Lecture 33: Loss of Heterozygosity
1/26
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
27 Terms
what is loss of heterozygosity
you have two copies of a gene but one is lost or inactivated so only one remains
no longer heterozygous
what does loss of heterozygosity make you more prone of
prone to inherent a mutant gene because it is no longer masked by the normal allele
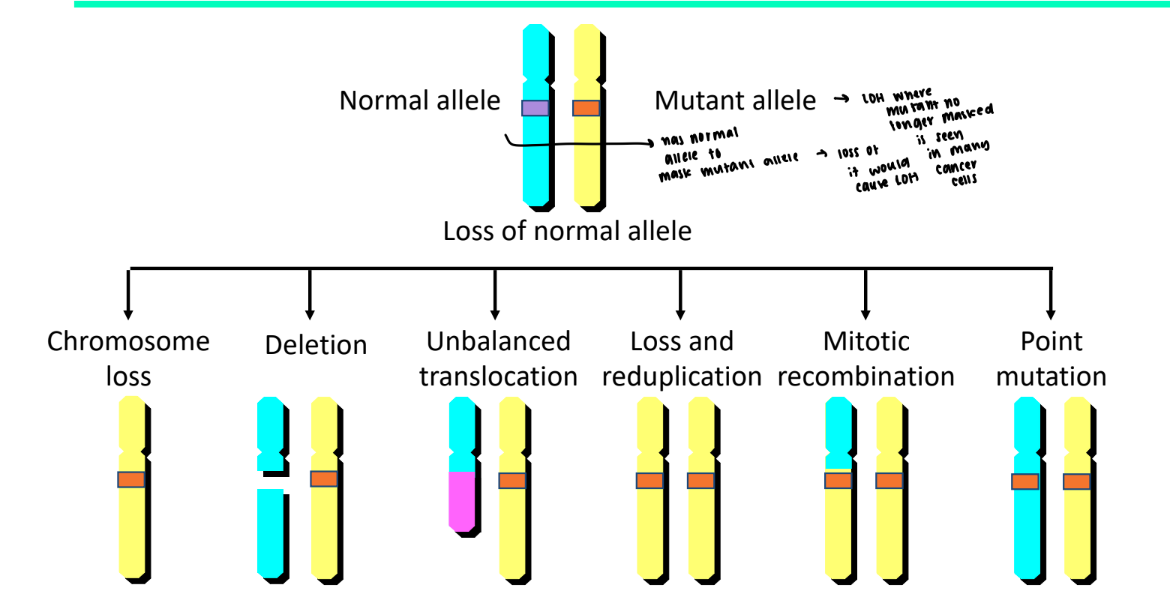
what is one example we’ve discussed that leads to loss of heterozygosity
crossover during homologous recombination because now both chromatids code for one frequency of the gene
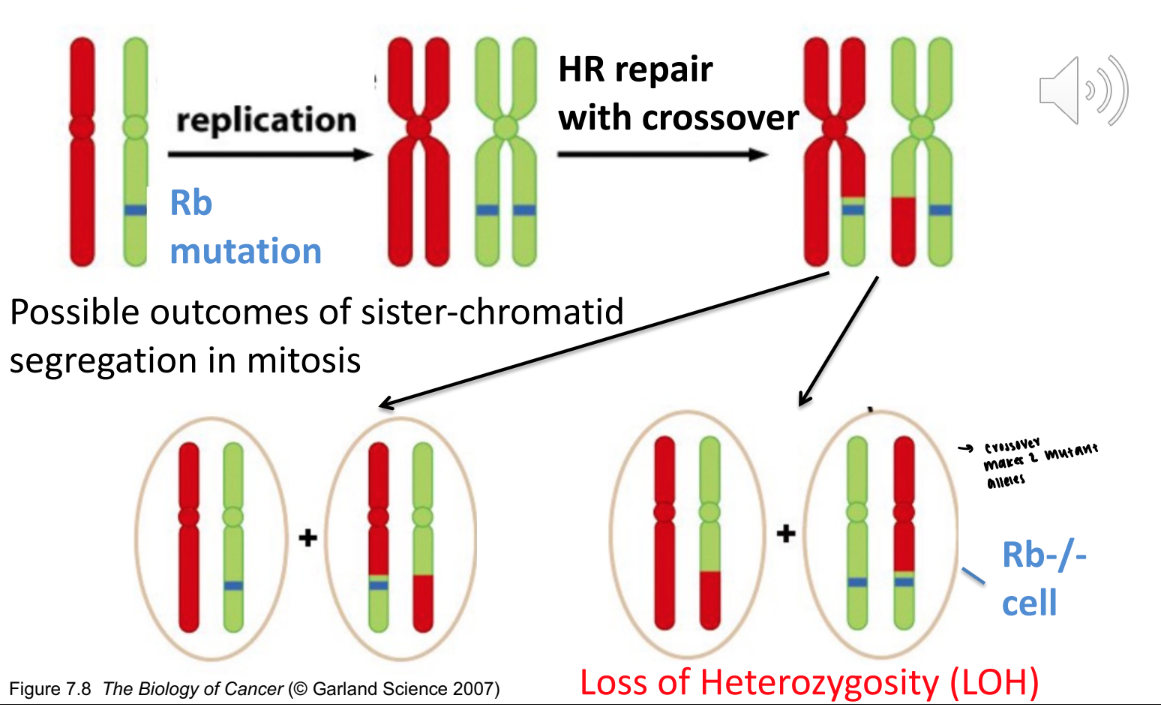
does loss of heterozygosity always lead to a malfunction
no, it just means that there is a loss of allele frequency, even if you aren’t losing function protein
red: function
