Bacterial Protein Synthesis
1/98
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
99 Terms
Gene expression
is the process by which genetic information flows from DNA to RNA to the protein.
Protein synthesis from RNA templates is called
translation
Translation is carried out by
ribosomes, which are large molecular complexes of ribosomal rna (rRNA) and proteins.
Protein Synthesis happens in 3 steps:
Initiation, Elongation, and Termination
Initiation requires 3 small proteins called
initiation factors
What are the three initiation factors
IF1, IF2, and IF3
Initiation step 1:
IF3 readily binds to the small ribosomal subunit, and its presence blocks the large and small subunits from prematurely associating. IF3 facilitates the binding of the mRNA to the small subunit of the ribosome.
Initiation step 2:
The binding occurs just 4 to 8 bases upstream of the AUG start.
What does the Shine-Dalgarno sequence do in initiation?
It positions the ribosome correctly on the mRNA so translation starts in the right place (P site) (remember it as a ribosome parking spot)
gene expression in eukaryotes
chromatin remodeling
transcription
RNA processing
translation
post-translational modifications
Initiation in bacteria step 1:
The RNA polymerase core enzyme and sigma subunit bind to the -10 and -35 promoter consensus sequences.
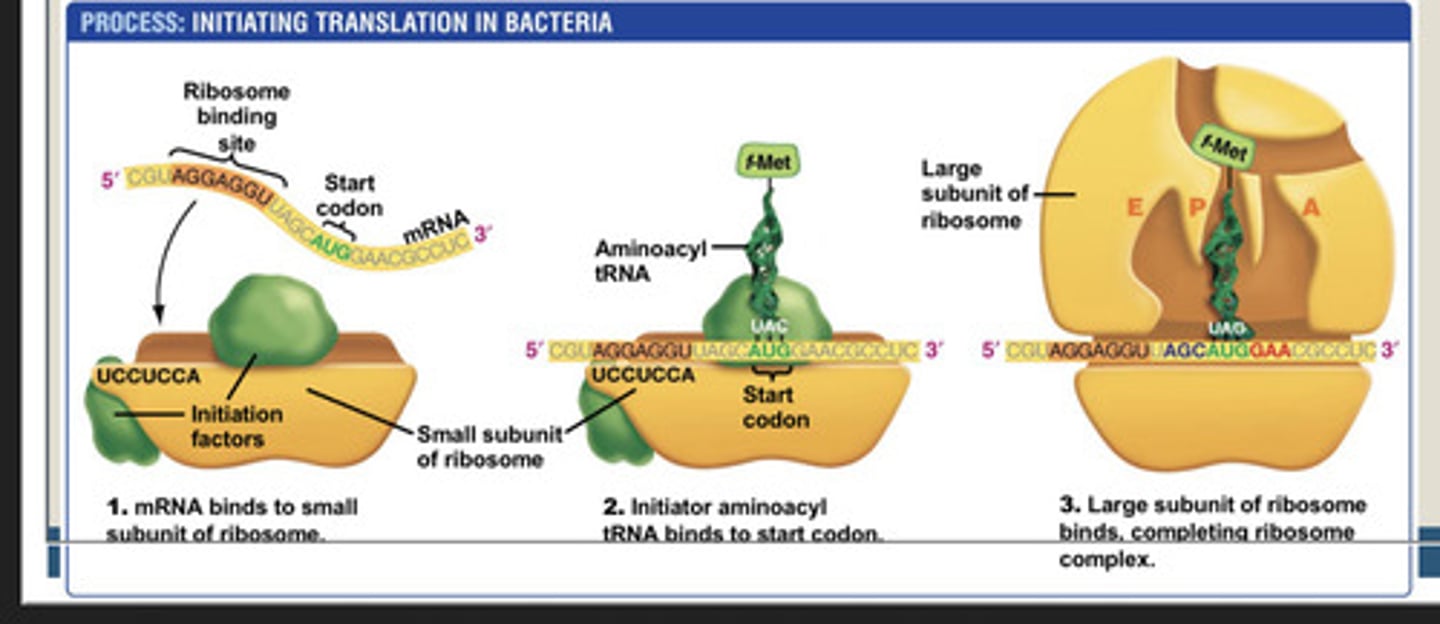
Transcription Initiation in bacteria step 2:
DNA unwinds near the transcription start site to form the open promoter complex
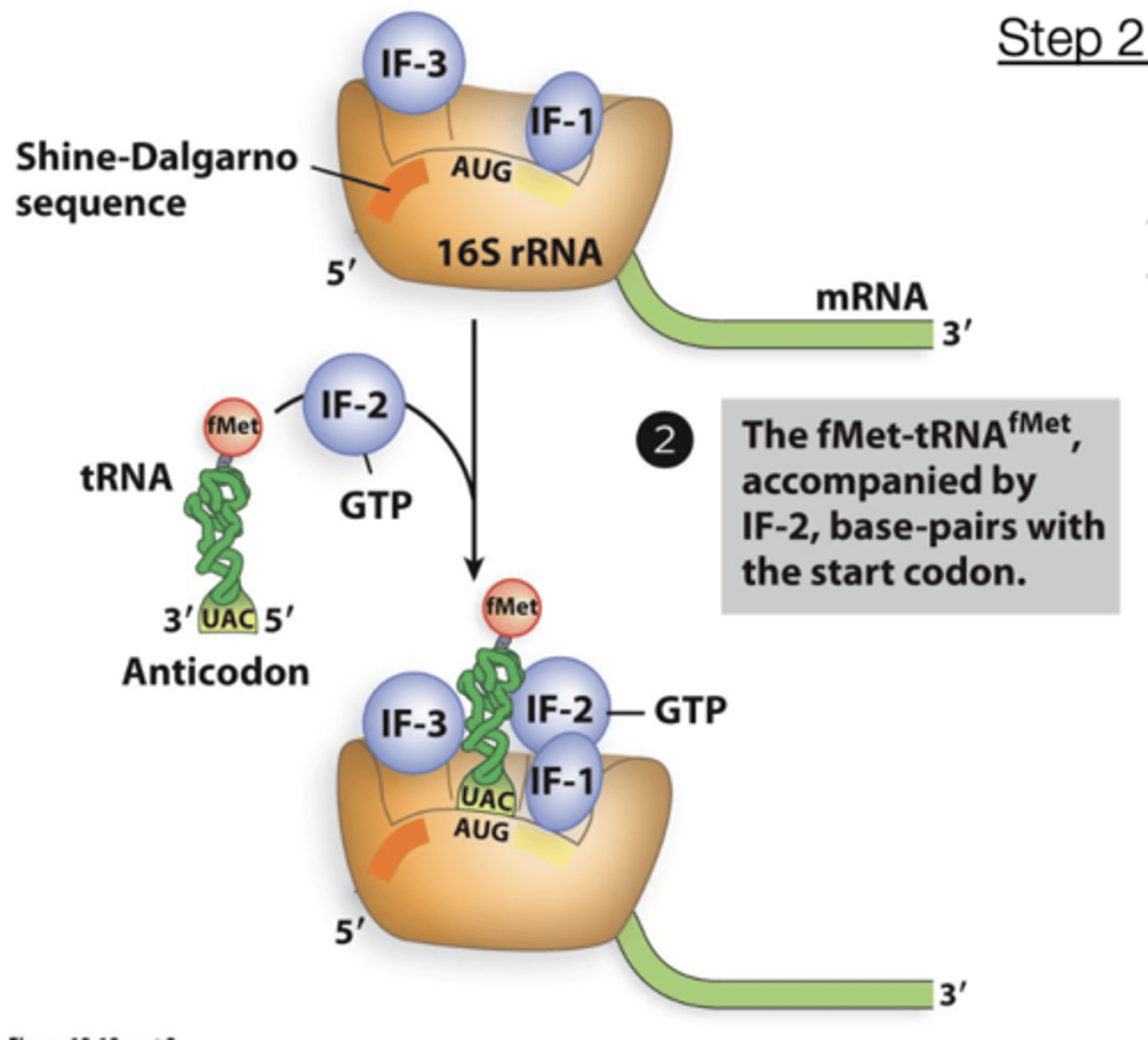
Transcription Initiation in bacteria step 3:
