BIOL 316 Exam #3
1/69
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
70 Terms
1. Explain how genetic variation arises. There are 4 main ways this happens.
1) Crossing over during during Prophase I. Non-sister chromatids from homologous chromosomes physically exchange portions of the chromosome
2) Independent assortment during Metaphase/Anaphase I. It is the idea that pairs of homologous chromosomes line up independent of each other (i.e, all paternal chromosomes aren’t on the same side of the cell, and neither are all the maternal chromosomes)
3) Random fertilization—ANY sperm can fuse with ANY unfertilized egg. This greatly increases the genetic variability among potential gametes
4) Mutations!—Can happen any time
1. In a diploid individual, one set of chromosomes comes from the mother and the other set comes from the father. That is, each chromosome has a homolog. When this diploid individual’s germ cells prepare to divide during meiosis I, homologous chromosomes line up on the Metaphase plate.
a. Do all the paternal chromosomes have to line up on the same side of the metaphase plate?
b. What is the phrase to describe the random way that homologous chromosomes can line up and separate during Meiosis I?
No, independent assortment
1. A dihybrid cross is one in which there are…
A. two (heterozygous) alleles being considered
B. two genes (both heterozygous) being considered
C. two genes (at least one heterozygous) being considered
D. two genes (both homozygous, but with different phenotypes) being considered
B
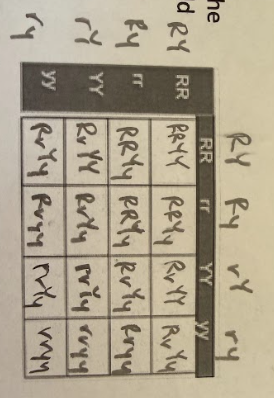
Suppose someone set the Punnett square up this way for the following cross: RrYy x RrYy. Is this correct? If not, how would you fix it?
Not correct because the gametes should carry only one allele for each trait and each gamete should have an allele for both traits. Each side of the Punnett square should have the following gametes: RY, Ry, rY, and ry
It is not correct. They need to put the potential genotypes of the gametes. They should have an allele for each of the genes, not two alleles for the same gene in each box.
1. In mice, dwarfism is caused by an X-linked recessive allele (Xd), and pink coat is caused by an autosomal dominant allele (P). If a dwarf female from a pure line is crossed with a pink male from a pure line, what will be the phenotypic ratios in the F1 and F2 in each sex?
The cross is female Xd/Xd ; p/p x male XD/Y ; P/P, where P = dominant allele for pink and d = recessive allele for dwarf.
F1 1/2 XD/Xd : P/p (pink female)
1/2 Xd/Y ; P/p (dwarf, pink male)
F2 3/16 XD/Xd : P/P (pink female)
1/16 XD/Xd : p/p (wild-type female)
3/16 Xd/Xd : P/P (dwarf, pink female)
1/16 Xd/Xd : p/p (dwarf female)
3/16 XD/Y ; P/P (pink male)
1/16 XD/Y ; p/p (wild-type male)
3/16 Xd/Y ; P/P (dwarf, pink male)
1/16 Xd/Y ; p/p (dwarf male)
1. Consider the following cross: A/a; B/b; C/c; D/d; E/e x a/a; B/b; c/c; D/d; e/e
a. What proportion of progeny will phenotypically resemble (1) the first parent, (2) the second parent, (3) either parent, and (4) neither parent?
Because each gene assorts independently, each probability should be considered separately and then multiplied together for the answer.
For (1), 1/2 will be A, 3/4 will be B, 1/2 will be C, 3/4 will be D, and 1/2 will be E.
1/2 x 3/4 x 1/2 x 3/4 x 1/2 = 9/128 = 7%
For (2), 1/2 will be a, 3/4 will be B, 1/2 will be c, 3/4 will be D, and 1/2 will be e.
1/2 x 3/4 x 1/2 x 3/4 x 1/2 = 9/128 = 7%
For (3), it is the sum of (1) and (2) = 9/128 + 9/128 = 9/64 = 14%
For (4), it is 1 – (part 3) = 1 – 9/64 = 55/64 = 86%
1. Consider the following cross: A/a; B/b; C/c; D/d; E/e x a/a; B/b; c/c; D/d; e/e
a. What proportion of progeny will be genotypically the same as (1) the first parent, (2) the second parent, (3) either parent, and (4) neither parent?
For (1), 1/2 will be A/a, 1/2 will be B/b, 1/2 will be C/c, 1/2 will be D/d, and 1/2 will be E/e.
1/2 x 1/2 x 1/2 x 1/2 x 1/2 = 1/32 = 3.1%
For (2), 1/2 will be a/a, 1/2 will be B/b, 1/2 will be c/c, 1/2 will be D/d, and 1/2 will be e/e.