green: nonfunctional
both are still considered loss of heterozygosity
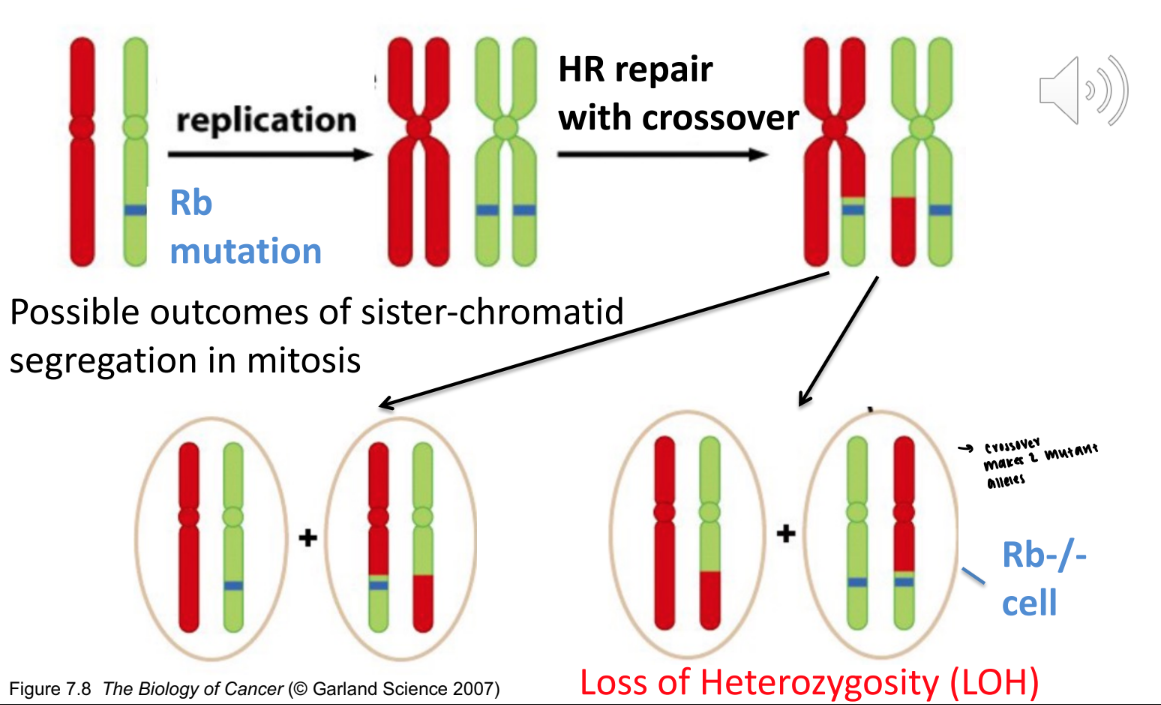
what is an example of loss of heterozygosity disease
retinoblastoma: eye tumor
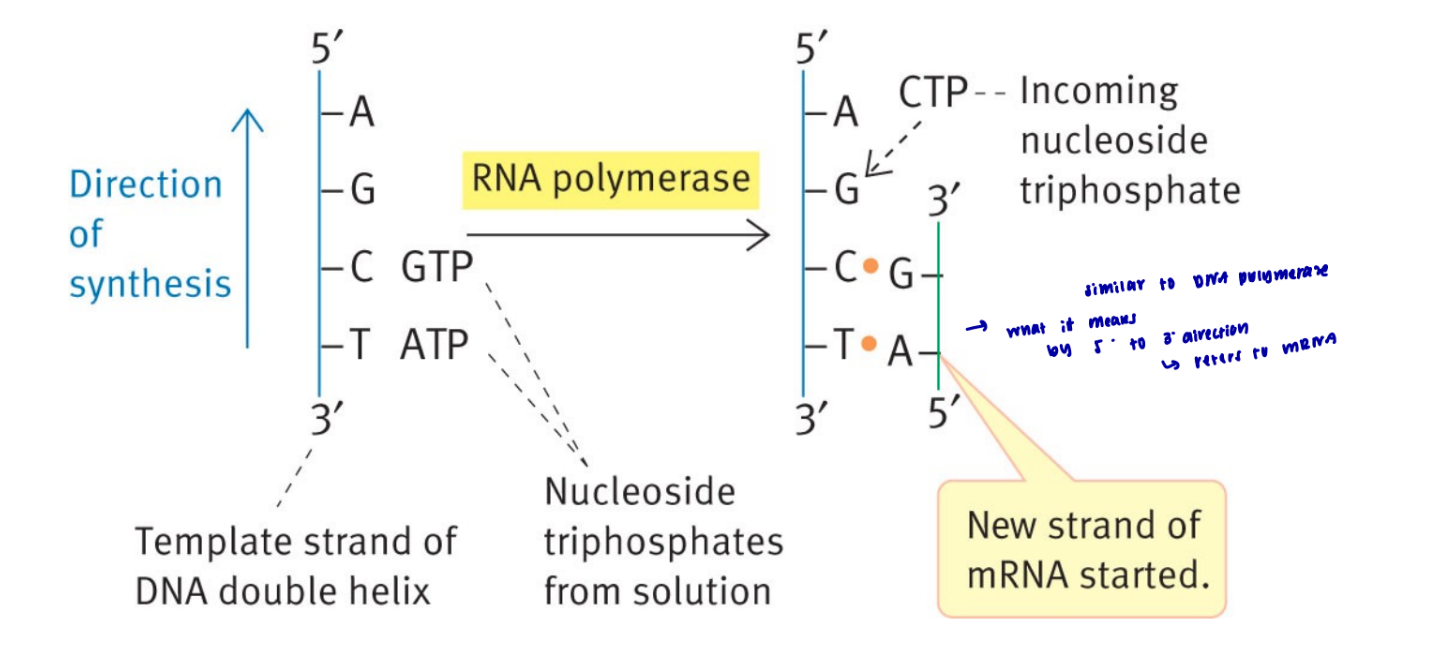
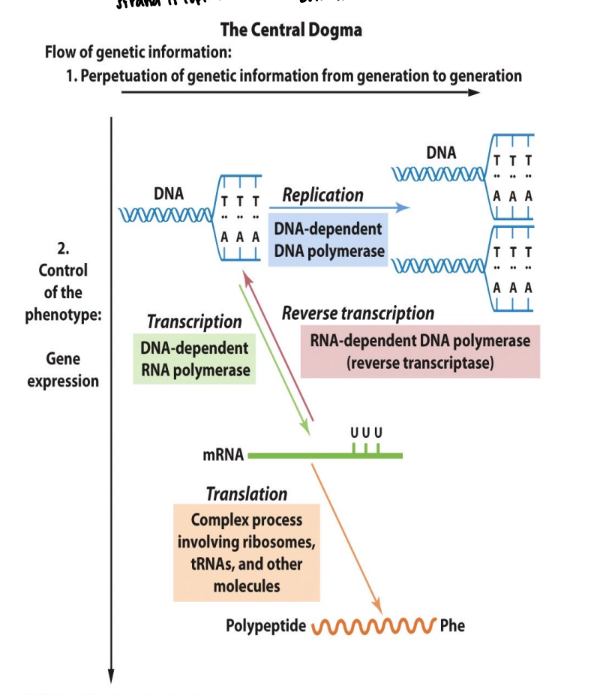
what is the difference between replication and transcription
replication requires both strands to be copied
transcription has only one mRNA made bc only one DNA strand is copied
what is transcription
the synthesis of a single strand RNA transcript complementary to one strand of DNA of a gene
what are snRNAs
small nuclear RNAs - structural components of spliceosomes
whare are miRNAs
short single stranded RNAs that block expression of complementary mRNAs
what is the main difference between ribose and deoxyribose
