2004: The Genetic Code and Protein Synthesis, Regulation of Gene expression in Bacteria
1/87
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
88 Terms
what are the possibilities for arranging/grouping three bases in an RNA chain?
Overlapping: 1 base overlap between 1 base in a nucleotide.
Punctuated: mRNA contains gaps
Unpunctuated: mRNA contains no gaps
What are the 3 characteristics of the genetic code?
Genetic experiments established that the code is a triplet (3 nucleotides), non-overlapping, unpunctuated code
Includes start code in DNA (ATG) which is complementary to AUG in mRNA, and stop codons
If you don’t have any information about where translation starts or stops, how many possible reading frames are possible for the ways DNA sequence can be decoded?
There are SIX reading frames in any DNA sequence.
3 on each strand.
If you keep going, sequences will repeat
Look for start sequence (ATG in DNA that codes for Met), long stretch, then stop sequence

What is the triplet code in mRNA? What does this mean for amino acids?
Groups of three consecutive nucleotides (= codons) that correspond to one amino acid.
The genetic code contains 64 different codons; with only 20 common amino acids, this leads to redundancy — some amino acids are specified by more than one codon
Deciphering the code: What was the first steps to discovering which triplet coded for which amino acid?
They pioneered the use of in vitro translation systems
They mixed:
RNA template
Ribosomes (and other necessary “stuff”)
Individual radio-labelled amino acids
Suitable reaction buffer (pH, [salt])
*Basically recreated protein synthesis in a test tube.
Proteins that are synthesized in the system can be precipitated with trichloroacetic acid (TCA) (usually) and analyzed.
Deciphering the code: What are the examples regarding how exactly did they use in vitro translation systems to establish which triplet coded for an amino acid in repeating RNAs?
When they used poly(U), which contains only uracils, a polypeptide containing only phenylalanine was synthesized. Ergo, the codon UUU corresponds to the amino acid Phe
They also found that poly-C directed synthesis of polyproline CCC = Proline poly-A had apparently been tried earlier by Watson and Ochoa in a similar type of experiment without success.
We now know that poly-A would direct synthesis of polylysine; however, polylysine (unlike most proteins) is soluble in trichloroacetic acid
Deciphering the code: How did Khorana Extended the Analysis of the Genetic Code beyond repeating RNAs? What was an issue with this?
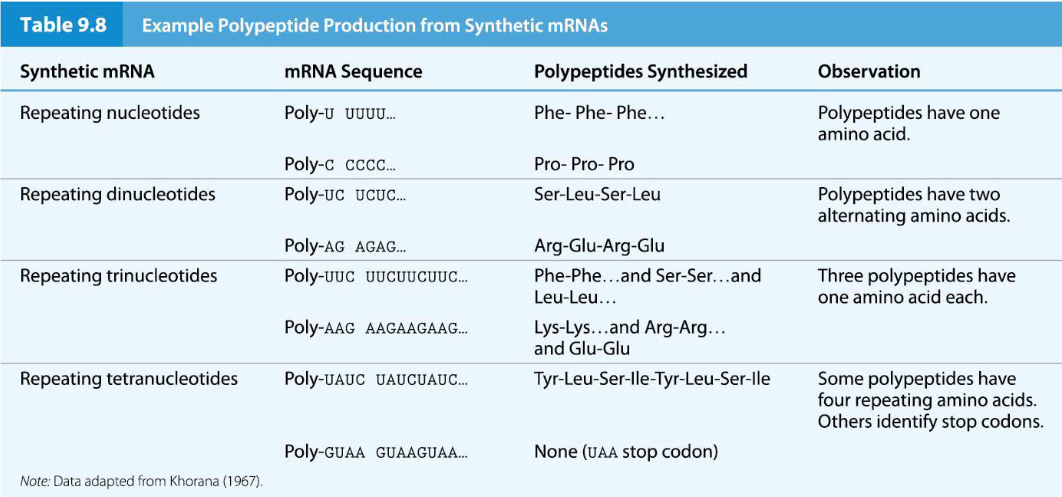
Har Gobind Khorana expanded the range of template RNAs by synthesizing mRNA molecules with repeating di-, tri-, and tetranucleotides,
Results with artificial mRNAs made from repeating nucleotides helped to narrow down the codons that might be associated with each amino acid, but could not show precisely which codon corresponded to which amino acid
Deciphering the code: How did they finally match codons with their correct amino acid?
They used trinucleotide mini-RNAs (one for each possible codon), as templates to resolve the ambiguities of previous experiments.
Their method involved:
Adding each synthetic mini-RNA trinucleotide (e.g; GUC) to an in vitro translation system.
Each system contained ribosomes and all twenty amino acids, but only one amino acid was radio-labeled with 14C at a time.
They then isolated the ribosome-tRNA-mRNA complexes. The complexes, if formed, would be trapped on the filter, while unbound tRNAs would pass through.
By determining which radio-labeled amino acid was associated with which mini-RNA trinucleotide, they could identify the specific codon-amino acid pairing. For example, the trinucleotide GUC bound valine; Therefore, GUC is the codon for valine
*They identified all 61 of the codon–amino acid associations and also identified the three stop codons, UAA, UAG, and UGA
What are synonymous codons? What are the exceptions to this?
All amino acids except methionine and tryptophan are specified by at least two codons, called synonymous codons.
What are the necessary structure for protein synthesis?
mRNA: Carries the genetic message to be decoded.
tRNA: Acts as an adaptor molecule, linking codons to specific amino acids.
Ribosomes: Catalyze the formation of peptide bonds.
Extra Factors: Various proteins that help regulate and facilitate the process.
tRNA: Describe tRNAs secondary structure
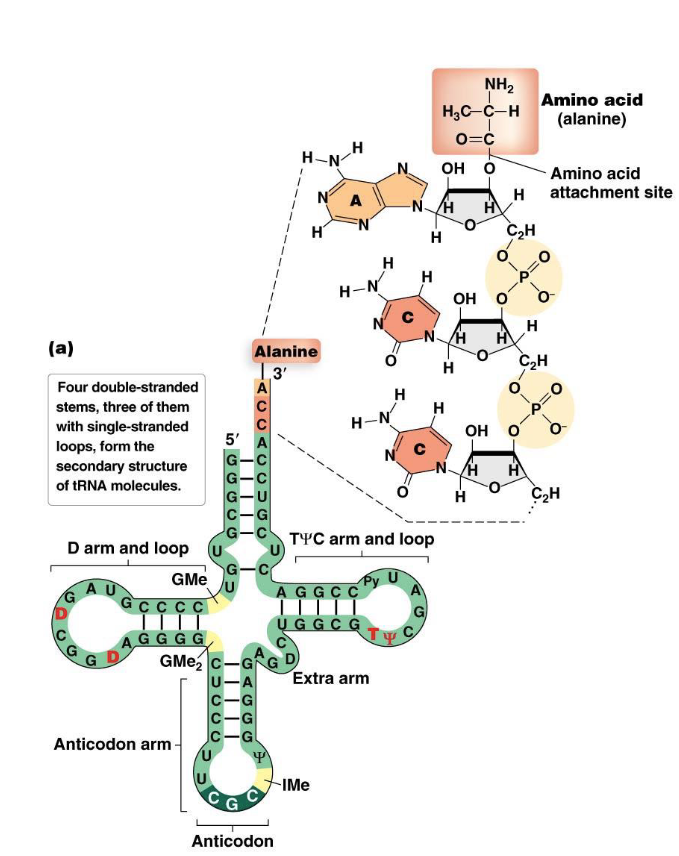
Often represented as a cloverleaf shape, formed by intramolecular base pairing that creates four double-stranded stems and three single-stranded loops.
One of these loops, the anticodon loop, contains a three-nucleotide sequence called the anticodon. This anticodon is complementary to and base-pairs with a specific codon on the mRNA.
tRNAs also contain a number of modified bases (e.g., dihydrouracil (D) and pseudouracil (ψ)) which are important for their structure and function.
The amino acid attachment site is at the 3' OH end of the tRNA. A specific amino acid is covalently attached here.
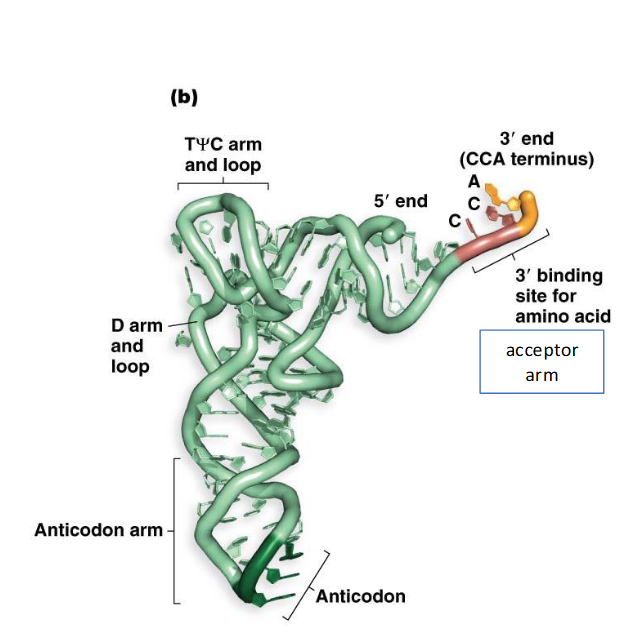
tRNA: Describe tRNAs tertiary structure
In its functional form and a better representation, tRNA folds into a compact L-shape.