1/2 x 1/2 x 1/2 x 1/2 x 1/2 = 1/32 = 3.1%
For (3), it is the sum of (1) and (2) = 1/16 = 6.2%
For (4), it is 1 – (part 3) = 1 – 1/16 = 15/16 = 93.8%
1. What is the chi-squared test/what is it used for in genetics?
The chi-squared test is a statistical test used to compare the observed allele frequency in a population to the expected allele frequency in a population, assuming the null hypothesis that inheritance of any allele is equally likely (unlinked genes on separate chromosomes, independent assortment of chromosomes, and no lethal alleles).
1. A chi-square test comparing observed and expected numbers of progeny is carried out, and the probability associated with the calculated chi-square value is 0.72. What does this probability represent?
A. Probability that the correct results were obtained.
B. Probability of obtaining the observed numbers.
C. Probability that the difference between observed and expected numbers is significant.
D. Probability that the difference between observed and expected numbers could be due to chance.
D
Given 130 coin flips, what is the expected number of heads up? What is the expected number of tails up?
What is the null hypothesis?
The outcomes of heads up or tails up on the coins are equally likely.
There is no difference between seeing heads and seeing tails. AKA seeing heads or tails is equally likely.
Given 130 coin flips, what is the expected number of heads up? What is the expected number of tails up?
Observed: 67 heads, 63 tails
Expected: 65 heads, 65 tails
Set up the chi-square equation for the class results and solve:
How many degrees of freedom are present? DF =( # of categories or progency classes – 1) = 2 – 1 = 1
f. Use the chi square table to determine p.
g. Using a p value of 0.05, was the null hypothesis met with respect to coin flips? What could it mean if the null hypothesis was not met?
d) X2 = 0.123
e) df = 1
f) between 0.9 and 0.5 0.9 > p > 0.5
g) Conclusion using p = 0.05
Your p-value is much greater than 0.05
Therefore:
You FAIL to reject the null hypothesis
Interpretation (this is the key part)
The null hypothesis = the coin is fair (50/50 heads and tails)
Since you failed to reject it:
The results are consistent with a fair coin
Any difference (67 vs 63) is likely due to random chance
1. From a presumed testcross A/a x a/a, in which ‘A’ represents red and ‘a’ represent white, use the χ2 test to find out which of the following possible results would fit the expectations:
a. What is the null hypothesis for all crosses? What are the expected phenotypic values? How many degrees of freedom exist?
The organism being tested is a heterozygote and that the A/a and a/a progeny are of equal viability. The expected values would be that phenotypes occur with equal frequency. There are two phenotypes, so 1 degree of freedom.
1. From a presumed testcross A/a x a/a, in which ‘A’ represents red and ‘a’ represent white, use the χ2 test to find out which of the following possible results would fit the expectations:
a. 500 red, 540 white
χ2 = [(500 – 520)2/520] + [(540 – 520)2/520]
= 1.538; p > 0.10, nonsignificant; hypothesis cannot be rejected
1. From a presumed testcross A/a x a/a, in which ‘A’ represents red and ‘a’ represent white, use the χ2 test to find out which of the following possible results would fit the expectations:
a. 5000 red, 5400 white
χ2 = [(5000 – 5200)2/5200] + [(5400 – 5200)2/5200]
= 15.385; p < 0.005, significant; hypothesis must be rejected
From a presumed testcross A/a x a/a, in which ‘A’ represents red and ‘a’ represent white, use the χ2 test to find out which of the following possible results would fit the expectations:
a. What do ‘b’ through ‘d’ suggest about sample size?
The larger the sample size, the more power the statistical test has.
1. A presumed dihybrid of two independently assorting genes in Drosophila, B/b; F/f, is testcrossed with b/b;f/f. (B = black body; b = brown body; F = forked bristles; f = unforked bristles.) The results are:
black, forked 260 brown, forked 200
black, unforked 210 brown, unforked 250
Use the χ2 test to determine if these results fit the results expected from testcrossing the hypothesized dihybrid.
a. What is the null hypothesis? What are the expected phenotypic values? How many degrees of freedom exist?
The hypothesis is that the organism being tested is a dihybrid with independently assorting genes and that all progeny are of equal viability. The expected values would be that phenotypes occur with equal frequency. There are four phenotypes so there are 3 degrees of freedom.
There is no difference between body color and bristle genes in flies. The genes are unlinked/independently assorting. 25% brown forked, 25% brown unforked, 25% black forked, 25% black unforked.
1. A presumed dihybrid of two independently assorting genes in Drosophila, B/b; F/f, is testcrossed with b/b;f/f. (B = black body; b = brown body; F = forked bristles; f = unforked bristles.) The results are:
black, forked 260 brown, forked 200
black, unforked 210 brown, unforked 250
Use the χ2 test to determine if these results fit the results expected from testcrossing the hypothesized dihybrid.
a. What is the χ2 value? Can you accept your null hypothesis?
