Applied Algebra C957 WGU Module 2 Review questions and answers 2026
1/112
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
113 Terms
Enzymes
Proteins that allow biochemical processes to occur
Function of Enzymes
1. Increase reaction rates of biochemical reactions
2. Not consumed in the reaction
3. Help processes that would otherwise occur slowly
Advantages of Enzymes
1. Milder reaction conditions
2. Higher reaction rates within an organisms lifetime
3. Regulation (on/off when needed or not)
4. No side products
Transition State
Transient higher energy configuration that can decay to substrate or product
Cofactors
"helper molecules" Non protein chemical compounds or metallic ions required for some enzymes to become functional
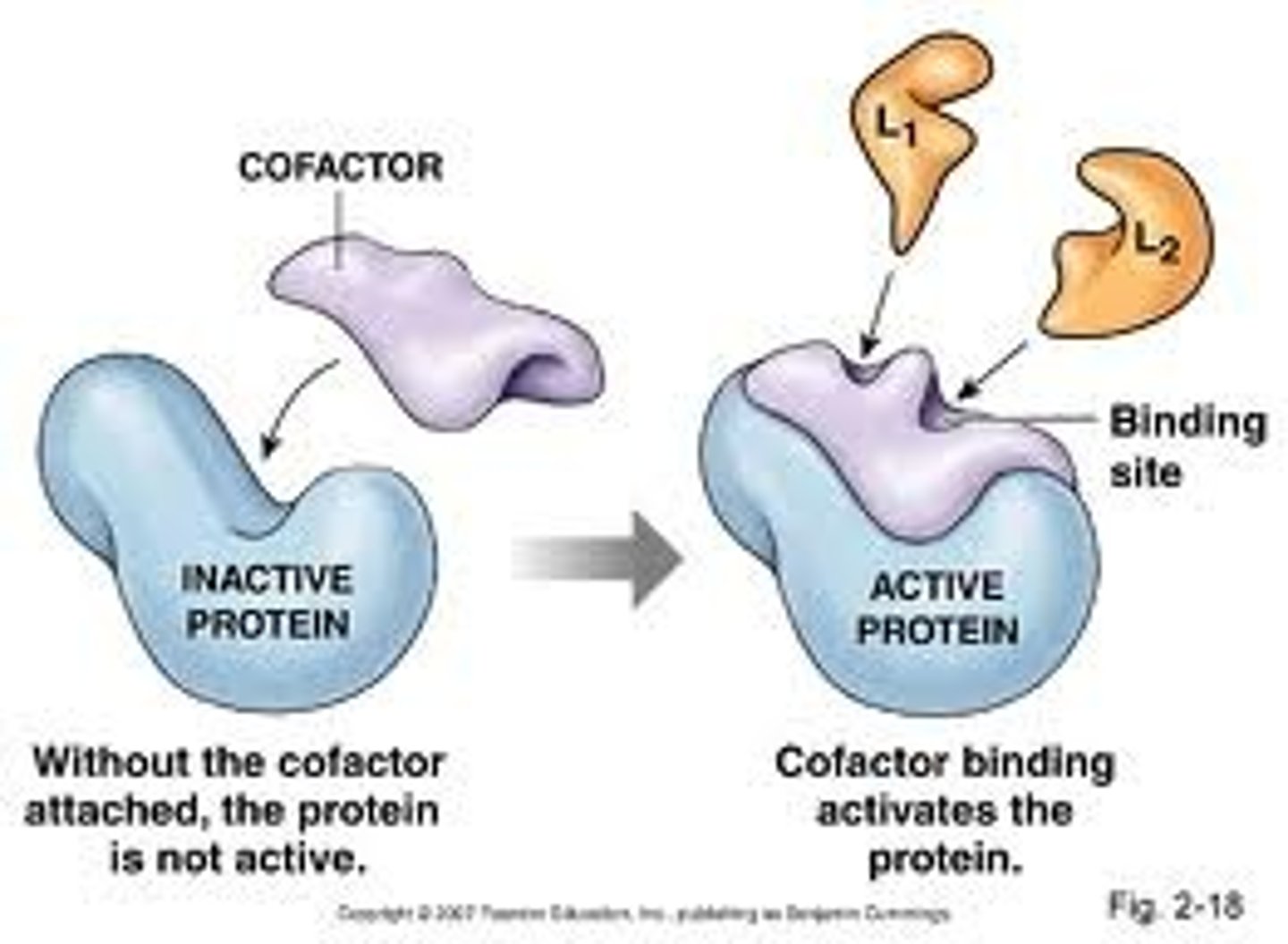
Inorganic Ions
Cofactor- metals that binds electrostatic-ally or participate in redox reactions
Coenzymes
Cofactor- organic molecules that "move" chemical substituents
Prosthetic groups
Cofactor- coenzyme that are permanently attached to an enzyme
HoloEnzyme
ApoEnzyme (protein without cofactor) + Cofactor = HoloEnzyme
Equilibrium Constant
K'eq= [P] / [S]
Free Energy Change
If G is negative, the reaction proceeds spontaneously, without any energy input (by itself)
If G is positive, the reaction proceeds not spontaneously, with energy input.