This L-shape exposes two critical regions: the 3' acceptor end (where the amino acid attaches) and the anticodon loop.
The exposed 3' acceptor end allows the amino acid to be delivered to the location in the ribosome where peptide bond formation takes place
The exposed anticodon loop facilitates base-pairing with the complementary mRNA codon.
Which enzyme facilitates the covalent bond formation between amino acids and tRNA? What must they recognize?
Amino acids are covalently attached to tRNA by special enzymes called aminoacyl tRNA synthetases (aaRS)
aaRS Enzymes must recognize both:
▪ the specific tRNA
▪ the matching amino acid
What’s the difference between a charged and uncharged tRNA? Why is this important?
tRNAs that carry an amino acid are called charged tRNAs, whereas tRNAs without amino acids attached are uncharged.
Correct charging of each tRNA molecule is crucial for the integrity of the genetic code
How does tRNA Nomenclature work?
The standard nomenclature for a tRNA indicates the specific amino acid it is supposed to carry using a three-letter superscript.
tRNA^Met: This tRNA has the anticodon that matches a methionine codon and is supposed to be charged with methionine.
tRNA^Ala: This tRNA has the anticodon that matches an alanine codon and is supposed to be charged with alanine.
*The actual amino acid attached to a tRNA would be indicated: Ala-tRNA^Ala
Is the genetic code completely universal? Or are their any exceptions?
Because the genetic code is universal, bacteria can be used to produce important proteins from plants and animals
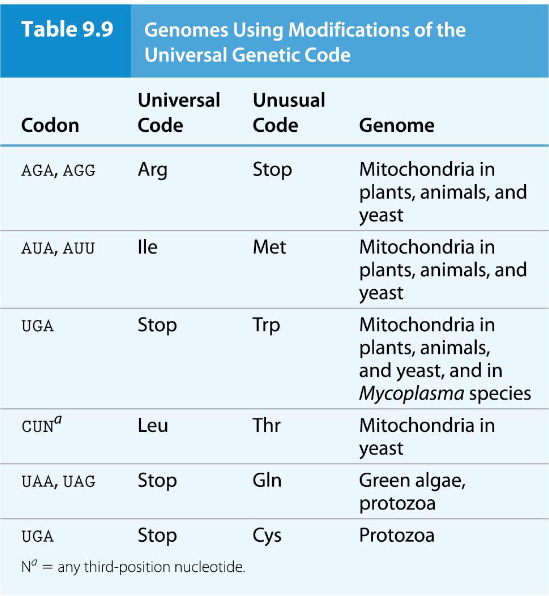
However, there are a few exceptions to the universality of the genetic code, found principally in mitochondria, though there are a few exceptions in living organisms
What are ribosomes, how do they function, and what is their composition?
Ribosomes are the “machines” that carry out protein synthesis
Ribosomes translate mRNA in the 5-to-3 direction, reading each triplet codon and joining the amino acids in the order specified by the codons
▪ They are ribonucleoprotein complexes containing both ribosomal RNAs
(rRNAs) and proteins
What are the subunits of ribosomes and how are they measured?
Ribosomes are composed of two subunits, the large ribosomal subunit and the small ribosomal subunit
The number and sequence of rRNA molecules and the number and type of proteins differs between bacteria, archaea, and eukaryotes
Ribosomal subunit size is measured in Svedberg units (S), a property based on size, shape, and hydration state, and determined by sedimentation rate in the ultracentrifuge
Describe the subunits of bacterial ribosomes
Small Subunit: 30S, containing 21 proteins and one 16S rRNA molecule.
Large Subunit: 50S, containing 32 proteins, a small 5S rRNA, and a large 23S rRNA.
Fully Assembled Ribosome: 70S.
Describe the subunits of eukaryotic ribosomes
Small Subunit: 40S, containing about 35 proteins and one 18S rRNA.
Large Subunit: 60S, containing 45 to 50 proteins and three rRNA molecules (5S, 5.8S, and 28S).
Fully Assembled Ribosome: 80S.
How many structures do the rRNA ribosomes fold into?
rRNA folds into stable secondary (and tertiary) structures
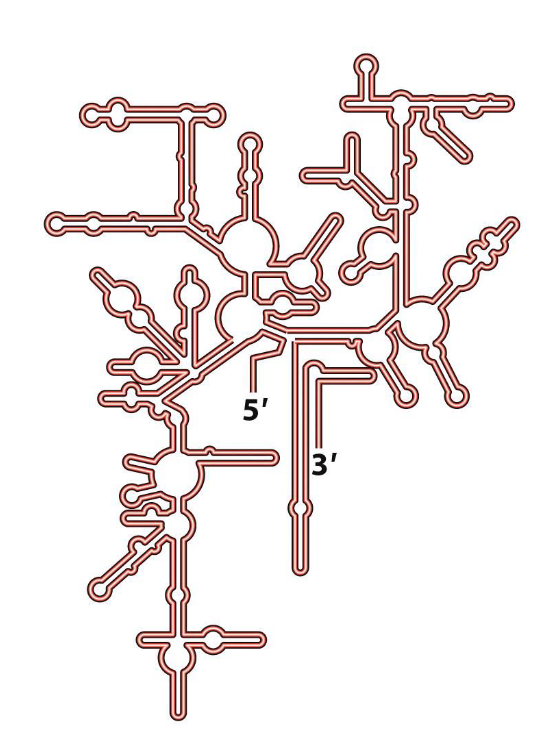
What are the binding sites for ribosomes?
A site (Aminoacyl site): This is where incoming charged tRNAs, carrying their specific amino acids, first bind and pair with the mRNA codon.
P site (Peptidyl site): This site holds the tRNA that is attached to the growing polypeptide chain.
E site (Exit site): This is where uncharged tRNAs (tRNAs that have delivered their amino acid) exit the ribosome.
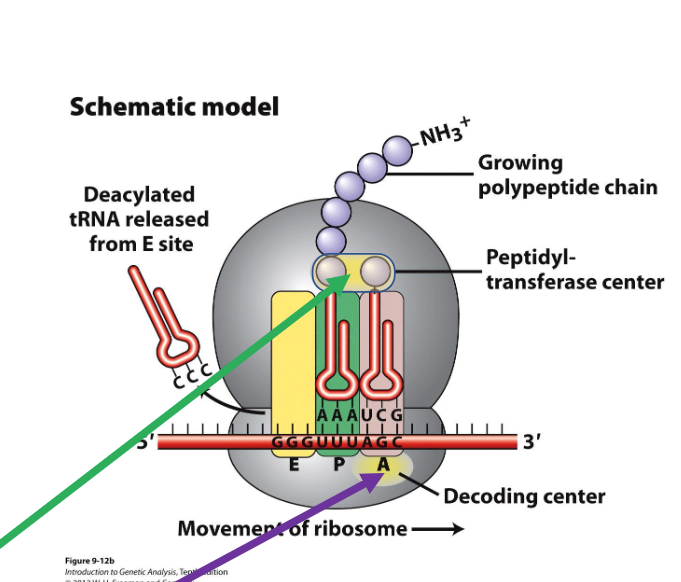
What are the 2 regions in the ribosomes cruicial for protein synthesis?
Peptidyl-transferase centre (where protein synthesis occurs)
Decoding centre (where tRNAs are matched to codons)
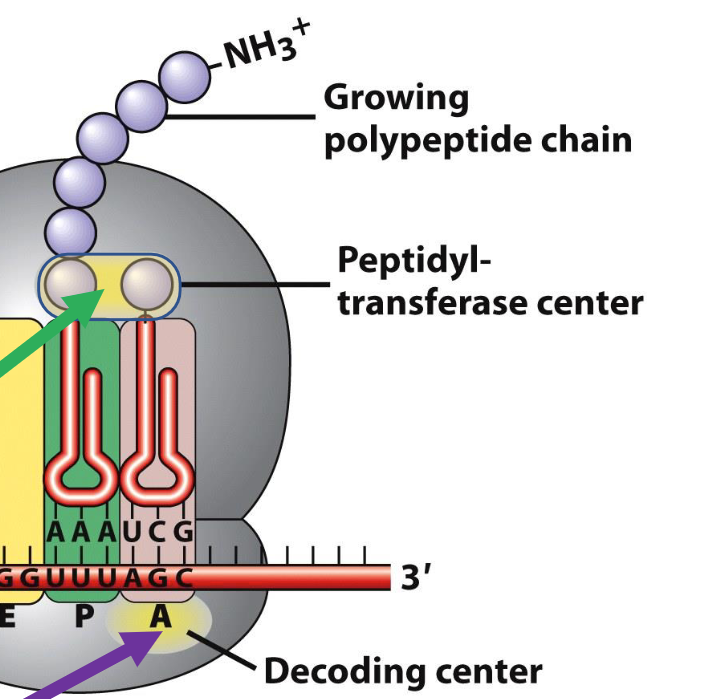
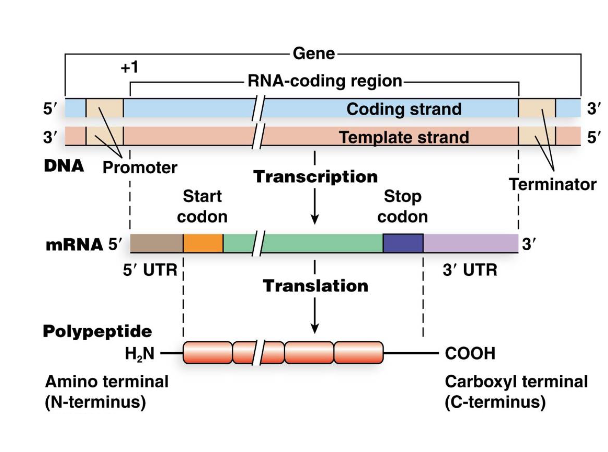
What are the boundaries of translation in mRNA and what are outside these regions?
