cells exam ii
1/17
Earn XP
Description and Tags
conn coll!
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
18 Terms
Reporter genes
ex. GFP (glow), luciferase (glow0, Lac Z (blue). They are used by scientists to identify certain segments of a gene and seeing if they get expressed. Replace target gene, such as FXR, and when it gets expressed, it glows.Reporter genes are genetic markers that produce measurable signals, like fluorescence or color, indicating the expression of a target gene. They help in studying gene regulation and the functionality of gene sequences.
What methods used to levels of transcripts?
Real Time PCR:
also uses reverse transcription polymerase reaction ; number of input DNA increases, the time to detect fluorescence, decreases.
—> transcript levels in different tissues
and Reverse Transcriptase PCR:
isolate desire mRNAm, TTTT primers to poly AAA tail, use reverse transcriptase to copy RNA into DNA, separate strands + second primer, now use PCR to amplify strand + use primers from gene being studied
Post Transcriptional Controls
Alternative splicing: allows one single gene to make multiple mRNAs and therefore proteins. Regulated by RNA binding proteins → how transcript will be spliced → can have very different results
mRNA localization and translation
RNA binding proteins
RNA binding proteins represses translation of its own RNA when found in excess
also high temp
miRNAs direct destruction of target mRNAs, regulate gene expression
these partially double stranded RNA Molecules first bind to Dicer (not shown) that cuts the ds RNA into ~21 bp pieces. The dsRNA then binds to proteins to form the RISC (RNA-induced Silencing Complex). One of the two strands of RNA are eliminated. . The remaining RISC searches for complementary mRNA sequences and either destroys them or inhibits their translation.
