🧬 Lecture 2: Eukaryotic Gene Expression + RNA Polymerases + Gene Promoters
1/13
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
14 Terms
Chromatin and Transcription in Eukaryotes
Chromatin Structure
Eukaryotic DNA is wrapped in chromatin
Chromatin is DNA wrapped around proteinsRequirement for Transcription
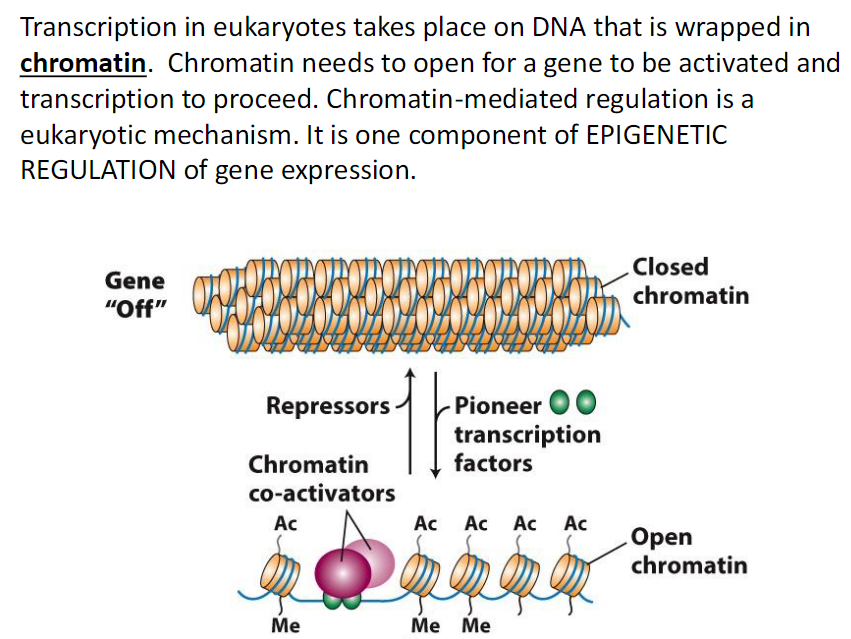
Chromatin must open for gene activation
Open chromatin allows transcription to proceedType of Regulation
Chromatin mediated regulation is eukaryotic specific
It is part of epigenetic regulation
Epigenetic regulation controls gene expression without changing DNA sequence
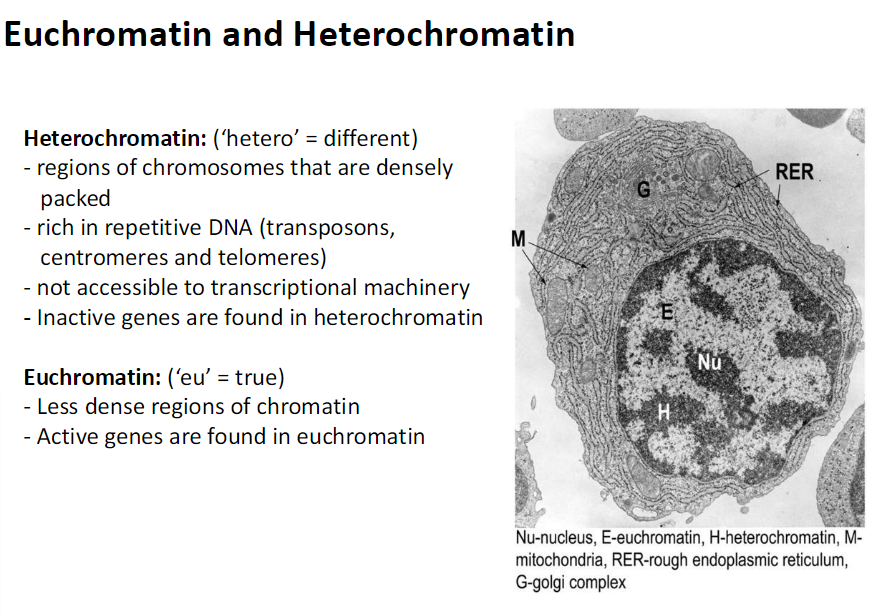
Euchromatin and Heterochromatin
Heterochromatin
Hetero means different
Regions of chromosomes that are densely packed
Rich in repetitive DNA
Includes transposons centromeres and telomeres
Not accessible to transcriptional machinery
Transcriptional machinery is proteins needed to transcribe genes