Boundaries of translation are defined by a start codon that corresponds to the N-terminus of the protein and a stop codon that is immediately after the C- terminus
The 5’ untranslated region (5’ UTR) and the 3’ untranslated region (3’ UTR) are segments of the mRNA outside of the translated regions
What are polycistronic mRNAs in bacteria?
Polycistronic mRNAs contain multiple polypeptide-producing segments, each with a translation-initiating region
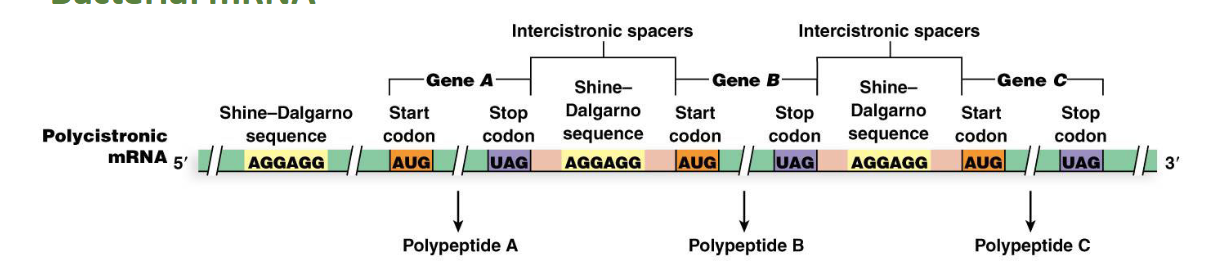
What is the Shine–Dalgarno Sequence in bacteria (location and function)?
Location: This is a purine-rich sequence of about six nucleotides, located three to nine nucleotides upstream of the start codon.
Function: It serves as a ribosome binding site. The translation-initiating region contains this ribosome binding site (called a Shine–Dalgarno sequence) and start and stop codons
What are the 3 tasks of ribosomes in bacteria, archaea, and eukaryotes? Which phases do ribosomes perform their requirements in protein synthesis?
Bind mRNA and identify the start codon, where translation begins
Facilitate complementary base pairing of mRNA codons and the corresponding tRNA anticodons
Catalyze formation of peptide bonds between amino acids on the growing polypeptide chain
These tasks correspond to the first two phases of protein synthesis (initiation and elongation). Last is termination that doesn’t use ribosomes
How is the three phases of translation different in eukaryotes and bacteria?
The big difference in translation between bacteria and eukaryotes is in:
How the codon is identified
How the ribosome is positioned
The second and third phases are similar in bacteria and eukaryotes though there are some differences
Initiation of Translation in Bacteria: What is required?
The messenger RNA (mRNA)
the small ribosomal subunit
the large subunit
the initiator tRNA – a special tRNA used to start translation
Initiation of Translation in Bacteria: What are the 3 initiation factor proteins and their roles
▪ IF1 – to prevent the initiator tRNA going into the incorrect site
▪ IF2 – to guide the initiator tRNA into the correct site
▪ IF3 – to keep the two ribosomal subunits apart
Initiation of Translation in Bacteria: Describe how the ribosome connects to the mRNA
How it begins: The 3’ end of the 16S rRNA (inside the small 30S subunit) base-pairs with the Shine–Dalgarno sequence on the mRNA
This positions the start codon in the correct place in the P site
Initiation of Translation in Bacteria: Describe initiator tRNA and its role in initiation of bacterial translation. How do the initiation factor proteins contribute?
The initiator tRNA must base-pair with the start codon
The initiator tRNA carries a modified amino acid, N-formylmethionine (fMet);
the charged initiator tRNA is called fMet•tRNA^fMet
▪ IF1 blocks it from going into the A site
▪ IF2 guides it into the P site (binds to GTP to facilitate)
Initiation of Translation in Bacteria: What’s the last step to initiation?
Finally, the 50S large subunit joins the 30S small subunit to form the full 70S ribosome
The three initiation factors (IF1, IF2, and IF3) dissociate at the same time as the 70S initiation complex is formed
The union of the two subunits is accompanied by the hydrolysis of GTP to GDP
Elongation of Translation in Bacteria: What’s the process of translational bacterial elongation that elongates the peptide chain?
Charged tRNAs associated with EF-Tu protein and GTP “inspect” the open A site
A tRNA with the correct anticodon sequence enters the A site and pairs with the codon on the mRNA
Hydrolysis of GTP releases EF-Tu (along with GDP) from the tRNA
The end of the tRNA carrying the amino acid enters the peptidyl transferase centre
Peptidyl transferase catalyzes peptide bond formation between amino acids at the P and A sites, extending the polypeptide by one amino acid as it is transferred to the tRNA at the A site
Elongation of Translation in Bacteria: How does the tRNA leave the ribosome and how can the elongation process continue?
The tRNA from the P site exits through the E site
Using GTP hydrolysis, EF-G protein translocates the ribosome along the mRNA
This moves the tRNA at the A site to the P site and opens the A site for the next charged tRNA carrying the correct anticodon
Termination of Translation in Bacteria: What triggers elongation to stop and termination to occur?
The elongation cycle continues until one of the three stop codons (UAA, UAG, UGA) enters the A site of the decoding center on the ribosome
Termination of Translation in Bacteria: What is used to recognize these stop codons (specific to certain codons)? How do they work?
In bacteria, the release factor RF1 recognizes UAG and UAA; and RF2 recognizes UAA and UGA
Function: Binding of the release factor(s) leads to hydrolysis of GTP and release of the polypeptide bound to the tRNA at the P site
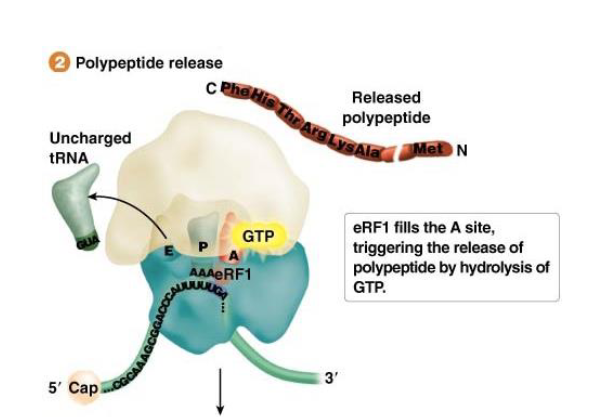
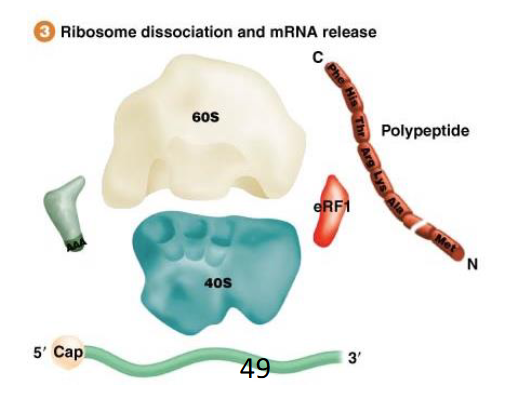
What occurs at the end of the bacterial termination process that allows translation to continue?
The ribosome subunits are dissociated and recycled for another round of protein synthesis; this does require some other protein factor
What are the antibiotics that target steps of protein synthesis and their mechanisms of action?
Inhibitor | Mechanism of Action |
|---|---|
Cycloheximide | Blocks translocation step |
Diphtheria toxin | Inactivates eEF2 |
Puromycin | Structural analog of aa-tRNA; triggers premature release of polypeptide |
Erythromycin | Blocks elongation step |
Streptomycin | Causes misreading during elongation |
Tetracycline | Prevents stable binding of EF-Tu·tRNA |
|
Is the central dogma simple? Does it go simply from DNA gene to RNA to Protein?