RNA polymerase holoenzyme initiates transcription and begins RNA synthesis. The sigma subunit dissociates shortly after transcription initiation, and the core enzyme continues transcription.
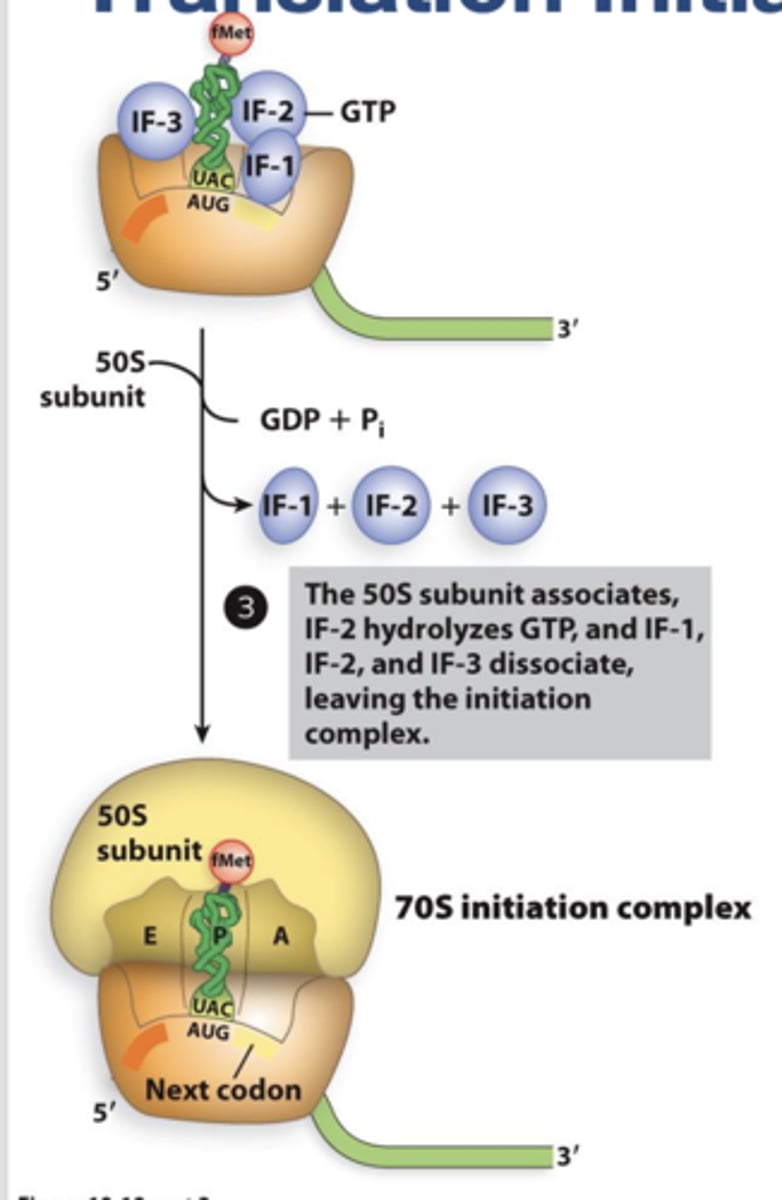
Transcription Elongation in Bacteria:
The core enzyme synthesizes until it encounters the termination sequence. As RNA synthesis progresses, the DNA duplex unwinds to allow the template strand to direct RNA assembly. The duplex closes following synthesis.
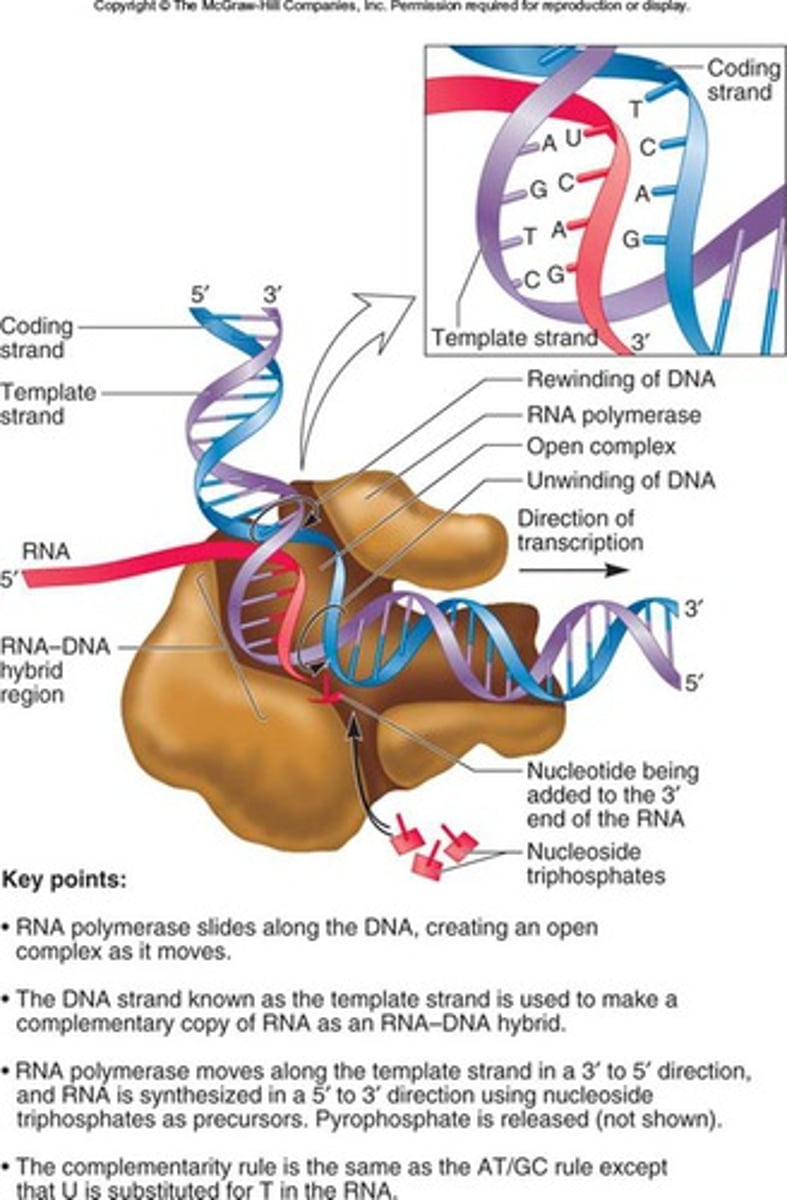
Termination in bacteria (intrinsic)
Inverted repeat sequences in the transcript fold into a complementary stem ending in a single-stranded loop. Then, hydrogen bonds between A-U base pairs break. Releasing the transcript and termination transcription.
Transcription in Eukaryotes
occurs in the nucleus and can bind to a DNA template on its own.
