BSCI222 Exam 2 RNA Transcription
1/65
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
66 Terms
RNA transcription
making RNA from DNA
Why is RNA transcription performed?
DNA stores information, but DNA cannot leave the nucleus (in eukaryotes)
So the cell makes a copy of the DNA as RNA = working copy of DNA instructions
This is so we can produce RNA that can be used to make proteins
mRNA carries instructions to ribosomes
Ribosomes build proteins
Different types of RNA
Cells have several classes of RNA molecules that serve different functions, including:
mRNA - messenger RNA; provides information to ribosome to build a polypeptide.
rRNA - ribosomal RNA; allows ribosomes to bind to mRNA.
tRNA - transfer RNA; binds to mRNA in ribosomes to deliver amino acids.
snRNAs - small nuclear RNAs; allows snRNPs to interact with pre-mRNA for splicing.
miRNAs, siRNAs - involved in regulation of mRNA through degredation.
mRNA
messenger RNA; provides information to ribosomes to build a polypeptide (protein)
rRNA
ribosomal RNA; forms the ribosome and allows ribosomes to bind to mRNA
tRNA
transfer RNA; “translator” between RNA and protein; binds to mRNA in ribosomes to deliver amino acids (brings amino acids to the ribosome, and matches codons on mRNA)
snRNAs
small nuclear RNAs; allows snRNPs to interact with pre-mRNA for splicing (Removes introns from pre-mRNA)
miRNAs, siRNAs
involved in regulation of mRNA through degradation (control how much protein is made)
What direction does RNA transcription occur in?
RNA transcription uses only one strand of DNA, synthesizing a single strand of RNA as the reverse-complement in the 5'->3' direction
DNA template: 3' - TAC - 5'
RNA made: 5' - AUG - 3'
Reverse-complement
RNA is built complementary to the template DNA strand and in the opposite direction
DNA → RNA
A → U
T → A
C → G
G → C
DNA template: 3' - TAC - 5'
RNA made: 5' - AUG - 3'
RNA polymerase only builds in the 5’ —> 3’ direction
DNA template (3'→5') → RNA made (5'→3')
What is used to make a specific RNA?
A transcription unit (aka, a gene) is used to make a specific RNA.
What kind of machinery do prokaryotes use for RNA transcription?
1) Prokaryotes have one RNA polymerase - Single enzyme complex with accessory proteins that reads DNA and uses it as a template to build RNA
2) Prokaryotes also have sigma factors = helper proteins that guide RNA polymerase to specific promoters (GPS)
What kind of machinery do eukaryotes use for RNA transcription?
Eukaryotes are more complex because they have multiple RNA polymerases that specialize in making different products
RNA polymerase I transcribes rRNA
RNA pol II transcribes pre-mRNA, small RNAs
RNA pol III transcribes tRNA, small RNAs
RNA polymerase I transcribes…
rRNA (ribosomal RNA)
small RNA
any short RNA (usually ~20–200 nucleotides) that does NOT code for protein
RNA polymerase II transcribes…
pre-mRNA, small RNAs (short RNA molecules that do NOT code for proteins aka miRNA, snRNA, etc.)
RNA polymerase III transcribes…
tRNA, small RNAs (short RNA molecules that do NOT code for proteins) including miRNA, snRNA. etc.
Prokaryotic RNA transcription steps
1) Initiation
2) Elongation
3) Termination
promoter sequence
A specific region of DNA located upstream of a gene that initiates transcription by acting as a binding site for RNA polymerase and transcription factors; it contains information for where the RNA polymerase should bind to the DNA, which strand to use as the template, and which direction to move in
What are the two key promoter sequences?
1) The Pribnow/TATA box (-10bp region)
2) the second consensus (average/common) sequence: -35bp region
Pribnow box
aka the TATA box; a 5'-TATAAT-3' consensus sequence at -10bp upstream
The second consensus sequence
5’-TTGACA-3'; at -35bp upstream
Holoenzyme
RNA polymerase core enzyme binded to sigma factor; this is the enzyme that actually builds the RNA
Prokaryotic RNA transcription 1) Initiation
RNA polymerase finding the correct start site on DNA (promoter recognition)
The sigma factor (a bacterial transcription initiation protein) complexed to RNA polymerase core enzyme (sigma factor binded to core RNA polymerase) binds to the promoter regions at -10 and -35 (the Pribnow/TATA box and second consensus box), and positions the holoenzyme (RNA polymerase + sigma factor) from -50 to +20 on the gene.