No, it requires it is the NET effect of several processes:
Product processing, Product degradation, and Regulation
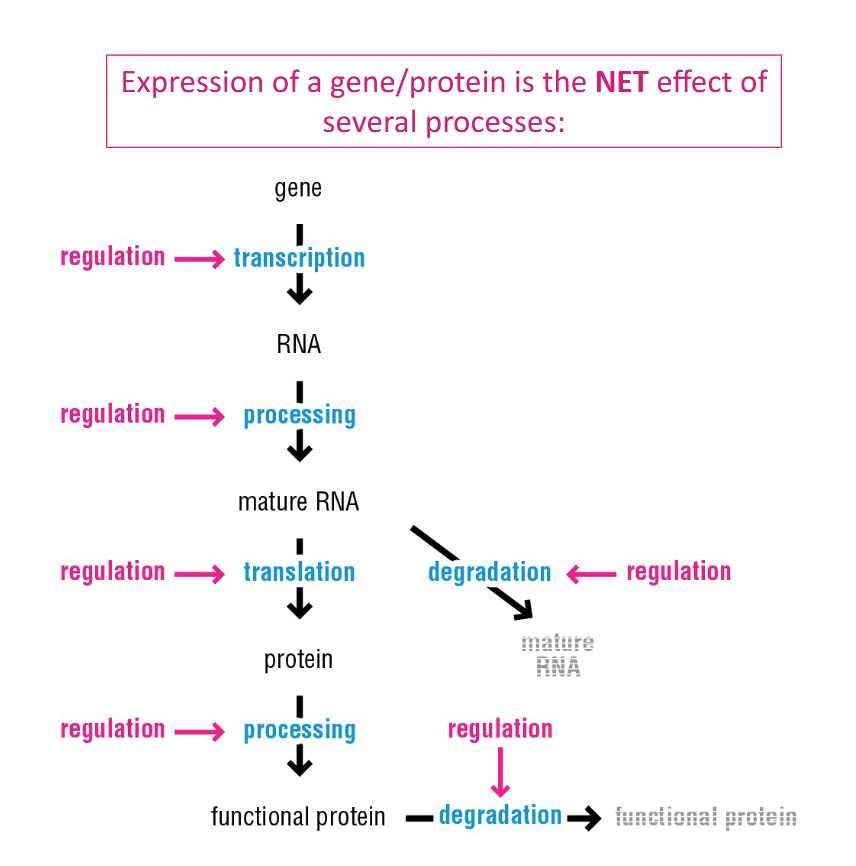
What’s the most fundamental way gene expression can be regulated?
Transcription: regulating the level of the initial RNA
Control synthesis of RNA: either ⬆ or ⬇ synthesis
Control degradation of RNA: either ⬆ or ⬇ degradation
What’s the difference between Constitutive vs. Regulated Transcription
Constitutive transcription/expression: Certain bacterial genes whose products are continuously needed for routine tasks are expressed all the time.
Regulated transcription: Other genes, whose products are needed only in response to changing environmental conditions, require regulated transcription.
For these genes, the cell must determine a default state of expression (ON or OFF) and then decide whether to activate or repress expression based on environmental cues.
What’s the difference between Negative and Positive Control of regulation of bacteria?
Negative control of transcription: Involves the binding of a repressor protein to a regulatory DNA sequence, which prevents transcription of a gene or gene cluster.
The default state can be ON (repressor inactive) or OFF (repressor active).
Positive control of transcription: Involves the binding of an activator protein to a regulatory DNA sequence, which helps RNA polymerase initiate gene transcription.
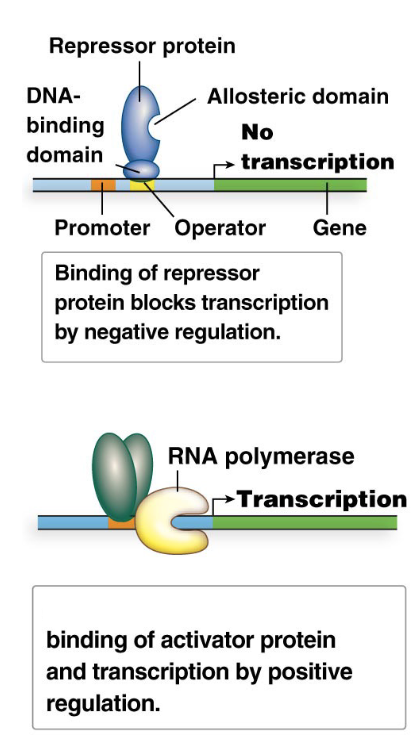
What are repressor proteins? Explain their mechanism and how they can be altered
Repressor proteins are a broad category of regulatory proteins that mediate negative control of transcription.
Operators: Repressors bind to specific DNA sequences called operators.
Mechanism: Repressor binding to the operator physically blocks RNA polymerase from initiating transcription, thus turning gene expression OFF.
Activation/Inactivation: Repressors can be activated or inactivated by interacting with other molecules (e.g., inducers or corepressors).
What are the 2 domains of Repressor Proteins?
DNA-binding domain: This domain recognizes and binds specifically to the operator DNA sequence.
Allosteric domain: This domain binds a small molecule (e.g., an inducer or corepressor) or another protein. Binding to the allosteric domain causes a conformational change in the repressor protein, which, in turn, alters the activity of the DNA-binding domain. This property is called allostery.
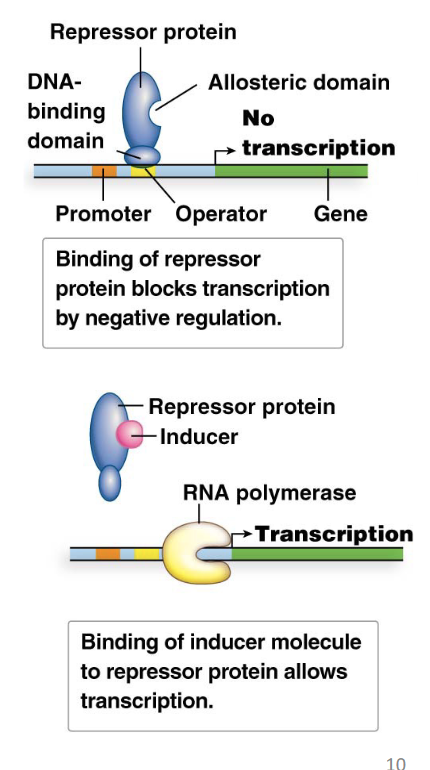
why do repressor proteins need allostery?
because their activity depends on the default state (ON of OFF) of gene expression.
Describe the two types of default states
Default State is OFF (Gene Not Needed):
When the gene product is not required (e.g., no lactose for the lac operon), the repressor protein is in its active form. In this state, it binds to the operator region of the DNA, physically blocking RNA polymerase from initiating transcription. This prevents the production of unnecessary proteins.
Default State ON (Gene needed):
Default State is ON (Gene Needed): When the gene product is required (e.g., low tryptophan levels for the trp operon), the repressor protein is in its inactive form. In this state, it cannot bind to the operator region of the DNA, allowing RNA polymerase to initiate transcription. This ensures the continuous production of essential gene products, like amino acids.
Explain how inducible and repressible systems and how they relate to the default states
Default State OFF (Inducible System):
If the specific inducer molecule (e.g., allolactose) is present in high concentration, it binds to the allosteric domain of the repressor protein.
This binding causes a conformational change in the repressor protein. This change inactivates its DNA-binding domain, meaning the repressor can no longer effectively bind to the operator or detaches from it if already bound, and transcription can occur (turns it ON)
Example: lac operon for lactose metabolism.
Default State ON (Repressible System):
If the specific corepressor molecule (e.g., tryptophan) is present in high concentration, it binds to the allosteric domain of the repressor protein.
This binding causes a conformational change in the repressor protein. This change activates its DNA-binding domain, meaning the repressor can now effectively bind to the operator region, and transcription is blocked (turns it OFF)
Example: trp operon for tryptophan biosynthesis.
What are operons, how are they related to bacteria, and what are two examples?
Operons: Clusters of genes that undergo coordinated transcriptional regulation by a shared regulatory region (an operator).
Common in bacteria: Genes within a particular operon nearly always participate in the same metabolic or biosynthetic pathway. This allows bacteria to efficiently turn on or off all the enzymes needed for a specific process simultaneously.
Two classic examples:
The lac operon (an inducible system)
The trp operon (a repressible system)
What is the lac operon responsible for in lactose metabolism and how does it relate to inducible systems?
The lactose (lac) operon in E. coli is responsible for producing the proteins needed to utilize lactose as an energy source.
Lactose Metabolism
Preferred energy source: Glucose is the preferred energy source for E. coli, metabolized via glycolysis.
Lactose as an alternative: Lactose is an alternative carbon source. Its utilization is controlled by the lac operon, an inducible system.
Inducible systems: These systems are turned ON only when an inducer compound is available. In the lac operon, the true inducer is allolactose.