Inactive genes are found in heterochromatinEuchromatin
Eu means true
Less dense regions of chromatin
Accessible to transcriptional machinery
Active genes are found in euchromatin
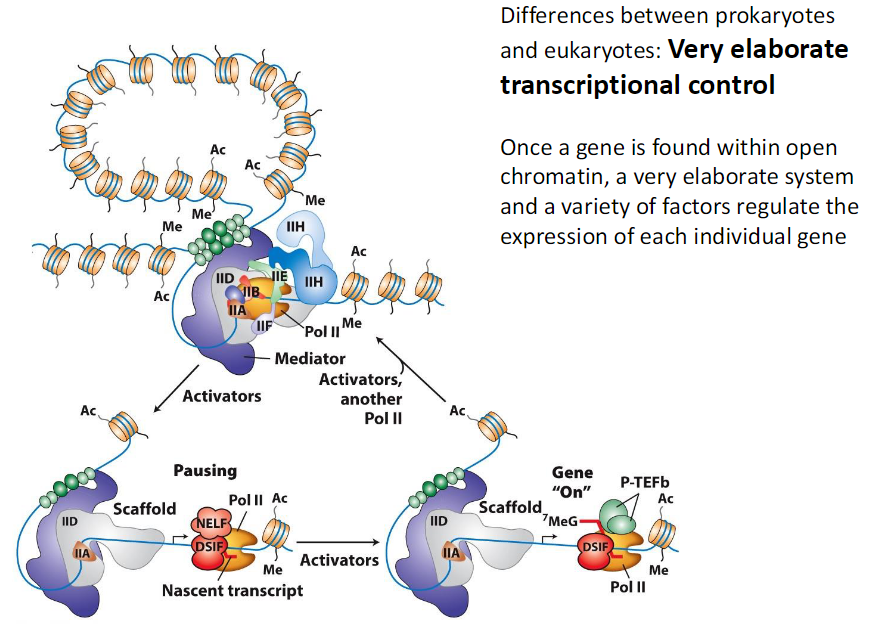
Differences Between Prokaryotes and Eukaryotes Transcription Control
Prokaryotes
Relatively simple transcriptional controlEukaryotes
Very elaborate transcriptional control
Gene must first be located in open chromatin
Open chromatin allows access to DNA
Once accessible a variety of factors regulate expression
Each individual gene is regulated separately
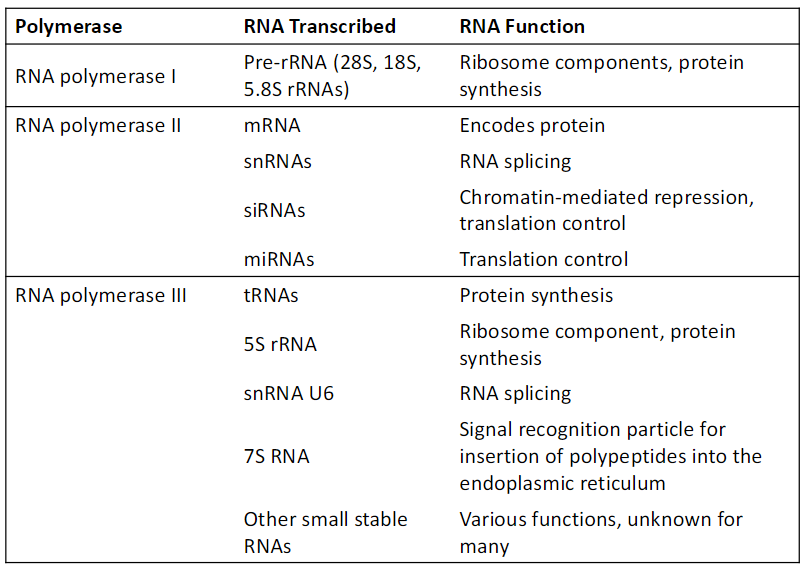
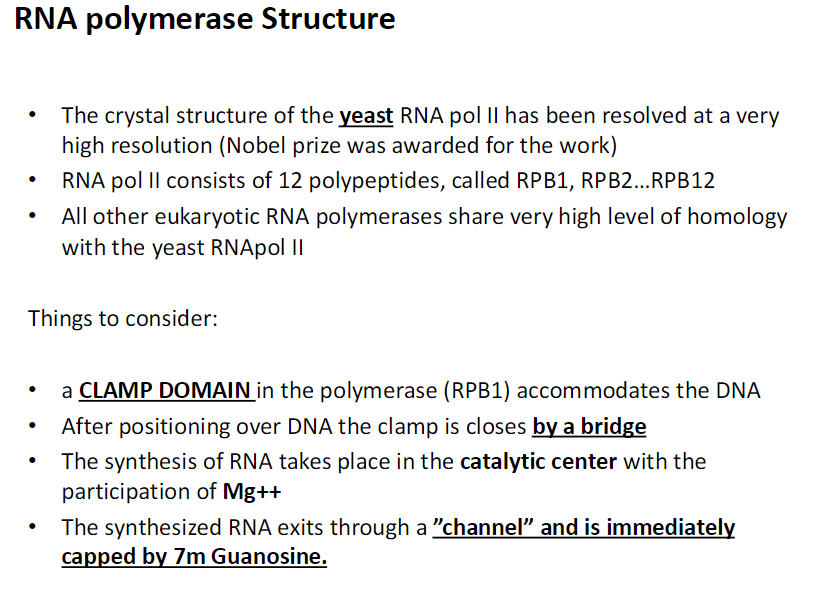
Eukaryotic RNA Polymerases
Don’t memorize chart
Know:
Pol I transcribes rRNA
Pol II transcribes mRNA
Pol III transcribes tRNA
RNA Polymerase Structure
Crystal Structure
The crystal structure of yeast RNA Polymerase II has been resolved at very high resolution
This work was awarded a Nobel prize