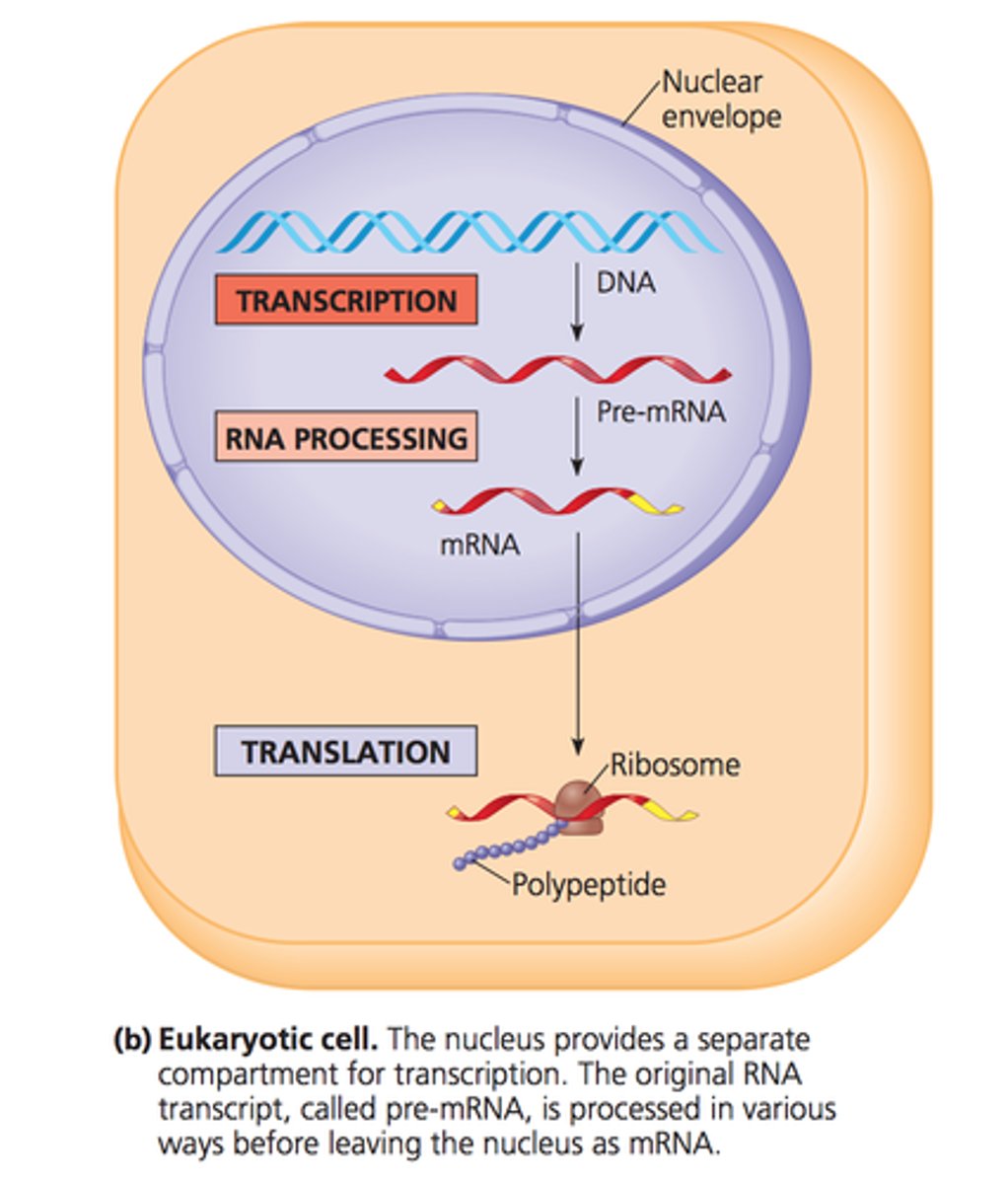
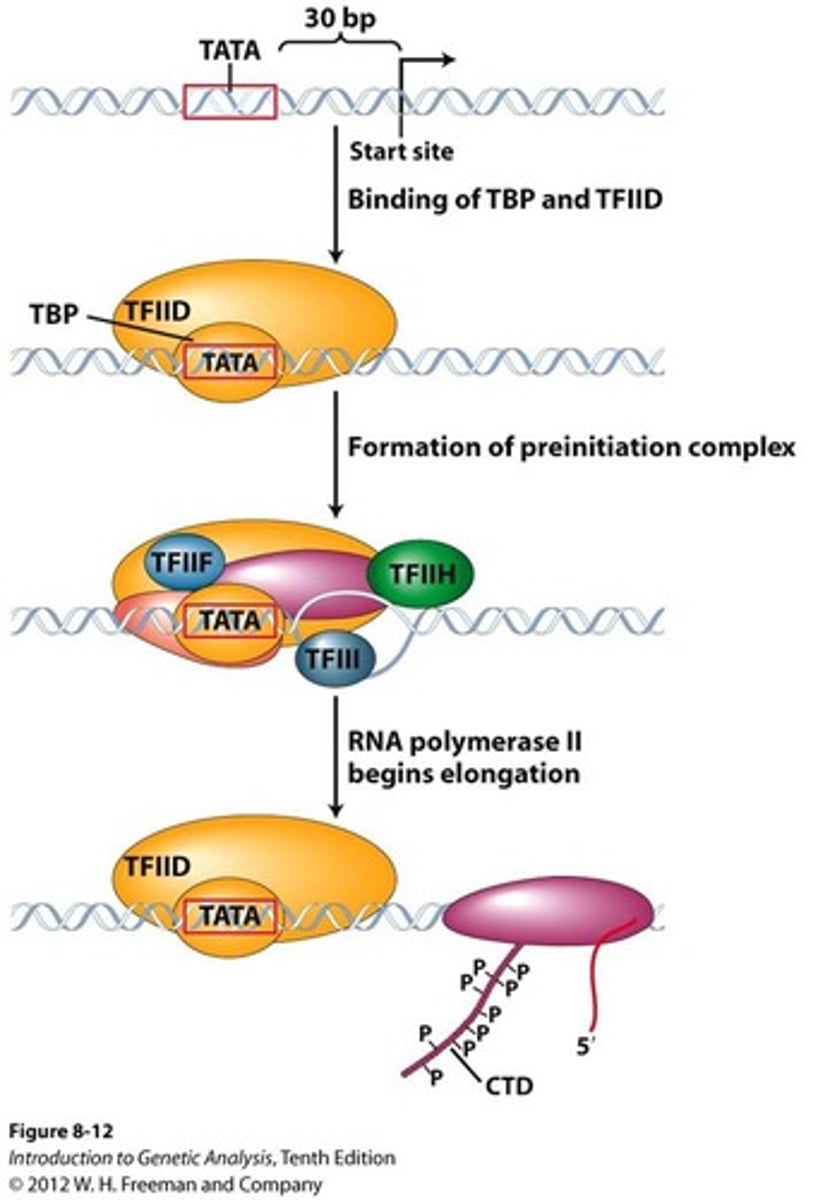
Initiation in Eukaryotes
Transcription factors bind to the promoter
RNA polymerase and additional transcription factors join to form the initiation complex
RNA polymerase begins to transcribe the template strand
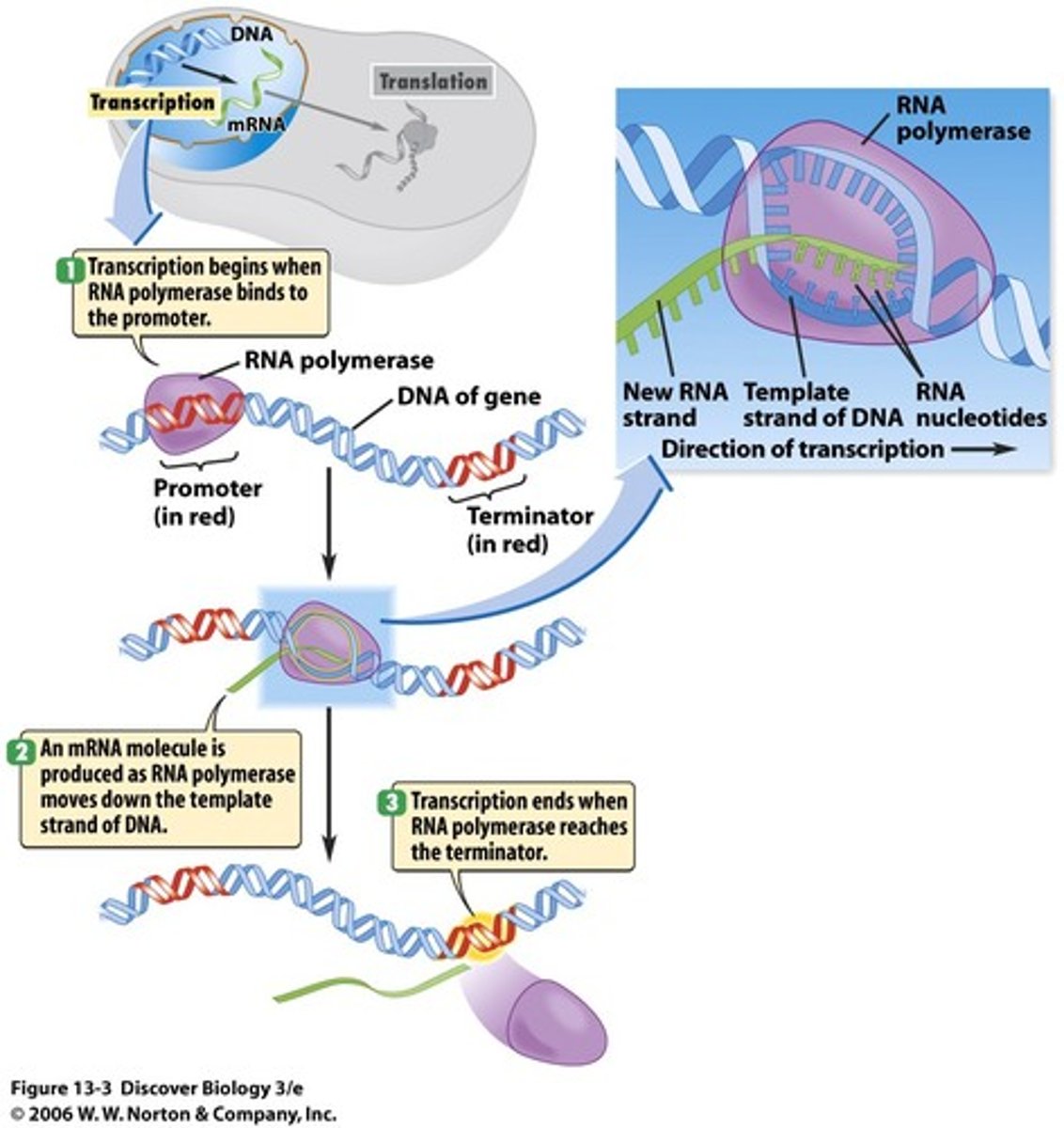
Elongation in Eukaryotes
RNA polymerase while moving along DNA unwinds the double helix and adds 40 nucleotides per second
RNA processing in bacteria
transcription produces an mRNA that can begin translation even before transcription is complete
RNA processing in Eukaryotes
Bacteria do NOT have RNA processing like eukaryotes; translation can begin while transcription is still happening
RNA processing has three steps
1. Capping the 5' end
2. Polyadenylation of the 3' end
3. Removal of introns
First step of RNA processing
A guanine is added to the 5' end with a 5' to 5' triphosphate linkage and is methylated (becoming the 5' cap)
Roles of the 5' cap
1. It protects the mRNA from being degraded
2. Facilitate the transport of mRNA into the cytoplasm
3. Facilitate intron splicing
4. Enhance translation efficiency
Second step of RNA processing
Part of the 3' end is removed, and then 20-200 called poly-A tail adenines (A's) are added to the 3' end
What does the Poly-A Tail Addition do?