Formation of transcription bubble: The holoenzymes undergoes a conformational change to bind to DNA more tightly, and binds tighter to the promoter, separating the DNA strands at the -10 TATAAAT sequence.
Initiation occurs at +1 site (the first nucleotide that gets transcribed, where RNA polymerase adds the first RNA nucleotide). Only the one template strand is used. RNA polymerase reads the strand that runs in the 3’ → 5’ direction so that RNA can be made in the 5’ → 3’.
Prokaryotic RNA transcription 2) Elongation
The core RNA polymerase moves downstream, building RNA polymer within the transcription bubble.
Prokaryotic RNA transcription 3) Termination
Types of termination
Rho-dependent termination
Rho-independent termination
It depends on the RNA sequence, which determines the structure (and whether proteins bind)
terminator sequence
a DNA segment that signals RNA polymerase to stop transcription
Rho-independent termination
At a Rho-independent terminator, the RNA folds back on itself (reverse complementary sequences on RNA) to form a hairpin loop structure. The hairpin causes RNA polymerase to pause, and disrupts the RNA:DNA duplex, releasing RNA (normally, RNA is base-paired to DNA inside RNA polymerase; but the hairpin pulls the RNA away from the DNA and causes RNA polymerase to stop).
Rho-dependent termination
The RNA sequence does NOT form a hairpin; instead, a specific sequence (rut site) recruits the Rho protein (Rho binds to the rut site, and moves in the 5’ —> 3’ direction aka while RNA polymerase is moving, Rho is chasing it). Eventually, RNA polymerase slows down at the terminator region and Rho catches up to it. When Rho reaches RNA polymerase, it pulls RNA polymerase away from the DNA template, which breaks the RNA-DNA base pairing inside the transcription bubble. Once RNA polymerase detaches, transcription stops.
basically Rho yanks RNA polymerase off
Rho-independent vs. Rho-dependent termination
Rho-independent → RNA structure drives termination (no rho protein needed)
Rho-dependent → protein (Rho) drives termination
Polycistronic DNA
Bacteria will group multiple protein-coding regions (genes) downstream of a single promoter. When transcribed, this will make one long RNA that can be used to transcribe multiple polypeptides (proteins).
Eukaryotic RNA transcription in eukaryotes is similar to prokaryotes, with what few key differences?
1) Chromatin remodeling
2) Promoter diversity
3) Termination
Chromatin remodeling
Changes in chromatin structure that regulate DNA accessibility for transcription
DNA has to be loosened on the nucleosomes to be available for RNA transcription in eukaryotes, not prokaryotes
What are nucleosomes and what do they do?
A nucleosome is the basic unit of chromatin, consisting of DNA wrapped around a core of histone proteins. They control how accessible DNA is to transcription machinery.
What is heterochromatin? Why does heterochromatin prevent transcription?
Tightly packed DNA that prevents transcription. DNA is tightly wound around histones, so proteins cannot access it.
What are histone modifications?
Chemical changes to histones that regulate nucleosome interactions and DNA accessibility.
How is chromatin remodeling done in eukaryotes?
Histone acetylation reduces positive charges on histones, loosening interactions with the negatively-charged DNA backbone. DNA becomes less tightly bound to the nucleosome and it is easier for proteins to access DNA for transcription
Histone de-methylation makes histones more soluble in water (removing nonpolar methyl groups = more polar), loosening heterochromatin to make DNA more available.
