BIOB11 midterm lec 1-6
1/238
Earn XP
Description and Tags
learning objectives and more
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
239 Terms
Identify differences in the process and function of mitosis vs meiosis (4 things that meiosis has that met doesn’t)
meiosis ends up with ½ the gen mat as mitosis
Mei includes recombination
Mei includes Synapsis: pairing of homo chromosome
Mei includes reduction division
Identify similarities in the process and function of mitosis vs meiosis (2)
they both begin with replication of chromosomes, and creation of sister chromatids
functions of the mitotic spindle in meiosis
organizes chromosome and pulls them to opposite poles (and is organized by centrosomes)

microtubules fxn in meiosis
main component of the mitotic spindle apparatus
kinetochore fxn meiosis
proteins that connect spindle fibres to chromo’s centromeres.
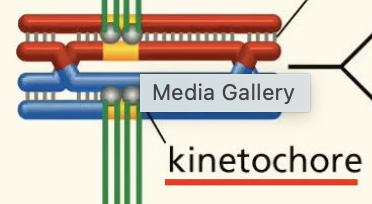
cohesions fxn in meiosis
all along the chromosomes ensuring sister chromatids stay together. (also at centromeres)
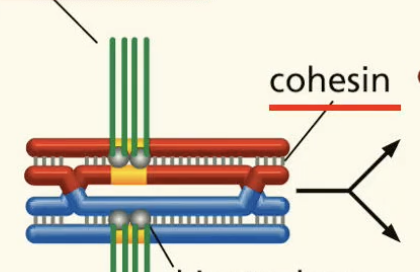
centromere fxn in meiosis
where the spindle apparatus connects to the chromosomes (by kinetochore Ps)
Law of segregation
an indv’s Mat and pat’s chromosome segregate from e/o during gamete formation
law of independent assortment
Segregation of a pair of alleles for one trait (gene) has no
effect on the segregation of alleles for another trait
what steps in meiosis allow laws of inheritance to apply
crossing over and synapsis
what is crossing over (genetic recombitiations) and when does it happen
allows for gametes to have new allele combos!
chromosome from one parent crosses over with chromosomes from other parent exchanging some bits.
prophase I
components of DNA ___+____+____=________
sugar, phosphate, nitrogen base= nucleotide
purines
A and G
pyrimidines
C and T
nucleotide vs nucleoside
Nucleosides:
Sugar + nitrogenous base (no phosphate!!)
Nucleotides:
Nucleoside + phosphate group(s)