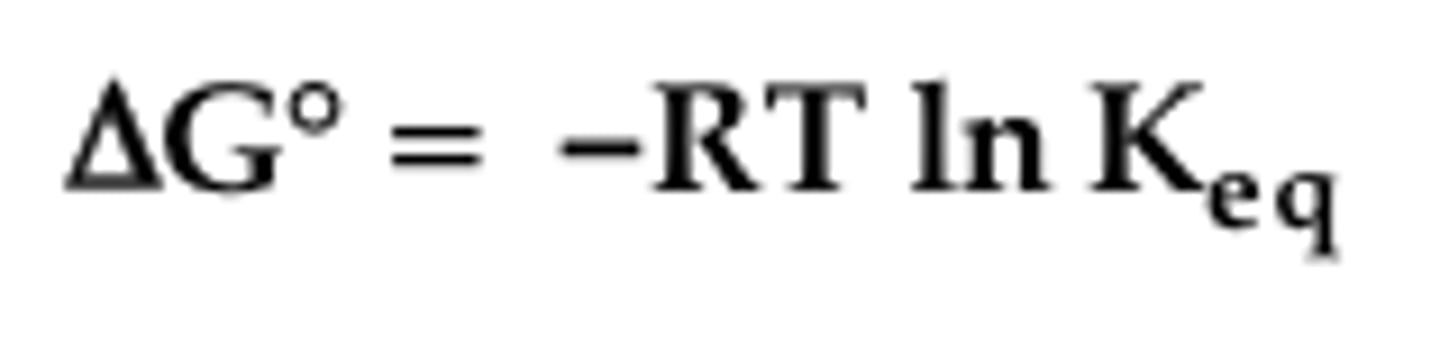
Desolvation
Displacement of water molecules that would otherwise impeed the reaction.
What affects the activity of Enzymes?
1. Temperature
2. pH
3. Ionic Strength
4. Presence of cofactors and prosthethic groups
Acid base catalysis
Give and take protons
Covalent Catalysis
Covalent bond forming between the enzyme and substrate
Changes reaction pathway
Metal Ion Catalysis
Use redox cofactors, pKa shifters
Electrostatic catalysis
Preferential interactions with Ts
Kinetics
Study of the rate at which compounds react
Rate
Amount of product accumulated or substrate consumed per unit of time
Rate is affected by
-Temperature
-pH
-Ionic strength
-Substrate and enzyme concentration
Inhibitors
Compounds that decrease enzymes activity
Competitive inhibitors
Mimics substrate and competes for active site.
Do not affect catalytic function
Vmax unchanged and Km increased
Noncompetitive Inhibitor
Alters conformation of enzyme so active site is no longer fully functional; Inhibitor binds to another site on the enzyme.
Inhibits catalytic function
Vmax decreased and Km unchanged
Uncompetitive Inhibitor
Substrate can normally bind to active site
Increase substate binding; do not affect catalysis
Vmax and Km decreased
Mixed inhibitors (noncompetitive)
Bind to enzyme without bound substrate
Inhibit catalytic function and substrate binding
Carbohydrates
Sugars that come from Photosynthesis
Energy source and storage
Structural component of cell walls in plants and bacteria
Covalently linked with proteins
Monosaccharides
-Class of carbohydrate, simple sugars, one unit only
- Aldehydes or ketones with two or more hydroxyl groups
Hexose
Most common monosaccharide in nature
Enantiomers
Stereoisomers that are non-superimposable images of each other
Diastereomers
Sterioisomers that are NOT mirror images of each other
- Different physical properties
Epimers
Type of diastereomers
- Monosaccharides that differ only in the configuration around 1 carbon atom
Pyranoses
6 membered oxygen-containing rings
Furanoses
5 membered oxygen containing rings
Mutarotation
Formation of alpha or beta sugars
Disaccharides
Two monosaccharides joined together by an O-glycosidic bond
-Simplest Oligosaccharide
Glycosidic bond
Anomeric carbon of one monosaccharide reacting with carbon carrying hydroxyl group from the second monosaccharide.