Chromatin remodeling proteins can alter nucleosome structure near promoters (Special proteins can move or remove nucleosomes near promoters to make DNA easier (or harder) to access for transcription - can also loosen/tighten DNA wrapping)
In eukaryotes, RNA polymerase cannot bind DNA by itself
It needs help from transcription factors + accessible chromatin
Promoter Diversity (promoter utilization)
Eukaryotes have multiple types of RNA polymerases, which requires having multiple types of promoters
Different RNA polymerases each recognize different promoters (RNA pol I → rRNA, RNA pol II → mRNA, RNA pol III → tRNA)
Basal transcription factors are required for RNA pol recruitment - these proteins bring RNA polymerase to the promoter
Regulatory transcription factors modify transcription (control HOW MUCH RNA transcription occurs)
Basal transcription factors
the required proteins that bring RNA polymerase to the promoter
Includes TATA binding protein, other box element proteins; Basal TFs bind to RNA pol and position it on DNA at +1
Regulatory transcription factors
In eukaryotes, not prokaryotes - Not required but control how much transcription occurs; proteins that interact with enzyme or alter DNA structure
Activators - make RNA polymerase work better, increase RNA transcription by helping RNA polymerase bind better, recruit machinery
Repressors - decrease transcription; interfere with RNA polymerase function
Eukaryotic Transcription Termination
Eukaryotes lack intrinsic terminators (stop sequence) like prokaryotes have; they stop using RNA processing + RNA degradation
RNA polymerase continues to transcribe RNA well past the 3' end of protein coding domain, into the 3’ UTR (untranscribed region), which includes the PolyA sequence (AAUAAA). NOT THE SAME THING AS THE TAIL (tail added after mRNA transcription). This signal tells the cell to prepare to stop transcription.
The RNA binding protein called Rat1 recognizes the PolyA signal, which enzymatically cleaves the transcript and degrades the remaining RNA in a 5'->3' direction.
The transcribed RNA upstream of the cleavage site goes free (the useful RNA/pre-mRNA is released after the cut)
RNA is cut at the poly-A signal, released, and Rat1 degrades the leftover RNA to stop RNA polymerase.
RNA processing
Eukaryotes modify transcript (pre-mRNA) into messenger RNA (mRNA), removing the unusuable parts, and leaving just the usable mRNA
What are the different ways eukaryotes perform mRNA processing?
Polyadenylation
Capping
Splicing
Alternative Splicing
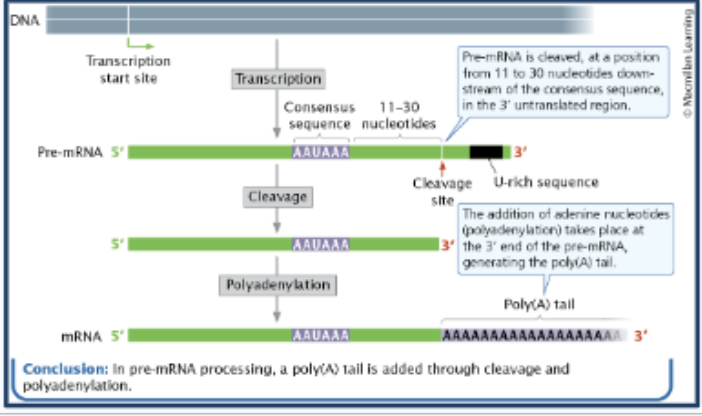
polyadenylation
Adds a 3' PolyA Tail (many As AAAAAAAA) to mRNA, protecting the 3'-end of the protein coding sequence from early degradation, assisting ribosome binding (PolyA Tail interacts with proteins that help ribosomes bind), and contributing to nuclear export (aids transport of mRNA out of the nucleus to the cytoplasm where translation occurs)
tail acts as a buffer
How does polydenylation influence the amount of protein made?
Long poly-A tail →
mRNA lasts longer (long poly-A tail = takes longer to degrade = mRNA lives longer)
ribosomes bind more efficiently (boosting translation efficiency)
translation repeats more times (because it helps with mRNA transport into the cytoplasm where translation occurs)
→ MORE protein made
Why is polydenylation necessary?
Pre-mRNA is cleaved at a position 11 to 30 nucleotides downstream of the consensus sequence, in the 3’ untranslated region.
