Prokaryote Biology Exam 3
1/74
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
75 Terms
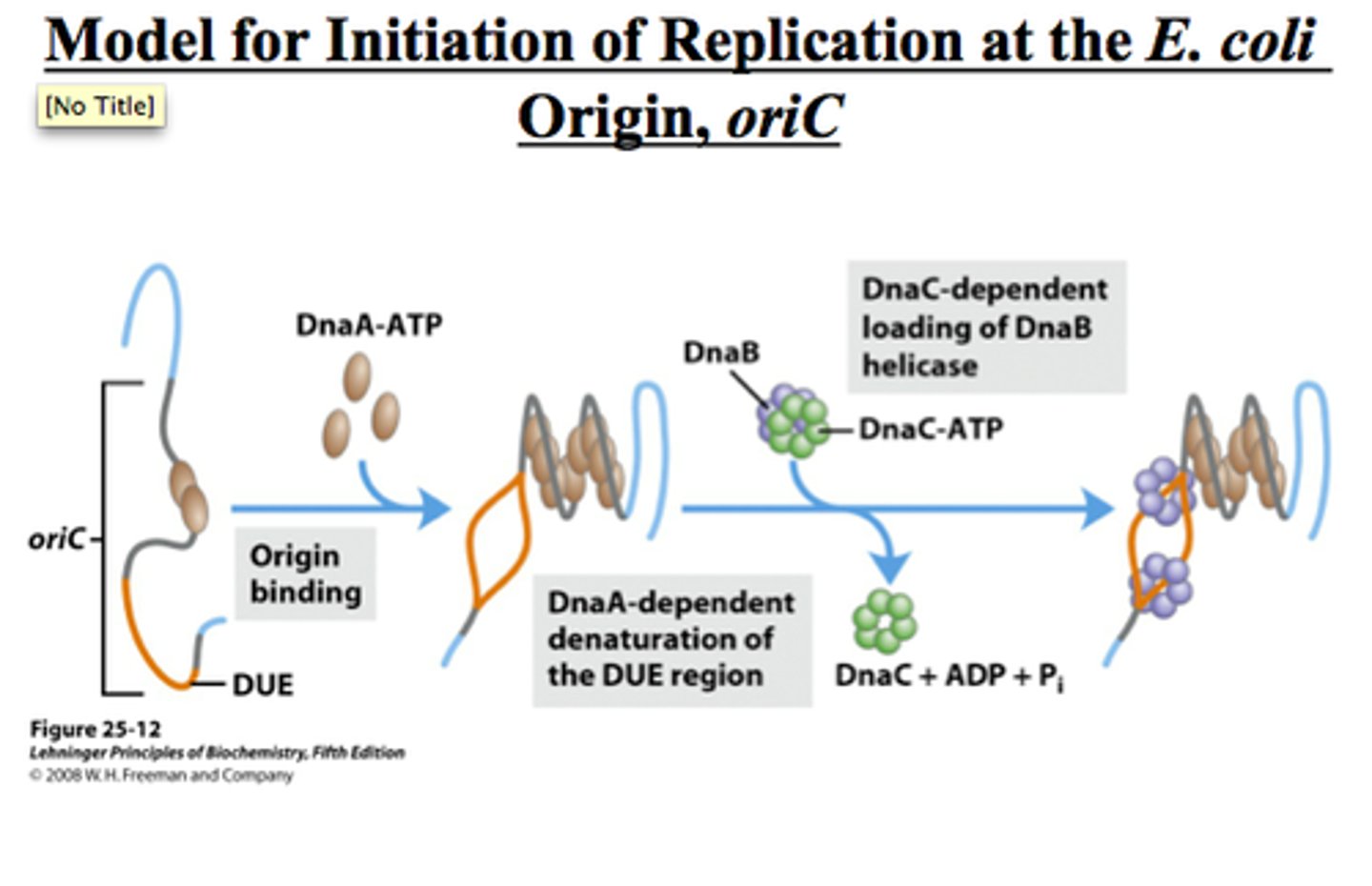
DnaA
- activates INITIATION of replication
- needs to be in its active form (DnaA-ATP)
- binds to the origin of replication (oriC)
regulation of levels of DnaA
-cell mass
-growth rate
-cellular health
gene
region of DNA encoding a particular polypeptide chain or functional RNA
operon
a collection of genes that are transcribed into a single RNA
promoter
the site on DNA where RNA polymerase binds and begins transcription
--> non coding DNA sequence UPSTREAM of structural gene (coding sequence) required for transcriptional initiation
cistron
functional unit of RNA containing information from a single gene
polycistronic
RNA produced from an operon containing information from more than one gene