Composition
RNA Polymerase II consists of 12 polypeptides named RPB1, RPB2 … RPB12 (RP = RNA polymerase, B = 2)
All other eukaryotic RNA polymerases share a very high level of similarity with yeast RNA Pol II
Key Features
Clamp Domain
Located in RPB1
Accommodates DNA during transcription
After DNA is positioned the clamp closes via a bridge
Catalytic Center
RNA synthesis occurs here
Mg++ ions participate in the reaction
RNA Exit and Capping
Newly synthesized RNA exits through a channel
RNA is immediately capped with 7m Guanosine to protect it and prepare it for processing

Eukaryotic RNA Polymerases
Complex Composition
Eukaryotic polymerases are complexes of multiple polypeptides
Similarity to Prokaryotes
Prokaryotic β and β’ subunits are similar to eukaryotic RPB1 and RPB2
Structural Conservation
All these enzymes share strong structural similarity across species
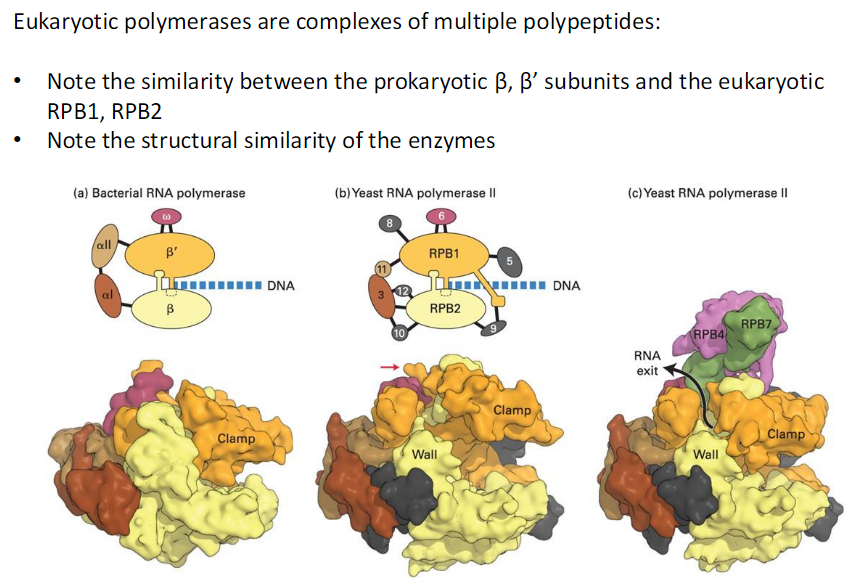