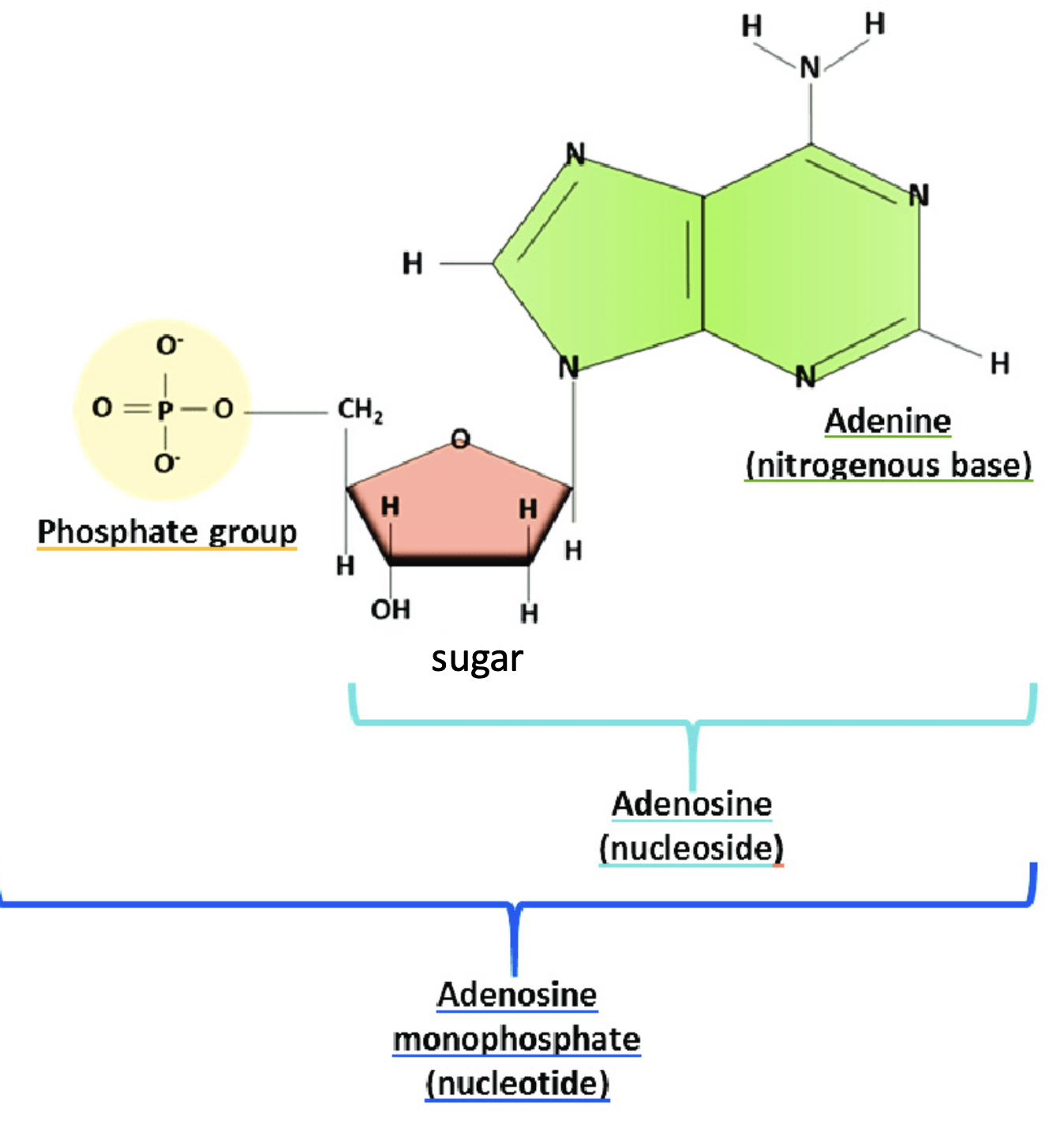
DNA 3 functions, and how structure allows them to happen
1. Storage of genetic information: linear bases sequences code for genes.
2. Replication and inheritance: strands can be pulled apart, and since strands can be templates.
3. Expression of genetic message: strands can be separated for RNA synthesis (transcription) to make proteins= expression
Identify which organisms use DNA as the keeper of their heritable information (3)
bacteria, archea, eukaryotes
Explain why mutations in germline vs somatic cells have different effects on offspring
only mutations in the gremlin cells will be passed on to offspring
Describe the flow of information in the Central Dogma of Biology (Directionality in the flow of information)
DNA, RNA, Protein
a gene is a:
vessel for heritability and change
Major and Minor grooves of DNA are:
protein binding sites
each spiral of DNA is how many bases?
10
phosphate is on what prime?
5’ phosphate (on carbon 5)
hydroxyl on what prime?
3’ hydroxyl (on carbon 3)
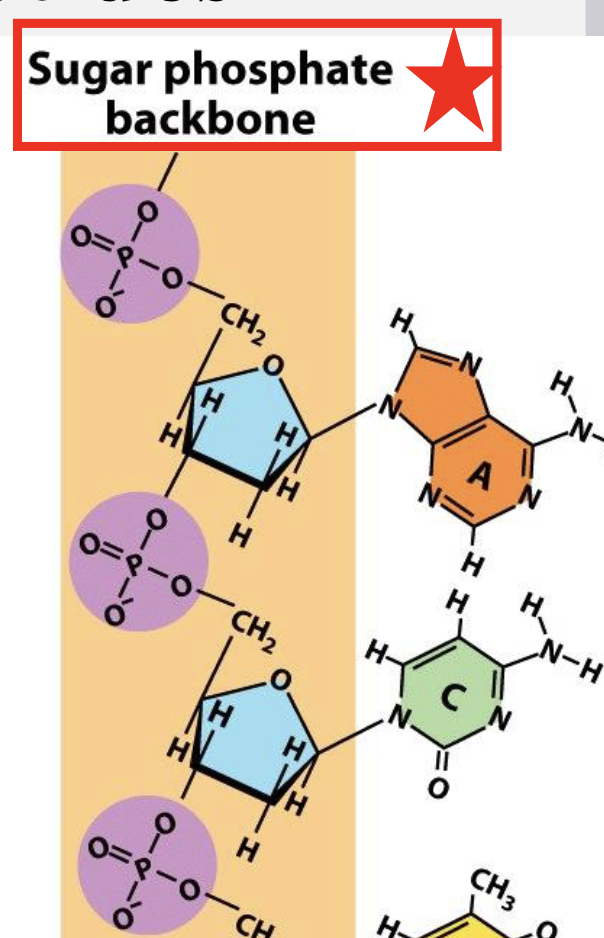
what end is the red box on?
5’
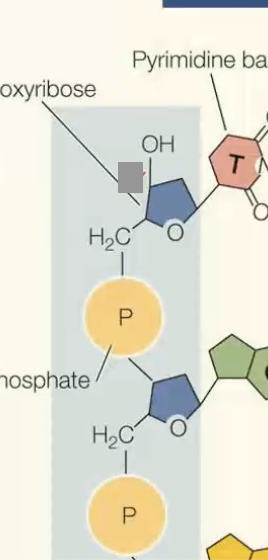
what end is grey box
3’
what are the 3 main genetics discoveries in order?
inheritance exists, chromosomes, DNA
what is chiasma
site of crossing over between 2 chromosomes during genetic recombination.
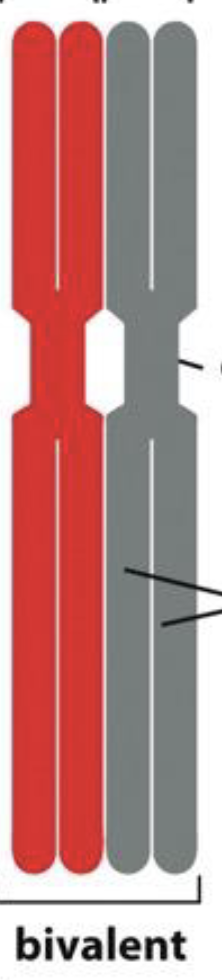

who are these cuties
homochromos/ bivalents/ a tetrad
prokaryote genome and a result
double stranded circular DNA in nucleotide region
results in lots of horizontal gene transfer= lots of external DNA
eukaryote genome shape
double stranded, in nucleus
pathogens, chloroplasts and mitochondria’s dan can be infused into a eukaryotes DNA T or F
true
explain viral DNA
singe or double stranded, needs to use host to replicate and protein synthesis (not alive!)
as gene density increases, gene coding % ____
increases too
gene density in prokaryotes vs eukaryotes
high in Process
low in euks
does genome size correlate with complexity?
no (called c value paradox apparently)
how can we have a larger genome size and smaller amt of genes encoding Ps?
lots of repetition!
is genome size correlated with number of genes?
no. most of it is noncoding DNA with other fxns, ex: regulatory (the actual gene section is so small and P coding regions even smaller!)
Identify the % of human DNA that encodes protein vs other types of sequences
1-2%
does the number of protein coding genes indicate complexity of an organism?
no! g-value paradox
an x shaped chomosome indicates that ____ has occurred
replication
genes are all the same size T or F
FALSA: can be all sizes
positive supercoiling (to relieve strain) occurs which direction of a DNA strand being unwound?
in front, overwinding
negative supercoiling (to relieve strain) occurs which direction of a DNA strand being unwound"?____
behind, underwinding
Topoisomerases
enzyme that uncoils DNA poisitive supercoiling, while it is being unwound.
DNA charge and why is it important
- because of the P group
can easily interact with histone’s positive charge
what are the 4 parts of the histone octamer
H2A, H2B, H3, AND H4, connected in a histone handshake
chromatin ARE MADE OF (3)
dna wrapped around histones, which then package themselves into nucleosomes
which histone serves as a linker of nucleosomes and what is the new shape it forms called?