Describe the steps of lactose metabolism
Lactose (a disaccharide of glucose and galactose) is transported into the bacterial cell by the enzyme lactose permease (a gated channel in the cell membrane).
Inside the cell, β-galactosidase breaks the β-galactoside linkage in lactose.
Lactose is primarily broken down into glucose and galactose. Glucose enters glycolysis directly, and galactose is further processed into glucose.
A small amount of lactose is converted into allolactose by β-galactosidase; allolactose acts as the inducer for the lac operon.
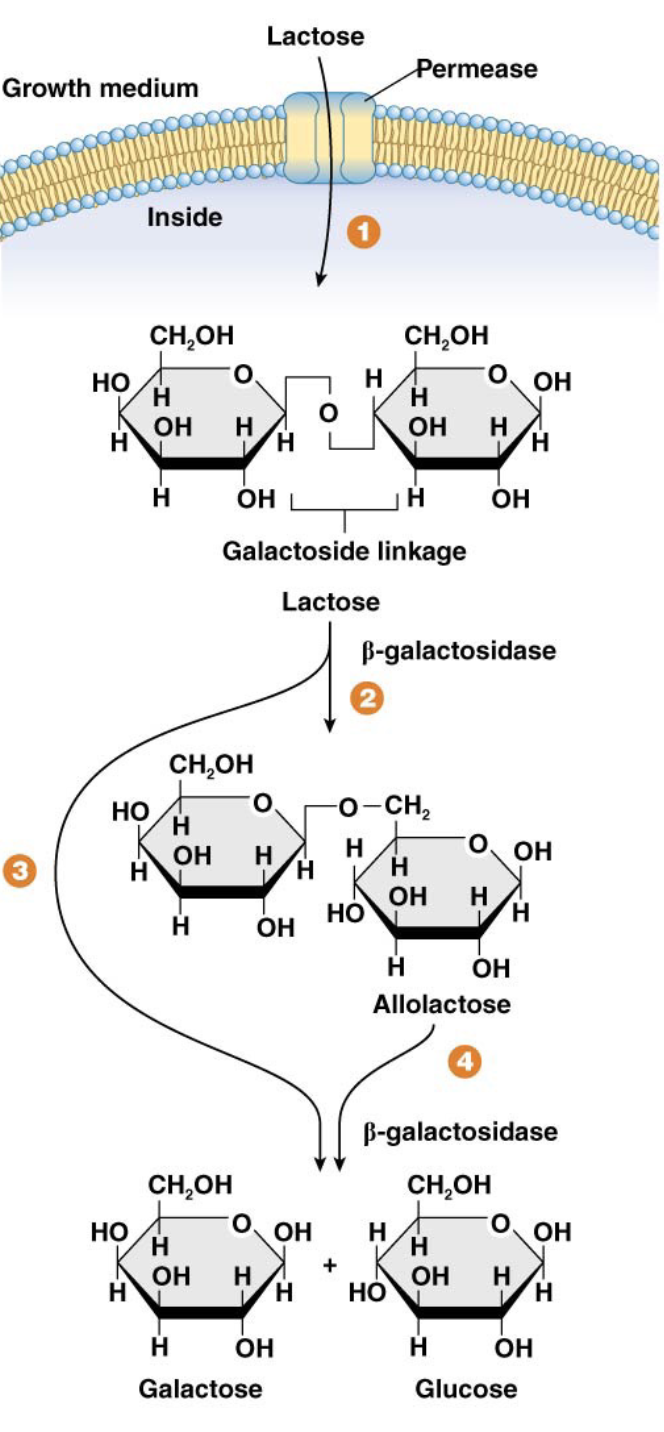
What happens if bacteria have the lac− phenotype?
Bacteria are unable to utilize lactose.
Describe the regulatory region of the lac operon
Promoter (lacP): Binds RNA polymerase.
Operator (lacO): Binds the lac repressor protein.
CAP-cAMP region: (Not discussed in detail here, but involved in positive regulation by glucose levels).
These regions partially overlap and are immediately upstream of the start of transcription for the structural genes.
What are the 3 genes in the coding region of the lac operon and which aspect of lactose metabolism each encodes for? What does this mean the entire lac operon?
The lac operon consists of a multipart regulatory region and three structural genes:
lacZ, which encodes b-galactosidase
lacY, which encodes the enzyme lactose permease
lacA, which encodes transacetylase
This means they are transcribed as a single, polycistronic mRNA, which is translated to produce the three distinct polypeptides
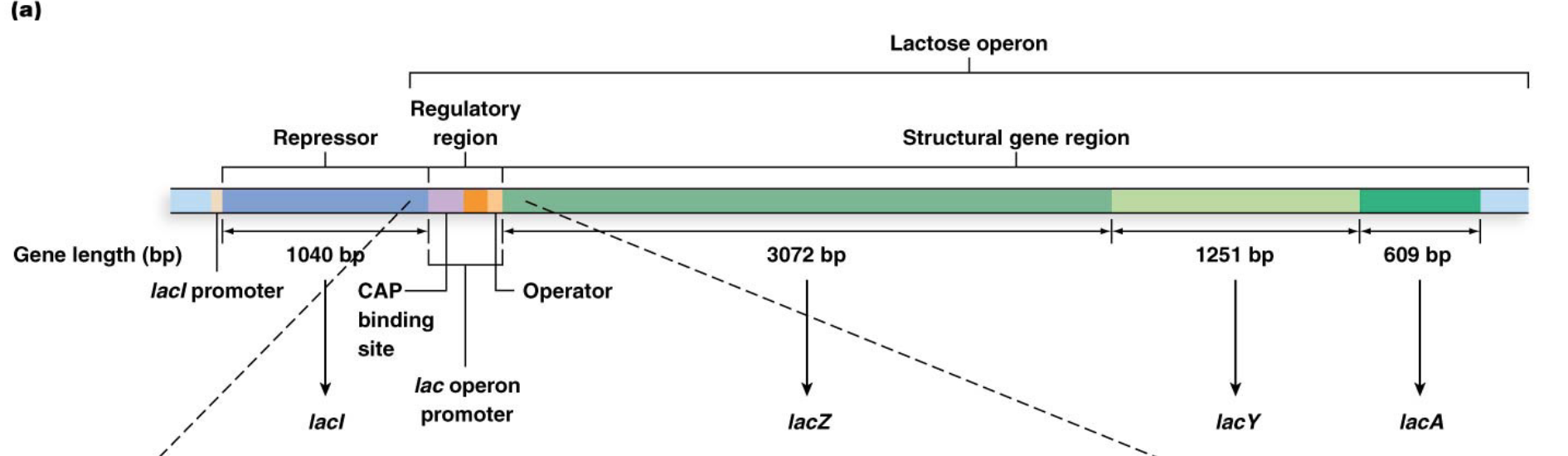
What is the lacI gene and its role?
Located next to, but not part of, the lac operon. It constitutively (but weakly) expresses the lac repressor protein.
The lac repressor protein is a homotetramer (made of four identical subunits). It has a DNA-binding domain that binds to the lacO sequence and an allosteric domain that binds the inducer, allolactose.
How does the lac operon work when lactose is absent or glucose is available. What’s this an example of?
The lac operon is transcriptionally silent when no lactose is available or when glucose is available
When no b-galactosidase is produced, there is no allolactose in the cell and the lac repressor protein binds to lacO, preventing transcription
This is an example of negative control
How does the lac operon work when lactose is available or glucose is absent?
When lactose is available to the cell and glucose is not, transcription of the lac operon is induced
With synthesis of b-galactosidase, allolactose is produced, and binds to the allosteric domain of the lac repressor
The formation of the inducer–repressor complex alters the DNA-binding domain of the repressor and prevents it from binding the operator
What is the trp operon responsible for in synthesizing the amino acid tryptophan and how does it relate to repressible systems?
The trp operon in E. coli contains five structural genes whose protein products are responsible for.
Preferred amino acid source: E. coli cells produce their own tryptophan when external sources are scarce.
Tryptophan as a regulator: Tryptophan biosynthesis is controlled by the trp operon, a repressible system.
Repressible systems: These systems are turned OFF only when a corepressor compound is available. In the trp operon, the corepressor is tryptophan itself.
Describe the regulatory region of the trp operon
Promoter (trpP): Binds RNA polymerase.
Operator (trpO): Binds the TrpR repressor protein.