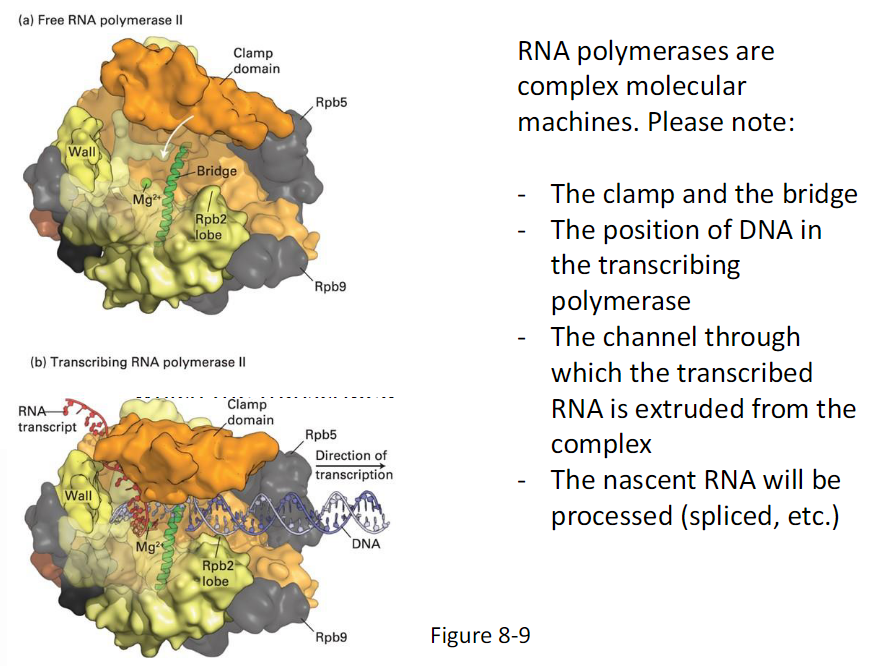
RNA Polymerase as a Molecular Machine
Key Features to Note
The clamp and the bridge
Position of DNA in the transcribing polymerase
Channel through which the transcribed RNA is extruded from the complex
Nascent RNA will be processed
Processing includes splicing and other modifications
Carboxy-Terminal Domain (CTD) of RNA Polymerase II
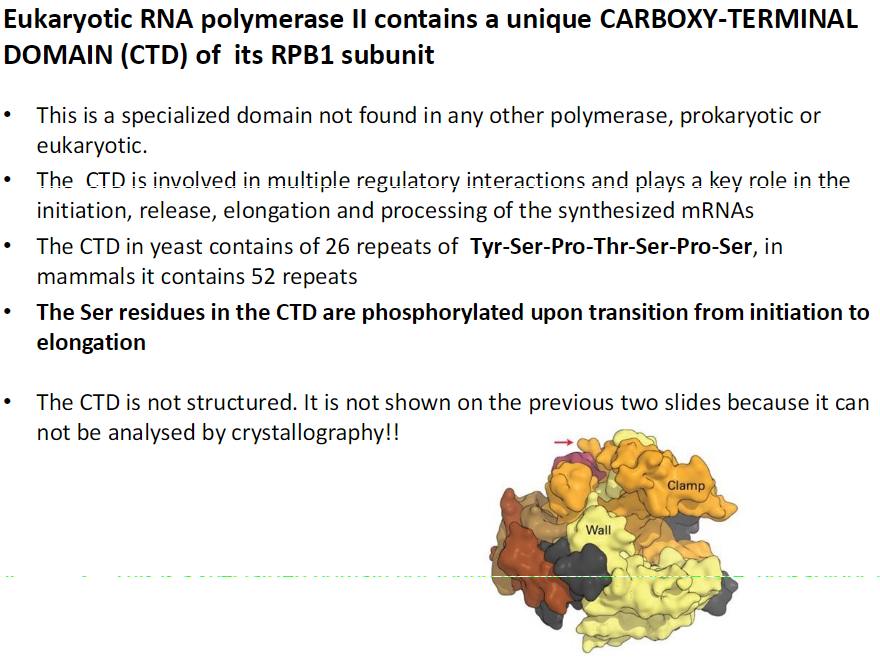
Unique Feature
CTD is a specialized domain of the RPB1 subunit (red arrow on image)
Not found in any other polymerase, prokaryotic or eukaryotic
Functions
Involved in multiple regulatory interactions
Plays a key role in initiation, release, elongation, and processing of synthesized mRNAs
Structure
In yeast, CTD contains 26 repeats of Tyr-Ser-Pro-Thr-Ser-Pro-Ser
In mammals, CTD contains 52 repeats
Ser residues are phosphorylated during transition from initiation to elongation
CTD is intrinsically unstructured