χ2 = Σ (observed – expected)2/expected
χ2 = [(260 – 230)2/230] + [(210 – 230)2/230] + [(200 – 230)2/230] + [(250 – 230)2/230]
= 11.3; p value is about 0.01, which means the differences observed are likely NOT due to chance. Reject the null hypothesis.
1. A presumed dihybrid of two independently assorting genes in Drosophila, B/b; F/f, is testcrossed with b/b;f/f. (B = black body; b = brown body; F = forked bristles; f = unforked bristles.) The results are:
black, forked 260 brown, forked 200
black, unforked 210 brown, unforked 250
Use the χ2 test to determine if these results fit the results expected from testcrossing the hypothesized dihybrid.
c. What does this result suggest about the color gene and the bristle gene?
That they might be on the same chromosome since they didn’t seem to follow independent assortment.
The color gene and bristle gene are linked and not independently assorted.
1. Write a null hypothesis regarding the color distribution of m&ms.
Any given m&m has an equal, random chance of being any of the colors. (nothing is causing an unbalanced ratio in m&m colors)
There will be an equal distribution of each color of M&Ms in the package (color: blue, yellow, green, pink, purple).
Observed = blue 5, purple 13, green 4, yellow 8, pink 15
Expected = 9 of each.
1. Calculate a chi-squared value for the color distribution of m&ms.
1. How many degrees of freedom are present in this test?
1. Using the chi-squared value, what conclusion can you draw about the color distribution of m&ms?

1. What is a gene?
A sequence of DNA that encodes a functional RNA or protein
A location on a chromosome that encodes for a functional RNA or protein that contributes to an individuals phenotypical characteristics and behaviors (it is inheritable).
1. What is an allele? How are they created?
An allele is a different version (in terms of DNA sequence!!) of that gene. They are created via random mutations.
An allele is a specific version of a gene at a specific locus (they have different DNA sequences). Different alleles are produced through spontaneous mutations that alter the DNA sequence.
How many different alleles can one person have for a given gene? Why?
Up to 2 if they are heterozygous, just 1 if they are homozygous for that gene. Each diploid person gets one allele of a gene from their mom and one from their dad.
A person can only have 2 alleles for a specific trait, they inherit one maternal copy and one paternal copy (these alleles are on two different chromosomes).
1. How many different alleles can exist for a gene in a given species? Why?
Unlimited! Different alleles result from DNA changes anywhere along the length of the gene. Therefore, there can be many, many alleles that exist in a population. They might not all cause a functional protein.
Several alleles can exist for a gene in a given species because of random mutations that lead to changes in the DNA sequence, which lead to alterations in the encoded proteins and phenotypical expression/observable traits, creating different alleles.
1. Researchers are studying a different STR locus with one possible sequence from an individual as follows: (showing only a top strand of DNA)
5’GGCACTTAGGGAACCCTCACTGAATGAATGAATGAATGAATGAATGAATGAA
TGTTTGGGCAAATAAACGCT 3’
What is the STR sequence?
How many repeats are present in this individual?
The part that repeats GAAT, 8
1. From the slide in class, 9 TCAT repeats at the TH01 locus results in a PCR product of 194 bases. How long would the PCR product be if a person had 3 repeats?
The difference between 9 TCAT repeats and 3 TCAT repeats is 6 TCAT repeats. Since TCAT is 4 nucleotides long, that is a difference of 6 * 4 = 24 nucleotides.
9 repeats is a PCR product of 194 bp
Therefore 3 repeats is 194 – 24 = 170 bp
1. Explain epistasis.
genotype (or allele) at one locus (gene) masks the phenotype of a gene at a second locus, regardless of second gene’s genotype
1. A dog breeder mated a yellow Labrador male and a brown Labrador female hoping for yellow and brown puppies. However, all the puppies produced in this cross were black. What are the genotypes of the parent and puppy labradors?
The yellow male must be ee to be yellow. The brown female must be bb to be brown. Since all the puppies from this cross were black, each puppy needs a B allele (black hair) and an E allele (not yellow). Male dog must be BB ee. Female dog must be bb EE. Puppies are Bb Ee.
1. A snapdragon plant that bred true for white petals was crossed with a plant that bred true for purple petals, and all the F1 had white petals. The F1 was selfed. Among the F2, three phenotypes were observed in the following numbers:
white 240
solid purple 61
spotted purple 19
Total 320
a. Propose an explanation for these results – does this suggest independently acting genes, or a epistatic relationship? How can you tell? If an epistatic relationship is suspected, which one and why?
The parents are both true-breeding (homozygous). The F1s are all double heterozygous (setting up a dihybrid cross). The F2s are in an approximate 12:3:1 ratio. Such a ratio deviates from a 9:3:3:1 ratio associated with the results of a dihybrid cross of independently-assorting and non-interacting genes, indicating dominant epistasis.