Leader region (trpL): Contains the attenuator region
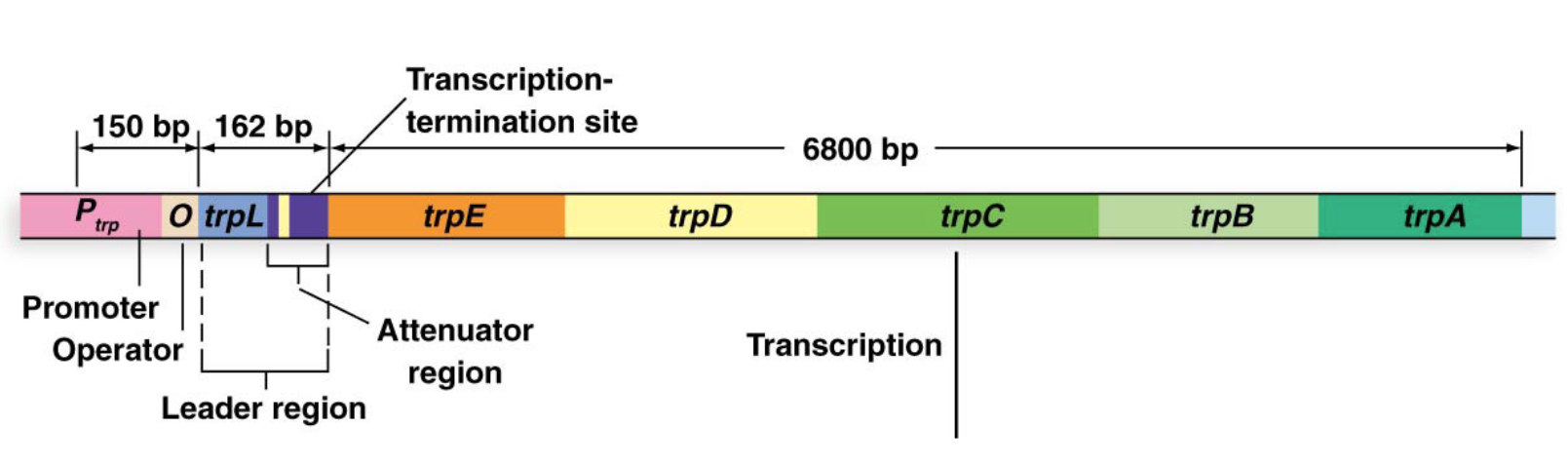
What are the 5 genes in the coding region of the trp operon?
trpE, trpD, trpC, trpB, trpA.
also polycistronic
What is the TrpR gene and its role?
A sixth gene, located outside the operon, encodes the TrpR protein, which is the trp repressor protein.
The TrpR protein is activated when bound to tryptophan.
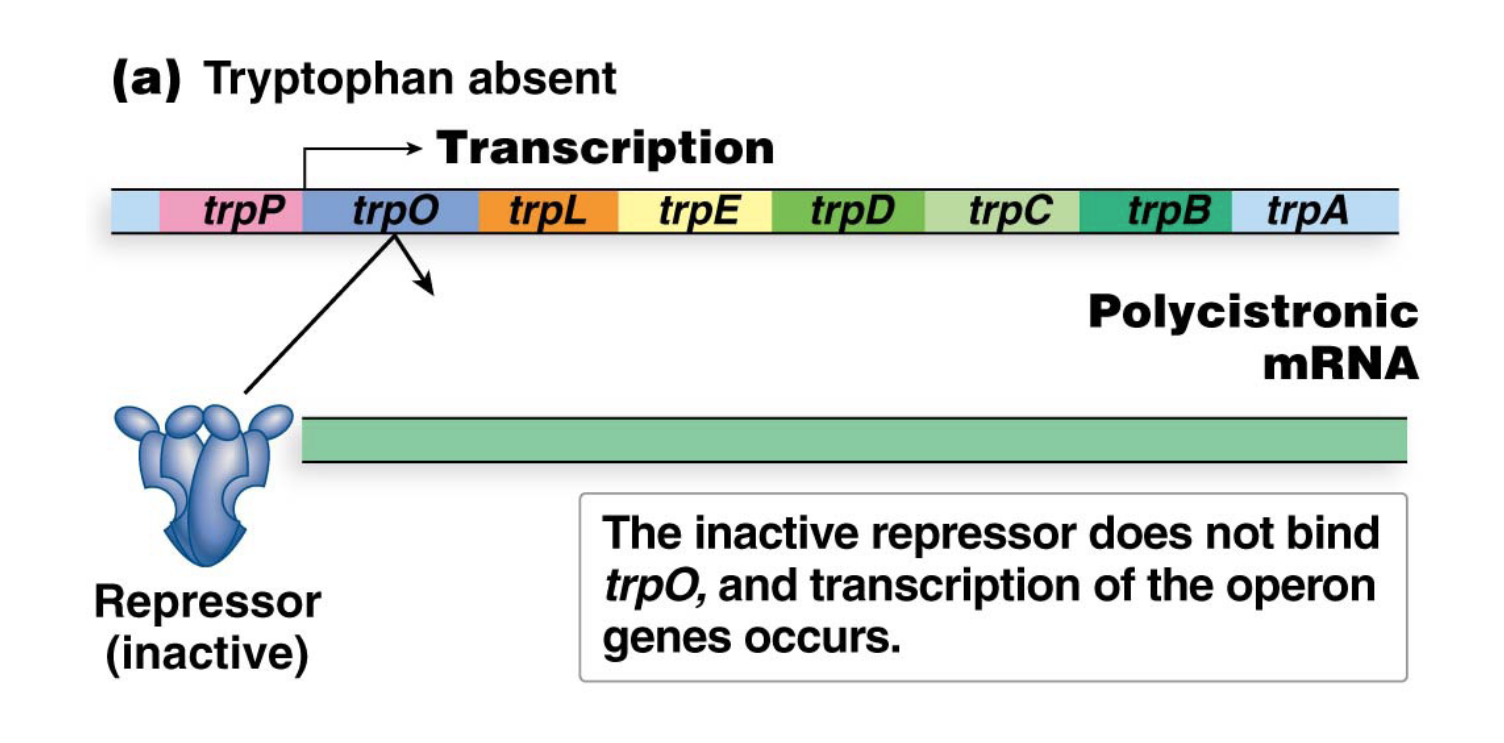
How does the trp operon work when Tryptophan is absent?
When tryptophan is absent from the cell, the repressor protein cannot bind the operator, allowing for transcription of the operon (and biosynthesis of tryptophan)
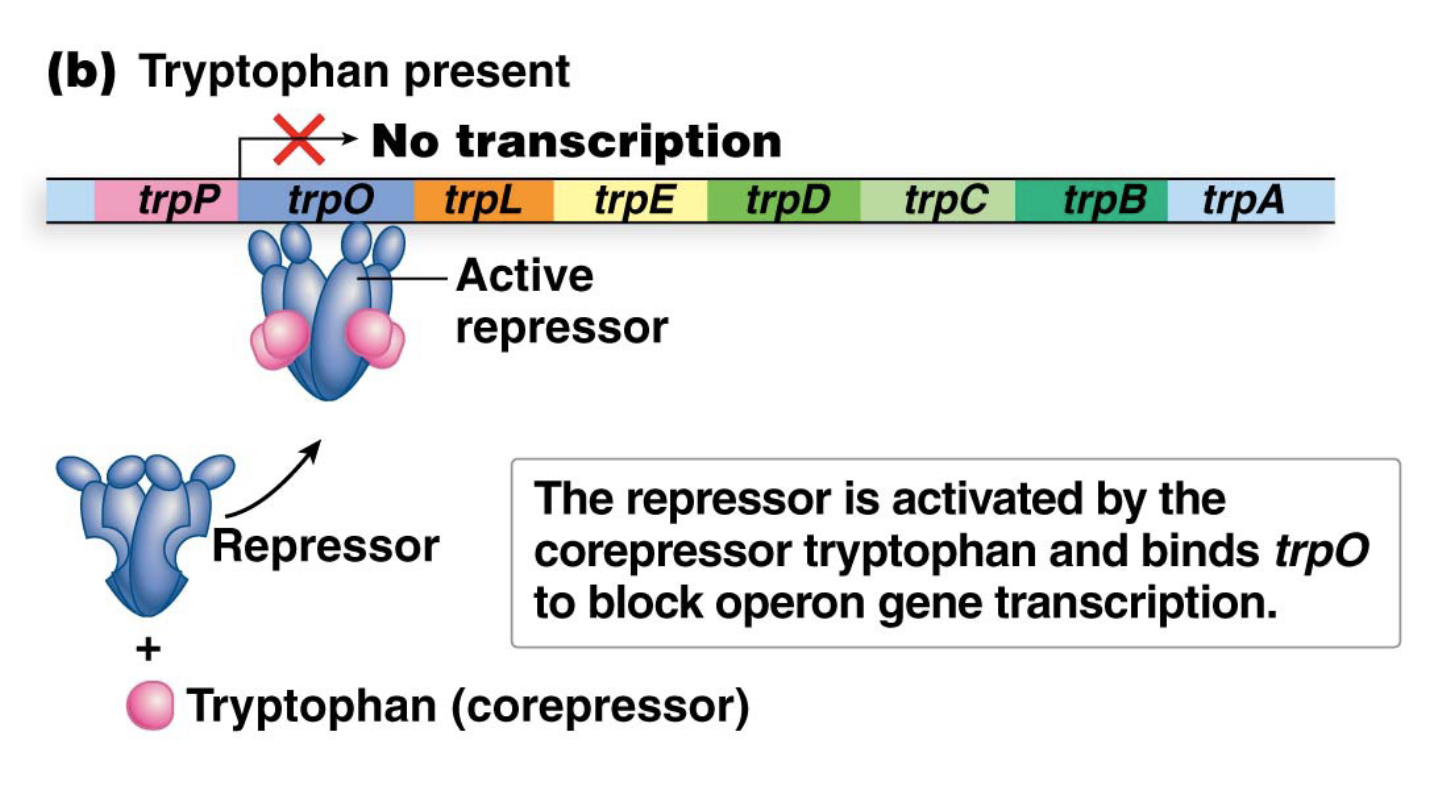
How does the trp operon work when Tryptophan is present?
