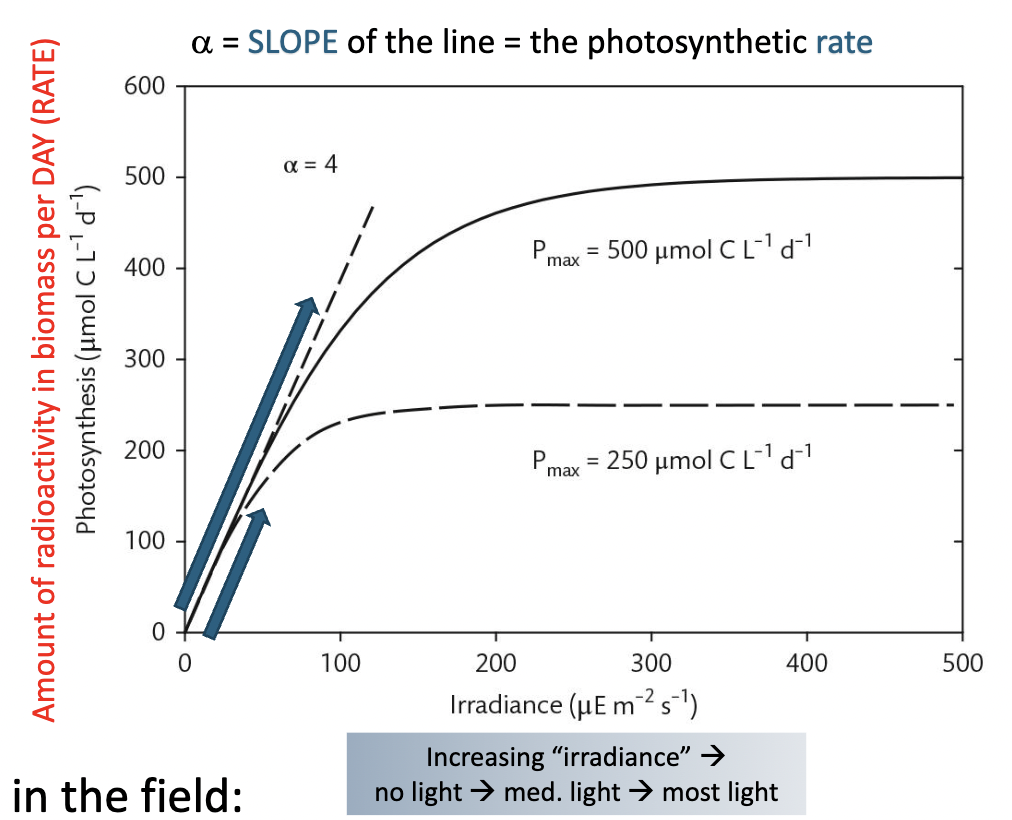
What is the gold standard for measuring photosynthesis rates in the field?
The “P vs I curve”
Another way is to measure rates of O2 production in the
light and compare to dark-incubated bottles.
What is the downside to measuring concentrations of chlorophyll a in water to collect data about primary production?
This method is at a fixed point in time and does not provide RATE data. Also, not all 1oP have significant amounts of Chl a in their cells, using alternative pigments as a primary photosynthesis pigment.
How should D.O. data be collected?
Dissolved oxygen should be collected as quickly and carefully
as possible. Ideally, samples should be measured in the field
immediately after collection.
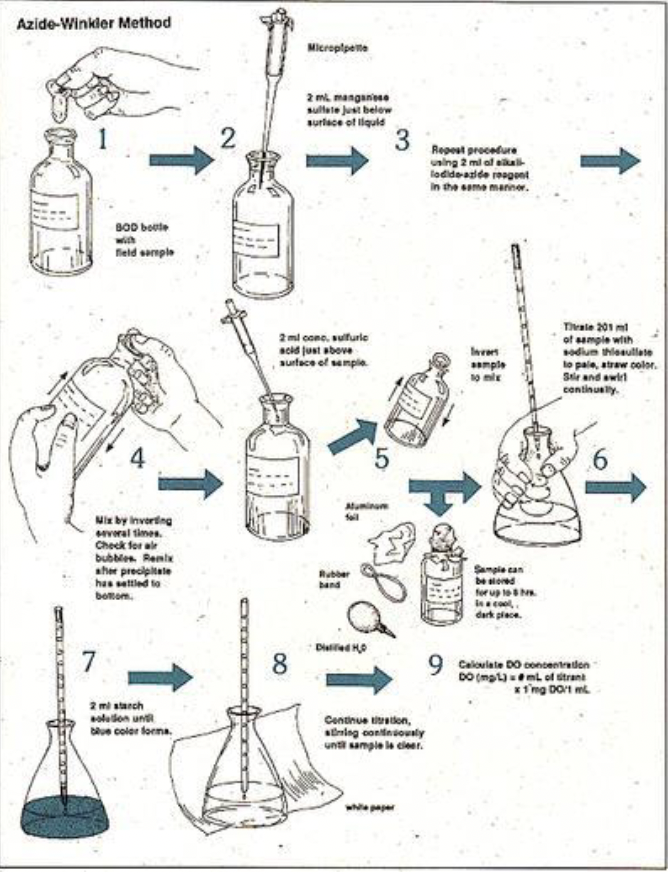
What are the problems with the Axide-Winkler method for measuring O2 production?