Enzymes degrade RNA from the ends - so poly-A tail acts as a buffer
capping
A modified guanine (G) is added to the 5′ end of mRNA (5’ cap)
Aids translation
Protects mRNA form degradation by 5’->3’ nucleases (enzymes that degrade nucleic acids from the 5′ end)
Aids mRNA transport to cytoplasm
splicing
before translation; removing introns (non-coding parts) and joining exons (coding parts that become protein) together
pre-mRNA: exon — intron — exon
↓
mature mRNA: exon — exon
Splicing required for export from nucleus
Delayed splicing keeps pre-mRNA in nucleus
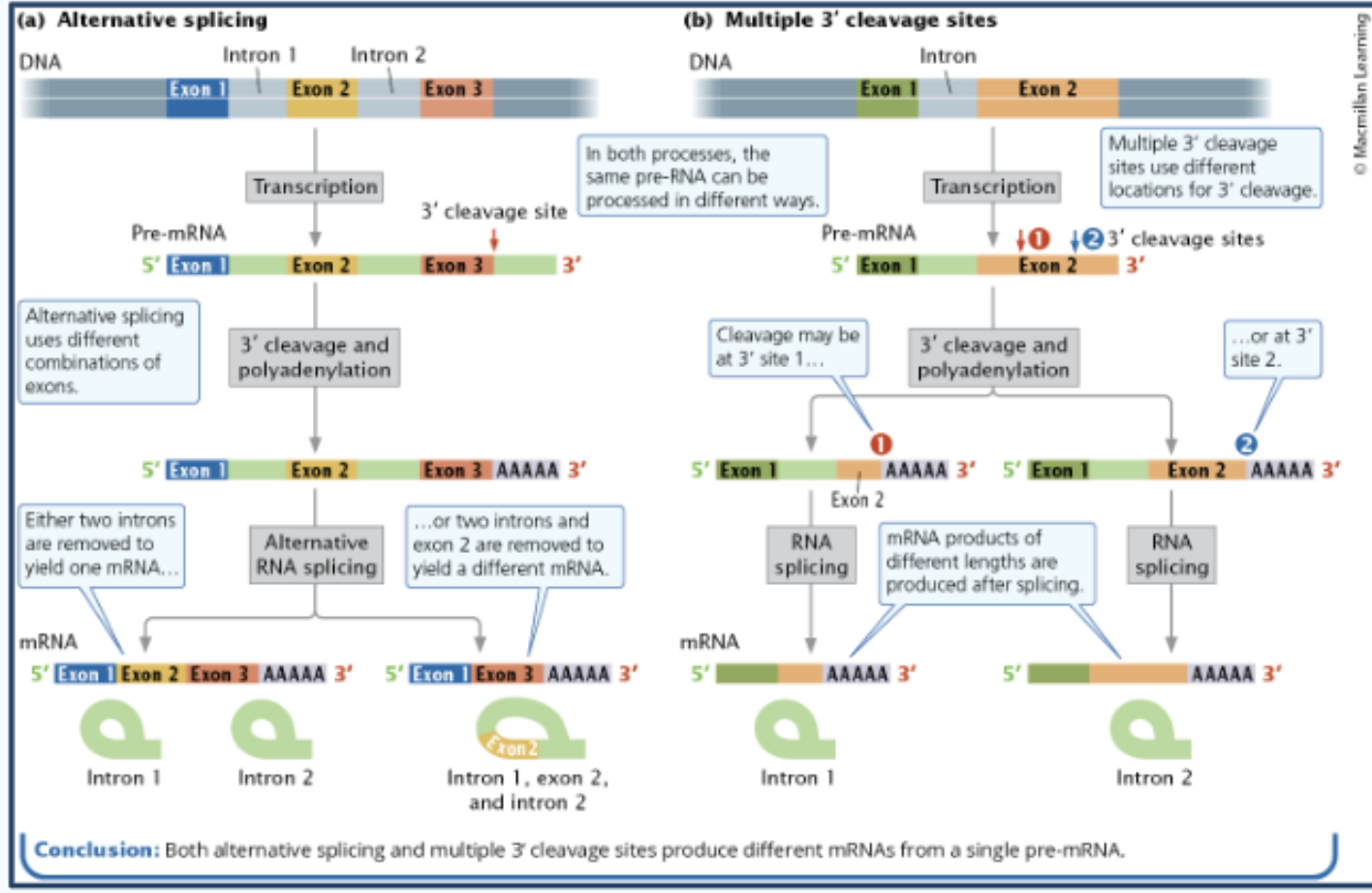
alternative splicing
multiple proteins form one pre-RNA; can shuffle exons making different mRNAs from the same transcript
tRNA processing
tRNA - transport RNA - small RNA molecule that transports specific amino acids to the ribosome
tRNA uses more than 4 nucleotides (Normally RNA = A, U, G, C, but in tRNA: some bases are chemically modified → “rare bases”) Ex. Uracil → Ribothymidine
tRNA-modifying enzymes chemically alter transcribed bases (after tRNA is made) to stabilize proper folding and aid secondary structure formation (how the RNA folds), ensuring the tRNA functions correctly during translation.