siRNAs DO NOT regulate gene expression - shut off viruses and function similarly
You identify a protein, named Camel, that affects the expression of the gene named Conn. Upon further investigation, you determine that the Camel protein sequence includes a section that is found in many RNA binding proteins. Name one way that the Camel protein might be affecting the expression of Conn based on these findings
1. The Camel protein could bind the Conn mRNA to affect it’s splicing such that the Conn mRNA is spliced differently in different tissues.
2. The Camel protein could bind the Conn mRNA to transport it for specific cellular localization.
3. The Camel protein could bind the Conn mRNA preventing it from being translated into a protein.
Protein Structure
Primary: amino acids with R groups linked together by peptide bonds (strong covalent)
R groups can be charged or uncharged, polar or nonpolar
3 letters, 20 in total - codon
determine structure and function
Secondary
alpha helices and beta sheets
held together by backbone to backbone H bonds (both beta and alpha)
both can be in one protein → form tertiary
H bond bonded to O of C=O bond in backbone
how membrane bound proteins cross the lipid as alpha helix
beta sheets can be parallel or antiparallel
first described!
Tertiary:
3D structure of entire polypeptide chain
Quaternary:
3D structure of multiple polypeptide chains
ex. neuraminadase is 4 polypeptides
symmetrical assembly of 2 different polypeptides
N terminus: amino
C terminus: carboxyl
polypeptide backbone does not include R chains!!
Bonds that help proteins fold?