Not shown in crystallography-based slides because it cannot be analyzed by this method

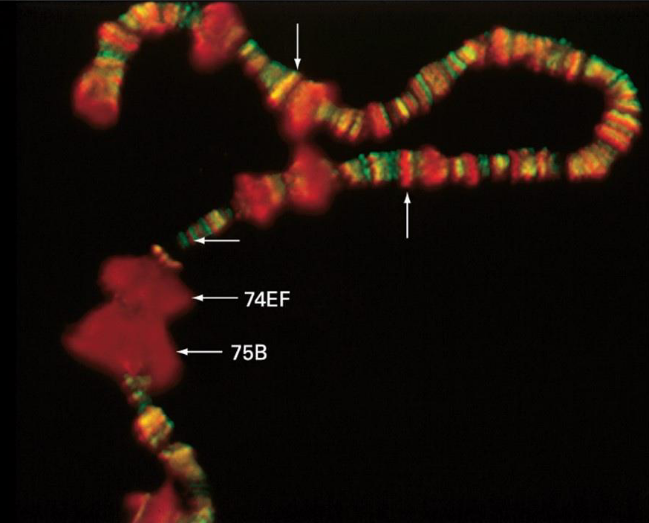
RNA Polymerase II in Drosophila DNA
Image Details
Drosophila salivary gland DNA
Red shows phosphorylated CTD of RNA Pol II
Green shows dephosphorylated Pol II
Key Points
Dephosphorylated Pol II (green) is at the transcription start site
Cannot start elongation yet
Actively transcribed genes can initiate transcription at multiple points
This produces multiple RNA copies from the same gene at the same time
Phosphorylated Pol II (red) is ready to start elongation
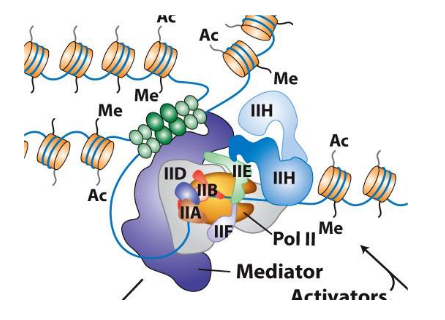
Regulation of Genes Transcribed by RNA Polymerase II (Descriptions from ChatGPT)
Basal Promoter Elements
Also called Core Promoter Sequences
Conserved sequences where the transcription machinery assembles
Promoter-Proximal Elements
Binding sites near the promoter for transcriptional activators
Help increase the efficiency of transcription initiation