H1, Histone H1 can line up nucleosomes end-to-end into two stacks called 30nm fibre (increases ration 6 fold)
how many nucleotides in a nucleosome
around 200
after the 30nm fibre is created, it can be folded even further with he help of what P? how?
• 30 nm fiber may further be gathered into a series of large loops
• These loops are attached to protein scaffolds
DNA-packaging ratio in a
mitotic chromosome is ————
fold!
10000
function of histone N-terminal tails
subject to covalent modifications and site of methylation and acetylation of histones
Euchromatin vs heterochromatin
Euchromatin: DNA that is less compacted and functionally active (accessible for
protein binding and transcription). Genes here can be expressed
Heterochromatin: DNA that is highly compacted and has little to no functional
activity
two types of heterochromatin and function
Constitutive heterochromatin
• Permanently silenced DNA
• Includes regions around telomeres and
centromeres
• Contains DNA repeats and few genes
Facultative heterochromatin
• Inactivated during certain phases of
organism’s life
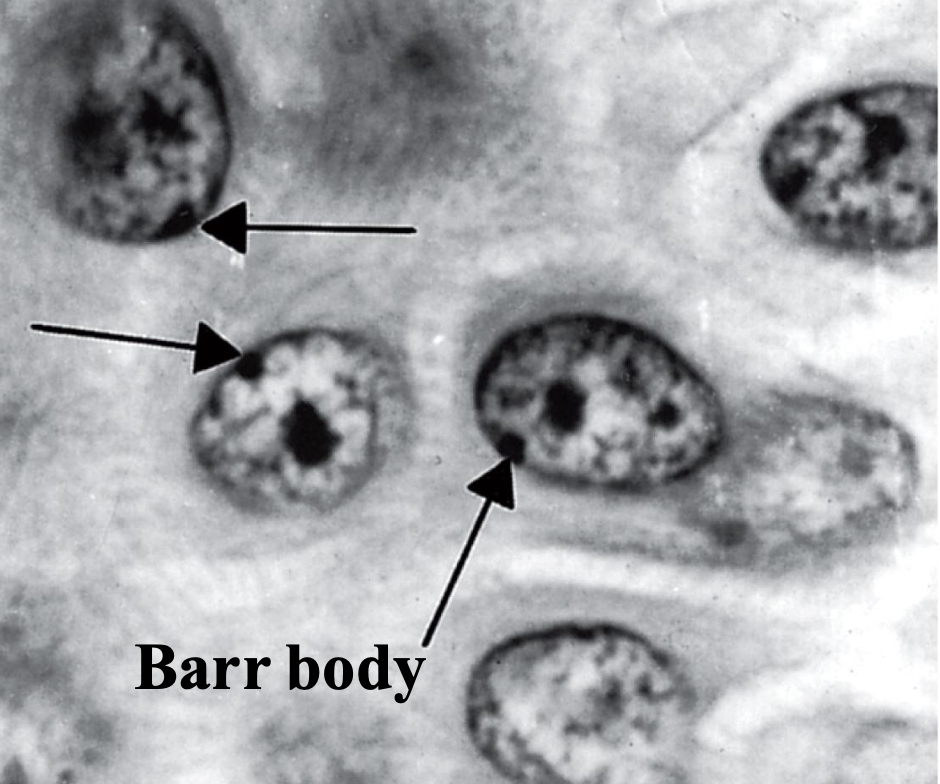
what’s this
The inactive X chromosome is condensed as
heterochromatin (Barr body)
what is X inactivation and when does X inactivation occur?
an example of facultative heterochromatin. when either the mat or pat chromosome is inactivated in almost all somatic cells.
early on in development(but when there are already a bunch of cells), and all daughter cells have the same one silencenced
explain x inactivation in calico cats and clones with identical genetics, but physical differences.
in heterozygous individuals, in some cells the orange colour can be silenced and in some the black colour will be silenced, and as these multiply, it results in different colour patches.
clones can have different chromosomes silenced.
epigenetics vs. genetics
genetics: permanent and inheritable changes in DNA that result in different traits.
epigenetics: reversible covalent modifications that change the expression of DNA without altering it. can be influenced by environmental factors.
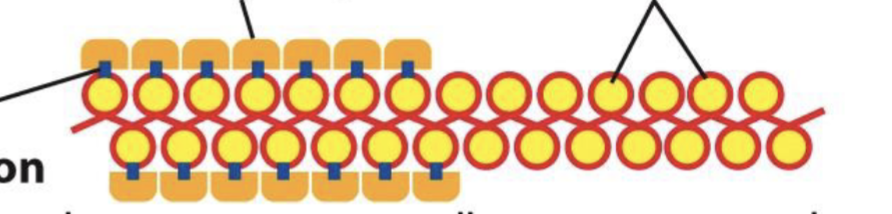
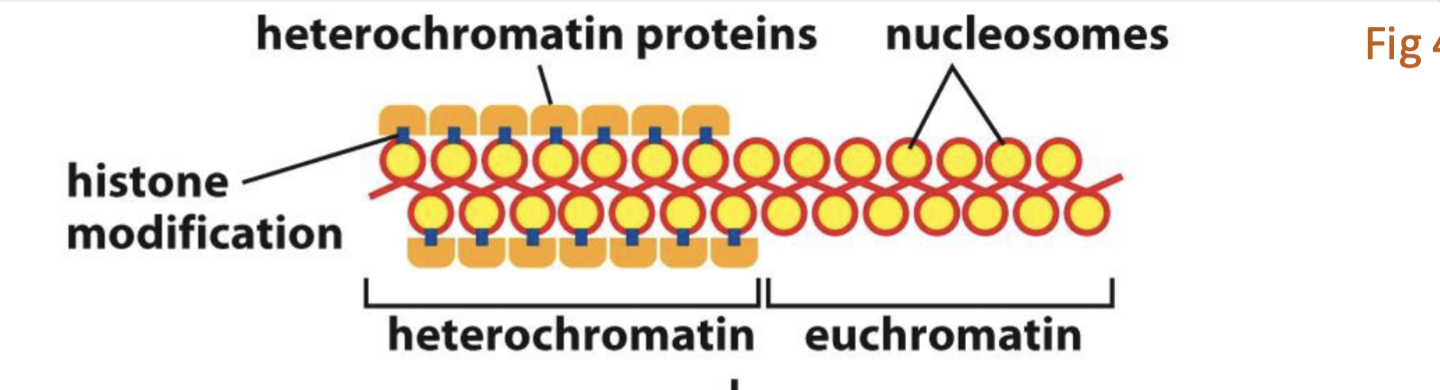
what’s euchromatin and what’s heterochromatin in the photo?
where’s the histone P, and the histone modification?
what are the three covalent modifications that can occur on the histone code on the N terminal tails?
acetylation, methylation, phosphorylation
acetylation does what
more open, increased transcription
methylation does what?
compaction, repression.
acetylate histone proteins by transferring acetyl group from
acetyl-CoA to specific lysine residues.