Covalent (peptide bonds) initial structure and then: hydrogen bonding,Van der Waals, electrostatic attractions, hydrophobic forces, disulfide bonds (covalent) form between two cysteine form keratin → stabilize 3D structure all together
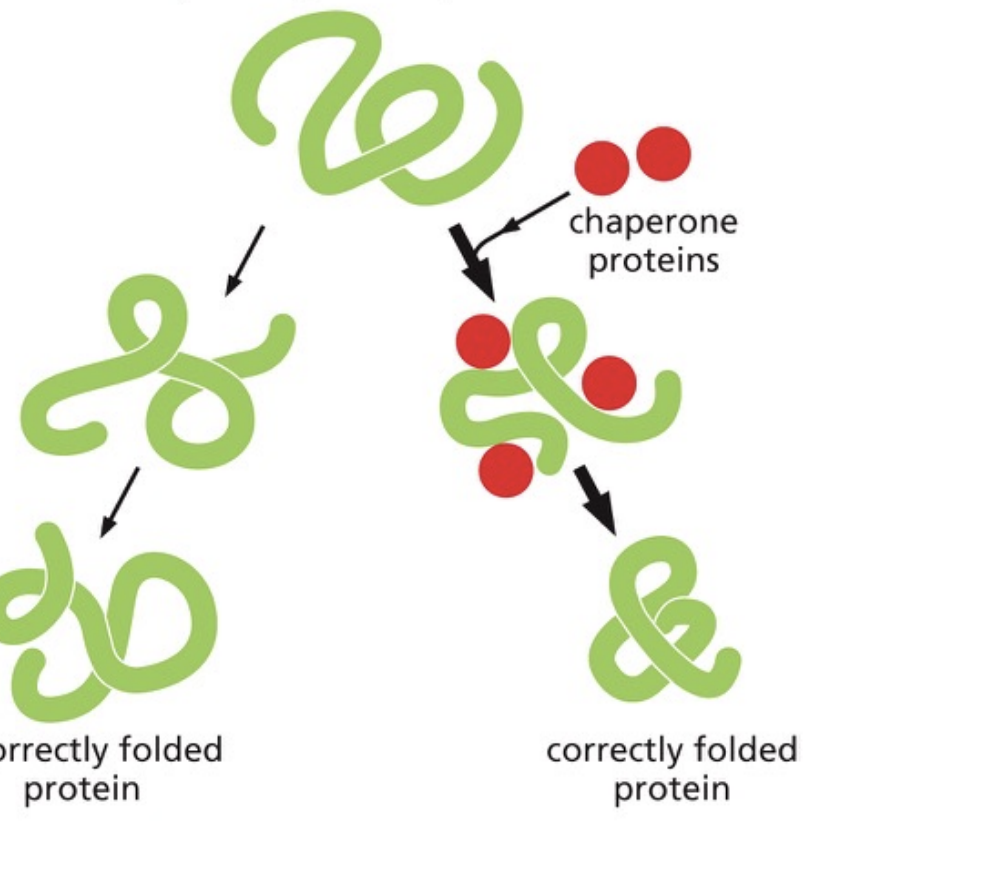
Protein folding
denatured proteins can offer recover shape
all info for 3D is in primary structure
ex. urea/denature remove/reform
Chaperone proteins
assist in protein folding
some bind to partly folded chains and help them form
others form an isolation chamber in which single polypeptide chains can fold correctly
Why is protein folding important?
improper folding leads to disease, such as prion diseases (neurodegenerative diseases)
Madcow/Creutzfeldt-Jakob disease
How do proteins work?
Ligands are any substance bound to a protein
the ability of a protein to bind selectivley to a ligand is due to the 4 noncovalent interactions
van der waals
hydrogen bonds
electrostatic attractions
hydrophobic forces
region of protein for ligand: binding site
allows proteins to interact with specific ligands - like cyclic AMP
made of particular amino acid (R) chains
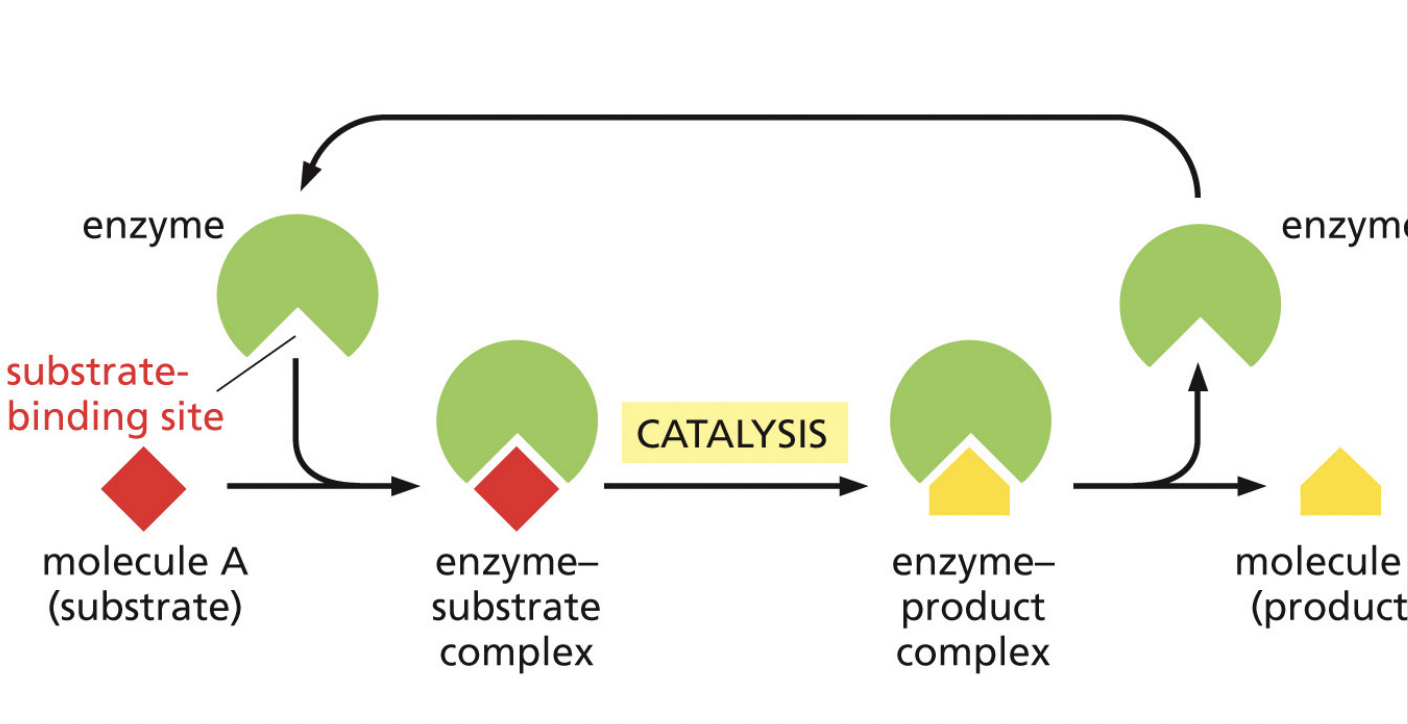
Enzymes
enzymes are proteins but not all proteins are enzymes
even energetically favorable reactions require activation energy to get started → ex. heat or enzymes
enzymes lower the activation energy of a reaction → can speed up rxns
a series of enzyme catalyzed reactions can create a linked pathway
energetically favorable - Δ G
energetically unfavorable + Δ G
reaction coupling to push unfavorable rxn
Enzymes convert substrates to products while remaining unchanged
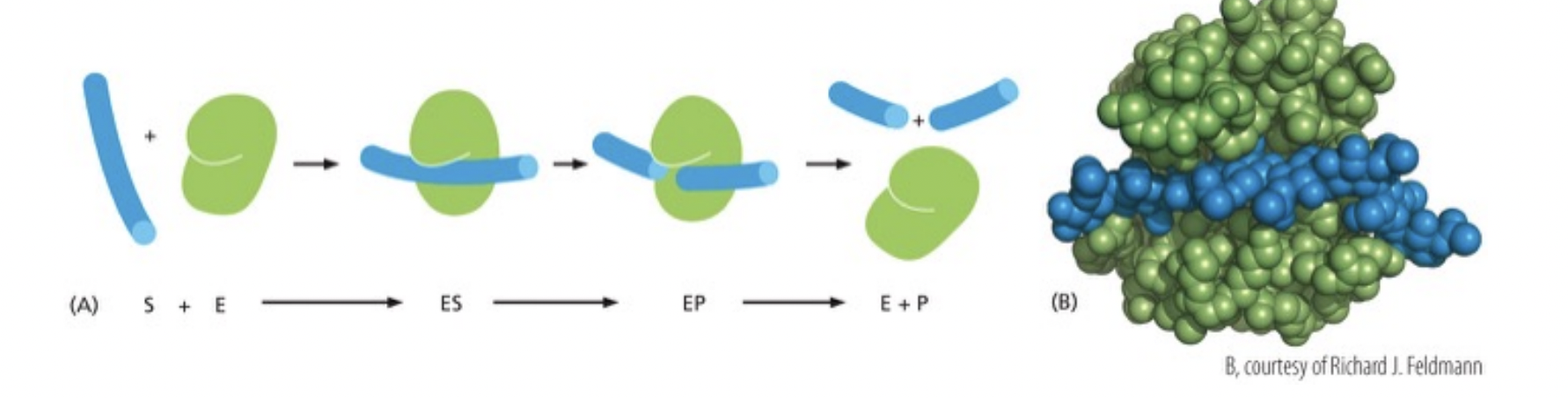
How do enzymes lower activation energy?