Polysaccharides
Natural carbohydrates found as polymers
Polysaccharides can be
-homopolysaccharides: same units
-heteropolysaccharides: different alternating units
-linear
-branched
-storage and structural function
Starch
Main storage polysaccharide in plants
-mixture of two homopolysaccharides of glucose
Glycogen
Main storage polysaccharide in animals
-branched homopolysaccharide of glucose
Dextrans
Bacterial and yeast polysaccharides
-Branched homopolysaccharides of glucose
Cellulose
Fibrous, tough, water insoluble substance
- found in plant cell walls
Linear homopolymer of glucose
Cellulase
an enzyme which allows fungi, bacteria, and protozoa to use wood as a source of glucose.
Chitin
Principal component of the hard exoseleton of arthopods and cell walls of mushrooms
Agar
Complex mixture of heteropolysaccharide containing modified galactose. Used for growing bacteria
Agarose
Component of agar, gel separation
Glycosaminoglycans
Heteropolysaccharides of extracellular matrix in animals
- linear polymers of repeating diasaccharide units
Heparin
Linear polymer
Glycoproteins
Proteins with small oligosaccharides attachd
Glycolipids
Lipids with covalently attached oligosaccharide
Proteoglycans
Sulfated glucosaminglycans attached to a large rod-shaped proteins in cell membrane
Nucleotides
Building blocks of DNA
- Energy for metabolism (ATP)
-Enzyme cofactors (NAD+)
-Signal transduction (cAMP)
Nucleic Acids Functions
-Storage off genetic info
- Transmission of genetic information (mRNA)
-Processing of genetic information (ribozymes)
Nucleotide
-Nitrogenous base (Pyrimidines) (C, U or T)
- Pentose sugar (Ribose or Deoxyribose)
- Phosphate (Mono-, Di-, and Tri-phosphate)
Nucleoside
-Nitrogenous base (Pyrimidines) (C, U or T)
- Pentose sugar (Ribose or Deoxyribose)
Nucleobase
-Nitrogenous base (Pyrimidines) (C, U or T)
Cytosine
Pyrimdine: Both in DNA and RNA
Thymine
Pyrimdine: Only in DNA
Uracil
Pyrimdine: Found in RNA
Adenine and Guanine
Found in both RNA and DNA
Glycosidic bond
Pentose ring (sugar) is attached to the nucleobase