Histone acetyltransferases (HATs)
what removes remove the acetyl group
ONLY Histone deacetylases (HDACs)
Histone methyltransferases (HMTs)
add methyl group(s) (1, 2, or 3) to lysine or arginine
residues.
remove methyl groups.
histone/DNA demethylases
chromatin has way better memory in ______ than in _______
eukaryotes, prokaryotes
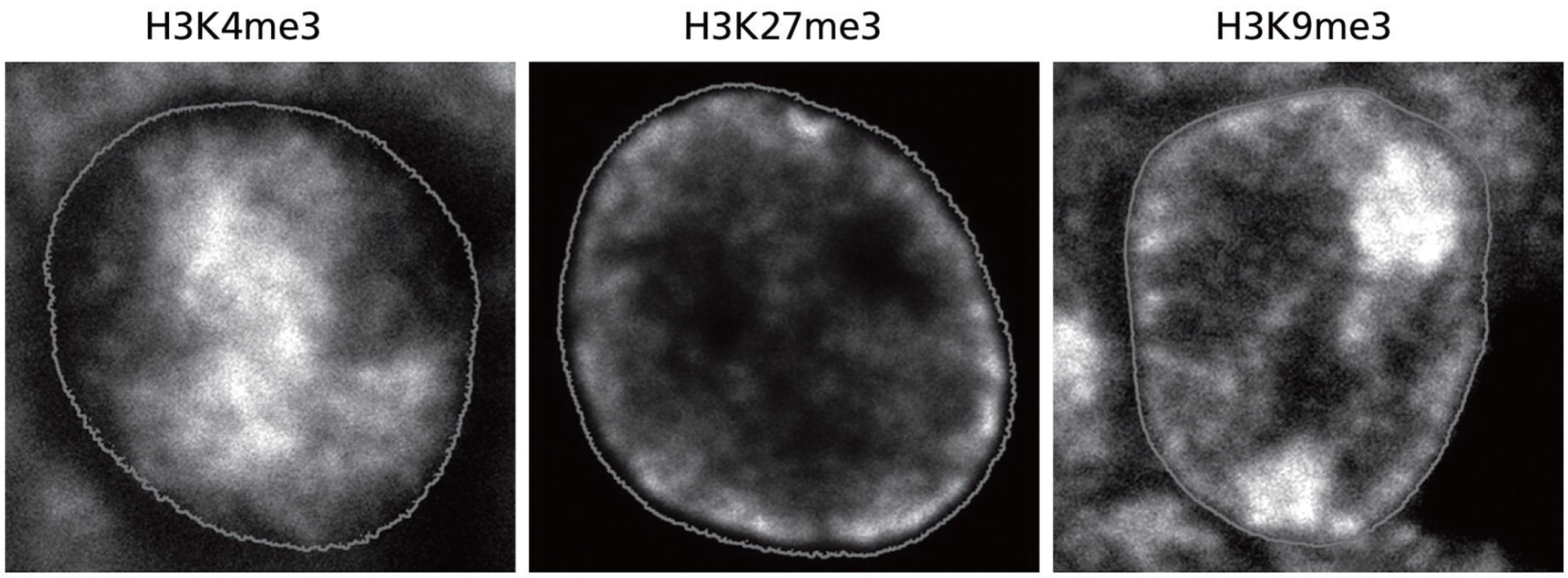
euchromatin or heterochomatin? (3)
euch, het, het
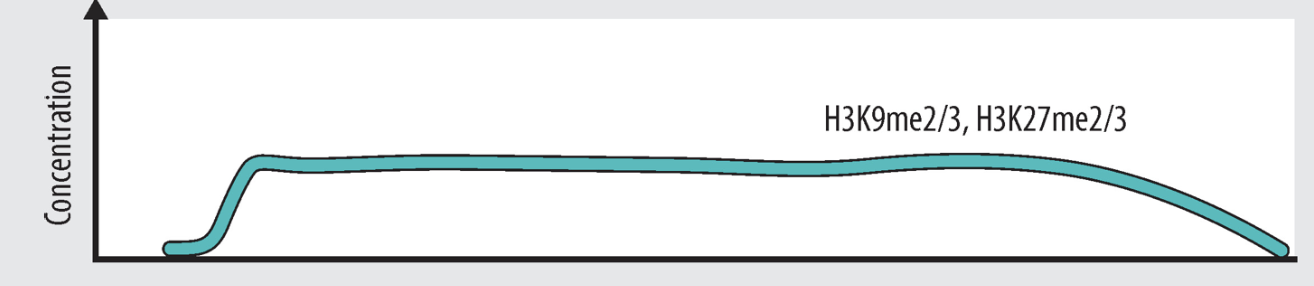
active or repressed gene
repressed

active or repressed gene
active
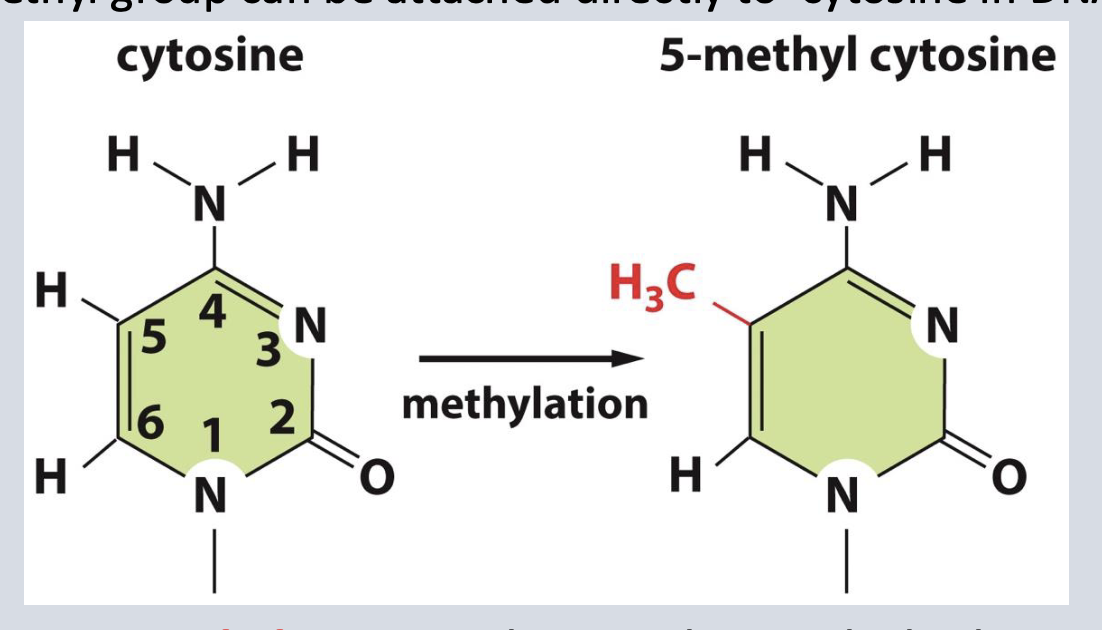
CpG
epigenetic marker that allows DNA methyltransferase to add a methyl group between the C and G (the p spot)
when is DNA methylated and why?
during replication so its methylation patterns can be passed on to its daughter strand.
do you inherit all your epigenetics from your parents?
no, zygote erases is all previous epigenetics and the blastocyst is fresh
exception: genomic imprinting, but thats only a tiny bit
role of reader-writer complexes
reads histone code, positions and activates “writer” enzymes, that attach it to other components in nucleus responsible for expressing, silencing, etc.