1. A snapdragon plant that bred true for white petals was crossed with a plant that bred true for purple petals, and all the F1 had white petals. The F1 was selfed. Among the F2, three phenotypes were observed in the following numbers:
white 240
solid purple 61
spotted purple 19
Total 320
a. showing genotypes of all generations (make up and explain your symbols).
Because a modified 9:3:3:1 ratio was obtained in the F2, the F1 had to be a double heterozygote. The F1s were also all white, meaning that AaBb individuals are white. Additionally, we know that aabb is the least likely outcome in the F2s and corresponds with spotted purple plants. In order to achieve a double heterozygote in the F1, the original white parent has to be A/A; b/b and the solid purple parent has to be a/a; B/B.
So, if the original cross was A/A ; b/b (white) x a/a ; B/B (purple). The F1 would all be A/a ; B/b. The F2 would be:
9 A/– ; B/– white, by definition
3 A/– ; b/b white, by deduction
3 a/a ; B/– purple, by definition
1 a/a ; b/b spotted purple, by deduction
Because a modified 9:3:3:1 ratio was obtained in the F2, the F1 had to be a double heterozygote. The F1s were also all white, meaning that AaBb individuals are white. Additionally, we know that aabb is the least likely outcome in the F2s and corresponds with spotted purple plants. In order to achieve a double heterozygote in the F1, the original white parent has to be A/A; b/b and the solid purple parent has to be a/a; B/B.
So, if the original cross was A/A ; b/b (white) x a/a ; B/B (purple). The F1 would all be A/a ; B/b. The F2 would be:
9 A/– ; B/– white, by definition
3 A/– ; b/b white, by deduction
3 a/a ; B/– purple, by definition
1 a/a ; b/b spotted purple, by deduction
a. Draw out a biochemical pathway to explain these data.
Under these assumptions, A blocks the expression of both B and b. The a allele has no effect on the expression of B and b. B results in solid purple, while b results in spotted purple.
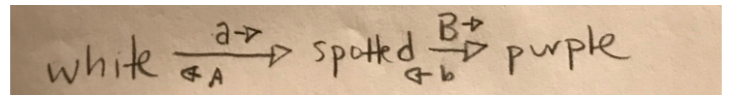
1. In roses, the synthesis of red pigment is by two steps in a pathway, as follows:
colorless intermediate (gene P) ——→ magenta intermediate (gene Q) ———> red pigment
a. What is the phenotype and genotype of a pure-breeding plant homozygous for a null mutation of only gene P?
gene P doesn’t work (nonfunctional enzyme)
phenotype: homozygous recessive (pure breeding) ppQQ
genotype: colorless intermediate (gene P nonfunctional, so can’t progress to magenta or red).
1. In roses, the synthesis of red pigment is by two steps in a pathway, as follows:
colorless intermediate (gene P) ——→ magenta intermediate (gene Q) ———> red pigment
a. What is the phenotype and genotype of a pure-breeding plant homozygous for a null mutation of only gene Q?
gene Q is nonfunctional
phenotype: PPqq, homozygous dominant for P, homozygous recessive for q
genotype: magenta intermediate (gene P works, Q doesn’t)
1. In roses, the synthesis of red pigment is by two steps in a pathway, as follows:
colorless intermediate (gene P) ——→ magenta intermediate (gene Q) ———> red pigment
a. What is the phenotype and genotype of a plant homozygous for null mutations of both P and Q?
phenotype: ppqq homozygous recessive for both genes
genotypes: colorless intermediate (genes to make magenta and red are nonfunctional).
1. In roses, the synthesis of red pigment is by two steps in a pathway, as follows:
colorless intermediate (gene P) ——→ magenta intermediate (gene Q) ———> red pigment
a. What F2 ratio is expected from crossing plants from parts a and b? (Assume independent assortment.) (2 pt)
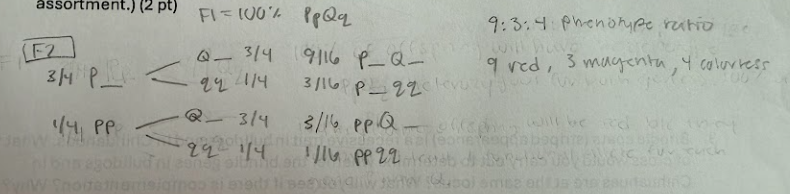
In roses, the synthesis of red pigment is by two steps in a pathway, as follows:
colorless intermediate (gene P) ——→ magenta intermediate (gene Q) ———> red pigment
a. What epistasis pattern does this suggest?
Recessive epistasis- the homozygous recessive condition of gene P (pp) is epistatic to the expression of gene Q.