Holding reactant substrates together in precise alignment to encourage rxn
binding of substrate to rearrange electrons in the substrate creating partial + or - to favor reaction
enzyme strains the bound substrate forcing to a transition state that favors rxn
a single enzyme might use a combination of these mechanisms
ex: lysozyme
Lysozyme lowers the activation energy by:
1. Straining the substrate
2. Altering the distribution of electrons in the substrate
hydrolysis +h2O rxn
binding site is referred to as the active site
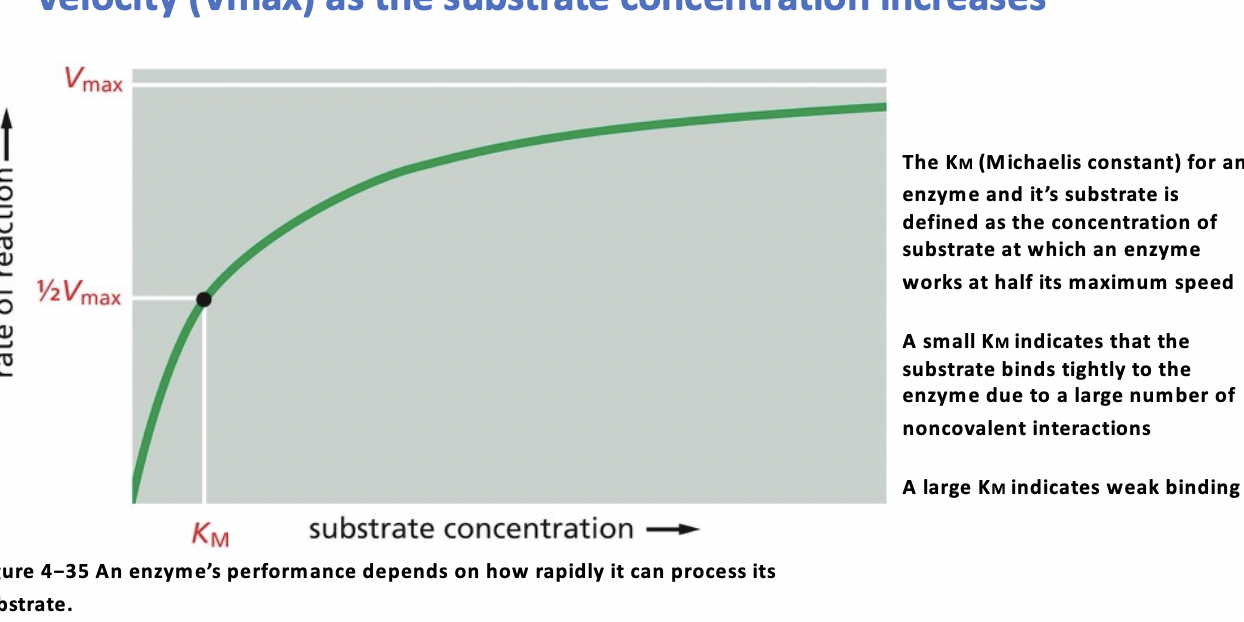
Vmax
maximum rate that an enzyme can convert a substrate to a product
Km
How well the enzyme binds to substrate
assay
way to measure the disappearance of a substrate and appearance of product
ex. using spectrophotometer
competitive inhibitor
blocks substrate from binding to desired enzyme, increases Km but Vmax stays the same
ex. Gleevec
What is the purpose of enzymes?
Enzymes speed up reactions by lowering the activation energy
Enzymes drive biological reactions in cells without the cells having to raise their temperature Enzymes usually catalyze the formation or breakage of a single bond
Enzymes bind to a substrate in their active site
Enzymes and substrates have a characteristic Vmax and Km that can be determined experimentally
Transcription Regulators
3 alpha helices of transcription regulator bind to sequence of DNA → homeodomain
sometimes ligand is DNA → these specific interactions with amino acids and DNA molecules demonstrate how it can bind to specific enhancers (DNA sequence)
How are proteins controlled?
allosteric proteins have two or more binding sites - saw with trp operon
feedback inhibitions: an enzyme that catalyzes an important reaction is turned off when a substance is present in high amounts
phosphorylation (covalent bonds): causes a conformational change which can either decrease or increase a proteins activity by adding a phosphate group
Groups that are covalently attached: multiple sites can be modified by other phosphates, acetyl (histones), fatty acids, sugars, and ubiquitin - can change conformation and mass
Non covalent GTP binding - molecular switches
Kinase: Enzyme that catalyzes the transfer of a phosphate group from ATP to a molecule
Phosphatase: Enzyme that catalyzes the removal of phosphate from a molecule