Bio 1A03 Test 3
1/149
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
150 Terms
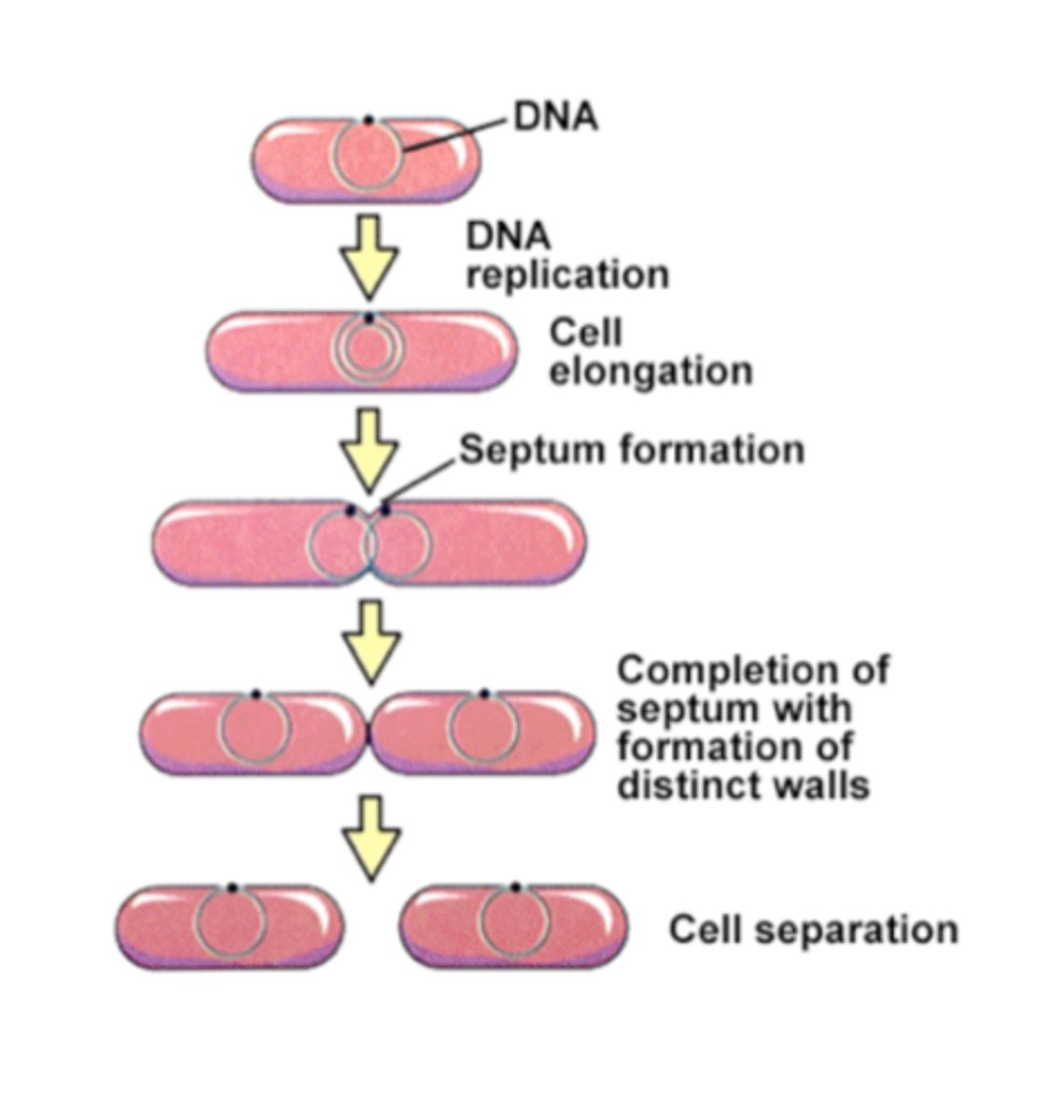
Binary Fission steps
-Initiated when proteins attach DNA to plasma membrane
-Begins along ORI region
-Newly synthesized DNA also anchored to membrane
-Cell elongates until ~2 times its original size and two has two chromosomes on opposite ends of the cell
-Cell splits in two
Quiescent stem cells, function
Permanently arrested in G0 phase. In adults: can replace non-reproducing specialized cells) ex adult skeletal muscle/myofiber)
Steps of quiescent stem cell to myofiber
-Activated (reenter cell cycle)
-Proliferative (look over this one)
-Commitment to differentiation
-Fusion into myotube (precursor of myofiber)
-Myofiber
steps of cell cycle
G1 phase
S phase
G2 phase
M phase
~G0 phase~ perhaps
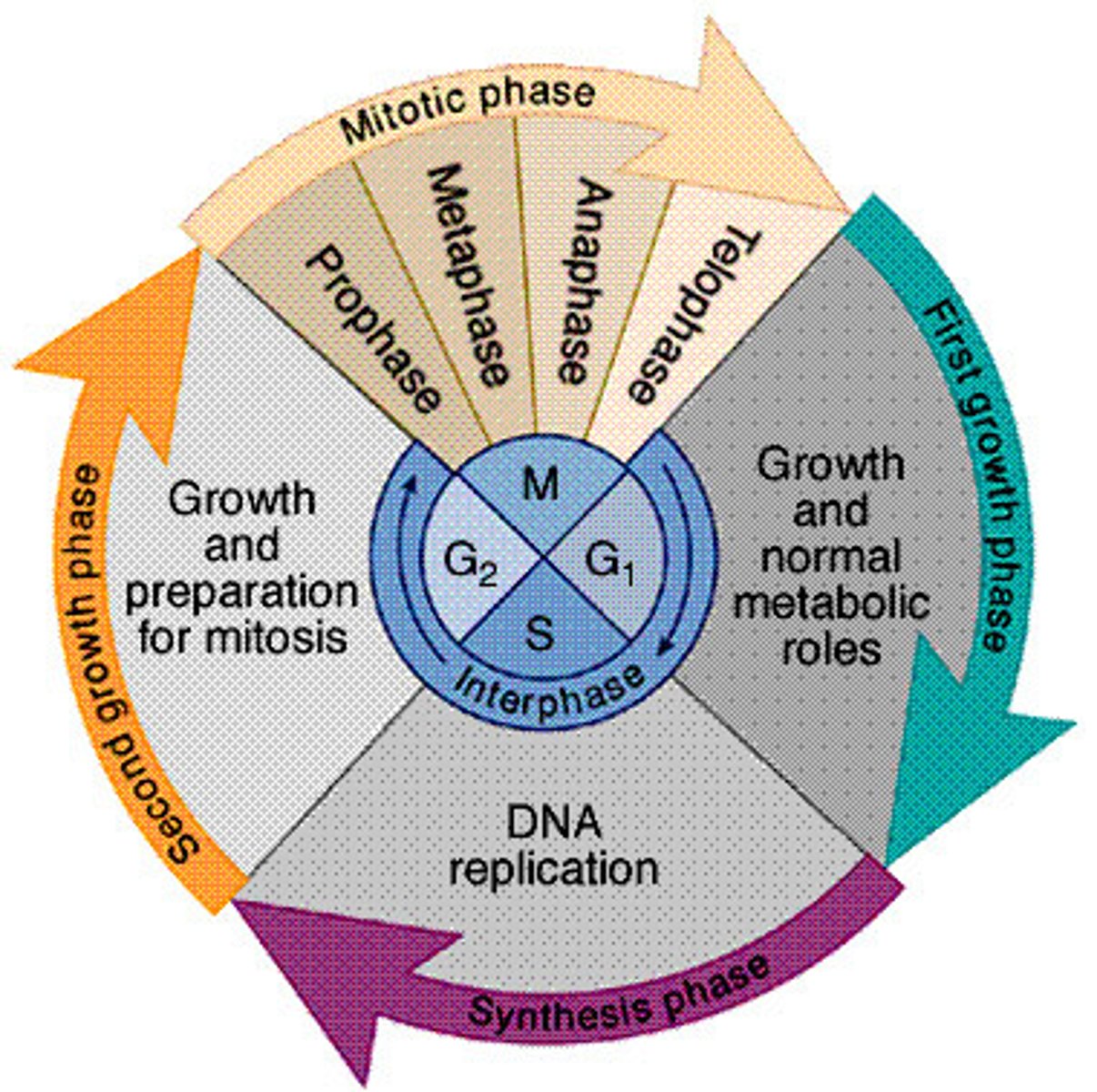
G1 phase
growth phase, prepares cell for dna synthesis
s phase
cell undergoes dna replication
g2 phase
growth phase, cell prepares for division
M phase
mitosis and cytokinesis
cells in permanent g0 phase
eye lens, nerve, mature muscle cells
binary fission is common to
bacteria, archaea, mitochondria, chloroplasts
Satellite Stem Cells
quiescent stem cells that specifically are activated to replace injured muscle fibers