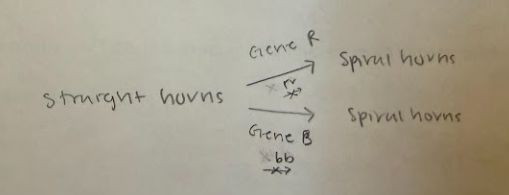
1. Spiral horns in dragons is caused by the presence of at least one dominant allele in each of two independently segregating gene pairs (e.g., R_B_). Dragons with rrbb genotypes have straight horns, and dragons with genotypes R_bb and rrB_ have spiral horns. You cross a dragon homozygous for spiral horns (RR BB) with a dragon homozygous for straight horns.
a. What are the relative proportions of the phenotypic classes expected in the F2 progeny after selfing the F1 progeny? (2 pts)
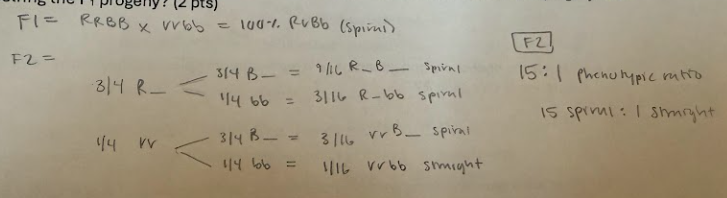
1. Spiral horns in dragons is caused by the presence of at least one dominant allele in each of two independently segregating gene pairs (e.g., R_B_). Dragons with rrbb genotypes have straight horns, and dragons with genotypes R_bb and rrB_ have spiral horns. You cross a dragon homozygous for spiral horns (RR BB) with a dragon homozygous for straight horns.
a. What epistasis pattern does this suggest?
Duplicate dominant epistasis
1. Spiral horns in dragons is caused by the presence of at least one dominant allele in each of two independently segregating gene pairs (e.g., R_B_). Dragons with rrbb genotypes have straight horns, and dragons with genotypes R_bb and rrB_ have spiral horns. You cross a dragon homozygous for spiral horns (RR BB) with a dragon homozygous for straight horns.
a. Draw out a basic biochemical pathway (like at the beginning of question 1) that could explain the results in the observed phenotypes. Make sure to correlate your gene/allele symbols with any potential enzymes in your pathway. (3 pts)
1. Brindle coats (striped appearance) is a recessive trait in bulldogs and in Chihuahuas. What type of cross would you carry out to determine whether the brindle genes in bulldogs and in Chihuahuas are at the same locus? What will you see if there is complementation? Why? (3 pts)
You would carry out a complementation cross to determine whether the brindle gene in bulldogs and chihuahuas are at the same locus, which involves crossing a homozygous recessive brindle bulldog with a homozygous recessive brindle chihuahua. If complementation occurs the F1 generation will all be heterozygous for each gene and have non-brindle coats b/c each parent provides one functional allele of a different gene (assuming they are at different loci), which restores the pathways/makes it functional. Brindle coats would only appear in F1 if genes was at same locus, in which each parent would be supplying a recessive allele making the gene nonfunctional (homozygous recessive).
1. Assume independent assortment and start with a plant that is A/a; B/b; CC
a. Using a branch diagram, what phenotypic ratio (and associated genotypes) is produced from selfing it?
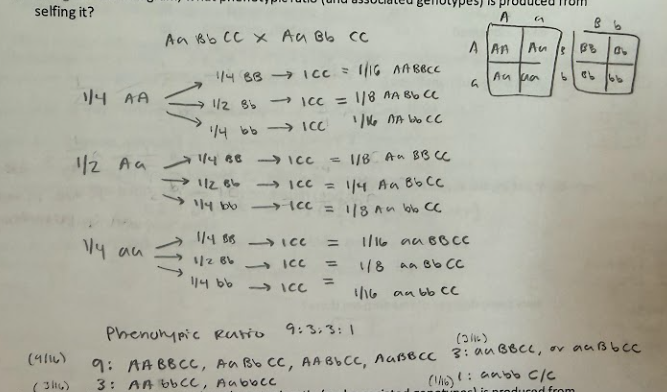

1. Assume independent assortment and start with a plant that is A/a; B/b; CC
a. Using a branch diagram, what phenotypic ratio (and associated genotypes) is produced from testcrossing it?

1. Are the following progeny numbers consistent with the results expected from selfing a plant presumed to be a dihybrid of two independently assorting genes, H/h; R/r? (H = hairy leaves; h = smooth leaves; R = round ovary; r = elongated ovary.) Explain your answer.
hairy, round 176 smooth, round 55
hairy, elongated 64 smooth, elongated 25
a. What is the null hypothesis?
There will be a 9:3:3:1 phenotype ratio for a dihybrid self-cross: 9 hairy round, 3 hairy elongated, 3 smooth round, and 1 smooth elongated. Assuming H and R are independently assorting genes and H, h and R, r are equally likely to be inherited.