open reading frames (ORF)
DNA sequence predicted to encode a protein, can be translated, and has start/stop codons
codon
sequence of 3 bases in mRNA that code for a specific amino acid
start codon
special codon that signals the start of translation (AUG)
stop codon
do not encode an amino acid and trigger that end of translation (UAG, UAA, UGA)
DnaB
-helicase enzyme
-requires ATP (energy)
Primase
-RNA polymerase
-synthesizes RNA primer
%GC and Tm
-Tm (midpoint temperature) is proportional to the %GC content in a thermal denaturation graph
-the absorbance at 260 nm increases going from double strands to single strands (*melting)
-sigmoidal shape graph
oriC
origin of replication (during initiation for DNA replication)
purine
adenine and guanine (A,G)
pyrimidine
Thymine, cytosine, and uracil (RNA) (T,C,U)
DNA ligase
-seals nick in DNA that have adjacent 5'-P and 3'OH (makes the last phosphodiester bond during the nick repair
-requires energy
-most ligases use ATP
Regulon
a group of genes/operons that are coordinately regulated
ter sequences
termination sequences that trap/block replication forks
Tus
(terminus utilization substance)
-binds to ter sequences
-blocks DnaB (helicase)
sigma factor
- transcription initiation
-ONLY in bacteria
-binds to RNA polymerase and recognizes the promotor
-key to initiation
-when bound to RNA polymerase = holoenzyme
TBP
"TATA binding proteins"
- recognizes TATA box sequence
TFB
"Transcription factor B"
-recognizes BRE (B recognition element) sequence
Two types of termination signals during transcription
Rho-dependent and Rho-independent