Tryptophan acts as a corepressor by binding to the trp repressor and activating
Activated repressor binds to trpO and prevents transcription of the operon
What are similarities and differences between transcriptional regulation in bacteria versus eukaryotes
Similarity:
Activator proteins bind their regulatory sequences to stimulate transcription
Repressor proteins bind their regulatory sequences to hinder transcription
Difference:
Unlike bacteria, eukaryotic regulation is generally through positive control mechanisms
Also, unlike bacteria, the regulatory proteins are found in large complexes in eukaryotes
How is regulation accomplished in eukaryotes
Expression of eukaryotic genes is controlled by a combination of regulatory sequences and factors that bind to those sequences:
These sequences can be located well away from the promoter – they can be several thousand base pairs away
Regulatory sequences can be downstream or upstream of the promoter
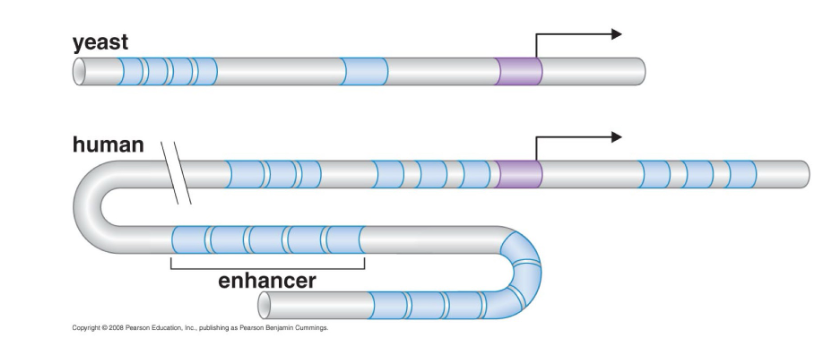
What are the three sets of regulatory DNA sequences are commonly involved in eukaryotic gene regulation?
Core Promoter Region:
Immediately adjacent to the transcription start site.
Contains the TATA box and other sequences.
Binds RNA polymerase II (Pol II) and its associated general transcription factors (GTFs).
Proximal Elements:
Located upstream of the core promoter (e.g., CAAT box, GC box).
Also regulate gene expression.
Enhancer and Silencer Sequences:
Located at greater distances from the core promoter.
Enhancers: Increase transcription.
Silencers: Decrease transcription.
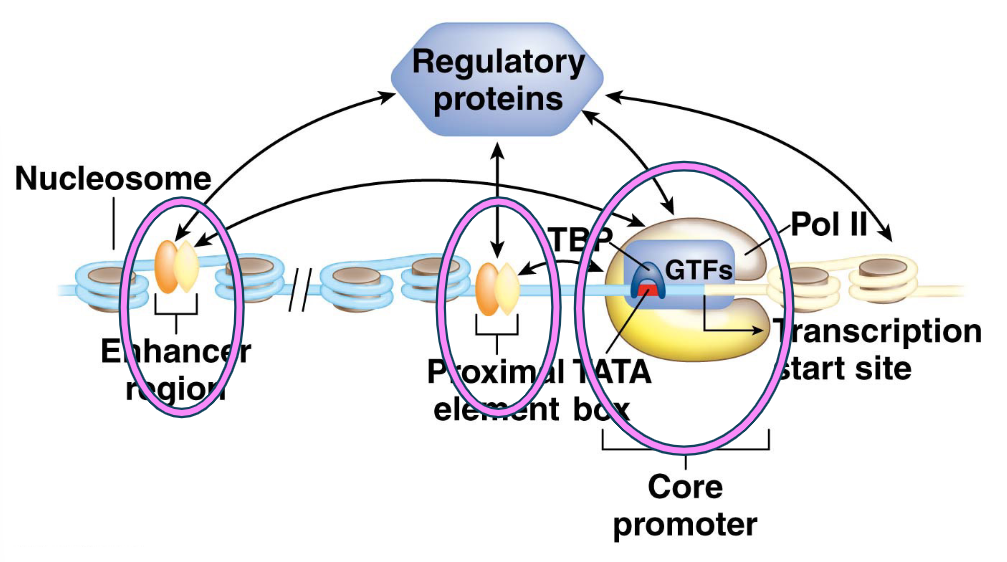
What are Cis-acting regulatory sequences and Trans-acting regulatory proteins?
Cis-acting regulatory sequences: All three types of regulatory regions contain cis-acting sequences, meaning they regulate transcription of genes on the same chromosome.
Trans-acting regulatory proteins: Pol II and GTFs are trans-acting regulatory proteins; they can bind to their target sequences on any chromosome.
How is the yeast Saccharomyces cerevisiae an example of Enhancer and Silencer Sequences
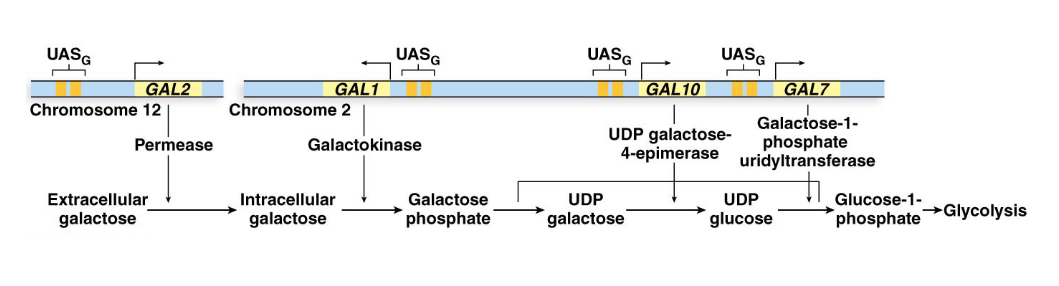
Galactose Utilization Pathway: When galactose is the only available sugar, wild-type yeast induce transcription of four enzyme-producing genes: GAL1, GAL2, GAL7, and GAL10. These genes collectively import and break down galactose.
What is the UASG Enhancer and what’s it bound to?
UASG Enhancer: Each GAL gene has its own promoter and similar enhancer elements. This enhancer is called the upstream activator sequence (UAS), or more specifically, UASG.
Gal4 Activator Protein: The UASG elements are bound by a regulatory activator protein, Gal4, encoded by the GAL4 gene. Gal4 is continuously present in cells.
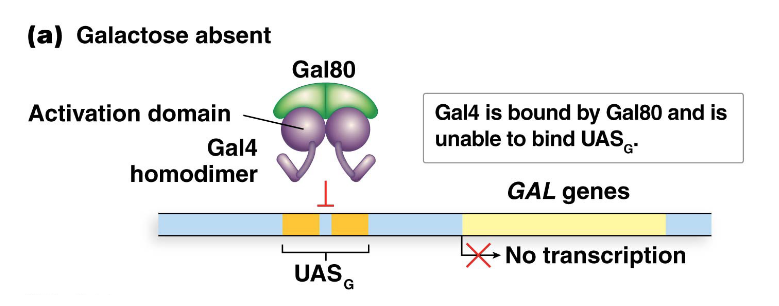
Describe Gal4 function in the absence of galactose
Each UASG element contains two 17-bp repeat sequences that serve as binding sites for Gal4.
Gal4 functions as a homodimer, with each polypeptide having two domains:
one for DNA binding (to the 17-bp sequence)
another for interacting with Gal80.
When galactose is absent, Gal80 interacts with Gal4, keeping it inactive. The Gal4 DNA-binding domain is unable to bind to UASG, resulting in no transcription of the GAL genes.
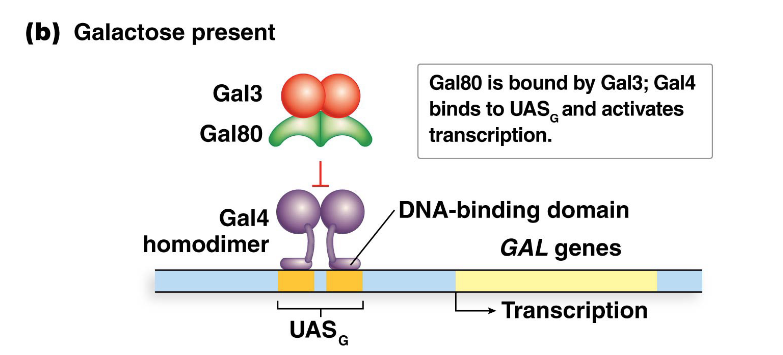
Describe Gal4 function in the presence of galactose
When galactose is present, galactose and Gal3 (encoded by the GAL3 gene) bind to Gal80.