ribose has a 2OH group whereas deozyribose has a 2H group, which makes ribose extremely unstable so it can’t code for anything genetic
what is the estimate amount of rRNA, tRNA, and mRNA in prokaryotes
more rRNA and tRNA becuase those can be recycled, mRNA is very small amount because it gets quickly degraded after it gets transcribed or else more proteins would constantly be made from the mRNA that we might now need
what other RNAs do eukaryotes have
small nuclear RNA that are part of the splicisome complex and splices introns from RNA
signal recognition particle: guides protein transport after translation to their designated areas
microRNA that bind to mRNA after transcription and translation to induce degradation
small interfering RNA as a result of viral infection and can bind to mRNA to induce degredation
telomerase RNA which guides the synthesis of telomere with telomerase if we need to extend certain cells' chromosomes
what makes RNA and DNA polymerase different from each other
RNA polymerase can start adding bases without the need of a 3’ OH group
where does DNA unwinding usually start at
AT regions because this is where there are 2 h bonds, making it weaker and easier to break apart
in what direction does RNA polymerase transcribe
5 → 3, in the direction of the opening of the fork → how it knows which strand to use as the template
what does sigma subunit in the prokaryotic holoenzyme do
recognizes the promotor region on the DNA to start transcription
it’s not part of the holoenzyme, it just guides the RNA polymerase to the promotor region and it knows where to go through cell signaling
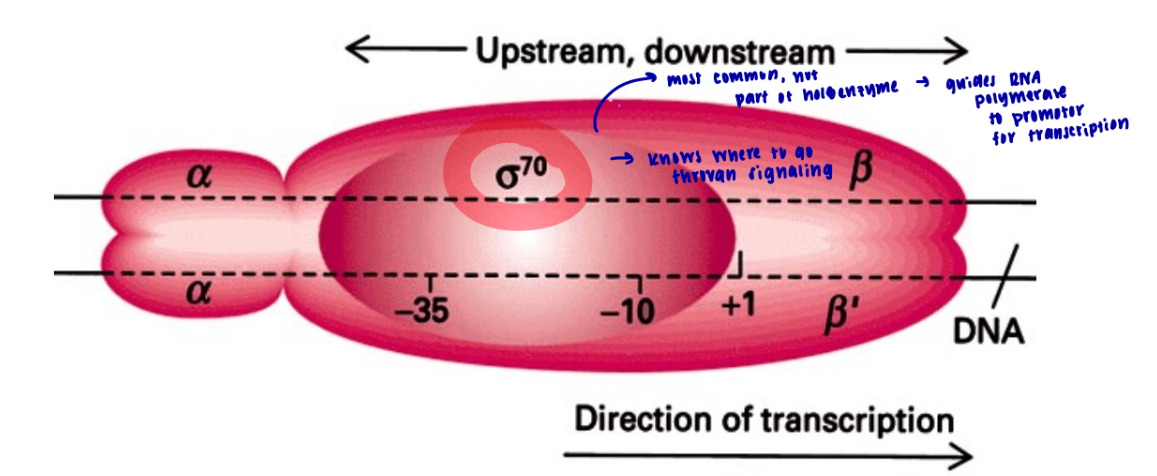