1. Increases the stability of mRNA
2. Helps with export from Nucleus into the cytoplasm
3. Helps with translation
Major types of Introns
Group I
Group II
Nuclear pre-mRNA
tRNA and rRNA
Third step of RNA Processing
RNA splicing. Introns (non-coding regions) are removed, and exons (coding regions) are joined together. This is done by spliceosome
Why is RNA splicing important?
1. It creates a functional mRNA
2. Allows for alternative splicing which means different proteins from the same gene
Where are the start and stop codons?
START:
- where the translation begins (AUG)
(codes for methionine)
STOP: (UAA)
- where the translation stops
Basic requirements for translation
mRNA, tRNA, ribosomes, amino acids, and energy.
Whats the kozak sequence?
Its a specific sequence in eukaryotic mRNA that helps the ribosome find the start codon (AUG) (think of it as the start signal for eukaryotes)
Ribosome structure
Two subunits made of ribosomal RNA and proteins; can be free in cytosol or bound to ER
Elongation in Translation
tRNA brings amino acids
Peptide bonds form
Ribosome moves (translocation)
Difference between Prokaryotic and Eukaryotes Translation Initiation
prokaryotes --> Shine-Dalgarno
Eukaryotes --> Kozak + scanning
Termination in Translation
Stop Codon enters A site
Release factor binds
Polypeptide released
Bacterial Transcription has 3 critical regions:
1. Promotor
2. RNA coding
3. Terminator
Promotor
DNA sequence that the transcription apparatus recognizes and binds. Specific binding sites within the promotor includes consensus sequences at -10 and -35. That is 10 and 35 base pairs upstream of the transcription start site. Some bacterial promoters contain a 3rd consensus sequence called the upstream element that takes part in the initiation of Transcription.
RNA coding
This is a DNA sequence that is copied into an RNA molecule
Terminator
A DNA sequence that signals where Transcription is to end. An example is a Rho-Independent terminator in bacterial cells.
Bacterial Transcription Step 1 Inititiation:
1. RNA polymerase holoenzyme (core enzyme + delta/sigma factor_ binds to the promoter
2. Promoter has -35 and -10 regions
3. DNA unwinds and forms open complex
4. RNA synthesis starts at the +1 site
5. After a few nucleotides are made, sigma factor leaves
Why does initiation happen in bacterial transcription?
1. Sigma factor = specificity; it ensures RNA polymerase binds to the correct gene
2. DNA must open so the template strand can be read
Bacterial Transcription Step 2 Elongation:
1. RNA polymerase moves along DNA
2. Addes Nucleotides 5' --> 3.'
3. DNA unwinds ahead and rewinds behind
4. RNA strand grows longer
Why does Bacterial Transcription Elongation happen?