Walter Flemming
Discovered stages of mitosis from salamanders
During interphase, the genetic material of a typical eukaryotic cell is
chromatin fiber
genetic material during s phase
sister chromatids are synthesized (including centromere) but chromosomes don't actually compact until M phase (46 X-shaped not linear chromosomes / 92 chromatids)
chromatids
chromosome in duplicated state
# chromosomes and chromatids in human before & after mitosis, meiosis I, and meiosis II
Mitosis
Before: 46, 92
After: 46, 46
Meiosis I
Before: 46, 92,
After: 23, 46
Meiosis II
Before: 23, 46
After: 23, 23
https://datbootcamp.com/biology-strategy/chromosome-and-chromatid-numbers-during-mitosis-and-meiosis/
genetic material during interphase
chromatin fiber, can't identify specific chromosomes
prophase (mitosis)
identical sister chromatids apparent
centrosomes (microtubules) form mitotic spindle and positioned at opposite ends of the cell
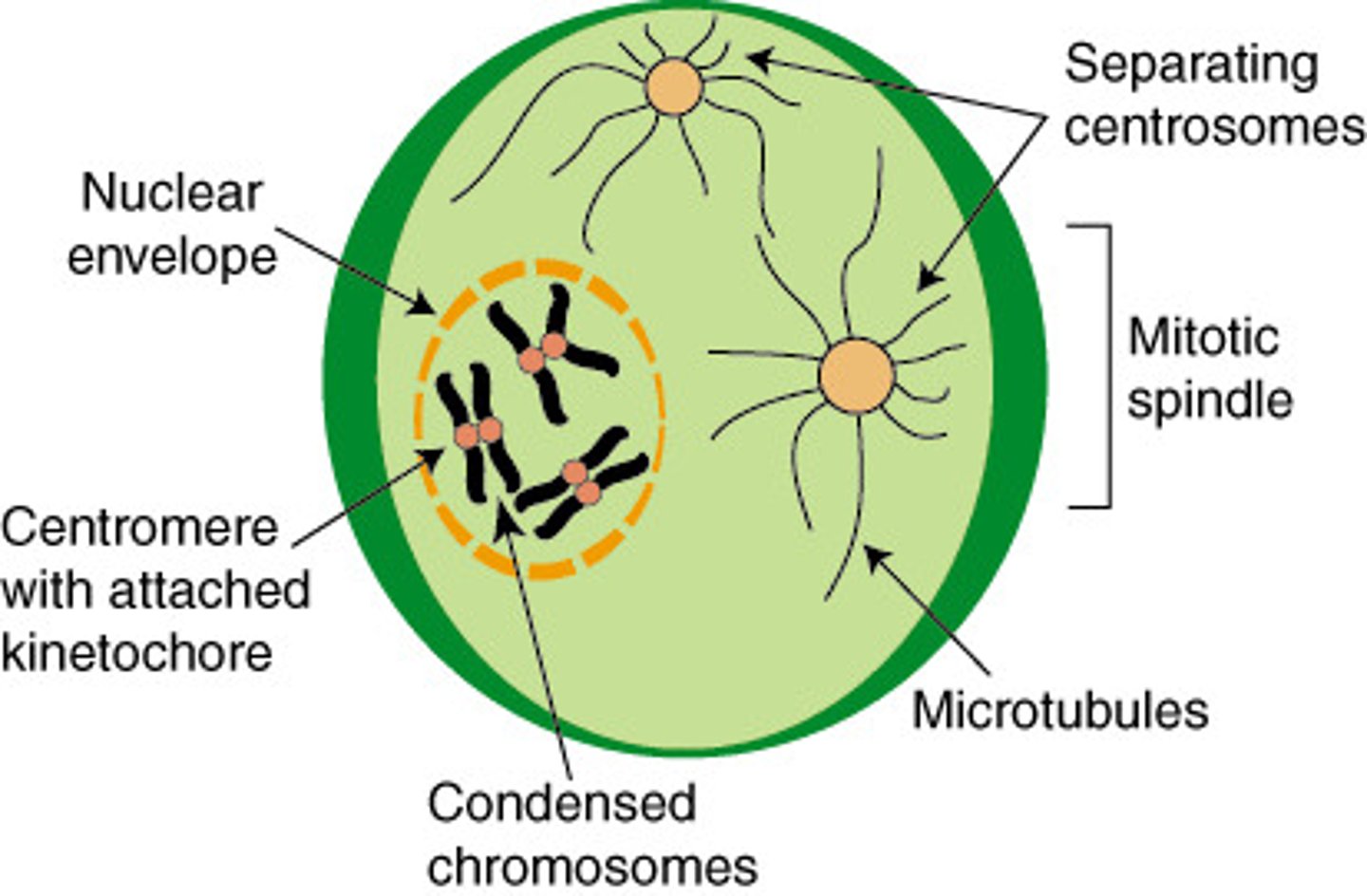
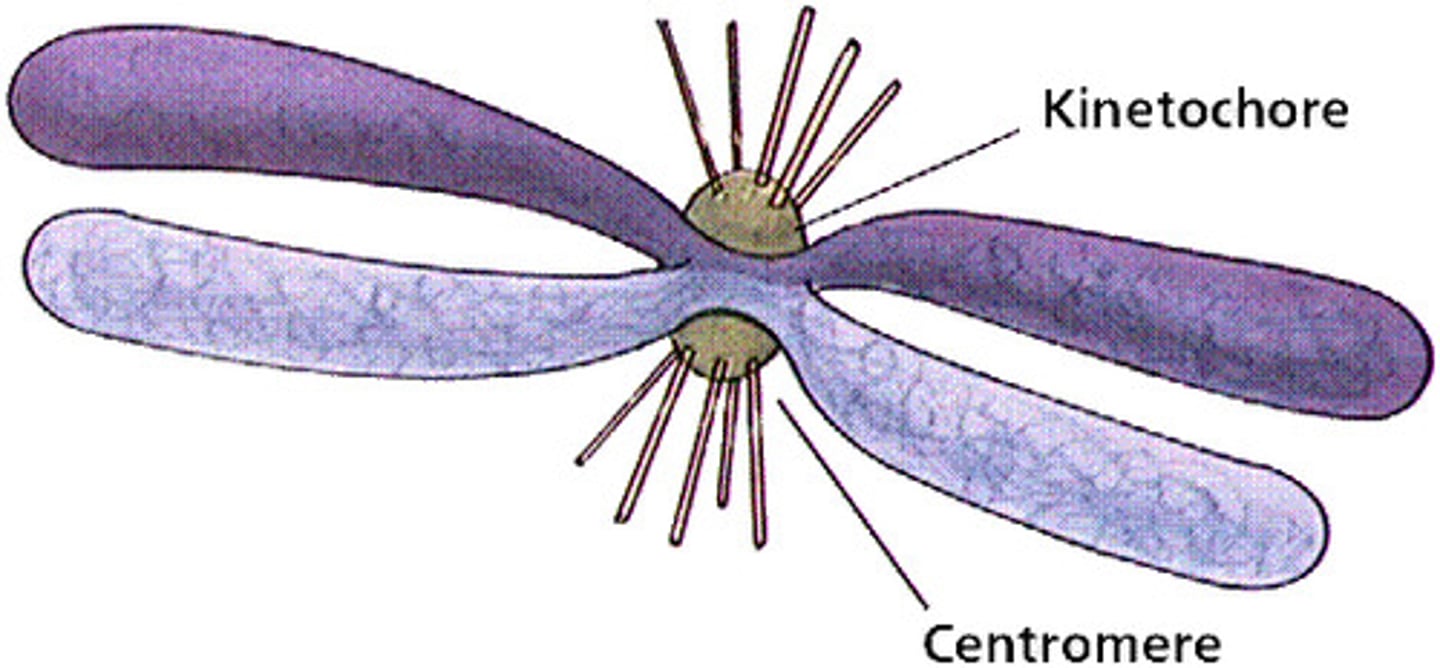
centromere vs centrosome
centromere: holds together two sister chromatids into 1 chromosome
centrosome: hold together mitotic spindles
prometaphase
fragmentation of nuclear envelope allows mitotic spindle to attach to kinetochores on centromeres