This binding causes Gal80 to release Gal4, freeing the DNA-binding domain of Gal4.
Gal4 can then recognize and bind to the UASG sites.
The transcriptional activation domain of Gal4 then activates transcription of the GAL genes.
What is epigenetics and its two types?
Epigenetics refers to heritable patterns or changes in gene expression that are not associated with any change in the underlying DNA sequence.
Types of Epigenetic Changes
Histone Modifications: Attachment or detachment of chemical moieties (methyl, acetyl, and phosphoryl groups) to specific amino acids (most commonly lysines) in the N-terminal tails of histone proteins.
DNA Methylation: Attachment or detachment of a methyl group (CH3) to DNA nucleotides, specifically to cytosines within CpG dinucleotides.
Describe chromatin, euchromatin, heterochromatin (and its two types)
Chromatin: The complex of nucleic acids (DNA) and proteins (histones) that compose eukaryotic chromosomes.
Euchromatin: Loosely compacted chromatin, rich in gene concentration, and generally transcriptionally active.
Heterochromatin: Highly compact chromatin that is transcriptionally silent.
Constitutive heterochromatin: Always compact and silent (e.g., centromeres, telomeres).
Facultative heterochromatin: Can switch back and forth between euchromatin and heterochromatin states, allowing for regulated gene silencing.
Explain the three Chromatin Modifiers: Writers, Erasers, and Readers
Writers: Enzymes that add chemical groups (e.g., HMTs, HATs).
Erasers: Enzymes that remove chemical groups (e.g., HDMTs, HDACs).
Readers: Proteins that recognize and bind to the modified histones, then recruit other proteins to alter chromatin structure or function.
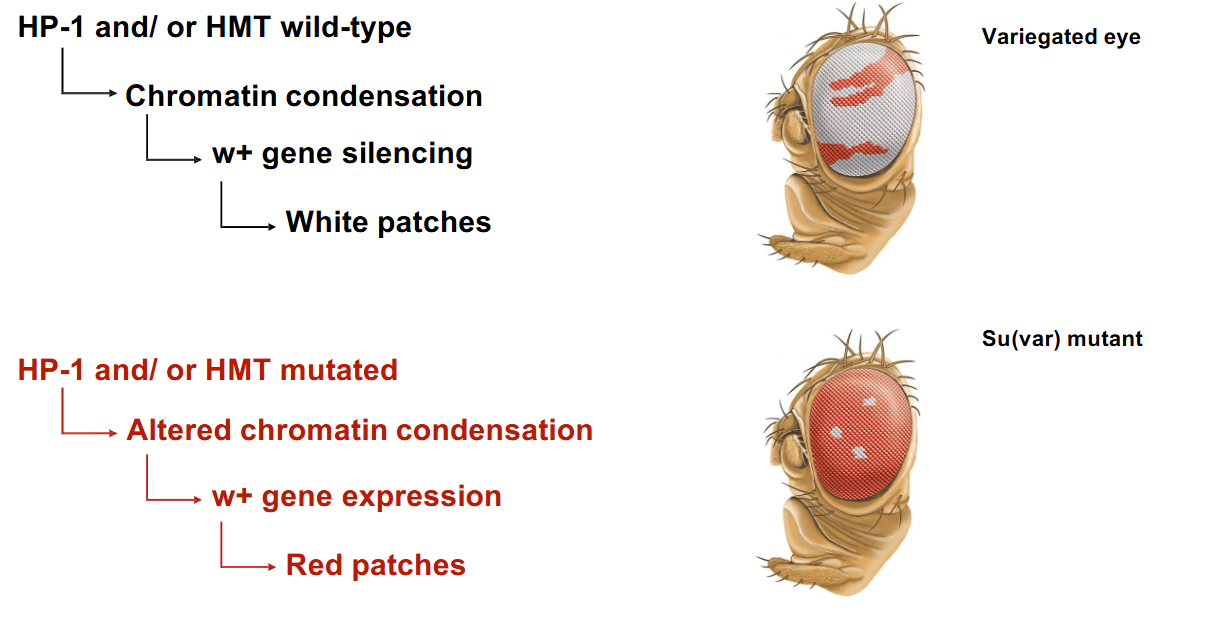
How is heterochromatin formed?
Histone methyltransferases (HMTs): These are "writers" that catalyze the addition of methyl groups (CH3) to specific lysine residues in histone proteins. A key modification for heterochromatin is the methylation of lysine residue 9 of histone H3 (H3K9me- histone epigenetic modifications that marks constitutively heterochromatic regions).
Heterochromatin protein-1 (HP-1): This protein specifically binds to H3K9me (reader) and plays a critical role in the formation and maintenance of heterochromatin, leading to chromatin condensation and gene silencing.
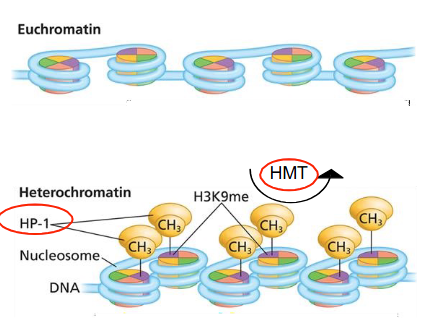
How do wild-type and defective heterochromatin actors in Drosophila (flys) affect the w+ gene for red eyes?
If HP-1 and/or HMTs are wild-type, the w+ gene can be silenced by chromatin condensation, resulting in white patches.
If HP-1 and/or HMTs are mutated, the chromatin condensation is altered, allowing w+ gene expression in some cells that would otherwise be silenced, leading to red patches.
The reversible transition between inactive heterochromatic DNA and active euchromatic DNA is mediated by what?
Chromatin modifiers: Enzymes that modify histones by adding or removing methyl or acetyl groups at specific amino acid residues (most commonly lysines).
Chromatin remodelers: Enzymes that catalyze changes in the distribution or composition of nucleosomes (histone octamers), altering chromatin accessibility.
Describe the Chemical Modifications of Chromatin (histone modifications)
Histone modifications that involve the addition and removal of acetyl groups and methyl groups at specific amino acids (usually lysine) in the N-terminal (amino-terminal) region of histones
These modifications alter the strength of association between nucleosomes and DNA,
Chemical modifications affect gene accessibility and gene regulation.
What are the primary targets of chemical modifications?
Acetyl and methyl groups are added/removed to/from specific lysine (K) residues in the N-terminal tail of Histone 3.
Three lysines are particularly important targets for writers and erasers: K4, K9, and K27.
What are the writers and erasers involved in Methylation and Demethylation?
Histone methyltransferases (HMTs): Writers that add methyl groups.
Histone demethylases (HDMTs): Erasers that remove methyl groups.
Describe the 3 specific chemical modifications, H3K4, H3K9, and H3K27, that are involved with Methylation and Demethylation
H3K9 methylation: Often marks constitutive heterochromatin (gene silencing).
H3K27 methylation: Often marks facultative heterochromatin (reversible gene silencing).
H3K4 methylation (often with H3K9ac): Associated with euchromatin and active transcription.
What are the writers and erasers involved in Acetylation and Deacetylation?
Histone acetyltransferases (HATs): Writers that add acetyl groups.
Histone deacetylases (HDACs): Erasers that remove acetyl groups.
Describe the H3K9ac chemical modifications involved with Acetylation and Deacetylation?
H3K9 acetylation (H3K9ac): Is strongly associated with an opening of the chromatin structure and active transcription. Acetylation reduces the positive charge of histones, loosening their grip on the negatively charged DNA.
What is DNA methylation, its target/location, and how is it catalyzed?
Modification: Involves the addition of a methyl group (CH3) to a DNA nucleotide.
Target: Occurs primarily to cytosines in CpG dinucleotides (where a cytosine is followed by a guanine). These CpG sites are often clustered in "CpG islands" near gene promoters.
Enzymes: Catalyzed by specialized DNA methyltransferases (DNMTs).
What are the functions of DNA methylation?
Silencing: Silences repetitive/transposable element sequences and noncoding sequences.
Regulation: Tight regulation of functional genomic regions, such as gene regulatory sequences (e.g., promoters).
How is DNA methylation related to the regulation of gene expression in eukaryotes?
Gene Silencing: Methylation of CpG islands in gene promoters is generally associated with gene expression repression. The methyl groups can physically block transcription factor binding or recruit methyl-binding proteins that in turn recruit chromatin-modifying enzymes (like HDACs) to condense chromatin.
Gene Activation: Conversely, demethylation of CpG islands can lead to gene activation, allowing transcription factors and RNA polymerase to bind to the promoter.
How does DNA methylation plays a dramatic role in determining the phenotype of female bees?
Female bee larvae are genetically identical. Their diet determines whether they become queens or workers, by altering their methylation patterns.
Larvae fed royal jelly are highly hypomethylated (less DNA methylation) and develop into queens.
Larvae fed worker jelly undergo extensive methylation (hypermethylated) and become workers.
This shows how epigenetic modifications can lead to wildly different phenotypes from the same DNA.