which basically means AFTER REPLICATION, histone modifications are scattered throughout, the RW complexes read them, and mark spots that need to be filled in order for it to be identical as parent. then the correct heterochromatin proteins are added to all the spots with midficwations.
how does the initial recruitment of these enzymes occur?
often occurs through a transcription factor protein binding to a specific DNA sequence
heterochromatin doesn’t go across whole chromosome because:
Barrier sequences on DNA that Barrier proteins bind to.
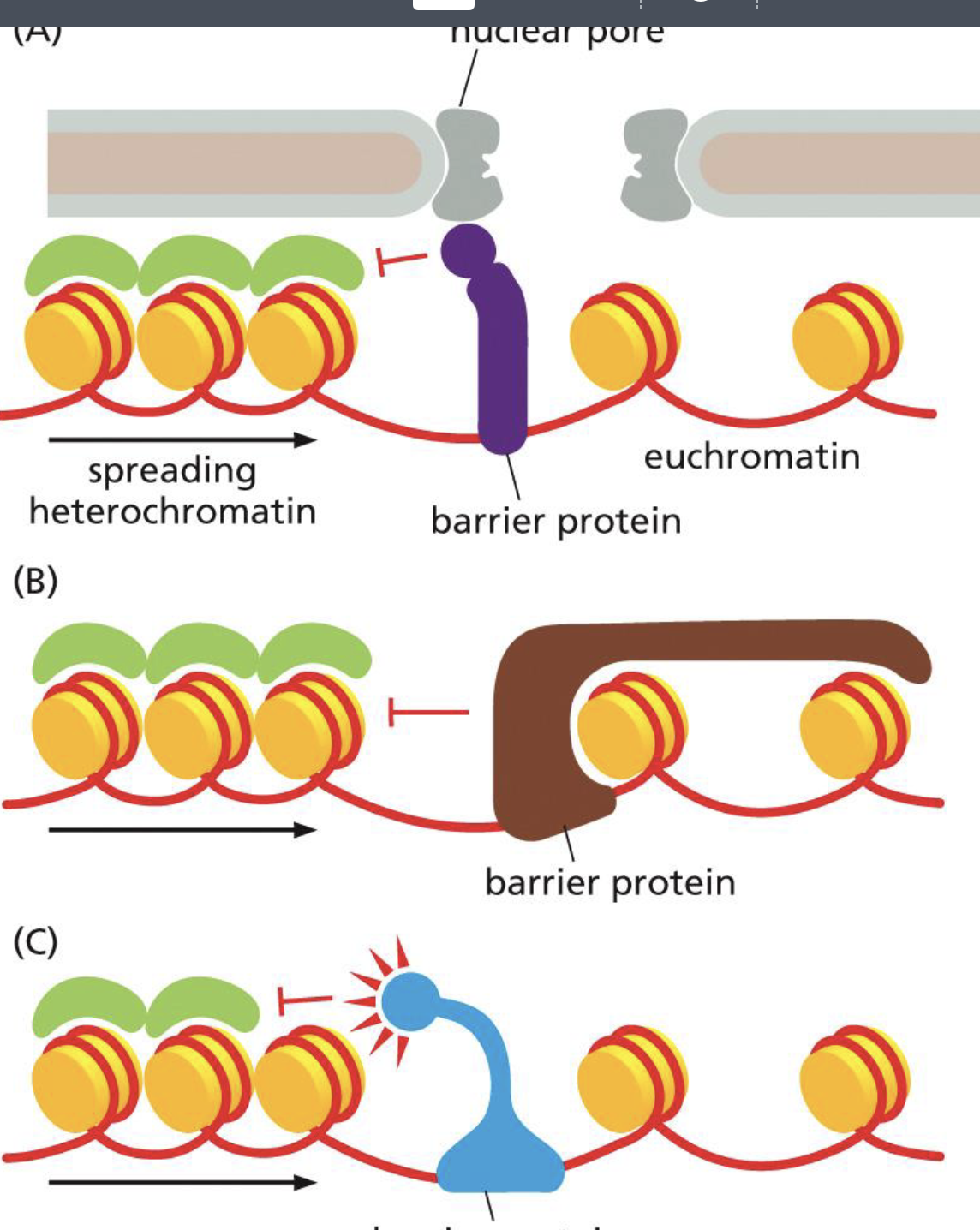
three kinds of barrier proteins
brown: physical protection
purple: linking scaffold P and nuc pore to make wall
blue: mechanism that directly opposed RW activity,
Explain why some regions/sequences of the genome are likely to be exhibit faster or slower rates of change
more important sequences are more conserved (less change). generally exons and protein coding sequences
point mutation, and explain an example
switching one nucleotide for another
(Changes in the nucleotide sequence of DNA can
be caused by errors in DNA replication or repair)
Ex: SNPs, a common version that usually is not detrimental=not selected against, persists.