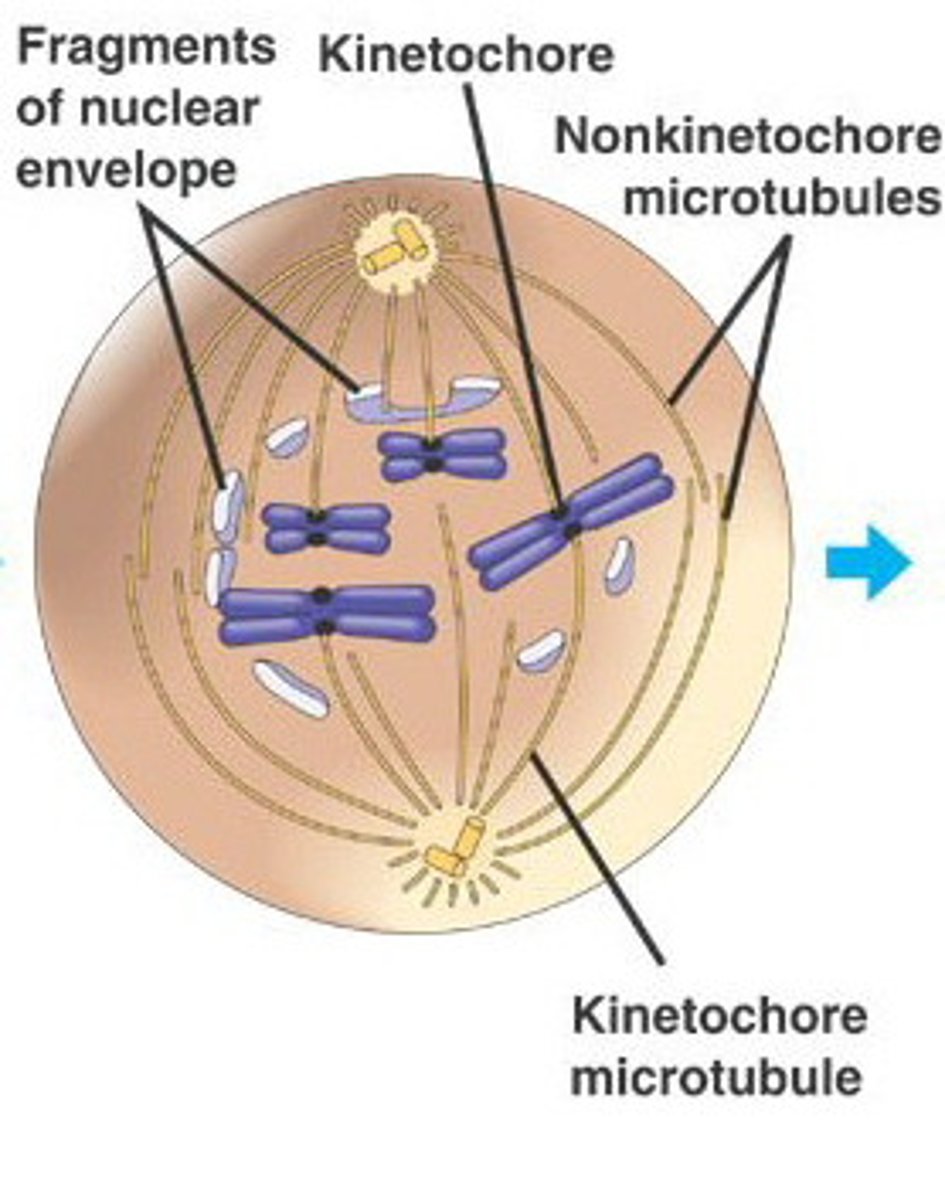
kinetochores
The structures on sister chromatids where microtubules attach
1) some microtubules attach directly and pull
2) some k.c are polar and repel the sides of the cell from each other
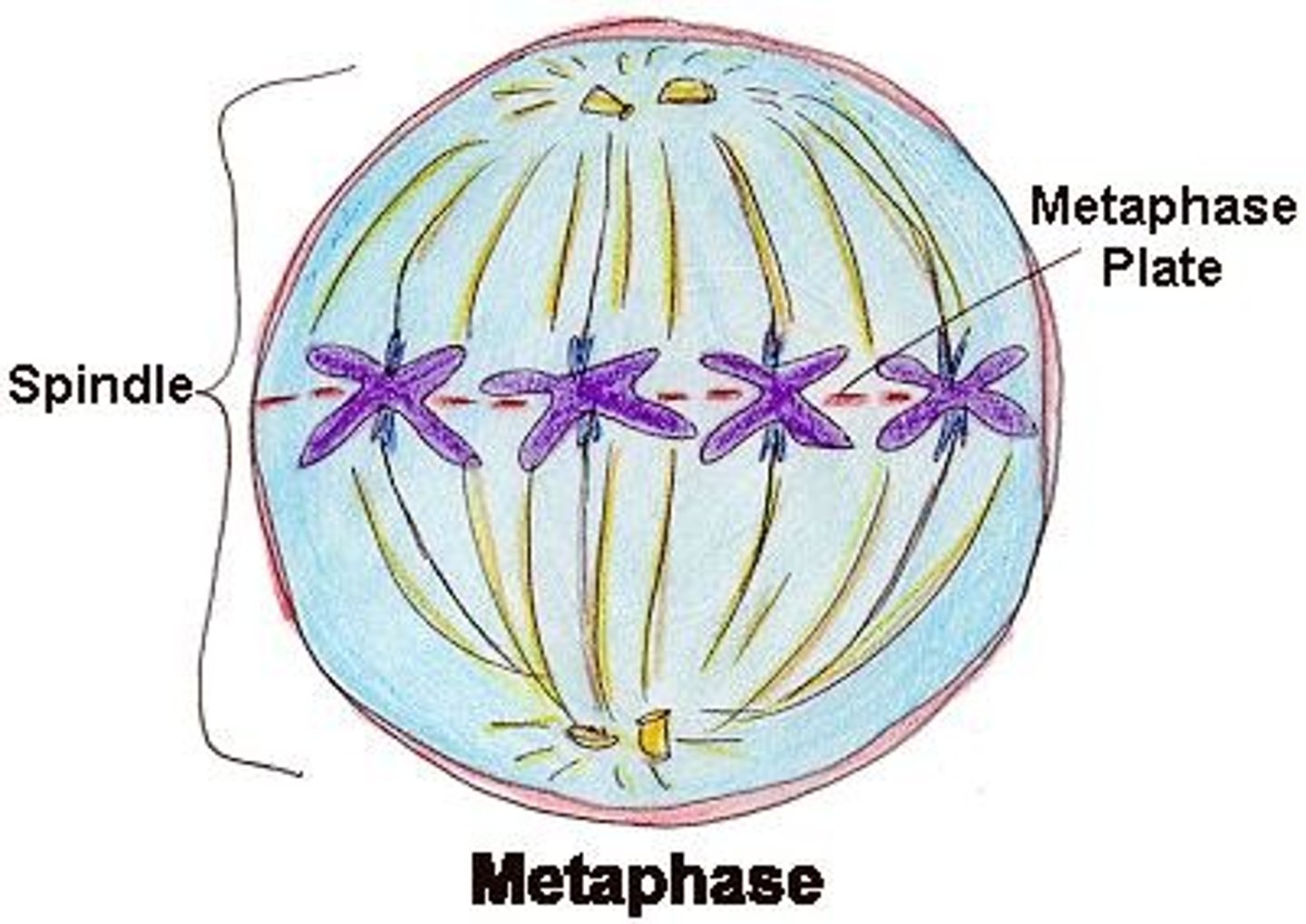
metaphase (mitosis)
chromosomes line up along middle of metaphase plate, facilitated by kinetochore microtubules attaching to the kinetochores of each sister chromatid
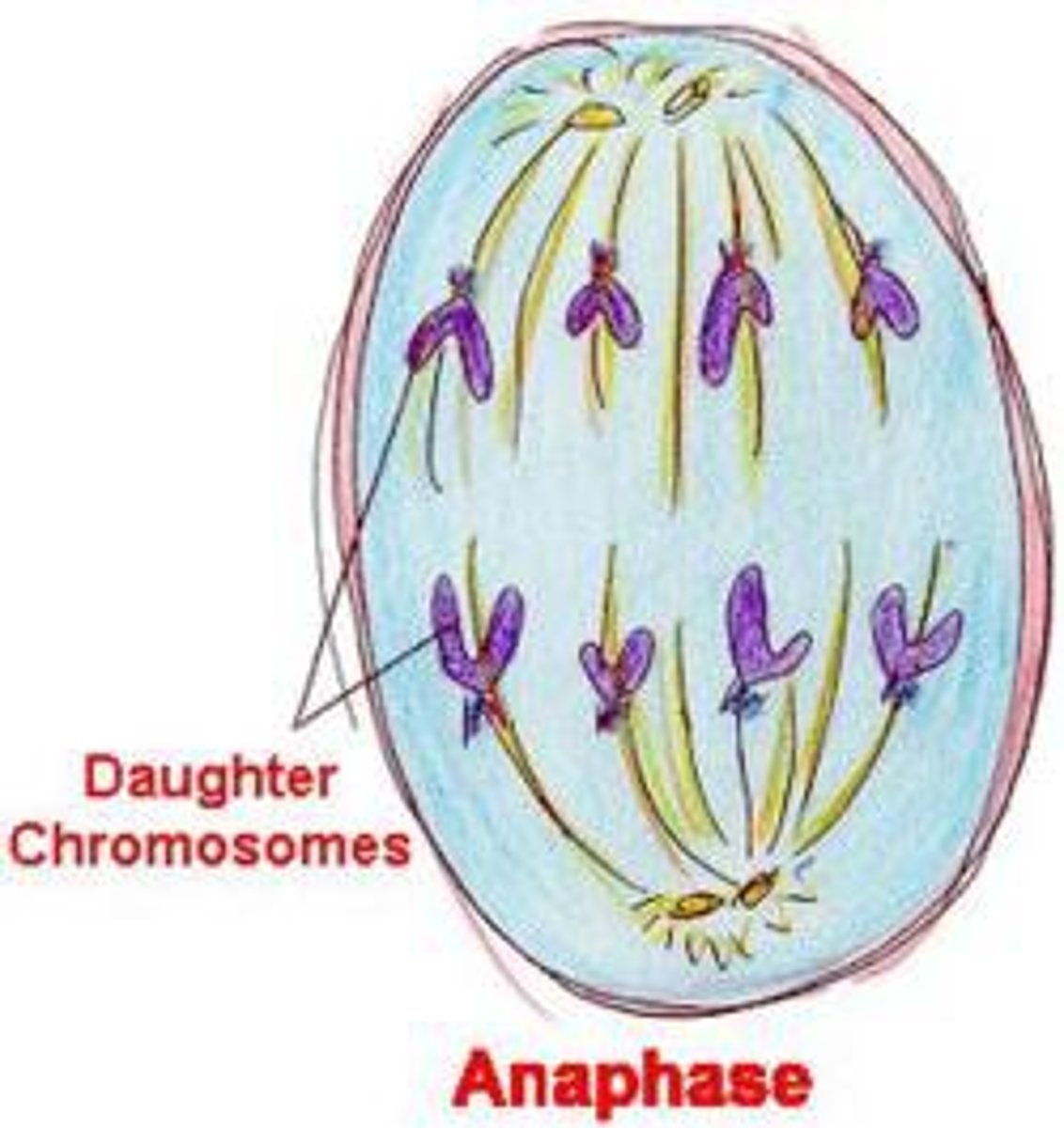
anaphase (mitosis)
kinetochore microtubules shorten and pull apart chromosomes
polar microtubules repel each other and help elongate the cell
equal segregation of chromosomes (or stop)
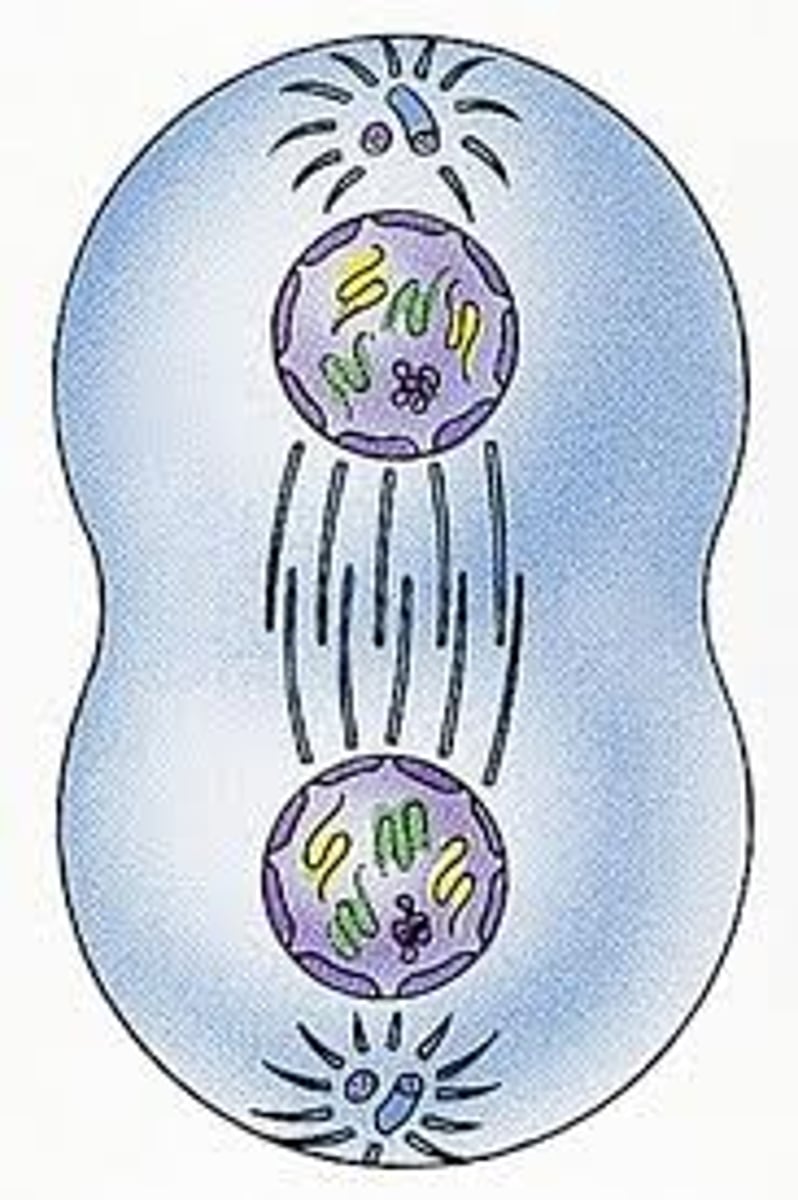
telophase (mitosis)
nuclear envelope reforms giving 2 new daughter nuclei within the cell
chromosomes decondense
spindle microtubules depolymerized
end of mitosis
cytokinesis
division of the cytoplasm
animals: begins with contractile ring, cleavage furrow separates cell
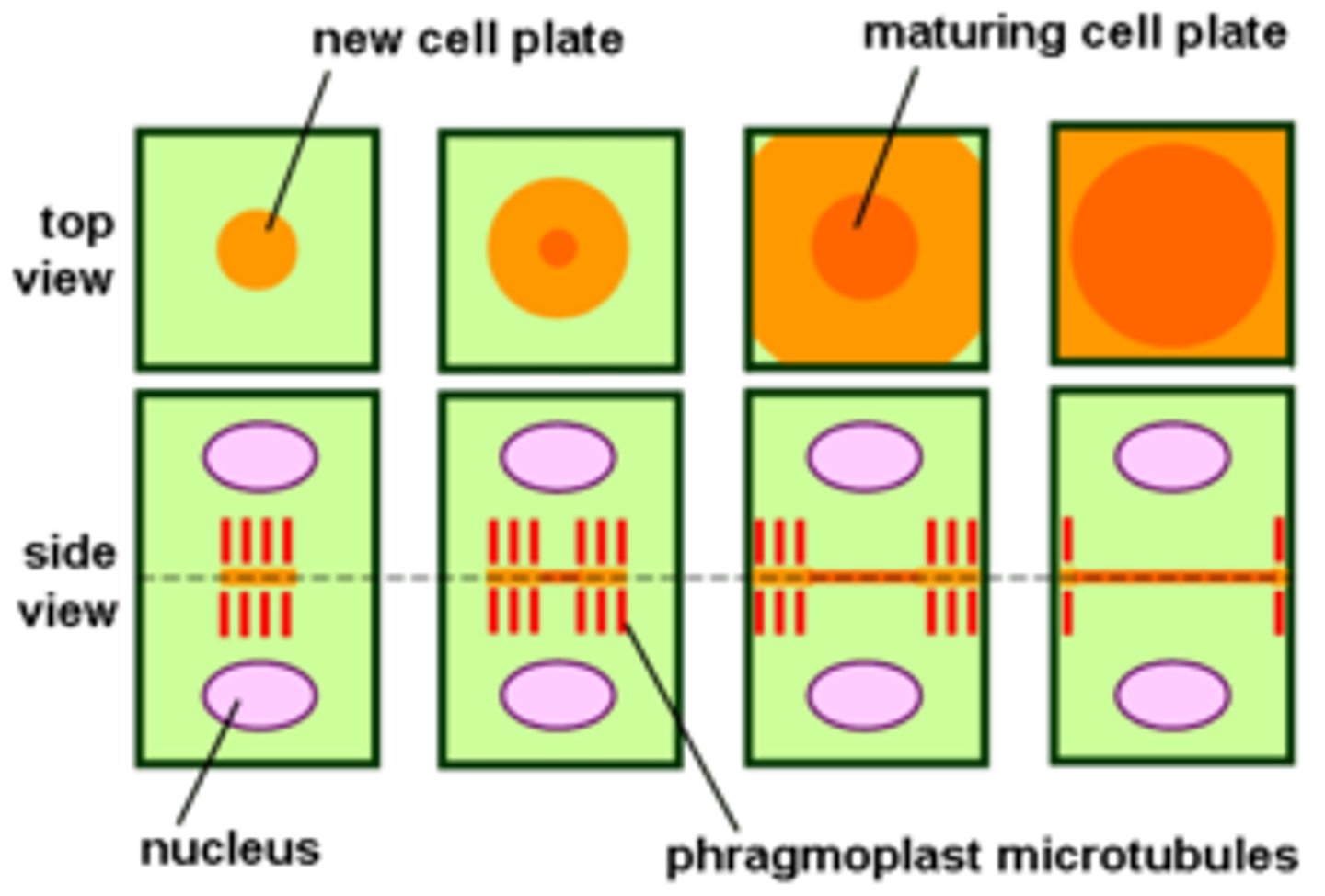
plants: new cell wall formed on cell plate region
MPFs
mitosis promoting factors : cyclin + CDK (cyclin dependent factors)
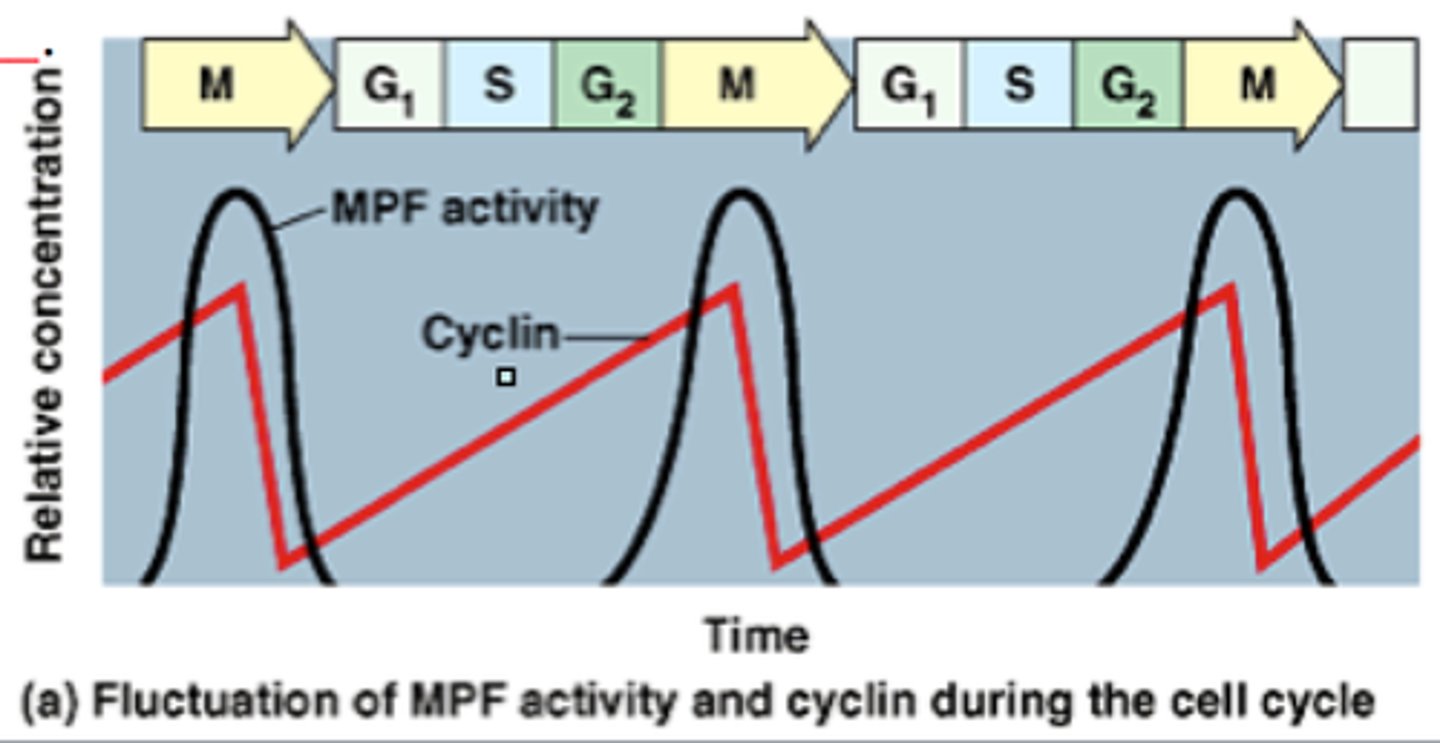
how does cyclin-cdk regulate mitosis?