1. Are the following progeny numbers consistent with the results expected from selfing a plant presumed to be a dihybrid of two independently assorting genes, H/h; R/r? (H = hairy leaves; h = smooth leaves; R = round ovary; r = elongated ovary.) Explain your answer.
hairy, round 176 smooth, round 55
hairy, elongated 64 smooth, elongated 25
What are the expected values?
(176 + 64 + 55 + 25) / 4 = 20
180 hairy round, 60 hairy elongated, 60 smooth round, and 20 smooth elongated.
1. Are the following progeny numbers consistent with the results expected from selfing a plant presumed to be a dihybrid of two independently assorting genes, H/h; R/r? (H = hairy leaves; h = smooth leaves; R = round ovary; r = elongated ovary.) Explain your answer.
hairy, round 176 smooth, round 55
hairy, elongated 64 smooth, elongated 25
How many degrees of freedom are there?
a. What is the χ2 value and the associated p value? Can you accept the null hypothesis?
df = 3

A dominant M allele in a loci in Jack-O-Lanterns (jols) results in an open mouth, while mm at that loci results in a closed mouth. At a different loci, the dominant T allele results in teeth, while tt at that loci results in a lack of (no) teeth.
An Mm Tt jack-o-lantern and an Mm tt jack-o-lantern mate.
What is your null hypothesis regarding the inheritance of the M/m and T/t alleles in jack-o-lanterns?
There is equal probability of inheriting the M/m and T/t alleles for each gene. No relationship between M and T genes, they are independently assorting.
M_Tt 3/8, M_tt 3/8, mmTt 1/8, mmtt 1/8
3:3:1:1 ratio expected
A dominant M allele in a loci in Jack-O-Lanterns (jols) results in an open mouth, while mm at that loci results in a closed mouth. At a different loci, the dominant T allele results in teeth, while tt at that loci results in a lack of (no) teeth.
An Mm Tt jack-o-lantern and an Mm tt jack-o-lantern mate.
The F1 generations are as follows: 22 jols with open mouths and teeth, 24 jols with open mouths and no teeth, 18 jols with no mouth (and no teeth).
Calculate X2 and determine how many degrees of freedom there are and what the associated p-value is…
3 degrees of freedom, p-value = < 0.005

A dominant M allele in a loci in Jack-O-Lanterns (jols) results in an open mouth, while mm at that loci results in a closed mouth. At a different loci, the dominant T allele results in teeth, while tt at that loci results in a lack of (no) teeth.
An Mm Tt jack-o-lantern and an Mm tt jack-o-lantern mate.
The F1 generations are as follows: 22 jols with open mouths and teeth, 24 jols with open mouths and no teeth, 18 jols with no mouth (and no teeth).
What is your conclusion about the X2 statistical test?
We reject the null hypothesis of equal probability of inheritance of M/m and T/t genes and the 3:3:1:1 expected ratio. The allele inheritance is not due to random change alone b/c the P-value is less than 0.005 (less than the statistically significant value of 0.05). Not independently assorting.
A dominant M allele in a loci in Jack-O-Lanterns (jols) results in an open mouth, while mm at that loci results in a closed mouth. At a different loci, the dominant T allele results in teeth, while tt at that loci results in a lack of (no) teeth.
An Mm Tt jack-o-lantern and an Mm tt jack-o-lantern mate.
The F1 generations are as follows: 22 jols with open mouths and teeth, 24 jols with open mouths and no teeth, 18 jols with no mouth (and no teeth).
What is an alternate explanation for the inheritance of the M/m and T/t alleles based on the observed F1 ratios? Include an illustration (biochemical pathway) of your proposed mechanism, labelled with all allele options and which form of epistasis you are covering.

What is responsible for most of the variation that arises in each generation? (origins of genetic variation)
The behavior of chromosomes during meiosis and fertilization.
What four mechanisms contribute to genetic variation?
1.Crossing over (Prophase I)- of meiosis 1
2.Independent assortment of chromosomes (Metaphase/Anaphase I)
3.Random fertilization (after completion of meiosis I and II)- after completion of meiosis 1 and 2- random combination of sperm and egg.
4.Mutations! (can happen any time) (can happen any time, changes in DNA sequence leads to different alleles).
Draw the two possible independent assortment outcomes for a cell that is 2n=2
Each of these arrangements are equally likely b/c the process is random.