- Bond formed between anomeric carbon
Phosphodiester bond
Bonds link successive nucleotides in linear polymers
5' of one end --> 3' of the other sugar
DNA Backbone
Phosphates + Sugar = Backbone
Chargaff's Rule
A=T and C=G
Forms of DNA
- B form : predominant structure of DNA
- A form: dehydrated conditions for RNA
- Z form: GC rich
Central Dogma of Molecular Biology
DNA -> RNA -> Protein
- DNA and RNA are reversible
-Once DNA passed onto protein it CANT leave
mRNA
Carries genetic information from DNA to the ribosome where proteins are synthesized
tRNA
Adaptor molecules which bring amino acids to the ribosomes to carry out protein translation
rRNA (ribosomal RNA)
Main part of the ribosome which synthesize all proteins
Denaturation of DNA
-High temperature or change in pH
-Reversible: Annealing
Mutations
Alterations in DNA sequencing that produce permanent changes in the genetic information encoded
- related to aging and cancer
Lipids
Hydrophobic, organic molecules that are
- low solubility in water
-good solubility in nonpolar solvents
Functions of Lipids
- membrane structure in cellular membranes
- Storage of energy
-water repellant
-enzyme cofactors
-antioxidants
-pigments
-signalling molecules
Types of lipids
1. Non-fatty acid: cholesterol
2. Fatty acids:
- Storage lipids
- Membrane lipids
Fatty acids
Carboxylic acids with hydrocarbon chains containing between 4-36 carbons
- Saturated: no double bonds between carbons
- monounsaturated: one double bond between carbons
- polysaturated: more than one double bond
Triglycerols
Simplest lipids constructed from fatty acids
Advantage of fats over polysaccharides
- Fatty acids carry more energy
- Carry less water per gram
Trans fatty acids
Forms during parial hydrogenation of unsaturated fatty acids as an unwanted byproduct
Waxes
Esters of long chained saturated and unsaturated fatty acids with long chain alcohols
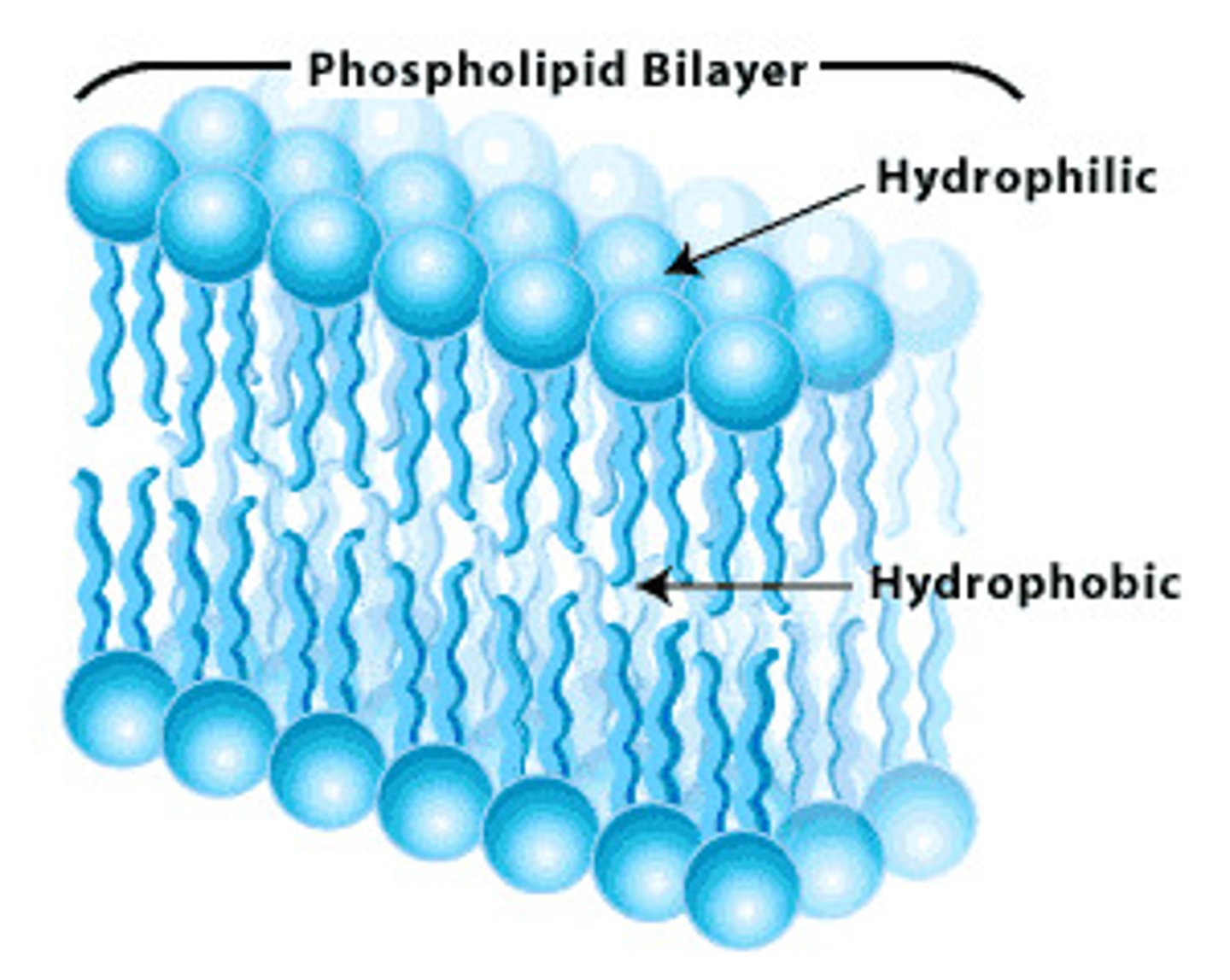
Structural Lipids in Membrane
Double layer of lipids that are amphipathic. One end is hydrophobic (inside) and one in hydrophilic.