cyclin-cdk binds to kinases (constant concentration) in order to activate them. these kinases then phosphorylate and activate proteins needed to break down the nuclear envelope, regulate assembly of microtubules, etc
S cyclin-CDK
helps initiate DNA synthesis (cyclin A + cdk 2)
G1/S cyclin-CDK complex
prepares cell for DNA replication
Mitosis checkpoints
G1/S checkpoint, if cell is ready for DNA synthesis (DNA Damage checkpoint)
G2 Checkpoint: Is everything replicated/cell ready for mitosis?
Pre-Anaphase checkpoint: All chromosomes attached to spindle fibers?
How does the G1/S checkpoint work?
If damage to DNA is present, kinases phosphorylate the gene for p53, increasing the production of its protein which accumulates in the nucleus and acts a transcription factor for the inhibitory proteins. These proteins bind to and block G1/S cyclin-CDK complex so that the cell can repair DNA.
How does the spindle-assembly checkpoint work?
Unattached kinetochores release a "wait" signal from the lack of tension that recruits checkpoint proteins. These remain until all kinetochores are attached to micotubules. Seperase breaks apart sister chromatids
semiconservative replication
each new DNA molecule consists of one new strand and one old strand
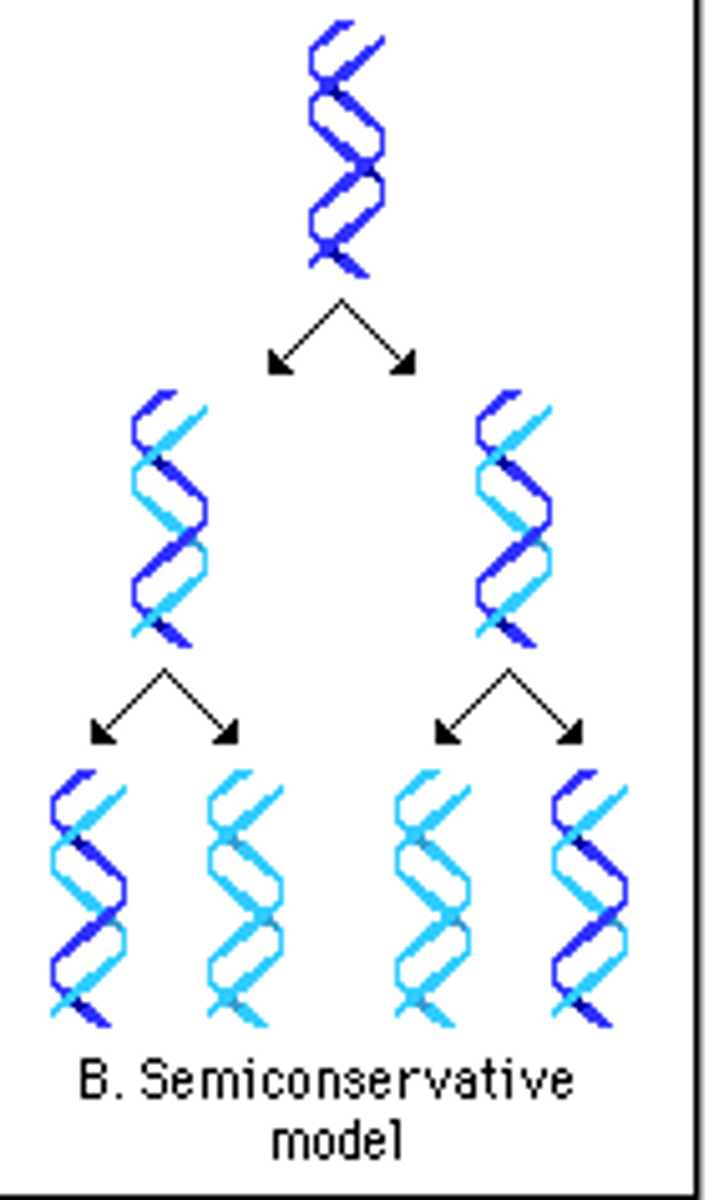
conservative replication
2 parental strands rejoin after replication
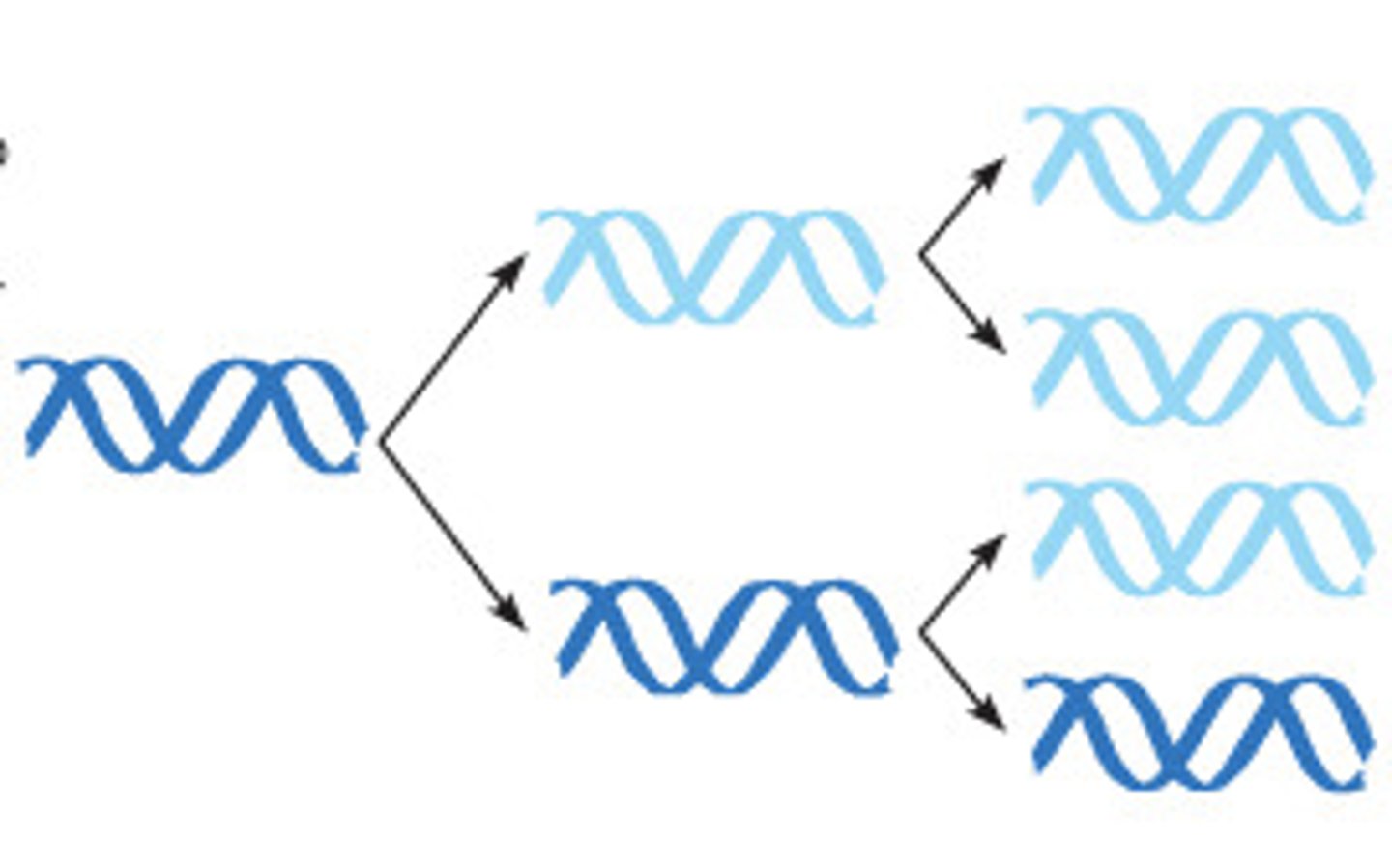
dispersive replication
all 4 strands mix together
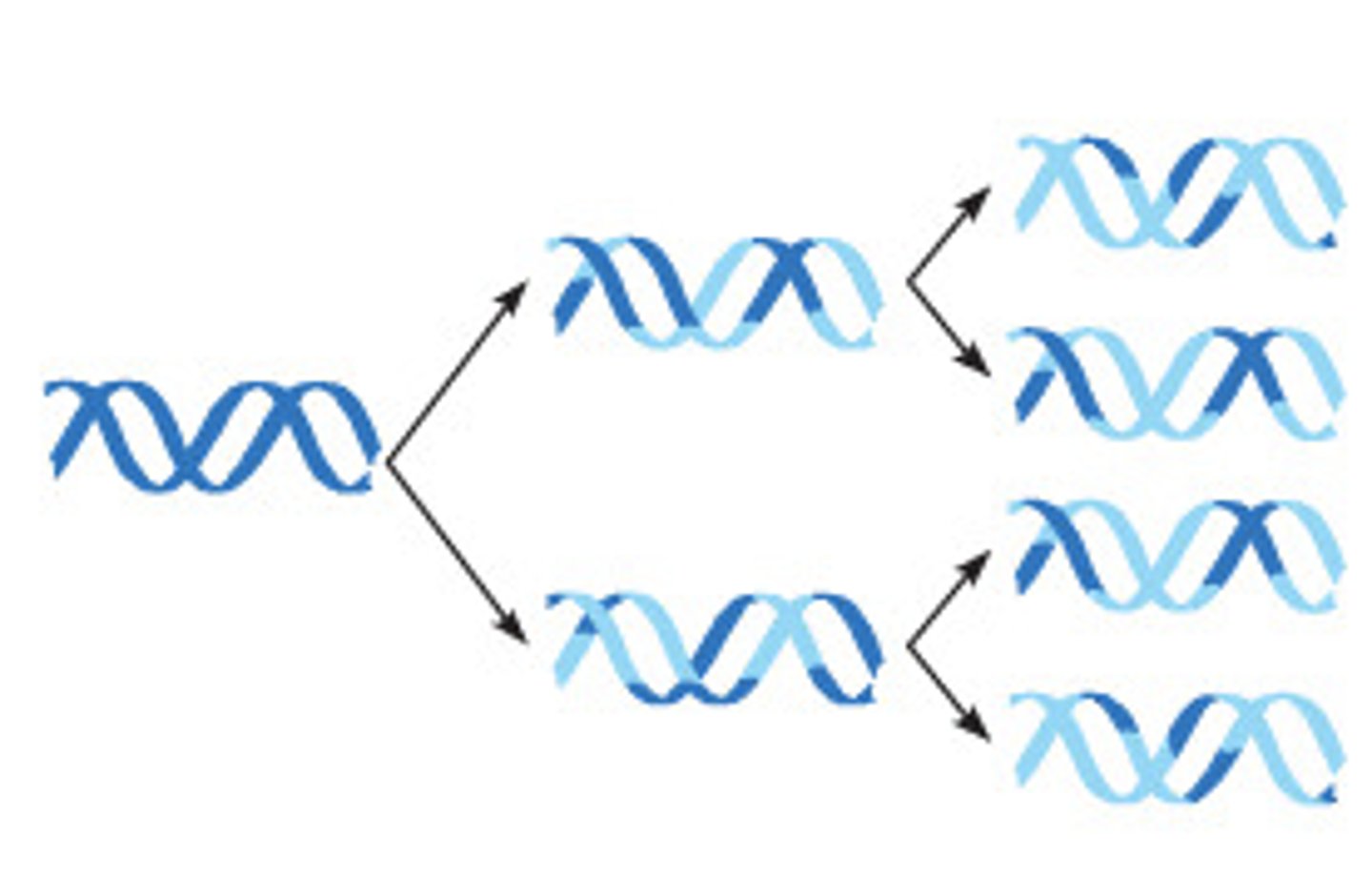
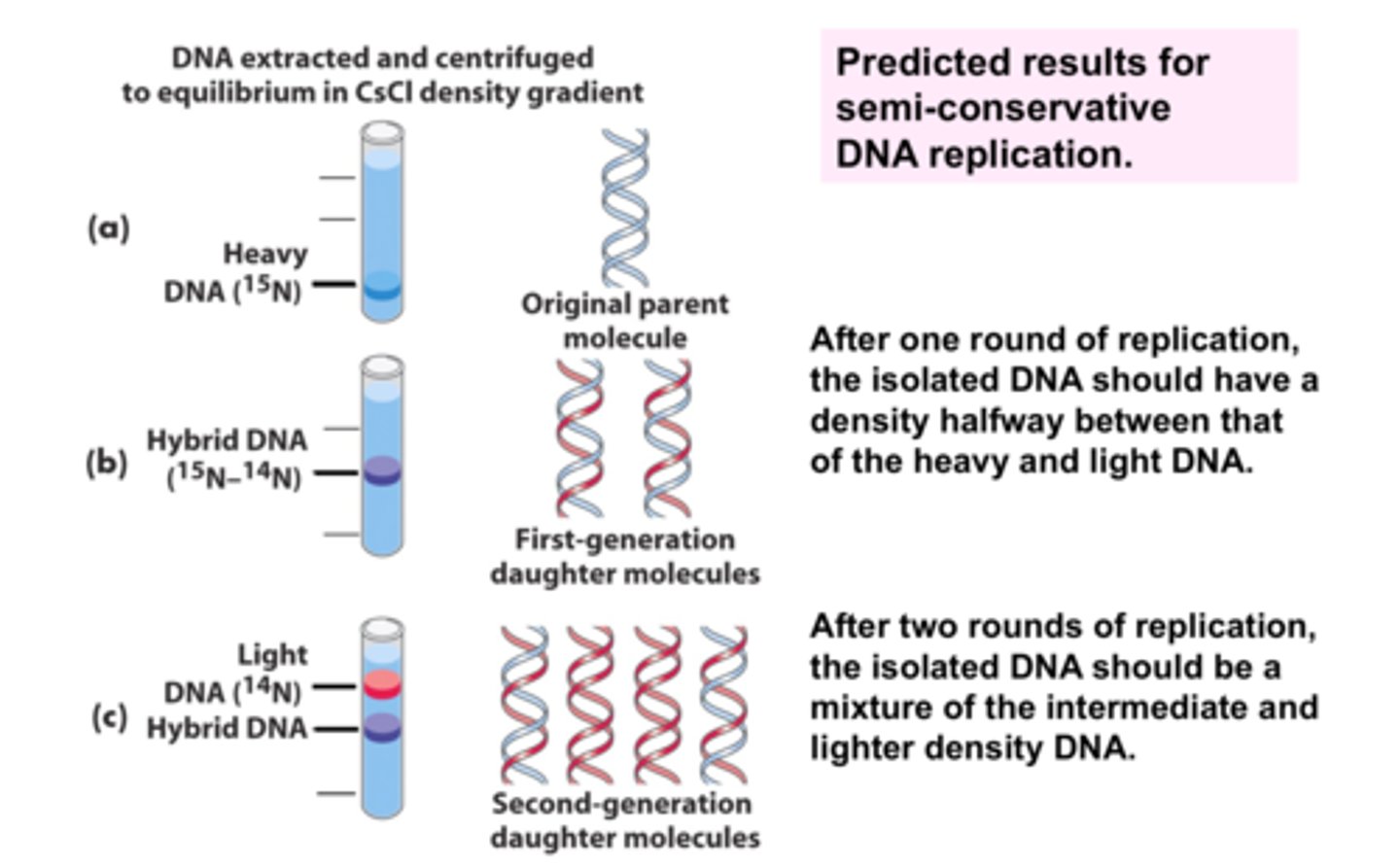
How was the semiconservative model confirmed?
Meselsohn & Stahl created DNA strand out of radioactive isotope of Nitrogen, replicated DNA, used centrifuge to separate the strands and used the placements of the new bands compared to the original strand to prove this model
prokaryotic vs eukaryotic DNA replication
Prokaryotes: only 1 ORI, replication moves in both directions due to circular DNA, no gaps in DNA
Eukaryotes: multiple ORI, okazaki fragments/dna ligase, free phosphate on 3' end of strands, telomeres (DNA shortening)
RNA primer