Of the five large scale rearrangements: Insertions, Deletions, Duplications, Inversions, Translocations
Which of these are likely to result
in loss/mutation of genes?
Deletion, by far!!
(not translocations because it would need to cut right in the middle of the gene, which is unlikely)
synteny
regions where genes are in conserved order, but are now at a diff position.
_____% of human genome is conserved across species
5, and 1.5% of that is P-coding
Apply an understanding of genome composition and rates of change to select the type of sequence a researcher should look at if identifying closely related individuals within a species vs constructing a phylogenetic tree
help
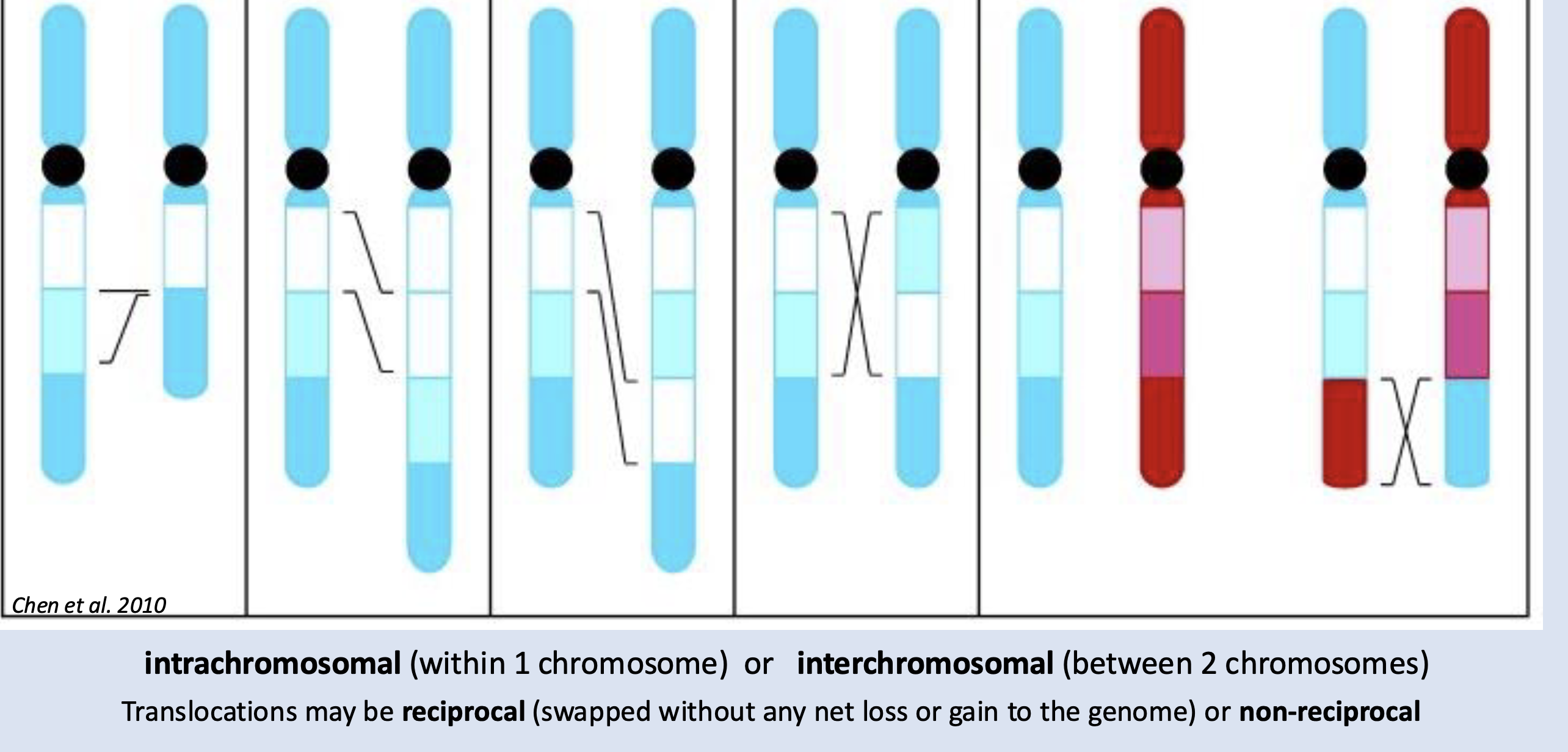
name them
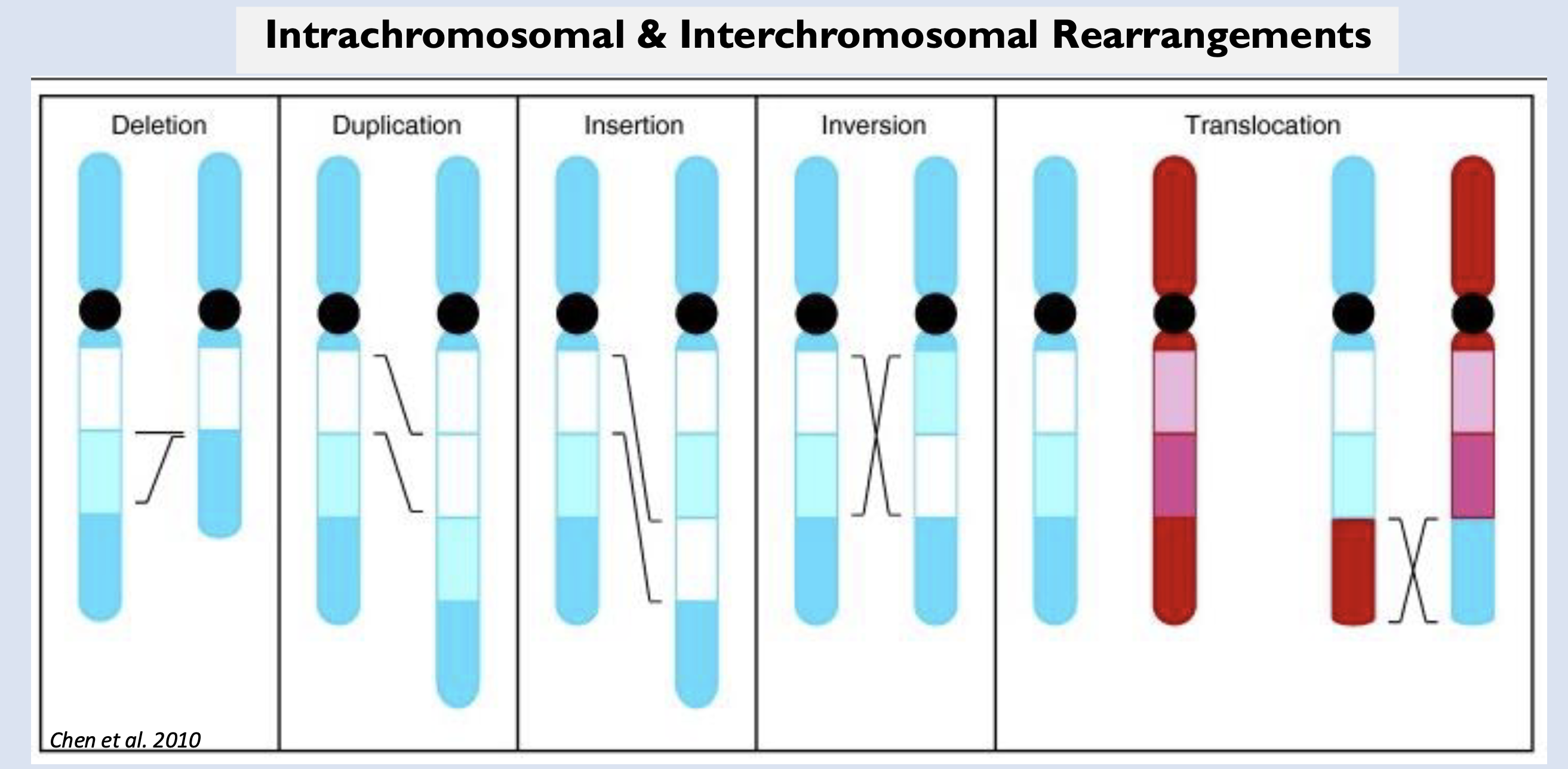
what do you think is