Rho-dependent
-relies on protein called Rho
-strong pause site at 3' end of the gene
-sequence specific
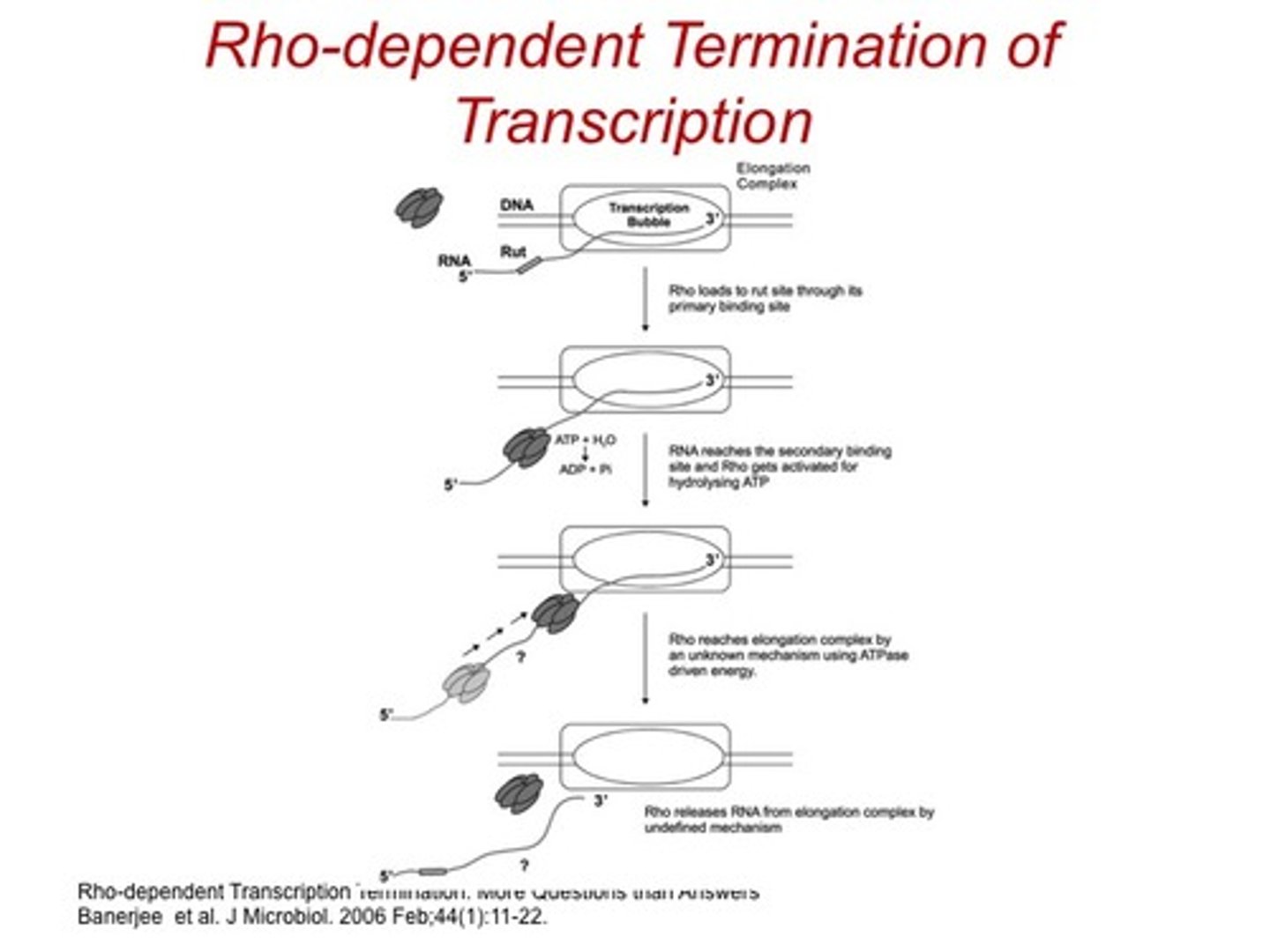
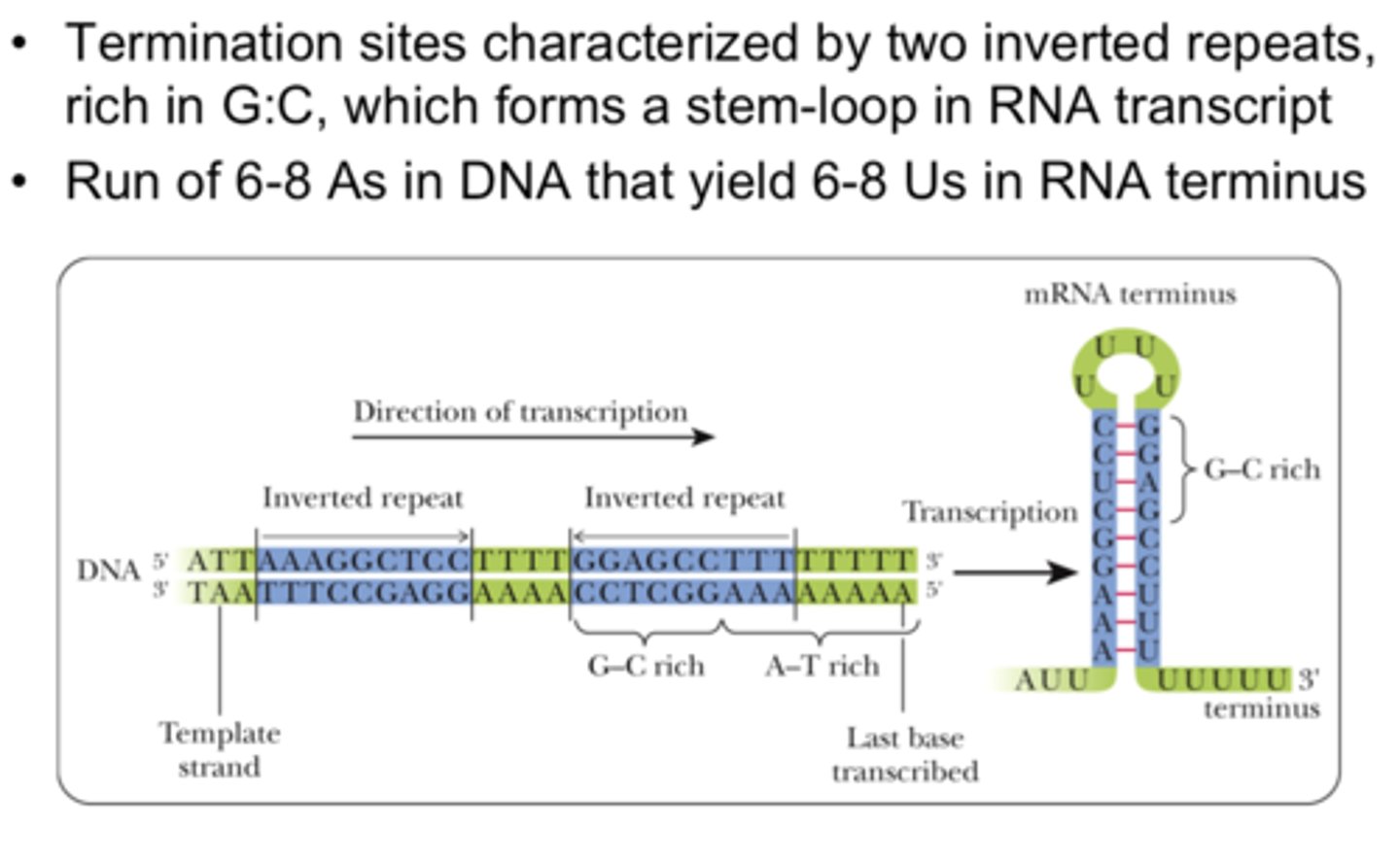
Rho-independent
-requires a GC rich region of RNA
-and 4-8 consecutive U residues
-sequence specific
peptidyl transferase
-in ribosome
-makes the peptide bonds between the amino acids to make a polypeptide
Shine Dalgarno sequence
-TRANSLATION initiation
-purine rich (A and G)
-Ribosomal binding site (RBS)
-in bacteria and archaea
fMet
-start codon in bacteria
-formylmethionine
Chaperons/chaperonins
assist in protein folding
(DnaK, DnaJ, GroES, GroEL --> helper proteins/heat shock proteins)
RecA
-protein that mediates recombination
-found in all three domains
-recognizes single stranded DNA