Lots of bottles (glass)
Tedious process, and care must be taken to
immediately process seawater once it comes out
of NiskinCan be confounded by respiration rates (see Fig
6.3).O2 evolution is the NET O2 production (comparing
dark vs dark bottles).If you want to measure rates, then you have to
have many bottle replicates to sample at various
time points – even more tedious, even more glass
bottles, and takes up a lot of space on tight
quarters ship lab spaces/benches.
What is net flux?
NET FLUX is the important aspect of examining CO2 & O2
concentrations, when we are modeling net heterotrophy
vs. net autotrophy in oceans.
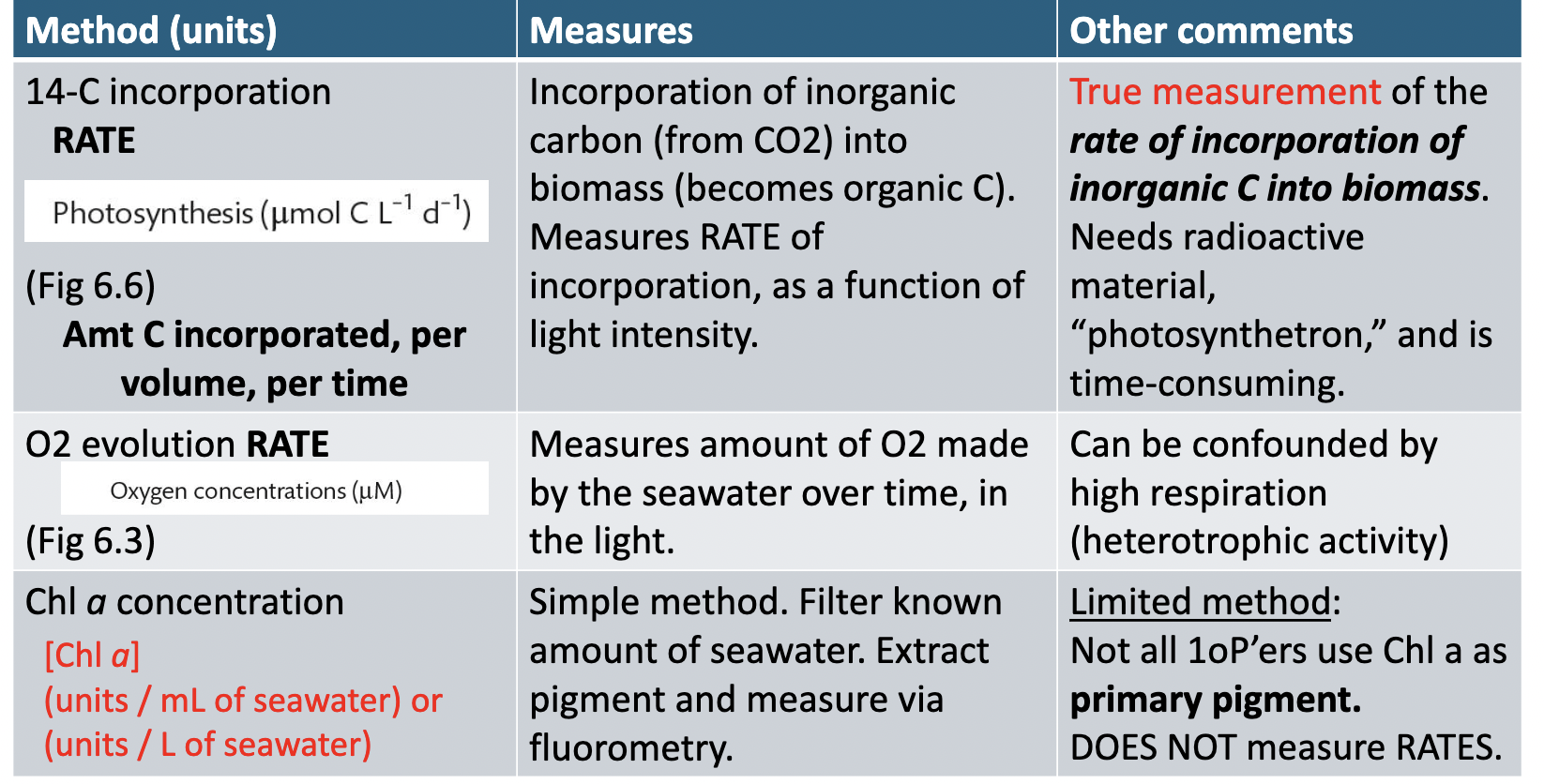
What are the 3 methods of measuring oceanic 1oP?
Conversion of C into biomass/org. C
Oxygen evolution rates
Concentration of chlorophyll a
What are ecotenzymes?
Enzymes outside the outer cell membrane, synonymous
with “extracellular enzymes”
What is catabolism?
The energy-producing parts of microbial metabolism.
What is anabolism?
Synthesis of cellular components and eventually growth.
True or False: Bacteria abundance in aquatic habitats >> fungal abundance
True.
• Not so for soil habitats, where both are abundant and important in OM degradation