5-10 nucleotide sequence that binds to BP with template DNA strands so DNA Pol can begin replication (can only add to an existing strand)
After fragment/strand is complete, another DNA pol replaces the primer with DNA nucleotides
leading strand
3'-5', only req's 1 primer
lagging strand
5'-3', multiple primers (one per okazaki fragment), replicates away from replication fork
Helicase
breaks Hydrogen bonds between base pairs
topoisomerase II
relieves the stress of unwinding the DNA for helicase by uncoiling/straightening it
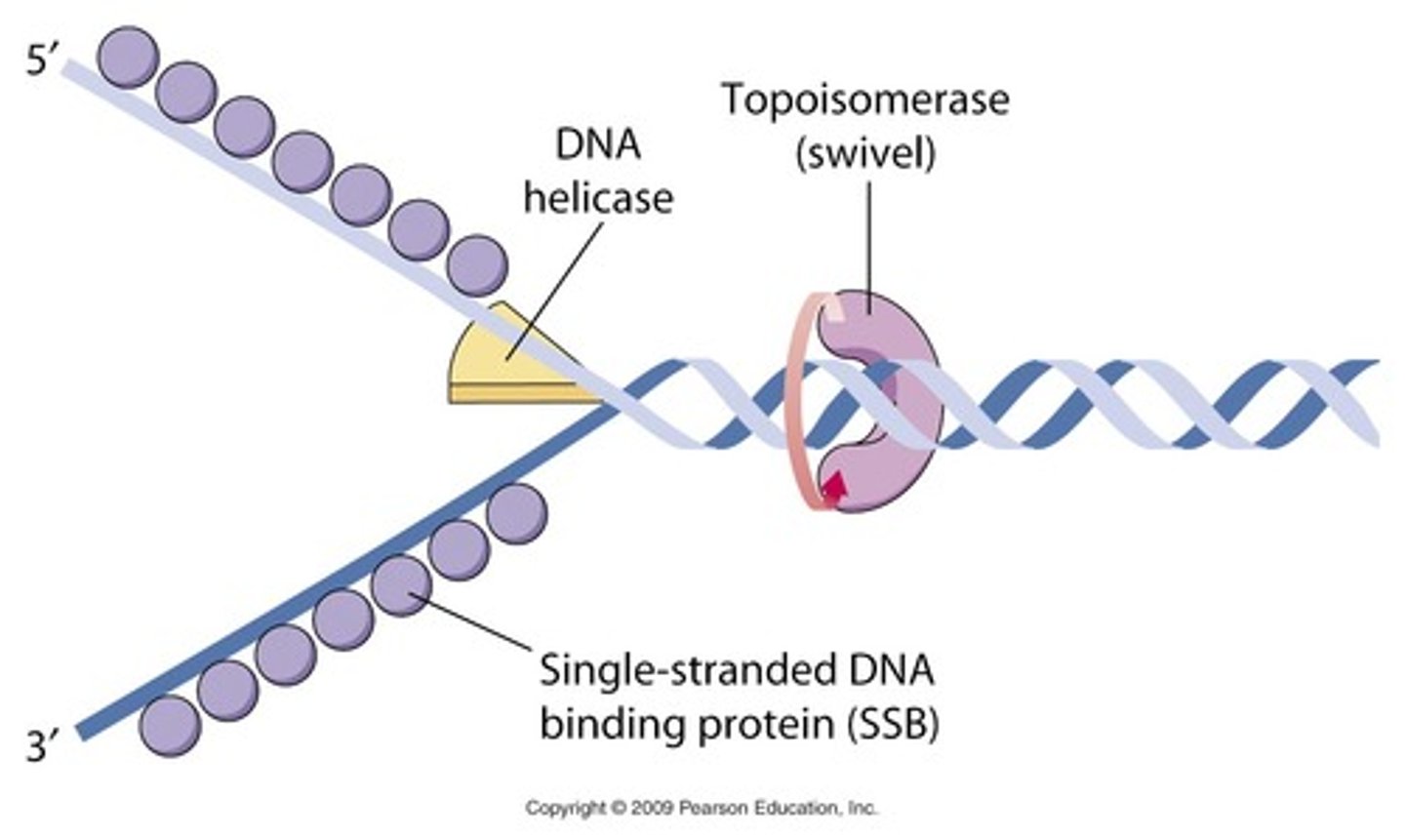
single stranded binding proteins (SSBs)
bind to each side of the DNA ladder keeping it from bonding back together during DNA replication
RNA primase
synthesizes RNA primers for DNA replication
Initiator proteins for DNA replixation
Helicase, SSBs, topoisomerase II
DNA polymerase I & III
dna pol I : elongation/synthesis/adding nucleotides
dna pol III : removes RNA primer & replaces with DNA
(prokaryotes only; eukaryotes different)
Reason for Telomeres and Telomere shortening
the last RNA primer on the lagging strand (3' end) has no 3' hydroxyl for DNA pol to attach to and replaxe these nucleotides once they are removed (unlike in prokaryotes) leading to shorter and shorter ends of DNA
What are telomeres?
ends of chromosomes designed to protect from DNA shortening. Made of of 100s-1000s of TTAGGG tandem repeats. get increasingly short with each division especially in somatic cells
telomere shortening does not occur in which types of cell?
gametes and stem cells. they contain telomerase, a reverse transcriptase that binds to the ends of the tails and adds repeats. potentially useful for aging/cancer
karyotype
A display of the chromosome pairs of a cell arranged by size and shape.
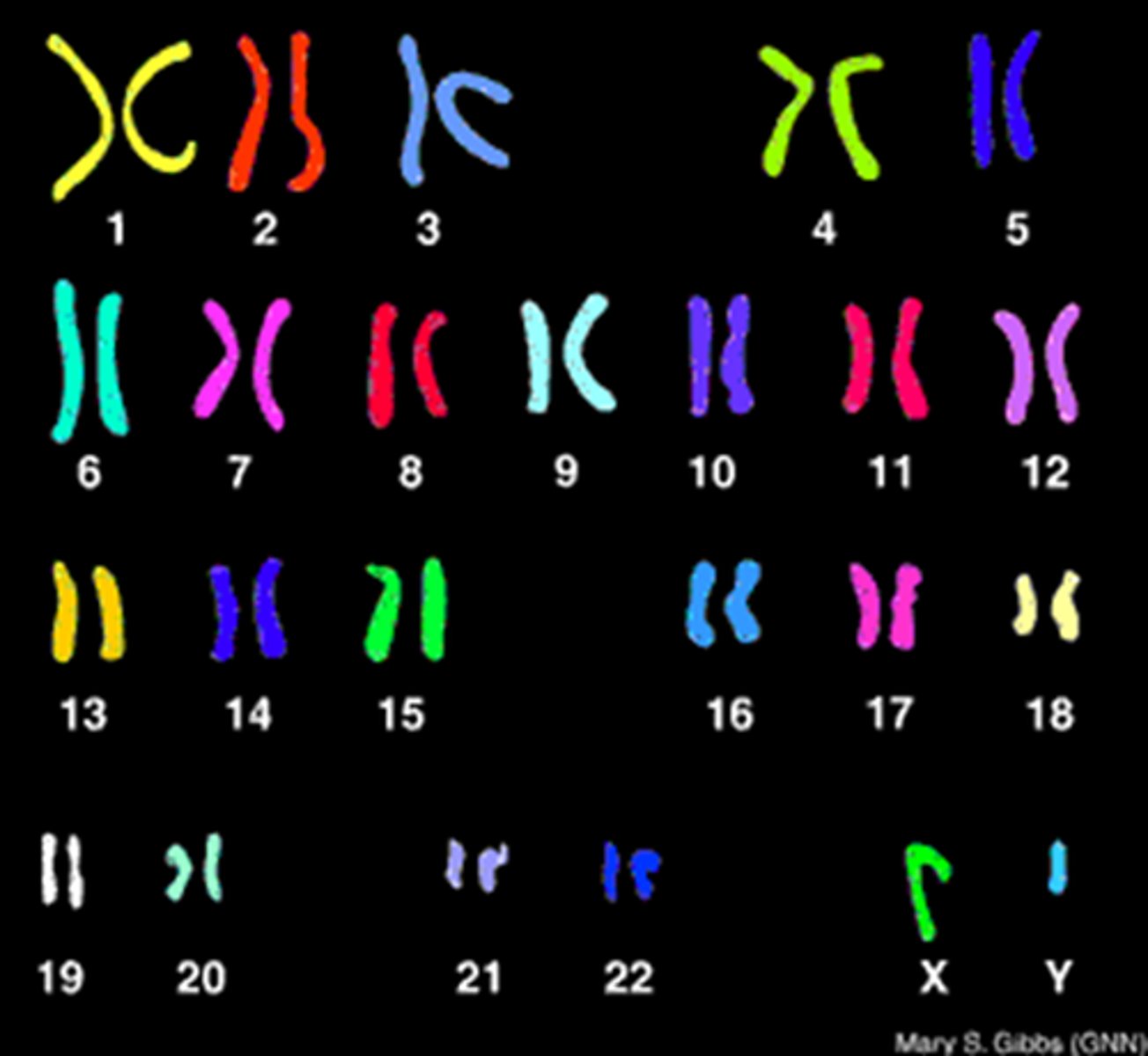
ploidy
number of sets of chromosomes in a cell
haploid vs diploid
one set of chromosomes vs two sets of chromosomes
mitotic spindle
a structure made of microtubules that controls chromosome movement during mitosis
unlike animals, during cell division, plant cells do not contain ____
centrosomes (though they do have mitotic spindle). final location defines the poles.
