Topic 4 – Small Sub-Unit analysis and Metagenomics
1/29
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
30 Terms
What is the role of SSU rRNA molecules in taxonomy?
SSU rRNA molecules help revolutionize taxonomy by providing a universal means to classify and understand evolutionary relationships among organisms.
Define metagenomics.
Metagenomics is the study of genetic material recovered directly from environmental samples, allowing for the analysis of complex microbial communities.
What is taxonomy?
Taxonomy is the classification of living forms, focusing on the commonalities and differences between different groups.
What does the term 'taxa' refer to?
Taxa refers to the categories that show degrees of similarity among organisms.
What is phylogeny?
Phylogeny is the study of the evolutionary history of organisms.
What are SSU rRNA molecules?
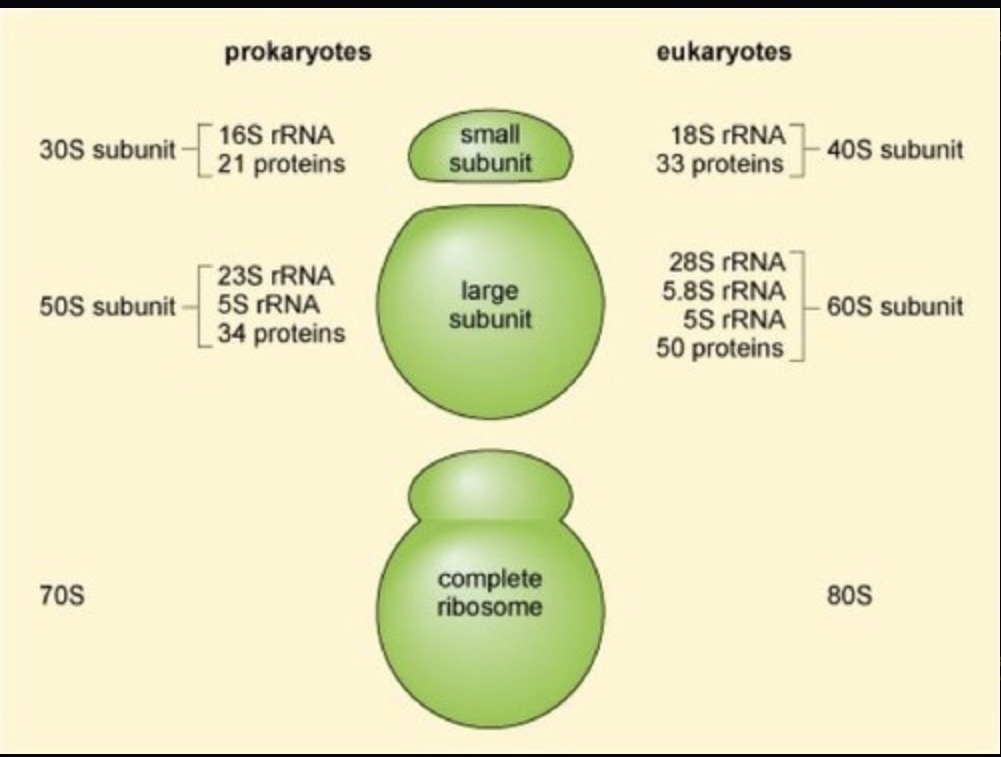
SSU rRNA molecules are components of the ribosome found universally in all domains of life, specifically 16S rRNA in prokaryotes and 18S rRNA in eukaryotes.
What are conserved regions in the 16S rRNA gene?
Conserved regions are sequences of DNA that are nearly identical across most bacterial species, essential for ribosome structure, and resistant to mutation.
What are variable regions in the 16S rRNA gene?
Variable regions are specific sections (V1-V9) of DNA where mutations can accumulate, allowing for species-specific identification.
How is the DNA sequence of the 16S rRNA gene amplified?
The DNA sequence is amplified using Polymerase Chain Reaction (PCR) with universal primers that bind to conserved regions of the gene.
What types of characteristics are used to identify bacteria?
Bacteria are identified based on cell wall composition, morphology, differential staining, oxygen requirements, and biochemical tests.
What is the last universal common ancestor (LUCA)?
LUCA is the origin point of the phylogenetic tree of life, representing the most recent common ancestor of all current life on Earth.
Why are SSU rRNA genes considered excellent candidates for molecular phylogeny?
Universally distributed
Functionally constant
Highly conserved
Of adequate length for analysis.
What is the purpose of using universal primers in PCR?
Universal primers are used to bind to conserved regions of the 16S rRNA gene, facilitating the amplification of the variable regions.
What is the function of ribosomes?
Ribosomes are complexes involved in translation, composed of a large subunit (LSU) and a small subunit (SSU) that contain proteins and structural rRNA.
What is the impact of mutations in conserved regions of the 16S rRNA gene?
Mutations in conserved regions can disrupt ribosome function, leading to cell death, as these regions are essential for ribosome structure.
What is 16S rRNA sequencing?
A method for amplifying variable regions of the 16S rRNA gene to identify bacteria.
Why is DNA preferred over RNA for sequencing?
DNA is more stable and less prone to degradation than RNA.
What does the 16S rRNA gene become after transcription?
It becomes part of the ribosome and is not translated into a protein.
What is the Great Plate Count Anomaly?
The discrepancy between the number of bacteria that can form colonies on agar and those countable by microscopy.
How can 16S rRNA gene sequencing help address the Great Plate Count Anomaly?
It allows identification of all bacteria present in a sample, even those that cannot be cultured.
What are the steps to identify bacteria using 16S rRNA gene sequencing?
Extract DNA from the sample.
Amplify the 16S rRNA gene using PCR.
Sequence the PCR products.
Compare sequences to databases.
What is one limitation of 16S rRNA gene sequencing?
It only detects bacteria and archaea, not fungi, viruses, or other microbes.
What is a major advantage of 16S rRNA gene sequencing?
It is the cheapest method for analyzing a large number of samples.
What does metagenomics analyze?
All genetic material in a sample, including bacteria, archaea, fungi, and viruses.
What technology is used in metagenomics?
Whole Genome Sequencing (WGS).
What is the output of Whole Genome Sequencing?
A taxonomic profile of all microbes and functional information about genes.
What is one limitation of metagenomics?
It requires more computational resources and is more expensive than 16S rRNA sequencing.
What question does 16S rRNA sequencing answer?
'Who is there?' regarding microbial taxonomy.
What question does shotgun metagenomics answer?
'What can they do?' regarding microbial function.
What is the Resistome?
The collection of all antimicrobial resistance genes in a specific environment.