-Polar head groups
- Nonpolar tails
Glycerophospholipids
Subtype of lipids
-Derivative of L-glycerol-3-phosphate
Sphingolipids
Subtype of lipids
-Backbone is made of amino alcohol sphingosine
Glycosphingolipids
Blood groups are determined by the type of sugars located on the head group of glycosphingolipids
Sterol: Cholesterol
Sterols: molecules that contain steroid nucleus
Cholesterol: Non fatty acid containing lipid
Functions of sterols
- modulate fluidity and permeability of membranes
-thicken the plasma membrane
-cholesterol bound to proteins, transported to tissues via blood vessels
- mammals obtain cholesterol from food or synthesize it in the liver
Steriod hormones
Steriods are oxidized derivatives of sterols
- More polar than cholesterol
Steriods are synthesized from cholesterol
Biological active lipids
play Active roles as signaling molecules between nearby cells
-Lipid soluble vitamins (A,D,E and K)
Biological Membranes
Cell membrane which separates interior parts of the cell from its surroundings
-composed of a variety of lipids and proteins
Functions of biomembranes
-Define cell boundaries
-Allow import and export of waste/nutrients
-retain metabolites and ions within the cell
-sense external signals and transmit information into the cell
-store energy as proton gradient
-support synthesis of ATP
Lipid Micelle
Form in the solution of amphipathic molecules that have larger head than tails