more highly conserved: sequences within
exons or the positions of genes on particular
chromosomes?
sequences within
exons
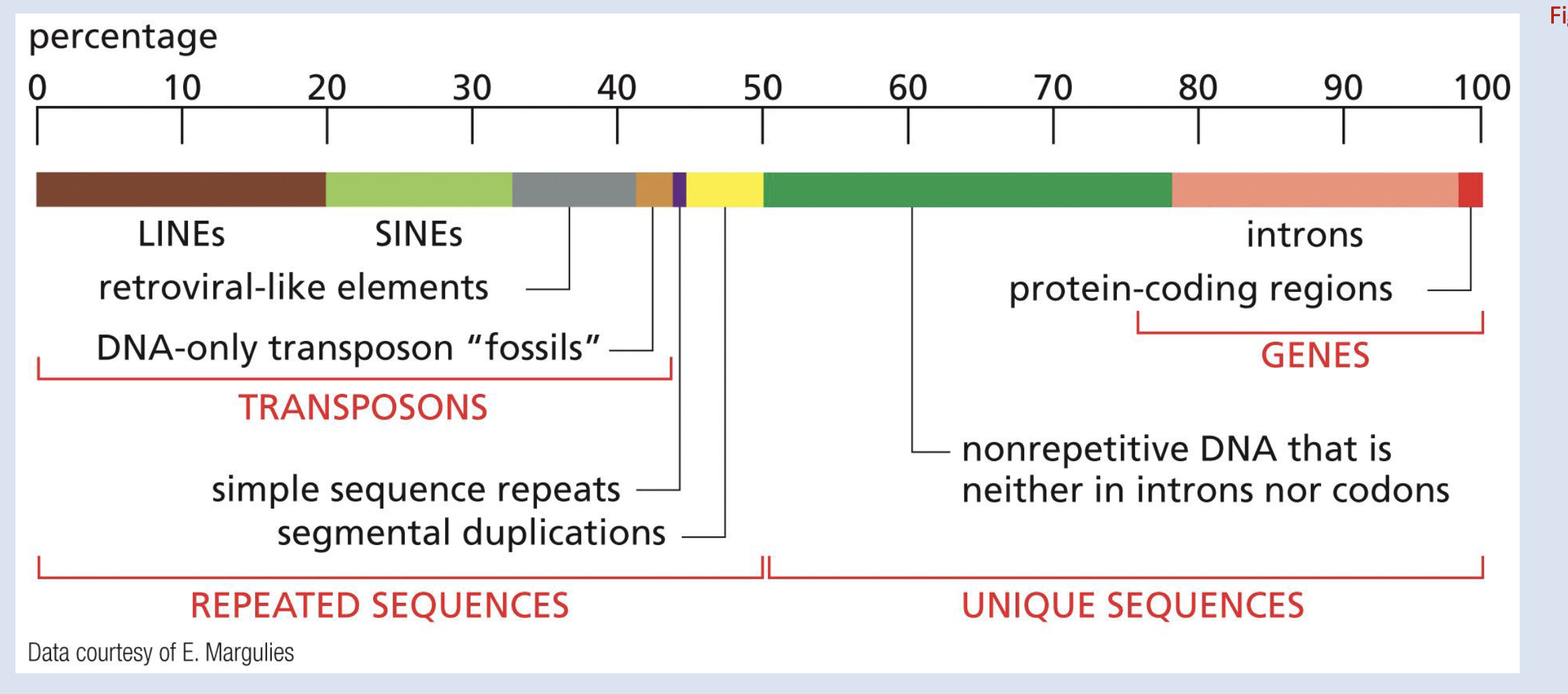
just know this
Satellite DNA
highly repetitive, (generally found at centromeres and telomeres) - 5-500 bp in tandem
repeats of up to 100 kb that can form very large clusters. Base composition is distinct from
bulk DNA
Minisatellite DNA
highly repetitive, 0-100 bp with up to 3000 repeats → highly variable (polymorphic),
therefore # repeats differs between individuals– basis for DNA fingerprinting in criminal
and paternity tests
Microsatellite DNA
highly repetitive, 1-5 bp in clusters of 10-40 bp scattered quite evenly throughout the
genome → highly variable (# of repeats changes fast!), can be used to compare closely
related populations.