what is the most common sigma subunit
sigma 70
what is upstream and downstream in transcription
going left, or toward the negative numbers is upstream (anything before start site of transcription) whereas going right toward positive number is downstream
explain the geography of prokaryotic gene
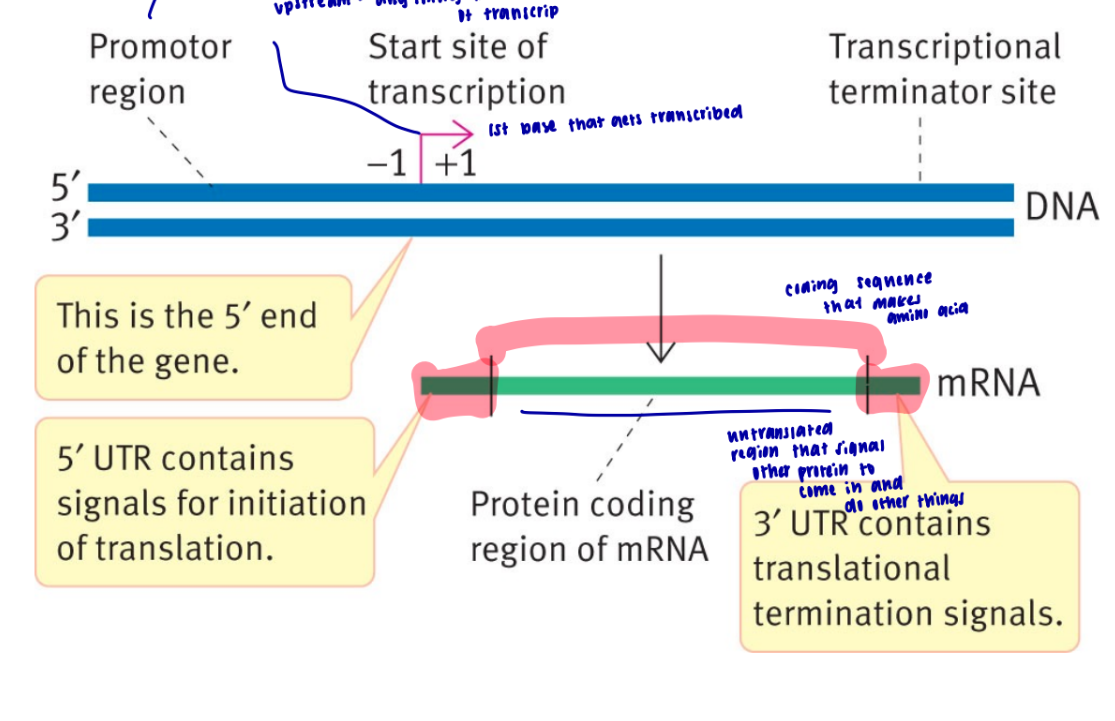
what does consensus sequence mean
it means it codes for the same thing, but has different variations
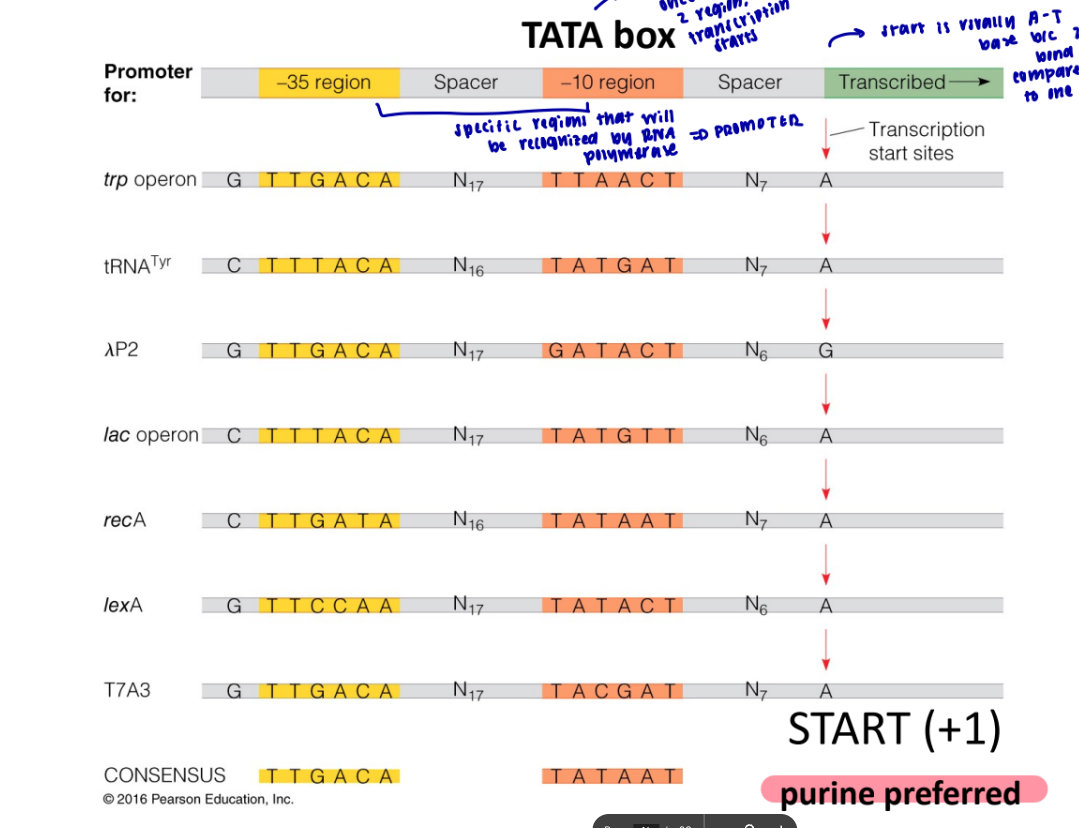

what are the two regions on the promotor that need to get recognized before transcription can occur for prokaryotes
-35 and -10, tata box
where is the TATA box in prokaryotes
in -10 region
what does transcription require
template (DNA)
rNTPs (what will build the mRNA through phosphodiester bonds linkage)
metal ion (to shield the negative charges from each other like DNA polymerase)
how many RNA polymerase do eukaryotes have and what do they code for
Pol I: makes rRNA
pol II: makes mRNA
pol III: makes tRNA and small RNA
how many RNA polymerase do prokaryotes have
only one
do eukaryotic cells have a sigma for promotor recognition
no, it has transcription factors, proteins that can bind to DNA and serve similar roles in recognizing promotoes and activating transcription
what does the transcription of mRNA look like
similar to DNA where it brings in rNTPs that are complement to the DNA bases and creates a phosphodiester bond between the bases so that it can link and become one chain
always occur in a 5 → 3 direction
only one strand if formed whereas in DNA, both strands are the template and then you end up with two