- Forms when the concentration of molecules is higher than a certain threshold
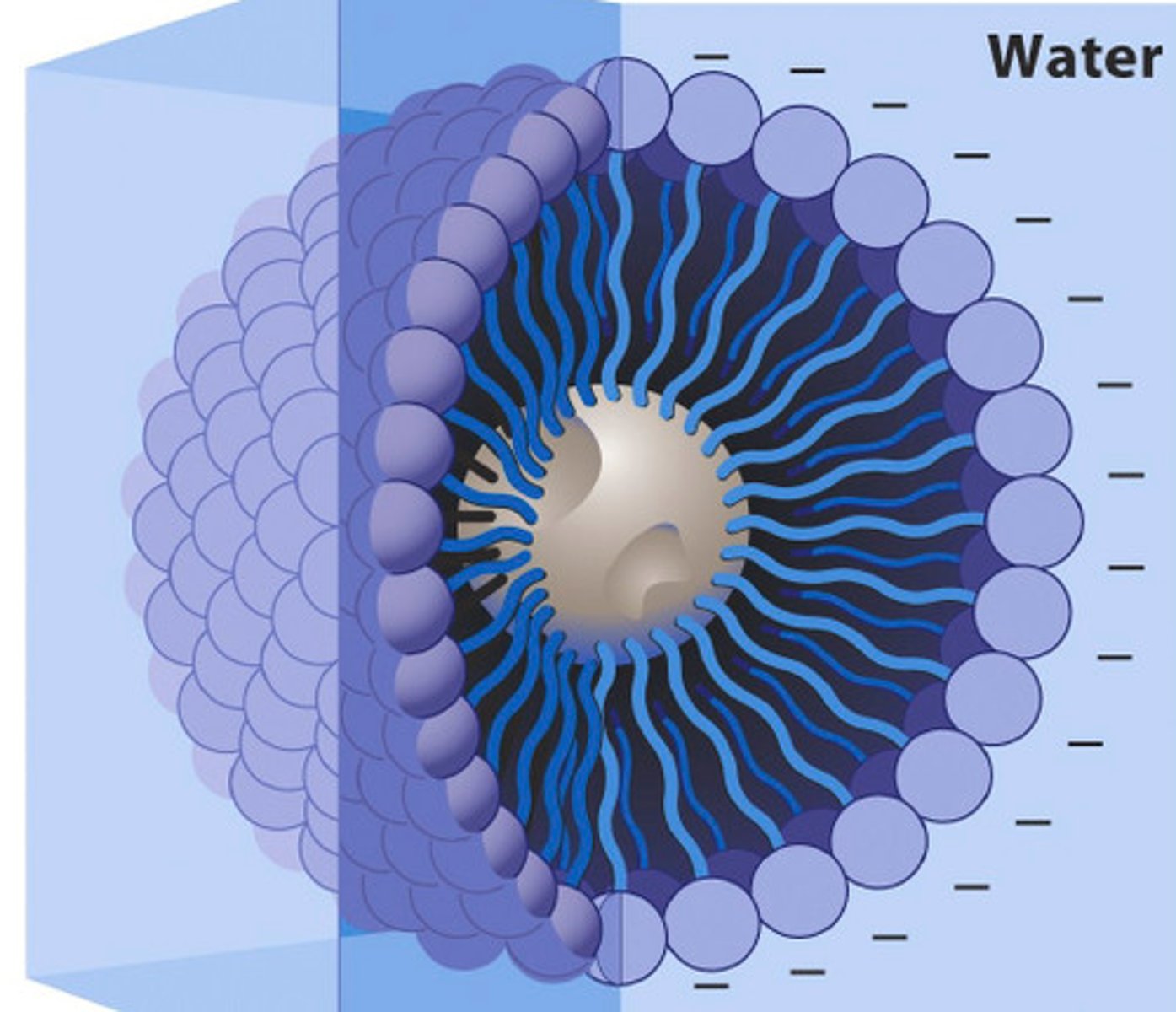
Lipid Bilayer
Forms in the solution of amphipathic molecules that have head and tail of equal cross section
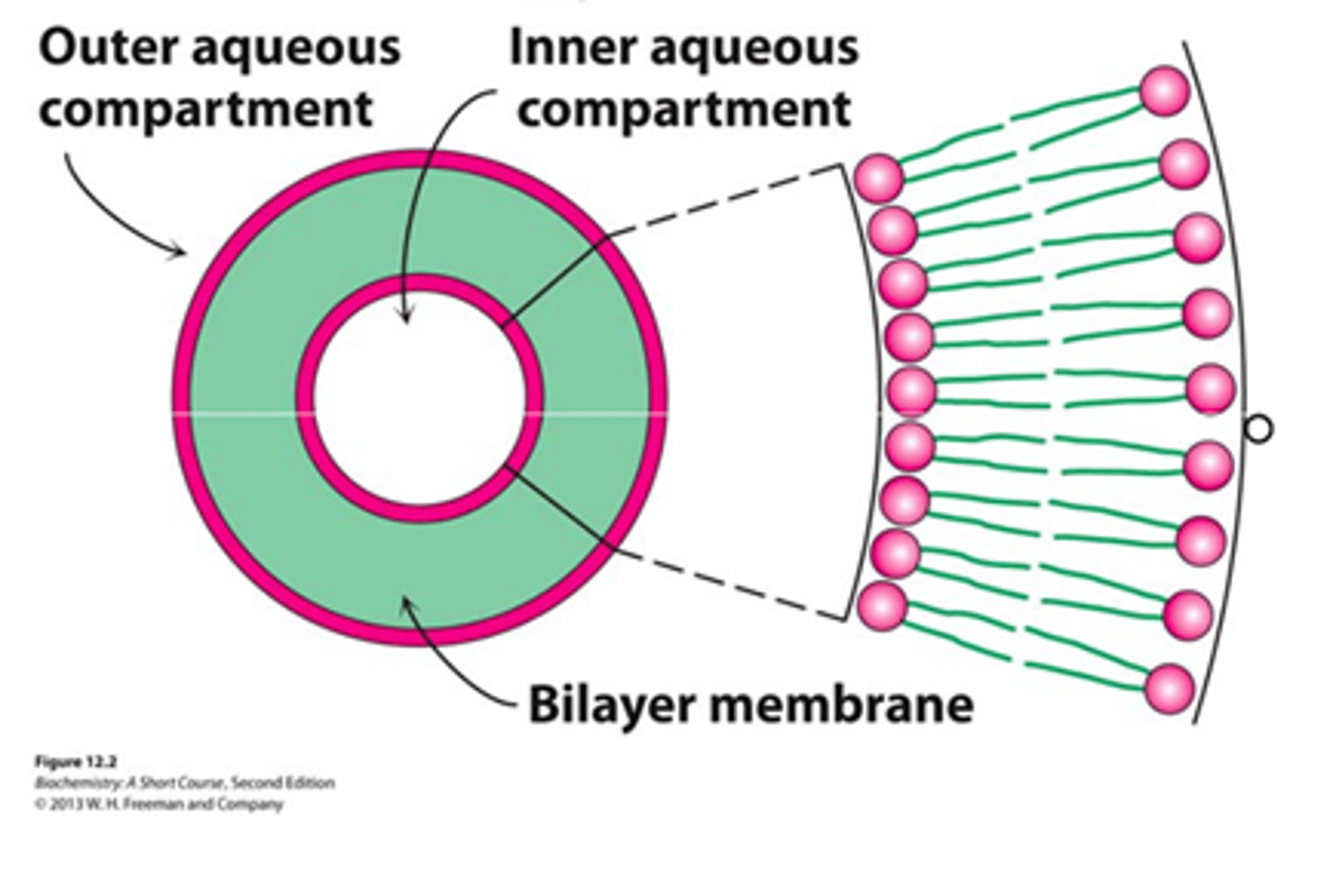
Lipid Vesicles - liposomes
Small bilayers spontaneously seal into spherical vesicles
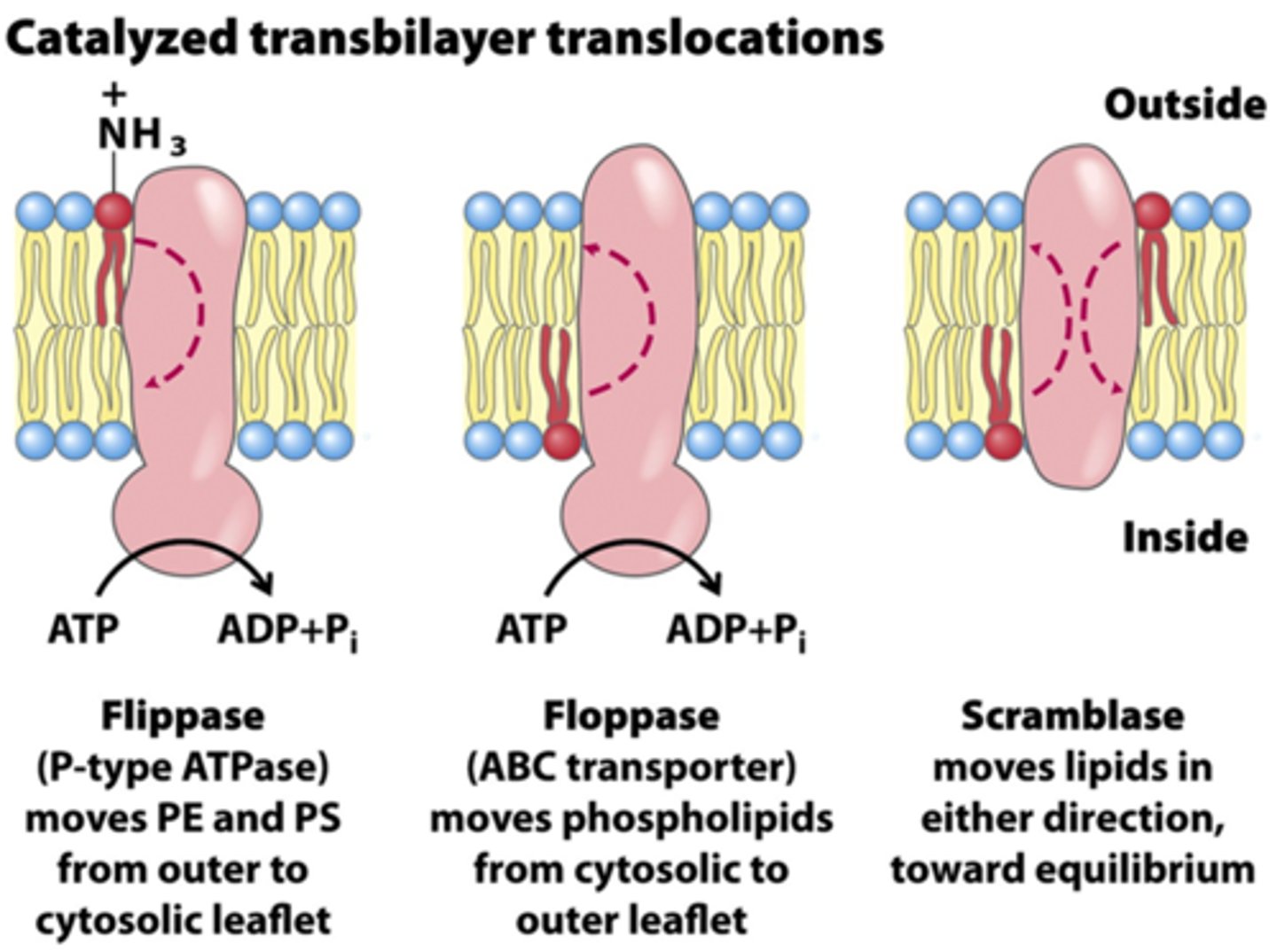
Flippases and Floppases
Enzymes that catalyze transverse diffusion of lipids from one leaflet to another
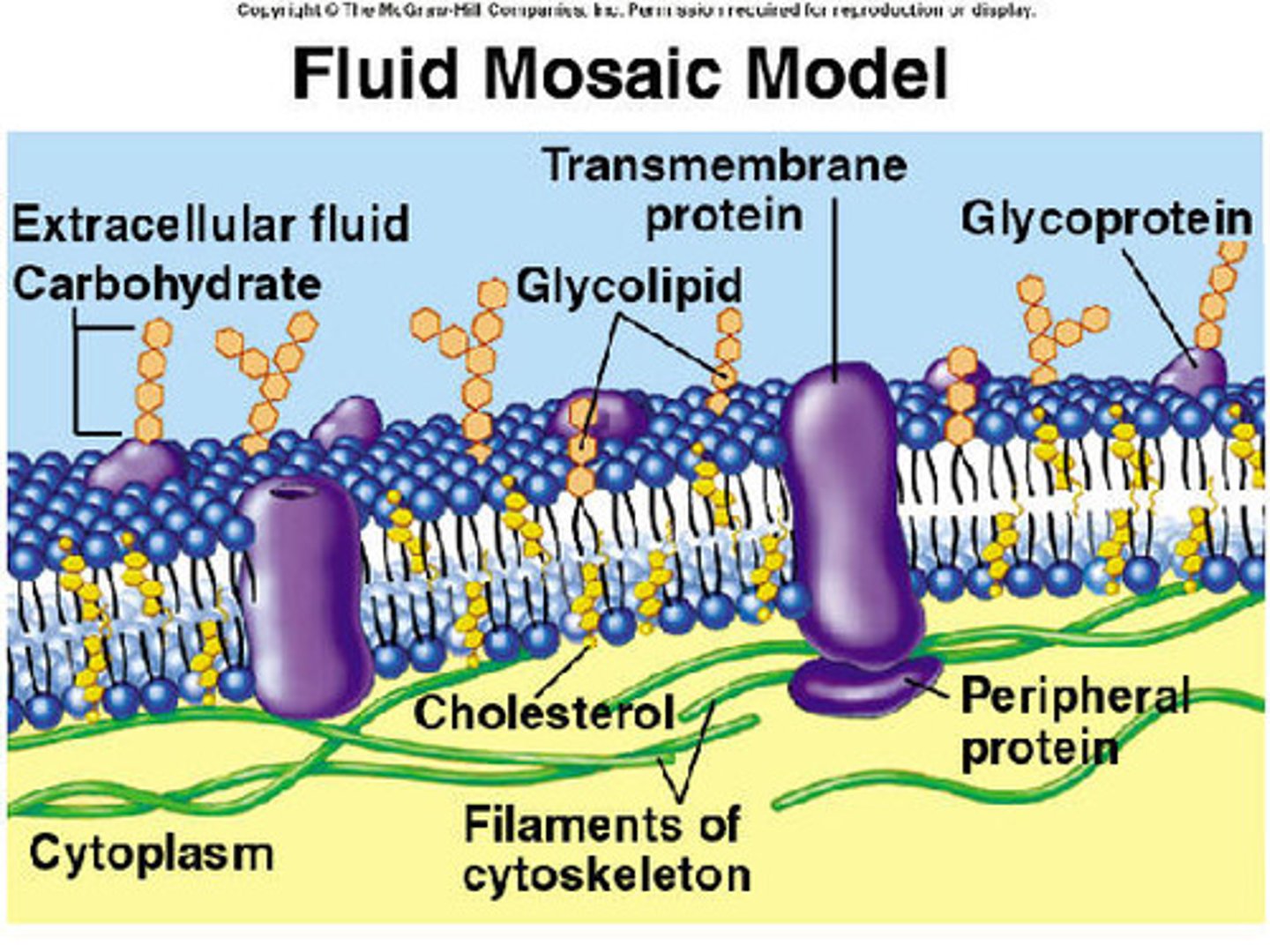
Fluid Mosaic Model
Structure of Biomembranes in Archaea
-Unique glycerol (D-glycerol-3-phosphate)
-Unique fatty acids (branched isoprene chains)
-Unique linages of fatty acids to glycerols (ether)
Functions of the proteins in the biomembrane
-Receptors: detecting signals from the outside (hormones)
- Channels, gates, pumps
-Enzymes (atp synthesis)
Three classes of membrane proteins
- Peripheral (Associate with the polar head group of membrane lipids)
-Integral (Span the entire membrane)
-Anchored (attached to membrane by fatty acid anchor)
Transporters (pumps)
-High specificity for particular substrates
- Behave like enzymes
-Gate at either membrane face
- move AGAINST a gradient
Ion Channels
-speficity for ion substrate
- single gate
-ions move DOWN a concentration gradient
Membrane transport systems