rRNA processing
rRNA = ribosomal RNA
Multiple rRNAs are transcribed together then processed (DNA —> one long pre-rRNA and then cut into separate functional rRNAs)
Transcript modification - Some bases of rRNA are methylated (methyl group is added) to stabilize RNA structure and help proper folding
Transcript cleaved - the long rRNA is cut into smaller pieces to multiple separate rRNAs
What pieces do we get?
5s, 16s, and 23s rRNAs in prokaryotes
5.8s, 18s, and 28s rRNAs in eukaryotes
These are the final products after they are cut
Translation
mRNA made by transcription is read by ribosomes to make polypeptides
mRNA —> protein
Protein structure is determined by…
interactions of amino acids
A change in amino-acid sequence of a polypeptide can alter protein function
Protein structure
Primary structure - sequence of amino acids
Secondary structure - hydrogen binding of backbone elements (alpha-helices or beta-sheets)
Tertiary, quaternary structure - interactions between R groups
Protein function is determined by…
Shape and amino acid R-groups
(Change structure = change in function)
What does the ribosome read?
mRNA is read in 3-letter codons, each coding for an amino acid
first 5’ AUG = start codon → Methionine (Met) - there can be other Mets on the protein but only the first one starts translation
This sets the reading frame
mRNA = nucleotide code (AUG, GCU, etc.)
Proteins = amino acids (Met, Ala, etc.)
tRNA is the translator between the two
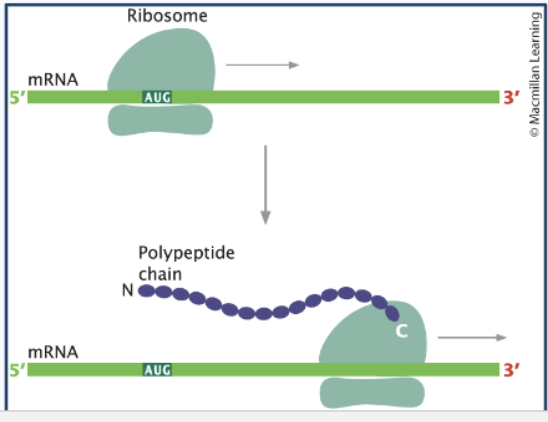
Structure of ribosomes
Ribosomes use codons in mRNA to direct polypeptide assembly
Small subunit - binds mRNA and reads codons (READER)
Large subunit - forms peptide bonds and builds the protein (BUILDER)
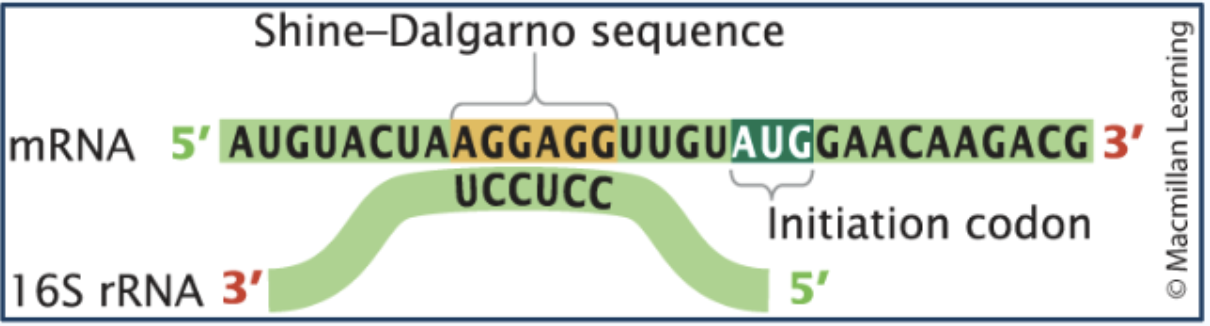
Initiation of prokaryotic translation
Prokaryotes recruit ribosomes to mRNA by having the ribosomes bind to a Shine-Delgarno sequence (5’-AGGAGG-3’) on mRNA, which positions the ribosome at the correct start codon.