Transformation
the process of importing free DNA into cells (ex: griffith experiment)
Conjugation
"bacterial sex"
-requires cell to cell contact
-requires pilus (sex pilus)
-F plasmid (plasmid encoded mechanism)
F plasmid
-fertility plasmid
-components: Tra region, oriT, IS3, IS2, oriV
oriT
origin of transfer during conjugation
IS
insertion sequences
phage
a virus that infects prokaryotes (bacteriophage)
lytic
a viral life cycle in which the virus lyses the cell releasing progeny
virulent phage
phage that lyses (kills) host cell after infection (nontemperate phage)
lysogeny
a viral life cycle in which the viral genome integrates into and replicates with the host genome (can become lytic)
temperate phage
phage capable of lysogeny
prophage
a phage genome integrated into the host genome
lysogen
a prokaryotic cell that harbors a prophage
phage conversion
alteration of a host cell by lysogenization
Examples:
-lysogens become immune to further infect by the same type of phage
-corynebacterium diphtheriae
transducing phage particle
-during generalized transduction
-does not degrade host DNA or lyse cell
-allows transfer of any gene
-low frequency and low efficiency
restriction endonucleases
allows enzymatic cleavage (restriction) of alien DNA
DNA methylase
protective methylation (modification) of host DNA
CRISPR
"clustered regularly interspaced short palindromic repeats"
- and adaptive system
-consist of repeats and spacers that do not encode proteins
-a primitive microbial immune system
three stages of CRISPR
1. spacer acquisition
2. crRNA processing
3. effector stage
translocasome complex