phragmoplast
In dividing plant cells, a structure formed by overlapping microtubules that guide vesicles containing cell wall components to the middle of the cell.
cyclin A
helps activate DNA synth
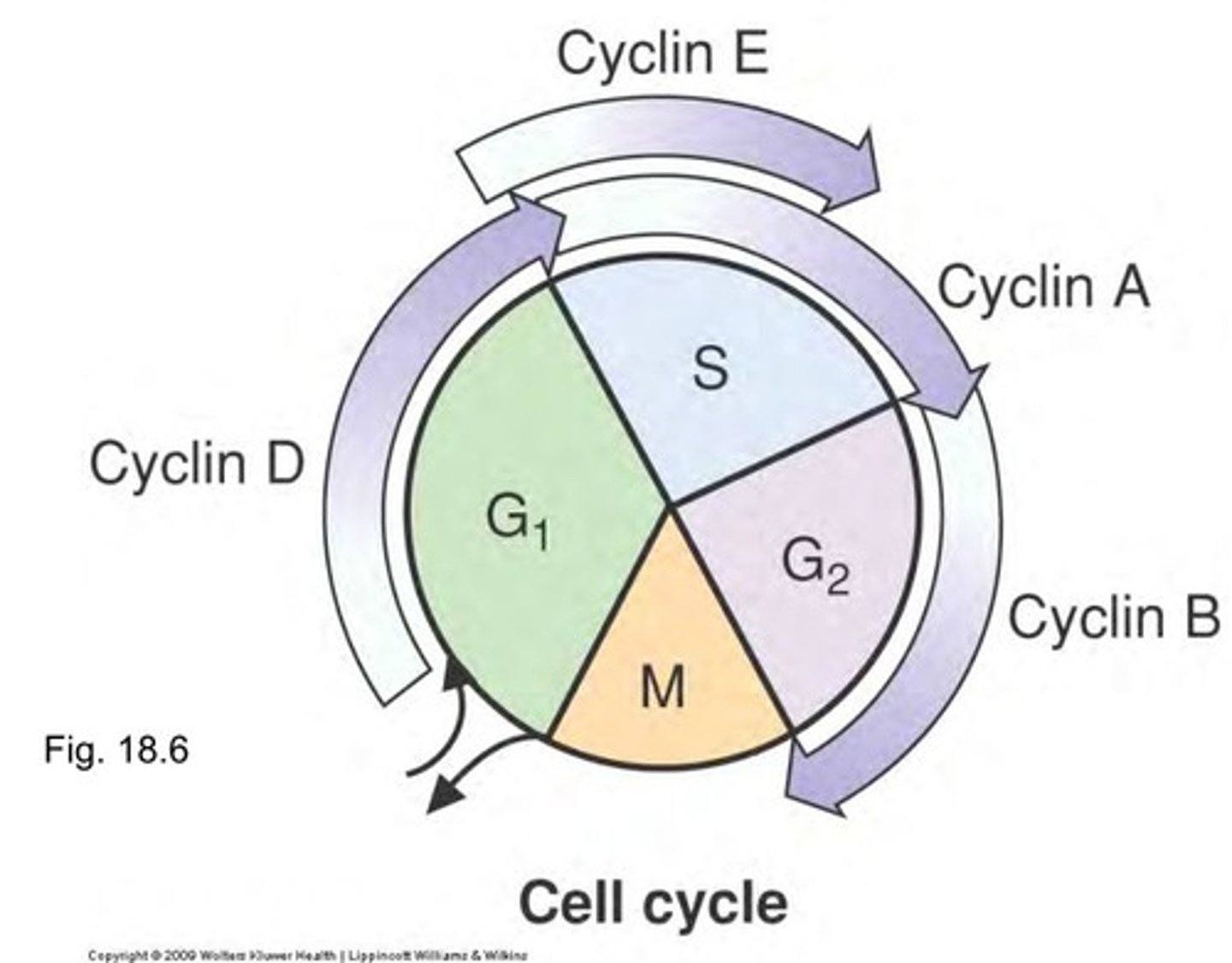
cyclin B
helps prepare for mitosis
Cyclin D & E
helps prepare for DNA replication (before A)
If one DNA strand is damaged during replication, what occurs in the other strand?
It slows down. DNA replication occurs simultaneously and the DNA polymerases interact together.
Amount of nucleotides lost per division
About 100. Adult cells can only divide under 50 times on average because of this, telomerase is inactive almost everywhere. Cells typically stop dividing when under 100 TTAGGG repeats are left. Telomerase reactivation has been associated with cancer.
Components needed for PCR (in vitro replication)
-Pair of primers to bind to specific point of DNA
-DNA polymerase (usually Taq)
-Template DNA
-essential ions and salts (buffered solution)
-free deoxyribonucleotides (dNTPs): dATP, dGTP, dCTP, dTTP
-topoisomerase and helicase not required due to thermocycler
Taq polymerase
heat-tolerant up to 95°C comes from thermus aquaticus
Stages of PCR with explanation
-Denaturation: unwinds DNA through heating
-Annealing: primers bind to DNA strands
-extension: DNA polymerase and dNTPS extends (/amplifies) DNA
gel electrophoresis
visualize DNA fragments, separate and compare lengths
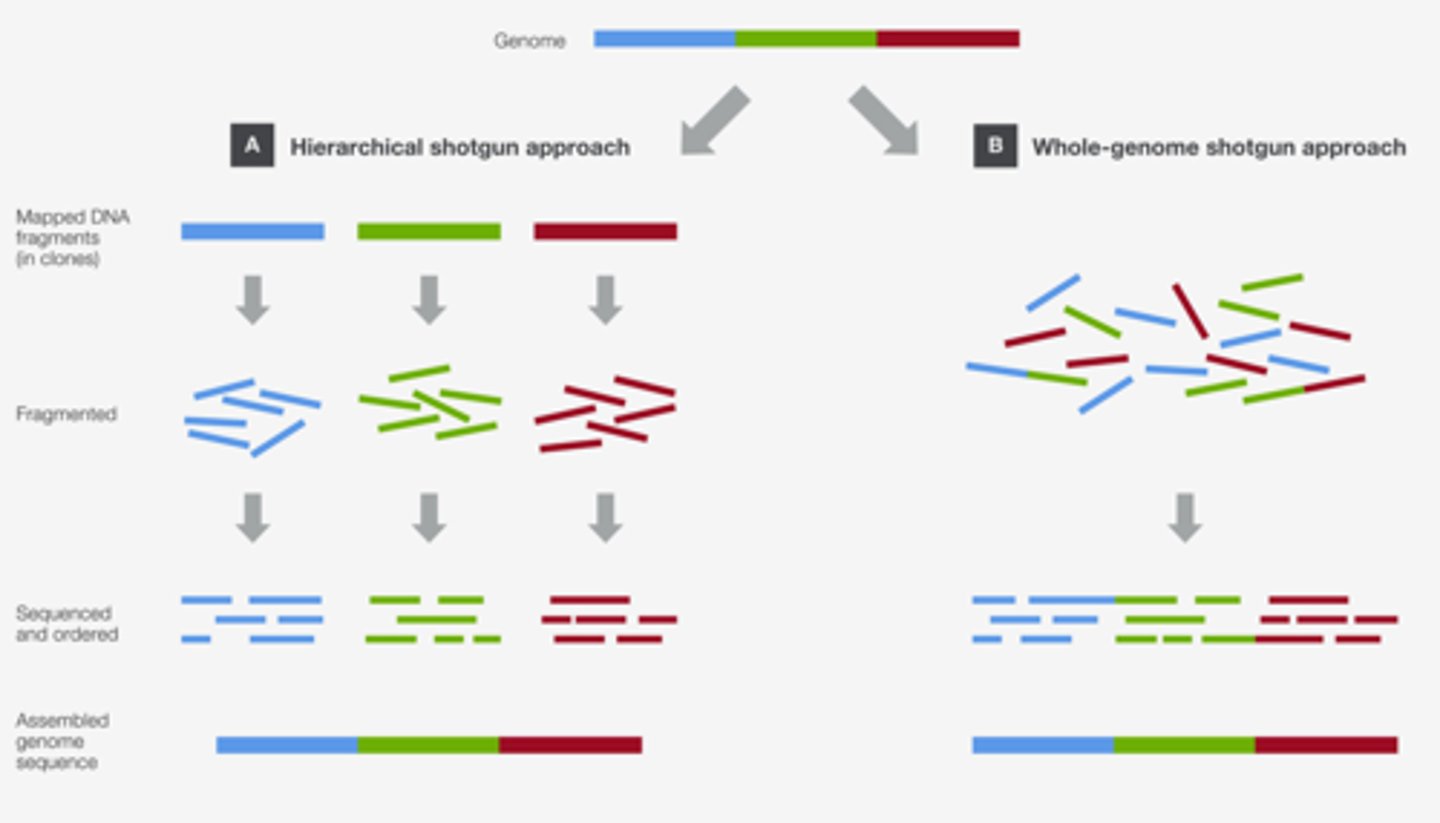
shot gun sequencing phases
1. Random sequencing of DNA in each fragment (10-50 copies of each region)
2. Identifying the regions of overlap and inferring / assembling each chromosome
3. annotating the sequences to best identify regions of coding/regulatory/non coding regions
sanger sequencing (dideoxy chain termination method)
-Same requirements as PCR
-work on the basis of modified fluorescent dNTPs missing 3' hydroxyl (ddNTPs) which stop elongation of DNA strand. DNA added to tubes with different ddNTPs leading to series of interrupted daughter strands with varying lengths.
-Only works on small fragments (few hundred nucleotides)
spectogram trace
compiles fragments from sanger sequencing into complimentary (because of the binding) sequence to be read left to right
contigs
Overlapping DNA segments that share the same sequence.
consensus regions
Regions of DNA having similar structure and function in different organisms.
Created from overlapping sequences/contigs
How many reading frames are there in a double stranded DNA and how do you decide which to use?
6 possible reading frames.
Look for long periods without stop codons, a start codon, possibly promoter or other regulatory site (usually done by a computer). easier in prokaryotes
sequence motif
A pattern of nucleotides or amino acids shared by different genes or proteins that often have related functions. (ex transcription factors, hairpin loops, ORFs)
____ % of eukaryotic genome is non coding
50%
What type of genetic information is most likely to undergo mutations and why?