Distal Regulatory Elements
Enhancers or repressors located far from the gene
Enhancers increase transcription
Repressors decrease transcription
Chromatin Structure
Organization of DNA and histone proteins affects gene accessibility
Open chromatin allows transcription
Closed chromatin inhibits transcription
RNA Polymerase II Promoters and General Transcription Factors
Promoters are recognized by general transcription factors
These factors recruit RNA Pol II and help initiate transcription
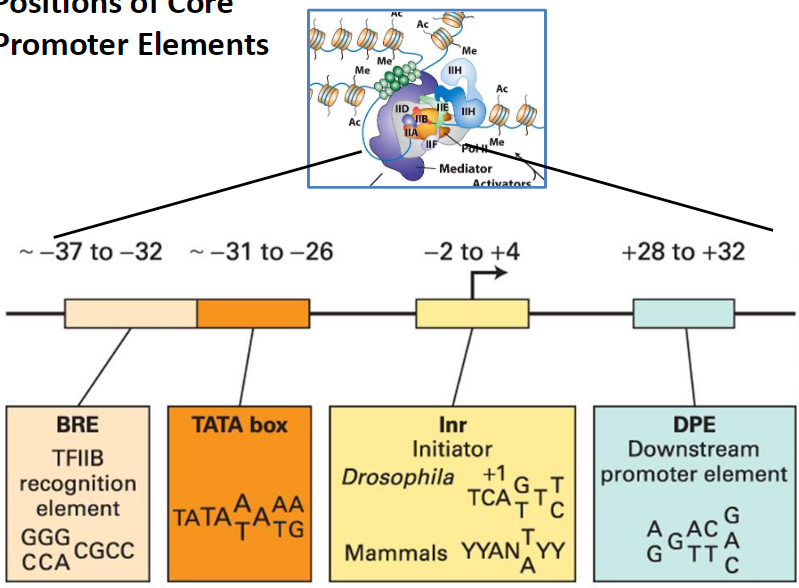
Positions of Core Promoter Elements
BRE
TFIIB recognition element
Located around -37 to -32
TATA Box
Located around -31 to -26
Inr (Initiator)
Spans -2 to +4
+1 is the transcription start site
Drosophila: TCAST
Mammals: YYANTYY
DPE (Downstream Promoter Element)
Located +28 to +32
Sequence example: GAGACAGTTC
Other Notes
Core promoter elements are binding sites for general transcription factors (Ac, Me, IIH, IID, etc.)