The small subunit of a ribosome contains 16s rRNA that has a complementary sequence (3' — UCCUCC — 5') which pairs with the Shine-Delgarno sequence through base pairing. This pairing positions the ribosome at the correct start codon.
An initiator tRNA carrying fMet binds directly to the start codon (AUG) in the P-site with the help of Initiation Factor proteins. Once this occurs, the large ribosomal unit joins the small ribosomal unit, forming the translation initiation complex.
GTP is hydrolyzed, providing energy. Initiation factors are then released, with fMet positioned in the P-site (where the growing protein chain is held). The ribosome assembles with fMet in the P-site.
Initiator proteins
helper proteins that make sure translation starts correctly and in the right order
What are the initiator proteins used for in prokaryotic translation?
Initiation Factor 2 (IF2) binds with fMet and GTP to form a complex (acts like a delivery protein that brings fMet-RNA to the start codon AUG); GTP provides energy
Initiation Factors 1 & 3 (IF1, IF3) complex with small subunit (they bind the small ribosomal unit; IF3 prevents the large subunit from joining too early, ensuring the ribosome only assembles after correct positioning. IF1 blocks the A-site, preventing tRNA from entering the wrong place.)
“Prevents ribosome assembly”: because IF1 + IF3 keep the ribosome in a “not ready yet” state
Sequence of how initiator proteins are used
IF1 + IF3 bind small subunit → block premature assembly
mRNA binds (via Shine-Dalgarno)
IF2 + GTP brings fMet-tRNA to AUG
Correct pairing = GTP hydrolysis (for energy)
IFs leave → large subunit joins
Shine-Delgarno sequence
5’-AGGAGG-3’; positions the ribosome at the correct start codon on a mRNA (tells the ribosome where to start on mRNA); upstream of the start codon
Shine–Dalgarno controls ribosomal positioning
NOT scanning like eukaryotes
Initiation of eukaryotic translation
Eukaryotes use 5’ cap to recruit ribosome
In eukaryotes, the small ribosomal subunit binds the mRNA at the 5'-cap and scans for the first 5' AUG to use as the start codon, which is part of a short consensus called the Kozak sequence.
As in prokaryotes, an fMet initiator tRNA binds at the start codon and the ribosomal initiation complex assembles (fully assembled ribosome sitting at the start codon, ready to start translation, including mRNA (start codon) + small subunit + initiator tRNA (fMet) + large subunit).
IFs dissociate when fMet binds AUG (start codon). Ribosome assembles with the fMet in the P-site.
Initiator tRNA
The initiator tRNA carries fMet (formyl-methionine), a modified form of methionine used only to start translation.
It base-pairs with the start codon (AUG), helping position the ribosome correctly
It associates with initiation factors (IFs), especially IF2 + GTP
Once correct pairing with AUG occurs, GTP is hydrolyzed and initiation factors dissociate
The large ribosomal subunit joins, forming the full ribosome
The initiator tRNA remains in the P-site, ready for elongation
What happens next after initiation for translation?
2. Elongation → build the protein (adding amino acids one by one to the growing polypeptide chain)
3. Termination → stop and release protein (the ribosome reaches a STOP codon and releases the completed protein