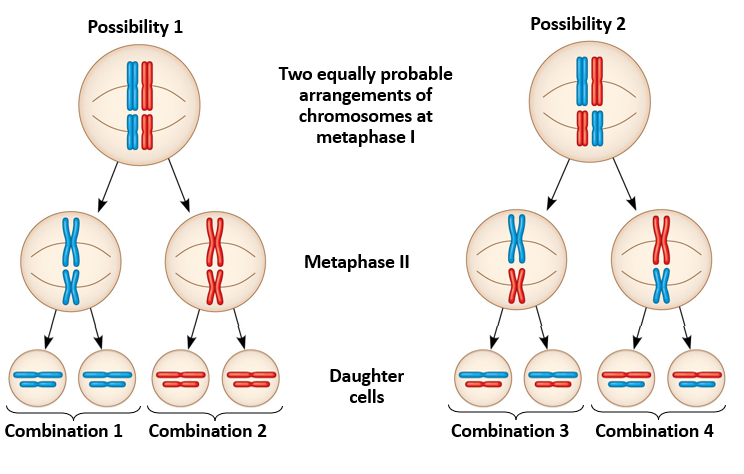
What is independent assortment of chromosomes?
Homologous pairs of chromosomes orient randomly at metaphase 1 of meiosis.
Each pair of chromosomes sorts maternal and paternal homologs (#’s of homologous pairs) into daughter cells independently of the other pairs.
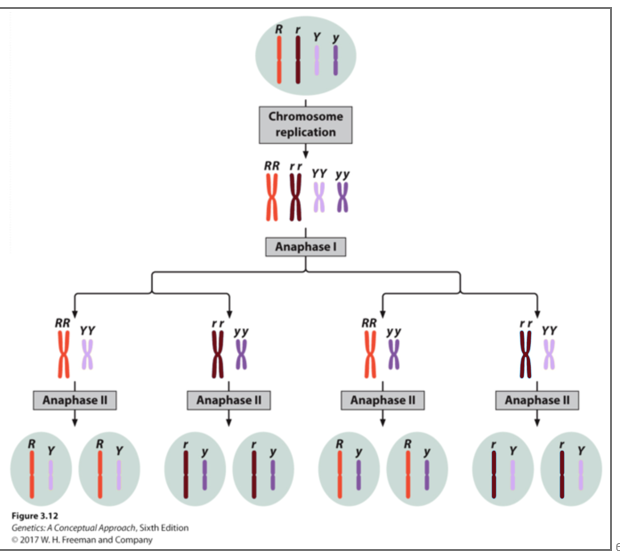
How do you determine the number of possible combinations of chromosomes in independent assortment?
The number of combinations possible when chromosomes assort independently into gametes is 2n, where n is the haploid number.
For humans (n = 23), there are more than ____ million (223) possible combinations of chromosomes that can occur in independent assortment. With random fertilization there is how many possibilities?
8 million: this is just for one person
Random fertilization (223 x 223) = 70 trillion possibilities!! (any given zygote: random sperm and egg)
This does not include crossing over, mutations, or recombination!!
What occurs with the independent assortment of 2 unlinked genes in a dihybrid cross?
A dihybrid cross is when you have two genes and each parent is heterozygous for both genes. (heterozygous gives the most combination of allele sin the offspring).
Produces a 9:3:3:1 ratio
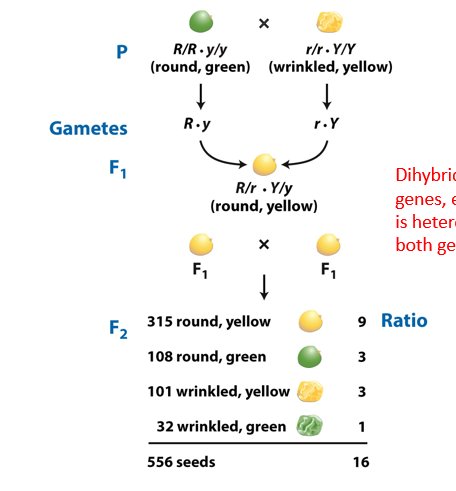
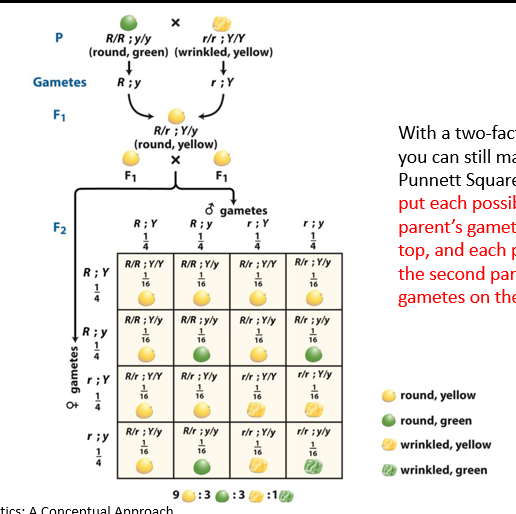
How do you draw a Punnett square for a dihybrid cross (genotypes underlying phenotypic 9:3:3:1 ratio)?
With a two-factor cross, you can still make a Punnett Square. Be sure to put each possibility for one parent’s gametes on the top, and each possibility for the second parent’s gametes on the side.