Mediator and activators interact with these elements to regulate transcription
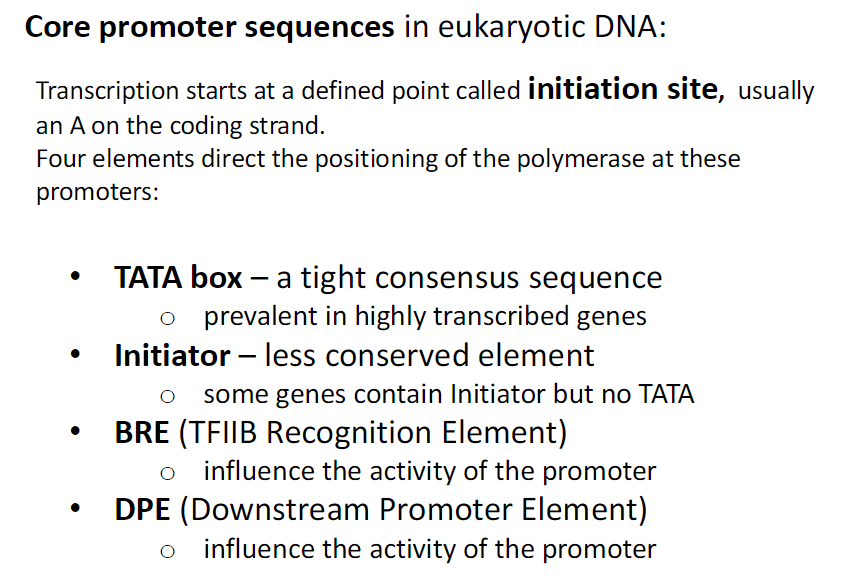
Core Promoter Sequences in Eukaryotic DNA
TATA Box
Tight consensus sequence
Common in highly transcribed genes
Initiator (Inr)
Less conserved element
Some genes have Initiator but no TATA
BRE (TFIIB Recognition Element)
Influences promoter activity
DPE (Downstream Promoter Element)
Influences promoter activity
Transcription Start Site
Defined point where transcription begins
Usually an A on the coding strand
Polymerase Positioning
Four elements (TATA, Inr, BRE, DPE) guide RNA polymerase to the promoter

RNA Polymerase II and Transcription Initiation
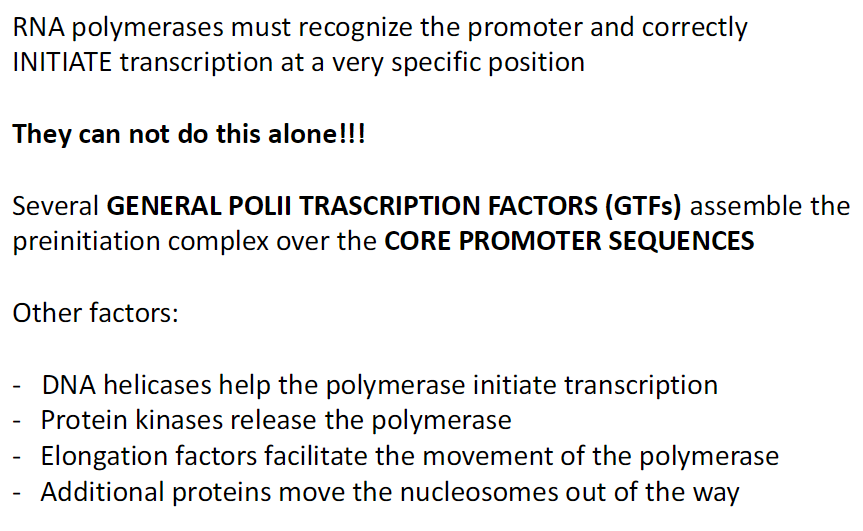
Promoter Recognition and Initiation
RNA polymerase must recognize the promoter
Initiate transcription at a very specific site
Cannot do this alone
General Transcription Factors (GTFs)
Assemble the preinitiation complex over core promoter sequences
Other Supporting Factors
DNA helicases – help unwind DNA so polymerase can start
Protein kinases – release polymerase to begin elongation
Elongation factors – help polymerase move along DNA
Chromatin remodelers – move nucleosomes out of the way

General Transcription Factors of RNA Polymerase II
RNA Polymerase I GTFs
Labeled as TFI
Examples: TFIA, TFIB
RNA Polymerase III GTFs
Labeled as TFIII
Examples: TFIIIB, TFIIIS
RNA Polymerase II GTFs
Labeled as TFII
Examples: TFIIA, TFIIB, TFIID, TFIIE, TFIIH