1. This is where the actual RNA transcript is built
2. Base Pairing ensures the correct sequence
Bacterial Transcription Step 3 Termination Intrinsic:
Rho-Independent
1. RNA forms a hairpin loop (GC-rich inverted repeats base-pair → stem loop)
RNA polymerase pauses (hairpin physically disrupts/stalls it)
Weak A-U bonds break (U-rich RNA paired with A’s in DNA = weak)
RNA transcript is released -→ transcription ends
polycistronic mRNA
molecules of mRNA that code for multiple proteins
isoaccepting tRNA
different tRNAs that accept the same amino acid but have different anticodons
third-base wobble
A relaxation of the strict complementary base-pairing rules at the third base of the codon
how does third-base wobble work
1. Most synonymous codons can be grouped into pairs that differ only in the third base; the pairs either both carry a purine (A or G) or both carry a pyrimidine (C or U)
2. Third-base wobble occurs through flexible pairing at the 3' most nucleotide of the codon and the 5' most nucleotide of the anticodon
3. A pyrimidine must still base-pair with a purine
Modification of Amino acids
Methylation, Glycosylation, Phosphorylation, Hydroliztion, Modififaction of Fatty Acid
What are signal sequences?
It's a short stretch of amino acids that is at the beginning of a protein that acts like an "address label" (think of it as a protein GPS). It's found at the N-terminus. It's often removed after the protein enters the ER.
Why are signal sequences important?
Sends proteins to secretion (outside the cell), cell membrane, and organelles (ER, Golgi, lysosome)
What is signal hypothesis?
It explains how proteins get directed to the rough ER during translation. (tells the ribosome to go to the ER)
Step-by-step Signal Hypothesis
1. Translation begins in the cytosol
- The ribosome starts making a protein
- The signal sequence (N-terminus) comes out first
2. SRP (signal recognition particle) binds the signal sequence
- Translation pauses
- Why? Prevents the protein from being made in the wrong place
3. The ribosome goes to the rough ER
- SRP brings ribosome to the SRP receptor on the rough ER
- Ribosome attached to a translocon (channel)
4. Translation resumes
- SRP leaves
- Protein is threaded into the ER as it's made
5. Signal sequence is removed
- An enzyme cuts it off
What does primase do and why?
- It makes a shorter RNA primer and adds a small RNA sequence at the start
- It does this because DNA polymerase cannot start on its own and it needs a 3' OH starting point
What does helicase do and why?
- It unzips the DNA double helix
- breaks hydrogen bonds between base pairs
- creates the replication fork
- It unzips the DNA because it has to be separated to be coopied
What does DNA polymerase do and why?
- It builds a new DNA strand
- Adds nucleotides 5' --> 3'
- Uses existing strand as a template
- This happens because it copies the DNA
Silent Mutation (synonymous)
Missense Mutation (conservative)
Adds the wrong amino acid
Nonsense Mutation (Point Mutation/Premature Stop)
Early stop codon
Frameshift Mutation
mutation that shifts the "reading" frame of the genetic message by inserting or deleting a nucleotide
What is a heterochromatin?
Its a tightly packed DNA in the nucleus. It's not easily accessible and is usually not actively transcribed. This controls gene expression, helps maintain cell identity, and proteins DNA
What are histones?
Are proteins that DNA wraps around to help package it inside the nucleus. They're like spools, and the DNA is the thread. They also help organize DNA and control gene expression.
Histone Modifications
What is a transversion?
A transversion is a type of point mutation where a purine (A, G) is replaced with a pyrimidine (C, T) or vice versa.
What is a transition?
Going from a purine to a purine (A <-> G) or pyrimidine to pyrimidine (C <-> T)
Template strand
3' --> 5'
New RNA strand
5' --> 3'
Deamination
the removal of an amino group from an organism, particularly from an amino acid
What happens: Changes one base into another, like Cytosine to Uracil.
Leads to point mutations
How it's repaired: BER
Type of Mutation: Spontaenous
What is Nucleotide excision repair (NER) or UV-damage?
It's often used to repair bulky UV-induced damage to DNA. An example is Thymine Dimers from UV light)
What are the steps of NER?
1. Damage recognition: UvrA + UvrB scan DNA and detect a distortion. The binding near the damage-causing UrvA to leave and UrvB to stay. Why? This happens because the cell needs to find the damaged spot.
2. Excision (cutting): UrvC joins UvrB and cuts out a short segment on both sides of the DNA around the damage. This forms the UVR BC complex; then UrvC cleaves the damaged DNA strand about 4 to 5 nucleotides to the 3' and 5' sides of the photoproduct. Why? To remove the entire damaged section.
3. DNA synthesis: DNA polymerase fills in the gap with correct nucleotides. Why? To restore the correct sequence
4. Ligation: DNA ligase seals the sugar-phosphate backbone. Why? To make DNA continuous again.
What are transposable genetic elements?
They are DNA sequences that can move within the genome by an enzyme-driven process, transposition.
- They vary in length, sequence composition, and copy number
- Movement occurs in 2 ways:
1. Excision of the element from its original location and insertion in a new location.
2. Duplication of the element and insertion of the copy in a new location.
Characteristics and Classification of Transposable Elements
1. The transposable element contains terminal inverted repeats on its ends
2. The inserted transposable element is bracketed by flanking direct repeats
What is depurination?