RNA viruses due to delicate RNA backbone, possible lack of proofreading
probability of a new mutation ________ with __________
decreases with increasing genome size
somatic mutations
occur only in cells of certain tissues and cannot be passed on (ex most cancers)
germline mutations
occurs in germ (sex) cells, can be inherited and cause mutation in all cells of offspring (ex, haemophilia)
joshua and esther lederberg experiment
"mutations are random"
1- bacteria grown on non selective agar plate
2- bacteria colonies brought to selective agar plate 2 containing penicillin -> can only grow if resistant (stamping)
3- only some colonies survive - contain mutation resistant to penicillin
mutagens
increase the likelihood of mutations in DNA at certain points (radiation, chemicals, smoking)
Types of DNA polymerase II proofreading methods
Mismatch repair
Base excision repair
Nucleotide excision repair
mismatch repair
kink in DNA from mismatch detected by proteins
Single stranded cleavage of DNA backbone by enzyme (usually nuclease) some distance away
Another enzyme removes successive nucleotides
DNA pol, ligase close the gap by redoing DNA synth
Base excision repair
-fixing incorporation of Uracil into DNA
-DNA uracil glycosylase cleaves uracil from backbone
-lack of nitrogenous base detected by AP endonuclease, cleaves backbone on either side of spot lacking a base
-DNA synth occurs at base (DNA pol, ligase)
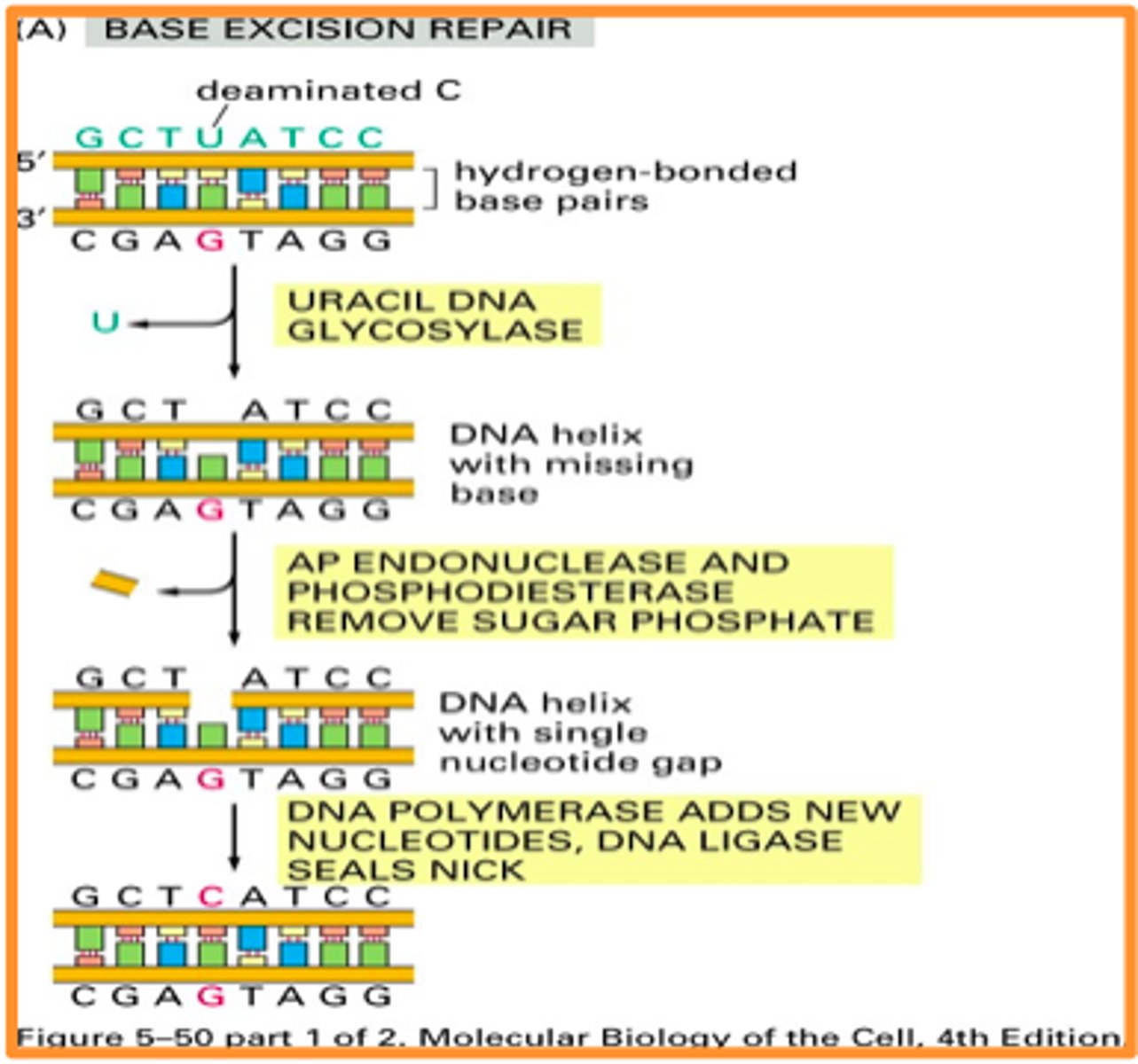
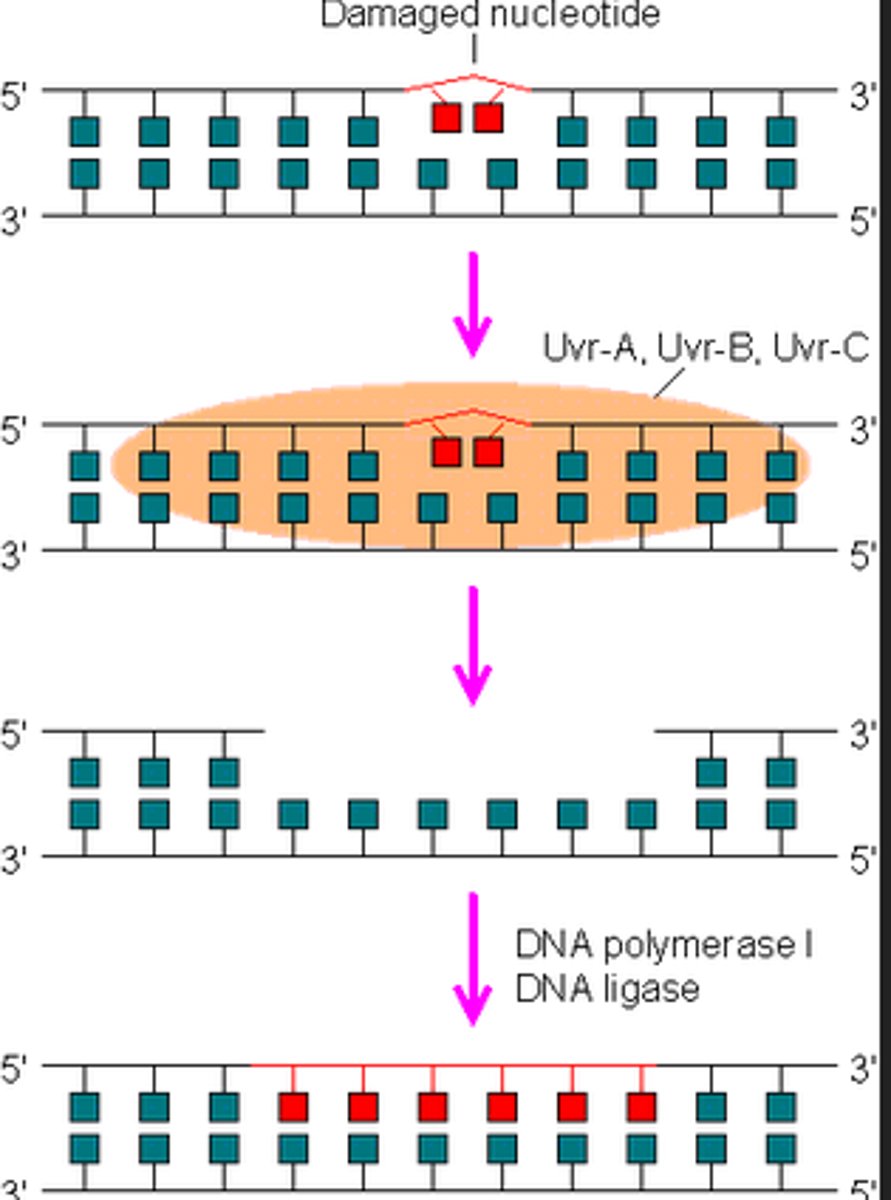
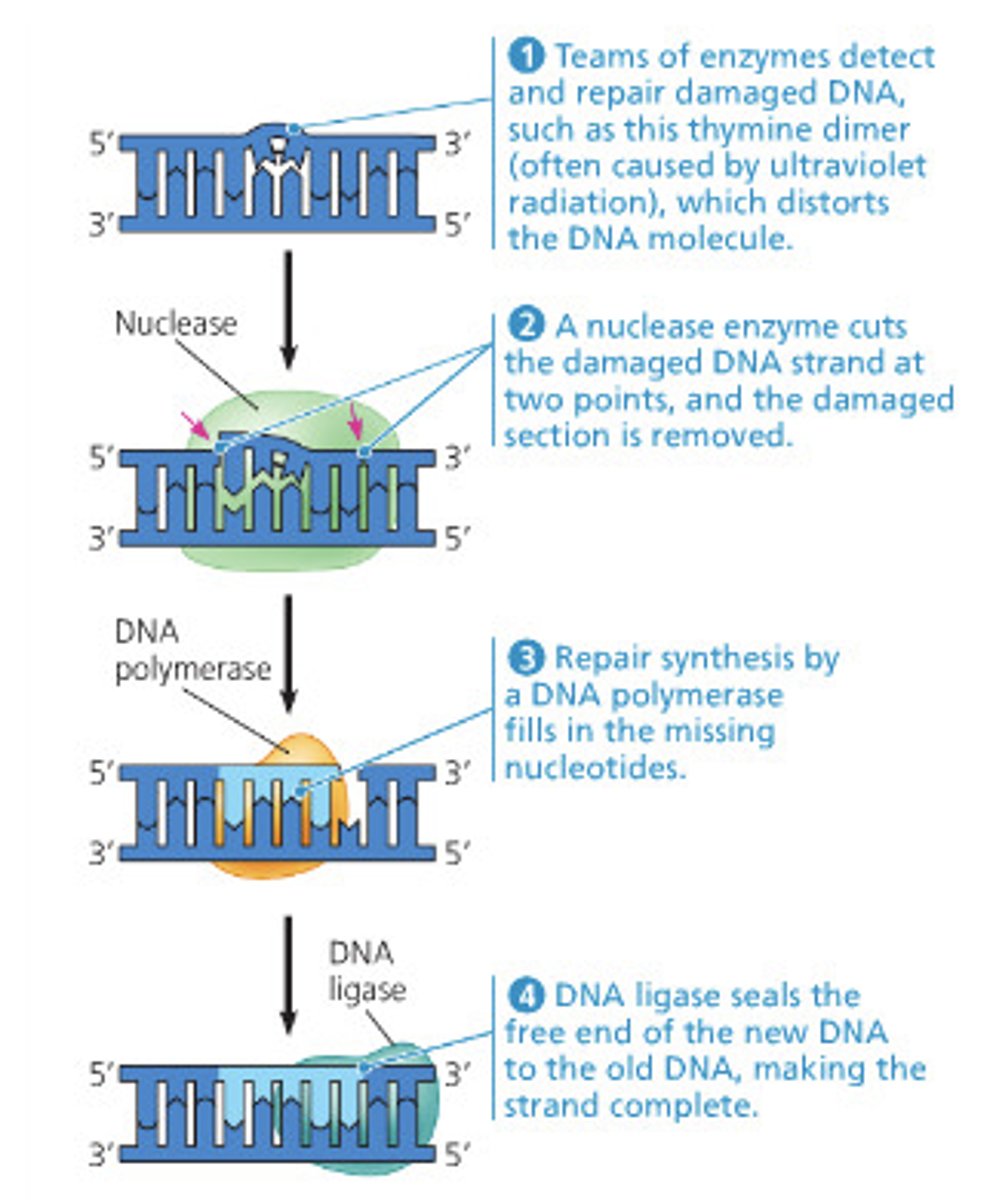
nucleotide excision repair
-similar to mimatch repair but with multiple NTPs
-damaged bases excised and replaced with re-DNA synthesis
single nucleotide polymorphism (SNP)
point mutation
synonymous mutation
silent mutation
non-synonymous mutation
missense mutation
Types of Large Scale Mutations
deletion, translocation, duplication, inversion
deletion of a centromere leads to
deletion of a chromosome (diploids might be able to make up for it with homologous chromosomes)
Duplication