transfers ssDNA into cell
-used by naturally competent gram positive bacteria
anticodon
hydrogen bonds with the mRNA codon specifying an amino acid (reading the codon)
3' (acceptor) end
binds the amino acid
charged tRNA
-each tRNA must be charged with the proper amino acid before it encounters the ribosome
-charged by using aminoacyl-tRNA synthetases (20 of these for each amino acid)
sense strand
coding strand (5' to 3")
antisense strand
template strand (3' to 5')
replisome
forms during initiation stage of DNA replication
-forms before DnaB unwinds DNA
Catenane
???
"catenane resolution"
-during termination stage of DNA replication
rolling circle replication
unidirectional plasmid replication
phosphodiester bond
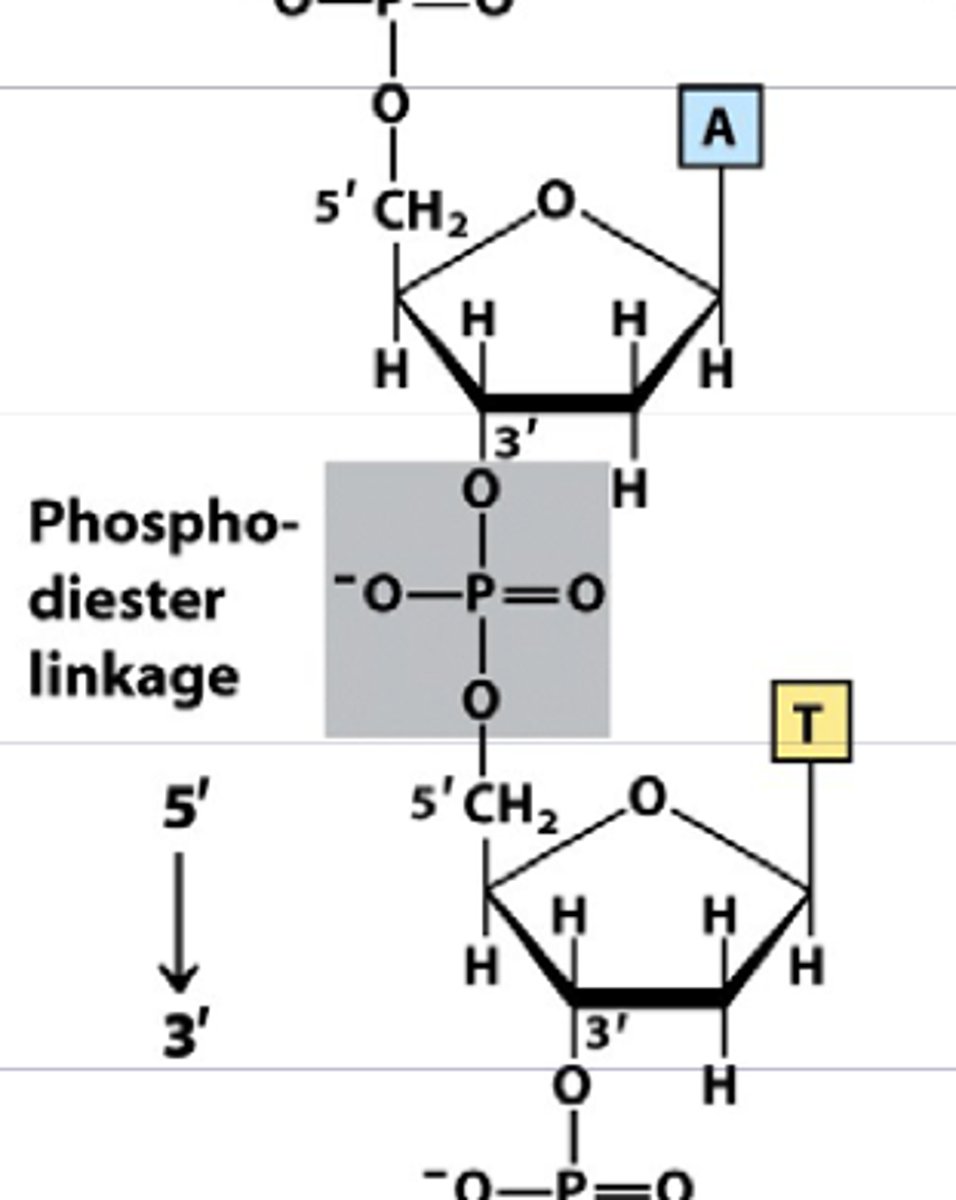
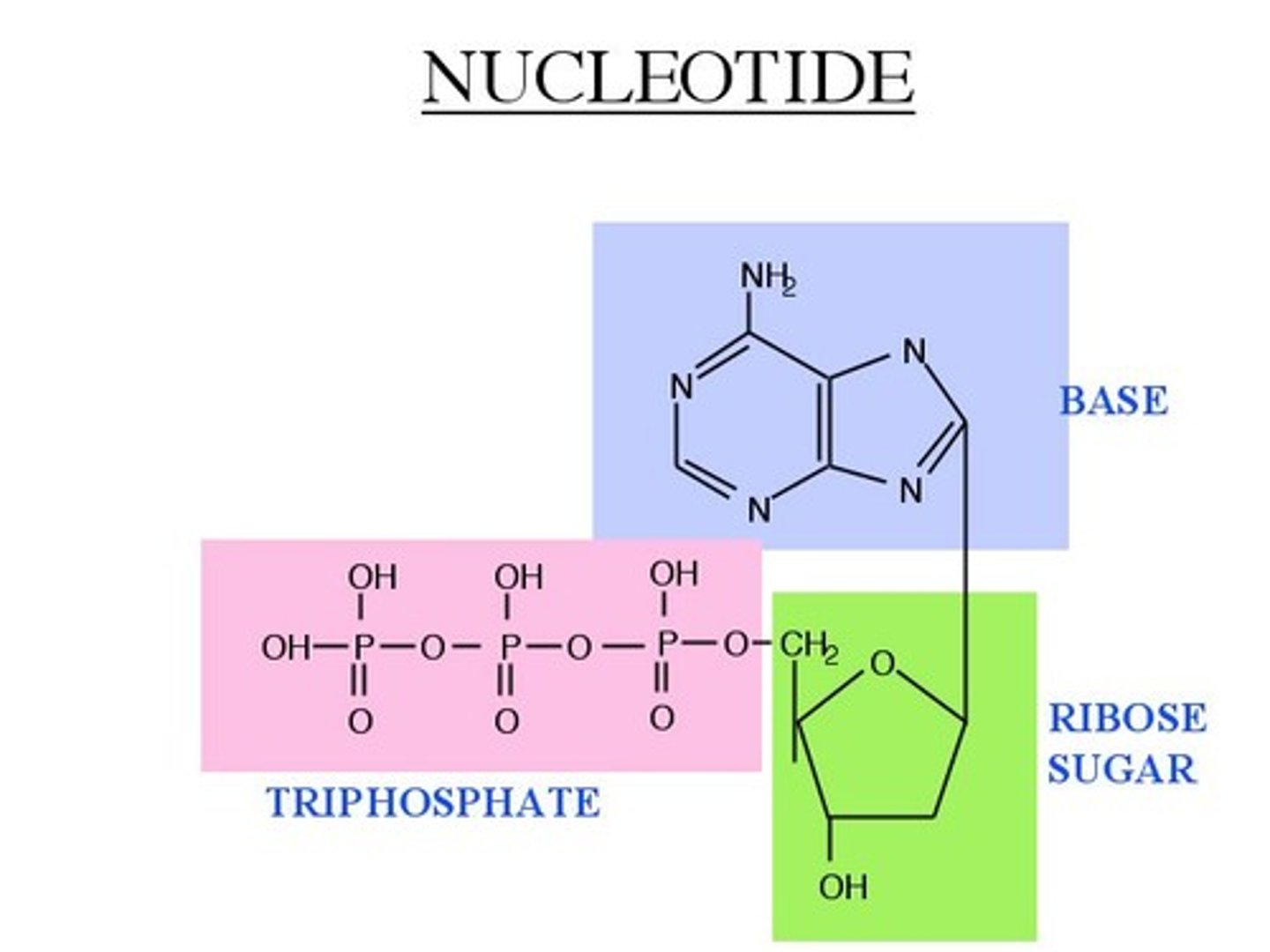
nucleotide