12
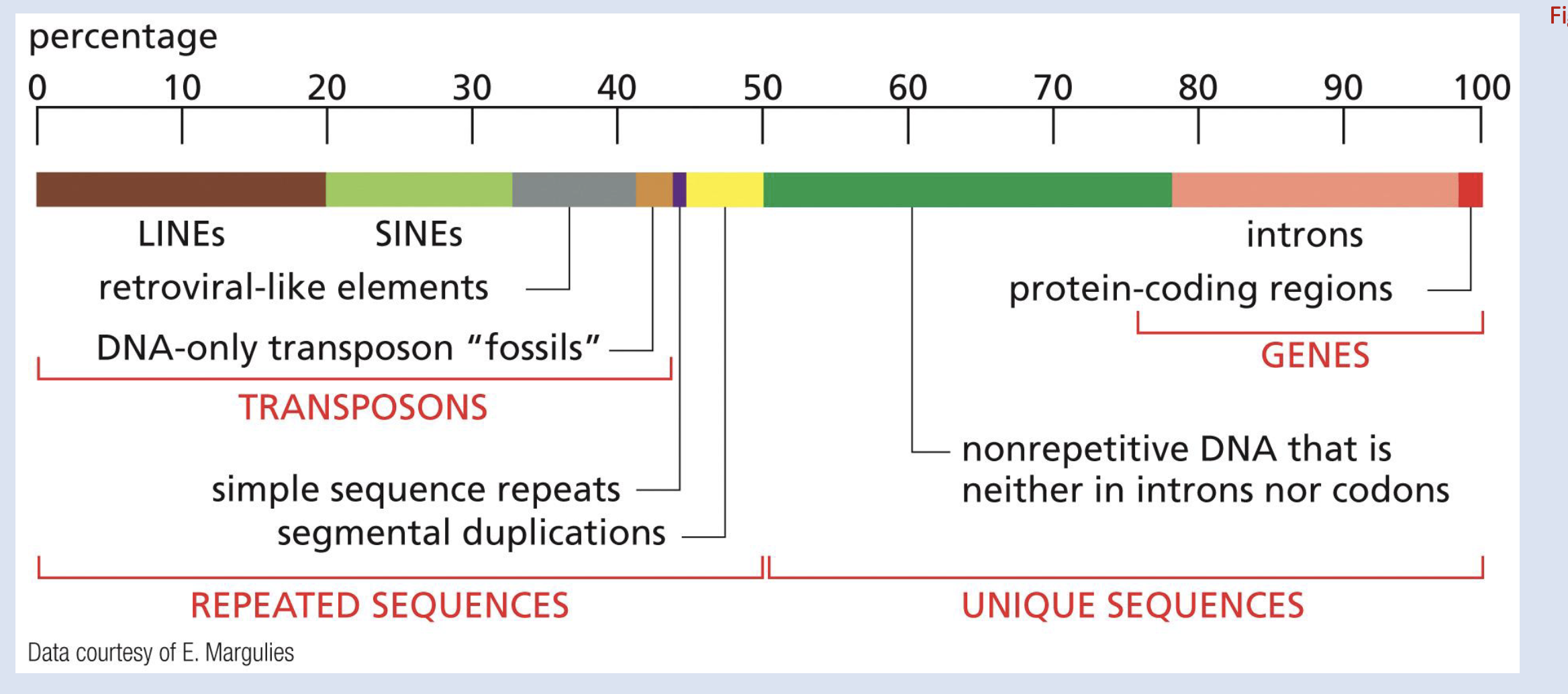
Name two kinds of Moderately repetitive DNA, 20-80% of the genome
genes: ribosomal RNAs and Histone genes
non-coding: SINES and LINES caused by transposons
Protein coding DNA Is likely
Non repetitive
what are the two forms of slippage, and what does it cause?
( what are the 2 forms of slippage??) expansion and deletion causes misalignment of DNA during replication
transposons vs retrotransposons method of locomotion, and details
transposon: cut and paste, by transposase enzyme results in a direct repeat behind it
retrotransposon: copy paste, by RNA intermediate, use and can encode a reverse transcriptase enzyme to catalyze production of
DNA from RNA.
Example: LINEs (long interspersed nuclear elements), SINEs (short interspersed nuclear elements)
what percentage of genetic change is due to transposons?
40%
why do humans have more transposon events than bacteria
because humans have less density, less of a chance of it hitting smog important
Explain 4 mechanisms that can contribute to creating new genes/alleles and explain how gene families and pseudogenes could form
segment shuffling, exon shuffling is when transposition includes exons
Horizontal gene transfer: viruses inserting their DNA
Gene duplication (ex: unequal crossing over ): different versions of the same gene!
Mutations: point mutations etc..
genes always come from pre-existing genes. T or F
T
what is exon shufflring?
when the exons from different areas combine due to transpositional activity. results in the creation of new proteins
what is the result of unequal crossing over
a way of gene duplication resulting in the expansion of multigenerational families. how? the chromosomes aren’t perfectly aligned during crossing over, so one gets a deletion and another gets extra. if it occurs consecutively, duplications and stuff.