aka amplification
little harm due to homologous chromosome
can be beneficial, give rise to similar but new gene (duplication & divergence)
Inversion
not harmful in somatic cells; harmful in germ if it causes gene breakage
-small inversions common in large populations, explains variation in gene order, long term genetic evolution/diversity
reciprocal translocation
two-way exchange of segments between the chromosomes
-mainly in non coding regions
-generally don't disrupt gene function
-can cause problems in gametes if homologous chromosomes don't pair during cell division, leading to vital genes missing
beta-globin gene family
-5 genes that arose from duplication and divergence event 200MYA
tandem repeats
up to 1000s of nucleotides in length, next to each other, identical or close
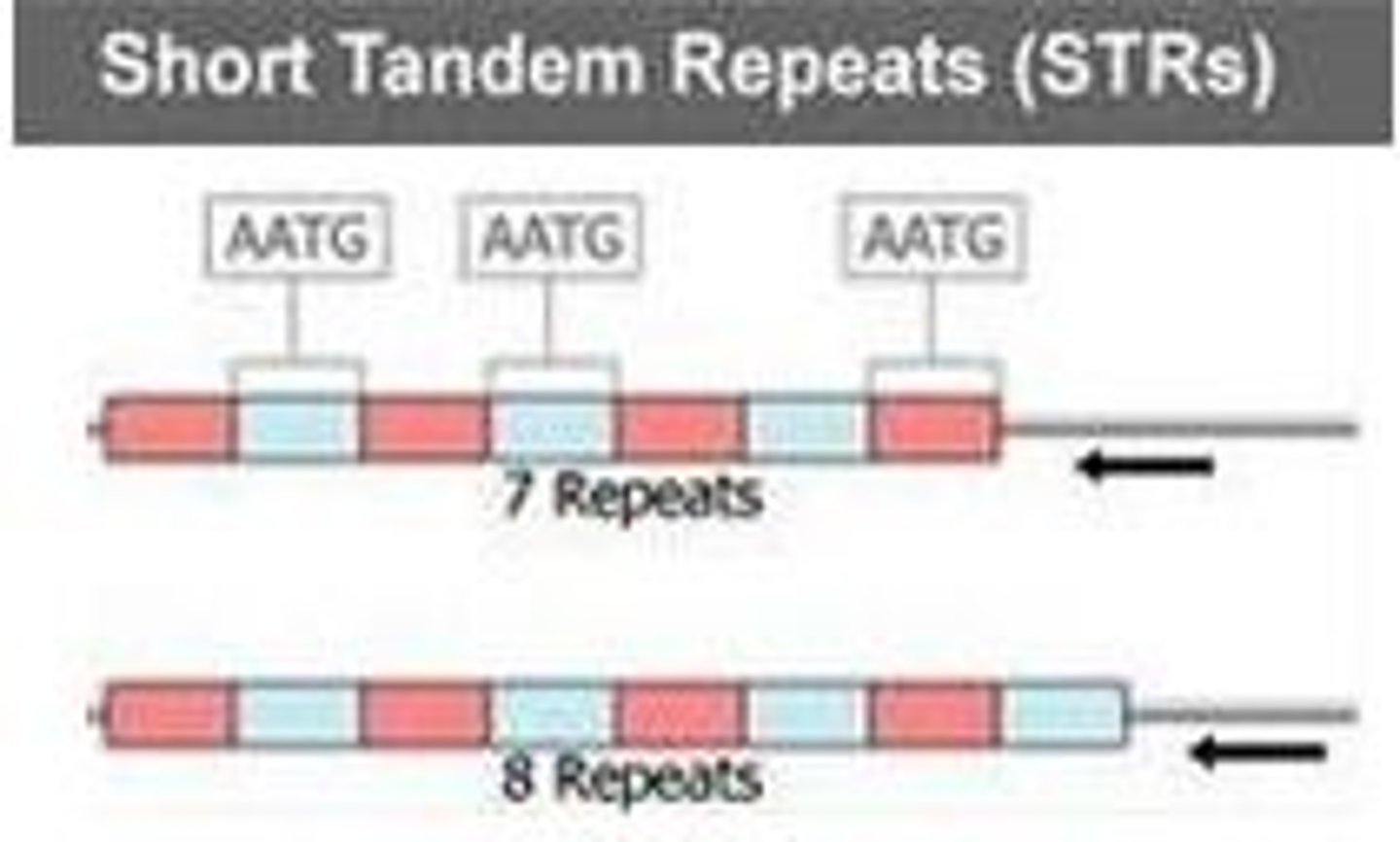
simple sequence repeats
short 2-3 base sequences repeated thousands of times
locus
position of a gene on a chromosome
DNA markers
Sequence variations among individuals in a specific region of DNA (usually non-coding)
Detected via microarray analysis, PCR, Southern Blot, DNA sequencing, used for identification/relatedness
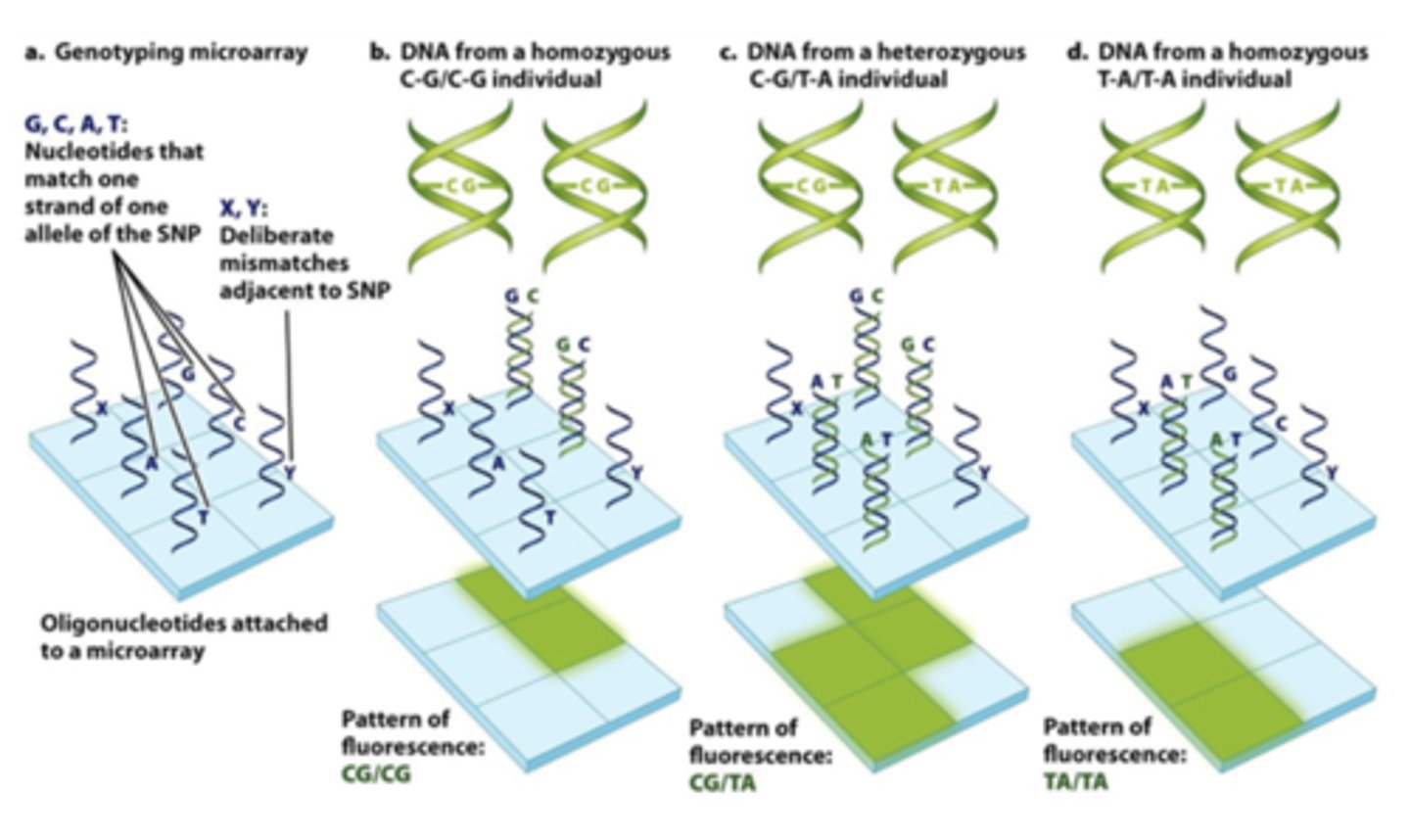
DNA microarray analysis of SNPs
-Common allele + SNPs of oglionucleotide (single strand, ofc) attached to glass
-Only matching alleles light up
-Identify homo vs heterozygous
variable number tandem repeats (VNTR) + identification
differences in copy number (I think?, identified using PCR and gel electrophoresis
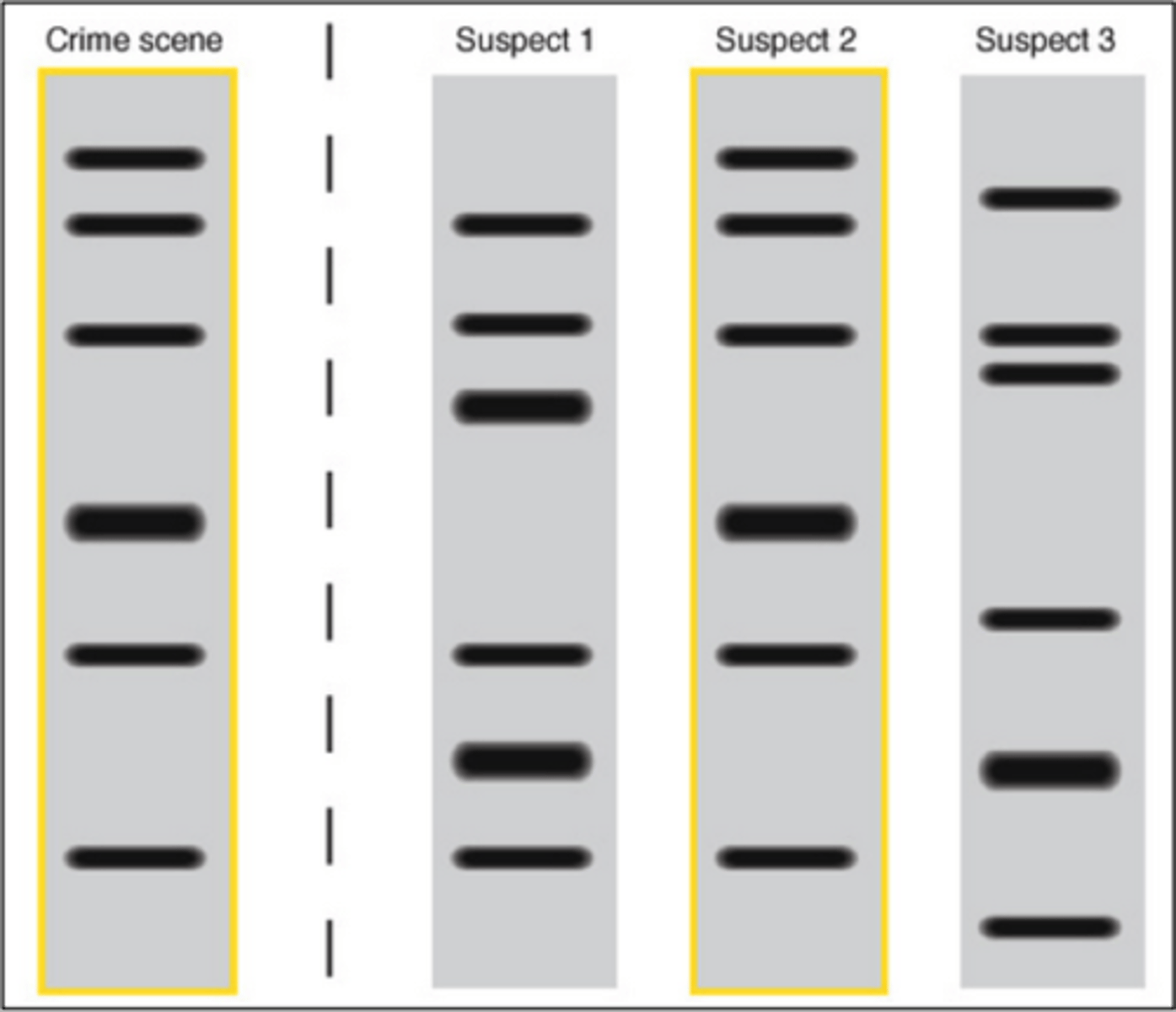
DNA fingerprinting use
forensics
relatedness
large scale population analyses
autosome
Any chromosome that is not a sex chromosome (somatic refers to cells)
sickle cell anemia alleles + symptoms
HbA: normal
HbS: sickle (glutamine substituted for valine)
aggregation of beta-globin, blockage of capillaries, anemia, acute body pain, organ/tissue damage
Heterozygotes will express both versions of the protein but will not display symptoms of sickle cell anemia
Provides advantage in fighting off malaria