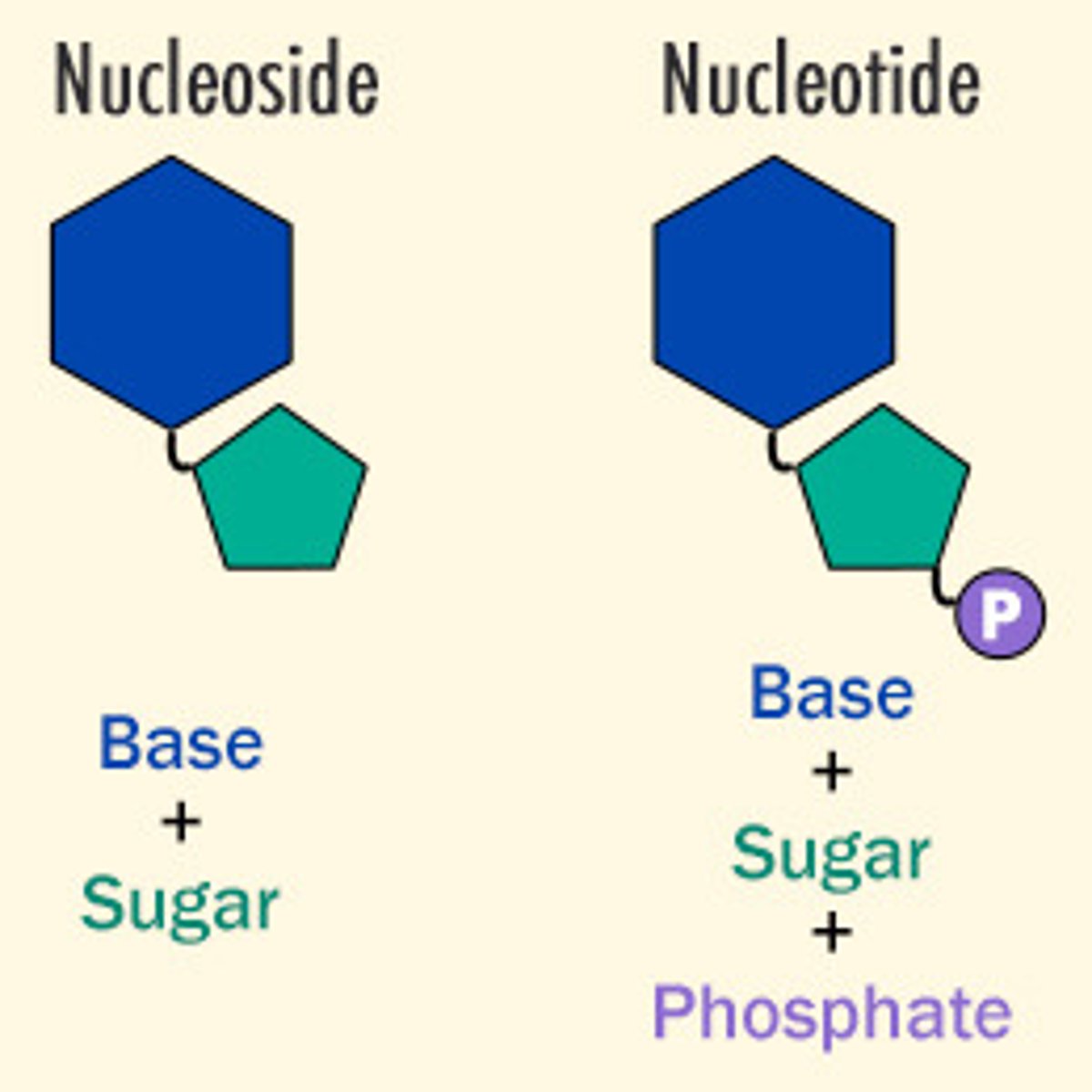
nucleoside
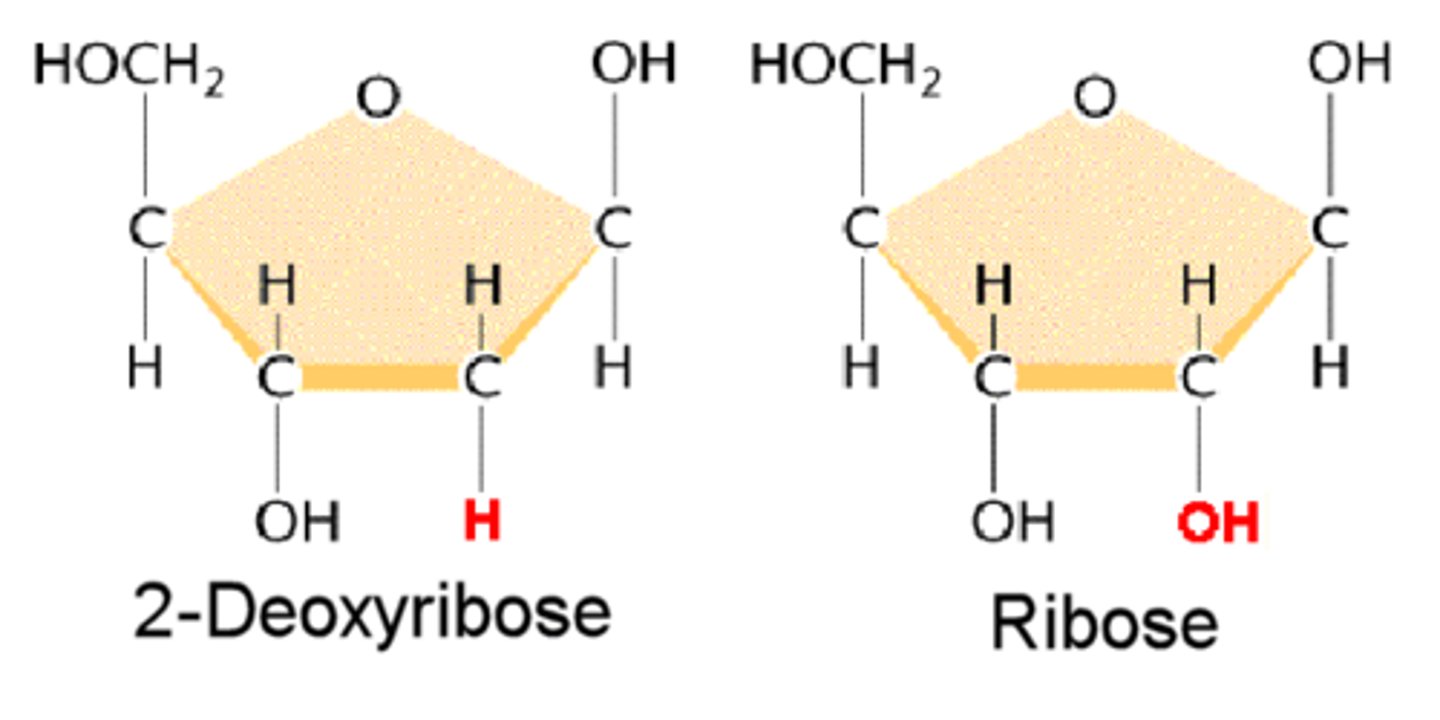
deoxyribose
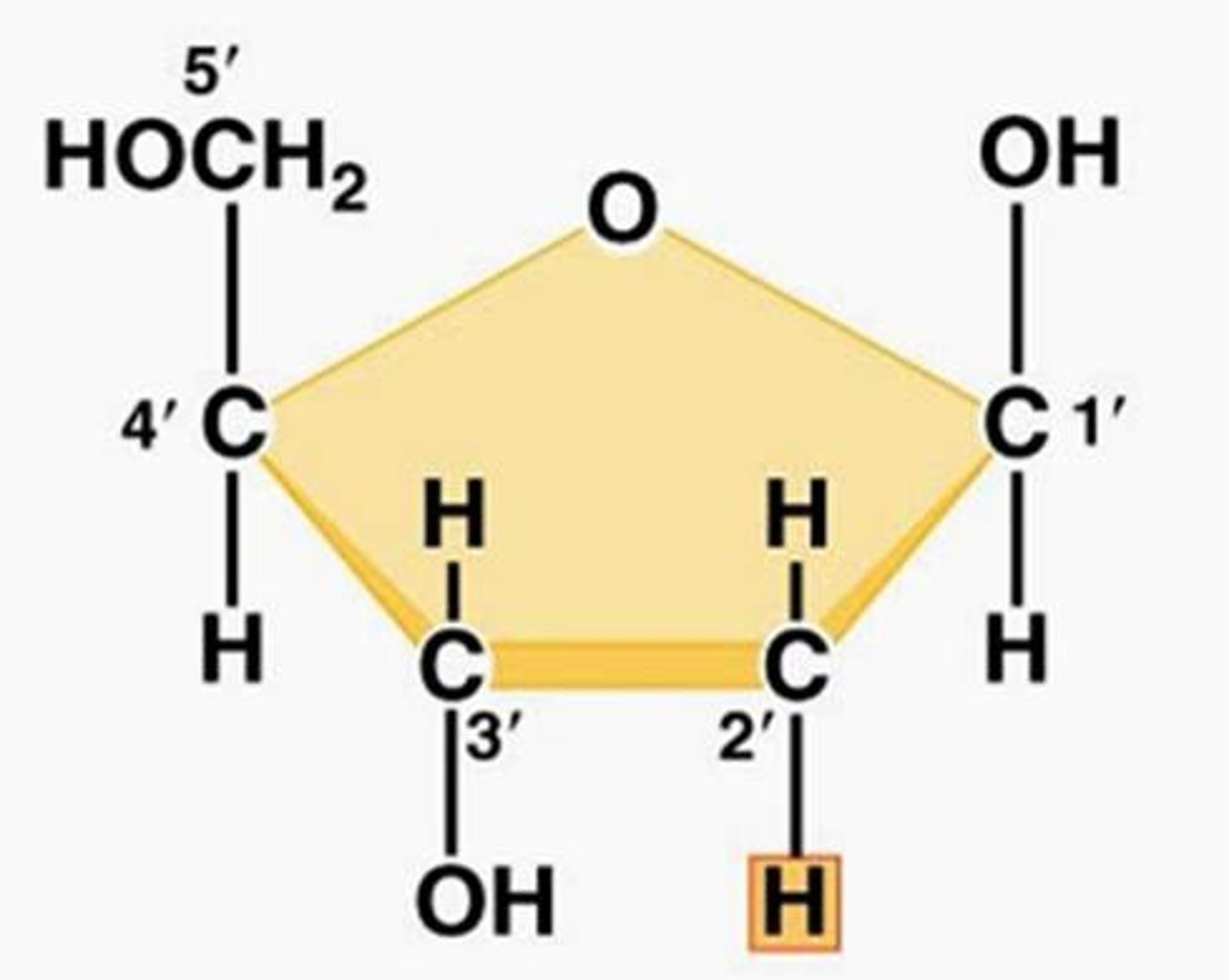
ribose
okazaki fragments
-in elongation stage of DNA replication (replication fork)
-present in the lagging/discontinuous strand (5' to 3')
integrase
Merodiploid
RepA
Inverted repeats
mRNA
-encodes protein
-thousands of types
-1500 nt
- 3 to 5 min
rRNA
-synthesizes protein
-three types
-5S = 120 nt; 16S = 1500 nt
-hours
tRNA
-shuttles amino acids
-27 types (set) --> 86 genes
-80 nt
-hours
sRNA
-controls transcription and translation
-20 to 30 types
-less than 100 nt
-variable half life
tmRNA
-free ribosomes on damaged mRNA
-about one type per species
-300 to 400 nt
-3 to 5 min
catalytic RNA
-enzymatic reactions
-size varies
-3 to 5 min half life