Need each possibility of alleles of parents on each side (gamete possibilities along the top and the side).
How do you set up a branch diagram?
It uses the law of multiplication
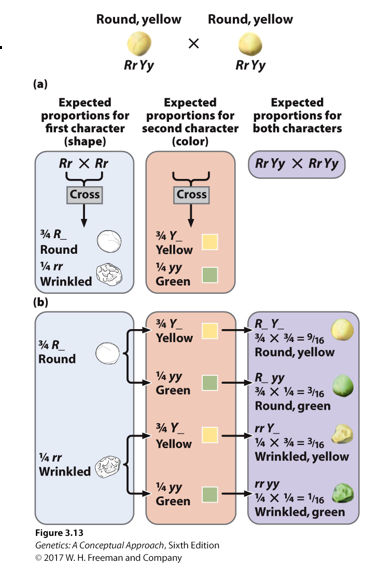
When should you draw branch diagrams?
If you have a 3-factor cross or more, use a branch diagram.
What is a test cross?
Crossing an individual (sometimes with unknown genotype) to a homozygous recessive individual.
What are the uses for test crosses? (3)
1) determination of an unknown genotype
2) determine whether genes are on the same chromosome or different chromosomes
3) also useful for certain mapping experiments
Chi square statistical test:
When you have a genetic hypothesis, you expect to see certain results. Usually framed using null hypothesis. What is a null hypothesis?
No differences or relationships exist between variables
(Ex. Null Hypothesis: An individual is Aa. Each allele is viable and equally likely to be inherited. Expectation: When test-crossed with an aa individual, ½ of the offspring will display the A_ phenotype.)
If you do this cross, and the observed numbers are not exact…is this just due to chance? Or is your hypothesis wrong? Enter: the Chi-Square test!!
What is the formula for the Chi square test?

What is P and df in a table of critical values of the X2 distribution?
P, probability value; df, degrees of freedom (# of classes or progeny clases-1). Most scientists assume that, when P < 0.05, a significant difference exists between the observed and the expected values in a chi-square test.
What is the p-value for statistical significance?
P of 0.05-5% change that the results aren’t random. P-value less than equal to 0.05 is the threshold for statistical significance.
What kind of p-value do scientists look for?
want a small p-value b/c it suggests your results are statistically significant and the results weren’t due to random chance (if they want their hypothesis to be supported).
What is the p-value or probability value?
The probability that the result you got was random. (translates to a percentage).
Mystery Dragon Practice Problem: SGSW FF? Use X2 to support or disprove hypothesis:
Null hypothesis SGSW alleles are equally likely to be inherited and the expected offspring (from a test -cross) is 50% 50%. If it had 10 babies we would see 5 G and 5 W babies. Actually observed: 7 green 3, white.

Describe the one gene, one enzyme experiment by Beadle and Tatum 1941
They took two media: 1) minimal medium- sugar, salts, and biotin, 2) complete medium- sugar, salts, amino acids, and many vitamins
They took spores and subjected them to X-rays, which caused random mutations in some of the spores.
They let the x-rayed spores reproduce and make offspring.
They then transferred each of the offspring spores to its own tube of complete medium and then transferred part of each colony to minimal medium. (The nutritional mutant can’t grow on minimal medium.
The nutritional mutant is transferred to a tube with minimal medium and a full set of vitamins, and doesn’t grow.
The nutritional mutant is transferred to a tube with minimal medium and 20 amino acids. The mutant is rescued by an amino acid mix→ mutation must block the synthesis of one or more amino acids.
The mutant is placed in minimal medium with one of the amino acids (20 total tubes). The mutant is rescued by arginine→ mutation must disrupt arginine biosynthesis.
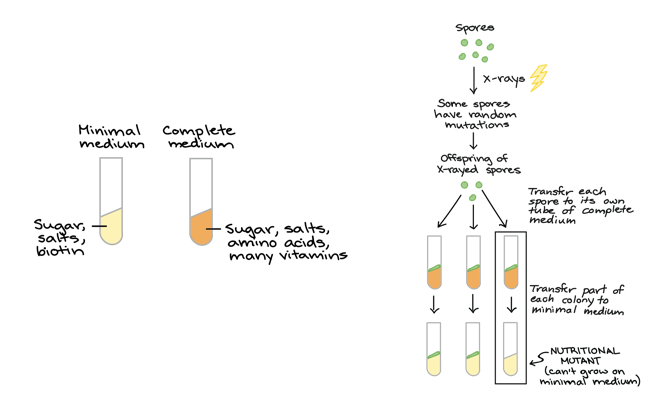
What were the key takeaways of the one gene one enzyme experiment done by Beadle and Tatum?
Individual genes were leading to individual enzymes (aka proteins)
Today we know some genes encode a protein that’s not an enzyme, a subunit of a protein, or don’t encode for proteins at all.