Loss of a purine base (A or G) from DNA
What happens during depurination?
The bone between the base and sugar breaks and leaves an AP site (abasic site) so no base is there
Depurination during G1 and why its important
G1 = before DNA replication so if this occurs it creates a missing base and if its not repaired then it leads to a mutation during replication (S phase)
- It matters because DNA polymerase will not know what base to add so it will insert the wrong nucleotide, leading to point mutations.
How is Depurination repaired?
With Base Exicison Repair (BER)
p53 repair pathway (ATM-mediated)
DNA damage (double-strand break) activates ATM kinase -> ATM phosphorylates and stabilizes p53 -> p53 acts as a transcription factor -> activates p21 -> p21 inhibits CDKs -> cell cycle arrest at G1 checkpoint
Why? To allow time for DNA repair
Result: If repaired -> cell cycle continues
If not repaired -> p53 triggers apoptosis
Whats apoptosis?
Its a programmed cell death; an organized and controlled way for a cell to kill itself. What happens is the cell shrinks, DNA is broken down, the cell breaks into small vesicles, and other cells clean it up. No inflammation occurs.
What's necrosis?
Retrotransposons
Transposable elements that move within a genome by means of an RNA intermediate, a transcript of the retrotransposon DNA.
Mismatch Repair
Fixes base-pair mismatches after replication. It recognizes methylated (old) vs unmethylated (new) strand.
Steps:
MutS detects -> MutL connects -> MutH cuts new strand -> exonuclease removes -> polymerase fills -> ligase seals
Why? To correct replication errors
Transposable Elements
DNA sequences that move within the genome
2 ways: Cut and Paste or Copy and Paste
Why: Can create mutations and genetic variation
Key features of Transposable Elements
Terminal inverted repeats (ends)
Flanking direct repeats (after insertion)
Why: Needed for movement and insertion
Transposition Process
1. Transposase makes staggeree cuts
2. Elements inserts
3. DNA polymerase fills and creates direct repeats
Mutagenic Effect of Transposition
Insertion can disrupt genets and cause nonfucntional protein
Retrotransposons
Move through an RNA intermediate
RNA -> DNA (reverse transcriptase ) -> inserted
Why: Allows copying of element
UV radiation
Physical mutagen --> Causes thymine dimers
Effect: Distorts DNA -> blocks replication
Repair: NER
Mutagens
Agents that cause DNA mutations
Types :
Physical (UV)
Chemical
Biological
What is Base Excision Repair (BER)?
A repair mechanism that removes and replaces damaged bases in DNA.
1. Damage recognition:
DNA glycosylase recognizes and removes the damaged base, creating an AP site (abasic site) Why? To remove the incorrect or damaged base
2. Backbone cleavage:
AP endonuclease cuts the DNA backbone at the AP site Why? To open the DNA so repair can occur
3. DNA synthesis:
DNA polymerase adds the correct nucleotide Why? To restore the correct DNA sequence
4. Ligation:
DNA ligase seals the sugar-phosphate backbone Why? To make the DNA continuous again
DNA polymerase adds the wrong nucleotide during DNA replication
During DNA replication, DNA polymerase may add the wrong nucleotide, creating a mismatch (incorrect base pairing)
What happens: Incorrect base is inserted, creating a mismatched base pair A paired with C)
Why its important: It leads to mutation after replication
Proofreading by DNA polymerase
DNA intercalating agents
mutagenic compounds of a size and shape that allow them to access the space between nucleotide base pairs, thereby distorting the DNA duplex and potentially causing insertion or deletion mutations
Radiation-Induced DNA Damage
- Photoproducts are aberrant structures with additional bonds involving nucleotides; caused by UV irradiation
- One common photoproduct is a thymine dimer, formed by covalent bonds between the 5 and 6 carbons of adjacent thymines
- A second is called a 6-4 photoproduct, formed by a covalent bond between the 6 carbon on one thymine and the 4 carbon on the other
Consequences of Photoproducts
- DNA repair systems of most organisms can identify and correct most pyrimidine dimers
- Those that are not repaired cause disruption of replication
- Such disruptions leads to mutations; these are the primary cause of the strong association between excessive UV exposure and skin cancer
Base Analog (5-Bromouracil, BU)
Acts like thymine but can change forms
Effect: causes transition mutations (A-T -> G-C)
Alkylating Agents (EMS)
Add chemical groups to bases
Effect: Alters base pairing -> transition mutations
Intercalating Agents
Insert between DNA bases
Effect: Causes insertions/deletions -> frameshift mutations
Photoreactivation Repair
Photolyase uses light energy to break thymine dimers
-Not in humans
Whats the other type of bacterial transcription Termination?
Rho-Dependent Termination which requires a protein called Rho (p factor) to stop transcription.
Steps:
Rho protein binds to RNA; Binds at a specific site called the rut site
Rho moves along the RNA; Uses ATP to travel 5’ → 3’ toward RNA polymerase
RNA polymerase pauses; Happens at a termination sequence
Rho catches up and separates RNA-DNA; Rho unwinds the RNA from the DNA
RNA is released → transcription ends